Abstract
Sclerotinia sclerotiorum is a notorious fungal pathogen with a broad host range, including many important crops. A previous study showed dill seed essential oil (DSEO) could inhibit S sclerotiorum pathogenicity and protect canola production. However, the molecular basis of DSEO anti-fungal activity is still not well studied. To investigate the mechanism of DSEO anti-fungal activity, RNA-sequencing was employed to identify differentially expressed genes (DEGs) of S sclerotiorum in response to DSEO treatment. A total of 2470, 3218, and 3793 DEGs were identified in S sclerotiorum after being treated by DSEO for 0.5, 1, and 2 h, respectively. These genes that express changes in the early stage are more likely affected directly by DSEO. Gene Ontology (GO) analysis revealed that these genes were mainly related to transmembrane transport, cell membrane, ribosome biogenesis, and proteasome complex. DSEO treatment primarily affected the membrane part of the fungal cell, particularly the endoplasmic reticulum (ER) membrane at 0.5 and 1-hour treatment. In addition, a bunch of DEGs associated with the proteasome pathway was markedly enriched at 2 h of treatment. It is speculated that DSEO achieves antifungal effects by influencing these targets or pathways. The information obtained in this study expanded the understanding of the antifungal mechanism of DSEO and enriched the resources available for interpreting its mechanism at molecular level.
Introduction
Fungal pathogens are seriously threatening natural ecosystems and food security. Sclerotinia sclerotiorum (Lib.) de Bary is a devastating soil-borne fungal pathogen that infects more than 500 host plant species and annually costs millions of dollars.1,2 The fungus forms sclerotia that can live for up to 4 to 5 years in the soil and infect plants by producing mycelium or ascospores.2,3 Canola production is threatened by this yield-limiting disease because there are no utterly resistant crop cultivars commercially available.4,5 In addition, mycotoxin production of sclerotia as a potential mutagenic and genotoxic agent and its contamination of crops or foods may lead to toxic effects on human health. 6 The use of chemical fungicides, procymidone, and boscalid are currently effective methods for controlling Sclerotinia diseases. However, overuse of chemical fumigants has led to several disadvantages, such as resistance strains, toxic residues, and negative environmental effects. 7 Thus, investigators seek effective plant-derived substances, such as plant extracts and essential oils, to control plant diseases. To date, essential oils, as potential natural products for fungal inhibition, have been widely used in combating diseases and food preservation.8,9
Dill (Anethum graveolens L.) is an annual herb in the celery family Apiaceae, which is native to the Mediterranean coast and grown widely in Eurasia and American countries. In China, dill is primarily cultivated in the southwest and northwest region. Essential oils extracted from dill seed have been proven to possess antibacterial and antifungal activities in the pharmacological and agriculture field.10‐15 According to a previous report, dill seed essential oil (DSEO) inhibits fungal pathogen (Aspergillus flavus) growth by affecting the plasma membrane and mitochondrial membrane, which causes reactive oxygen species (ROS) generation in fungal cells. 15 DSEO can also remarkably suppress S sclerotiorum, causing morphological alterations of hyphae and sclerotia. 13 Within the cell, DSEO inhibits ergosterol synthesis, succinate dehydrogenase, and malate dehydrogenase activities. 13 Ergosterol is an indispensable component of the cell membrane, regulating membrane fluidity, asymmetry, and integrity of a fungal cell. 16 Malate dehydrogenase and succinate dehydrogenase are mitochondrial enzymes involved in adenosine triphosphate (ATP) biosynthesis.17,18 Thus, the plasma membrane and mitochondria were considered potential targets of DSEO. However, no studies have been reported regarding the precise molecular target of DSEO in fungal cells. No report has been found on fungal cells’ global gene expression pattern in response to DSEO. The molecular mechanism by which DSEO inhibits fungal cells remains unknown to date.
In this study, we performed genome-wide gene expression analysis of S sclerotiorum treated with/without DSEO through RNA-sequencing. DSEO regulated genes related to coding membrane integral protein, which involved material transmembrane transport, protein synthesis, and degradation. We also suggested disruption of the ubiquitin/proteasome function by DSEO may lead to programmed cell death (PCD) in fungal cells. Our study first provided the view of the molecular events in S sclerotiorum underlying the response to DSEO. It could provide the foundation for research into the molecular-genetic antifungal mechanism of DSEO.
Materials and Methods
Pathogen and Chemicals
S sclerotiorum strains (1980) 19 were gifted by Dr Jiatao Xie from Huazhong Agricultural University. Dill (A graveolens L.) seeds were purchased from an herbal material market in Yunnan Province (China) and identified by pharmacognosy with reference to those in Chinese Pharmacopoeia (2020). The extract of DSEO was obtained following the method used in previous studies.13,20 In brief, 200 g dill seeds were grounded by a grinder and hydrodistilled for 5 h using a Clevenger-type apparatus (SS85-1000). DSEO was dried and stored in a glass vial with sealing film at 4 °C. The oil recovery of dill seeds oil was 1.71%.
Gas Chromatography–Mass Spectrometry Analysis
The analysis of the chemical composition of DSEO was performed in the gas chromatography–mass spectrometry (GC/MS) system (Agilent 7890/5975) and separated by the HP-5ms column (30m × 0.25mm × 0.25μm) (Agilent Technologies, USA). 20 μL DSEO was dissolved in 480 μL of ethyl acetate and 1 μL of this solution was injected into the GC-MS system. Helium was used as carrier gas at a flow rate of 1 mL/min. The parameter of chromatographic separation of compounds was the GC oven temperature from 80 °C (1 min) to 180 °C at a rate of 5 °C/min, increased to 260 °C at a rate of 10 °C/min. The temperatures of quadrupole, ion source, interface, and inlet were 150 °C, 230 °C, 250 °C, and 250 °C, respectively. The mass spectrometry (MS) scans were acquired in the 20 to 500 m/z range (full scan mode) for 35 min with a solvent delay of 3 min. The combination of the correspondence of the linear retention index (RI) was calculated using a standard mixture of C8-C30 n-alkanes (Shanghai Yuanye Bio-Technology Co., Ltd) according to the Van den Dool and Kratz formula. The mass spectra with respect to those reported in the National Institute of Standards and Technology (NIST17) database was used as an additional parameter for peak assignment.
S sclerotiorum Culturing and DSEO Treatment
S sclerotiorum was cultured on potato dextrose agar (PDA) medium in an incubator set up at 25 °C until further use. Different concentrations of DSEO were dissolved in PDA medium (containing 0.1% Tween 20) to obtain series final concentrations of 2, 1, 0.5, 0.25, 0.125, and 0.0625 μL/mL. PDA medium without DSEO was used as a control. The medium containing essential oil was transferred to glass Petri plates and solidified half an hour later. Then, an agar block (6 mm diameter) containing mycelial material was cut from a 3-day cultured S sclerotiorum and transferred to the center of the newly prepared Petri plate with DSEO. The Petri plates were sealed and incubated at 28 °C ± 2 °C. Control plates without DSEO were set up in the same procedure. The hyphal diameter was measured after incubation for 24 h. The lowest concentration of DSEO that is sufficient to 100% inhibits the visible in vitro growth of S sclerotiorum in PDA is the minimum inhibitory concentration (MIC).
To test the effect of DSEO against S sclerotiorum in infecting plants, Arabidopsis detached leaves were sprayed with 0.5 μL/mL DSEO dissolved in 0.1% Tween 20 solution. Control plants were sprayed with a mock solution containing 0.1% Tween 20. The plants were inoculated with S sclerotiorum after treatment. DSEO sprayed and control plants were grouped in different boxes and sealed to maintain humidity.
Microscopy Imaging
Fungal images were captured by a Confocal Laser Scanning Microscope equipped with an argon/krypton laser (Leica TCS SP5). To visualize the DNA of fungal cells, samples were treated with 100 ng/ml 4',6'-diamidino-2-phenylindole hydrochloride (DAPI) in phosphate buffered saline (PBS) for 30 min. DAPI fluorescent signals were imaged at the excitation wavelength of 358 nm and emission wavelength of 461 nm.
RNA Extraction, Library Preparation, and Sequencing
For each treatment, S sclerotiorum was cultured in 100 mL flasks containing 50 mL of potato dextrose broth (PDB) medium and grown at 25 °C on an orbital shaker at 140 rpm for 24 h before DSEO treatment. The concentration of DSEO for treatment was 0.5 μL/mL. The mycelia were harvested at 0.5, 1, and 2 h after DSEO treatment. The mycelia without DSEO treatment were considered the control. Both control and treatment groups performed 3 biological repeats.
RNA of control and treatment groups was extracted using the Trizol method. 1 μg RNA was used as input material for the library preparations. The RNA quality was determined by Agilent 2100 Bioanalyzer (Agilent Technologies). Moreover, messenger RNA (mRNA) was purified using oligo(dT) coupled beads. First-strand cDNA was prepared using M-MuLV reverse transcriptase mixed with random primers, and the second-strand cDNA synthesis was subsequently performed by employing DNA polymerase I. After adaptor ligation, cRNA libraries were amplified from purified cDNA fragments by Phusion High-Fidelity DNA polymerase. Library fragments (370-420 bp) were purified with the AMPure XP system (Beckman Coulter) and then sequenced on an Illumina Novaseq platform (150 bp paired-end).
RNA-Seq Data Analysis
Raw data were processed to obtain clean data (clean reads) by removing adaptor sequences, reads containing N base, and low-quality reads. Then, clean reads were mapped to the S sclerotiorum reference genome in the National Center for Biotechnology Information (NCBI) database using HISAT2 v2.0.5. Gene expression levels were estimated using feature Counts v1.5.0-p3 by counting the reads mapped to each gene. Gene expression level was calculated by Fragments Per Kilobase of exon per Million fragments mapped (FPKM).
Differentially expressed genes (DEGs) between the control and treated groups was calculated using the DESeq2 R package (1.20.0). Genes with an adjusted P value less than .05 and the absolute value of fold change more than 2 identified by DESeq2 were considered DEGs. Gene Ontology (GO) enrichment analysis of DEGs was performed using the clusterProfiler R package.
Quantitative RT-PCR Validation and Analysis
1 μg RNA was used as input material for the cDNA preparations. First-strand cDNA was synthesized from DNase I treated RNA with a HiScript III-RT SuperMix for qPCR (Vazyme). The reaction mixture (10 µL) contained RNA (2 µL), 5 × gDNA wiper Mix (2 µL), and RNase-free water (6 µL). After DNA was removed at 42 °C for 2 min, 10 × RT Mix (2 µL), HiScript III Enzyme Mix (2 µL), Oligo(dT) (1 µL), and RNase-free water (5 µL) were added into the reaction mixture. cDNA was synthesized at 37 °C for 15 min, and then the reaction stopped at 85 °C for 5 s.
Gene-specific primers for quantitative RT (qRT)-PCR analysis were designed through the NCBI primer blast tool (https://www.ncbi.nlm.nih.gov/tools/primer-blast/). The S sclerotiorum Ubiquitin gene was used as the reference gene. The reaction mixture (20 µL) contained a cDNA template (2 µL), 2 primers (10 µM, 0.4 µL each), SYBR Green Supermix (10 µL) (Bio-Rad), and RNase-free water (7.2 µL). Three technical repeats were performed for each sample. CFX384 touch real-time PCR detection system (Bio-Rad) was employed to run qRT-PCR. PCR was conducted via thermal cycling (95 °C for 10 s, 60 °C for 30 s, and 40 cycles). Gene expression levels were analyzed using the CFX Manager software (Bio-Rad) by the ΔΔCT method.
Results
Identified Components of DSEO
A total of 26 different components of the DSEO were identified by the GC-MS analysis (Table 1). The results showed that the main components of the DSEO consist of D-limonene, 4-allylanisole, dihydrocarvone, D-carvone, anethole, and apiole with content varying between 4.73% and 27.00%, representing 96.68% of the total oil. The other 20 identified components were minor in the oil, representing 3.27% of the total oil.
Chemical Constituents of the dill Seed Essential oil (DESO) Determined by the Gas Chromatography–Mass Spectrometry (GC/MS) Analysis.

Abbreviation: RI, retention index. Bold indicates the main components of DSEO.
Interference of S sclerotiorum Hyphal Growth by DSEO
To investigate the MIC of DSEO against S sclerotiorum for transcriptome analysis, we tested the hyphal growth of S sclerotiorum after different concentrations of DSEO treatment. DSEO completely inhibited fungal growth at the concentration of 0.5 μL/mL after 24 h of incubation (Figure 1A), indicating that the MIC of DESO against S sclerotiorum at 24 h was 0.5 μL/mL. Then, we detected the effect of 0.5 μL/mL of DSEO on the hyphal morphology of S sclerotiorum after 4 h of treatment (Figure 1B). We found that cell morphology changed significantly, including evidently reduced vacuole structure, cell narrowing, and cytoplasmic separation (Figure 1B). Next, we examined the nuclear morphology of cells with or without DSEO treatment by DAPI staining. In S sclerotiorum cells, nuclear condensation was detected within the control cells, whereas nuclear material was loosed and distributed throughout the cell after DSEO treatment (Figure 1C). We also tested the DSEO antifungal activity on plant leaves (Figure 1D). Detached Arabidopsis leaves were sprayed with 0.5 μL/mL of DSEO before inoculation with S sclerotiorum mycelial agar plug. Figure 1D showed significantly suppressing lesion development in DSEO group after a 3-day infection. These results proved that 0.5 μL/mL of DSEO inhibited S sclerotiorum proliferation and pathogenicity.
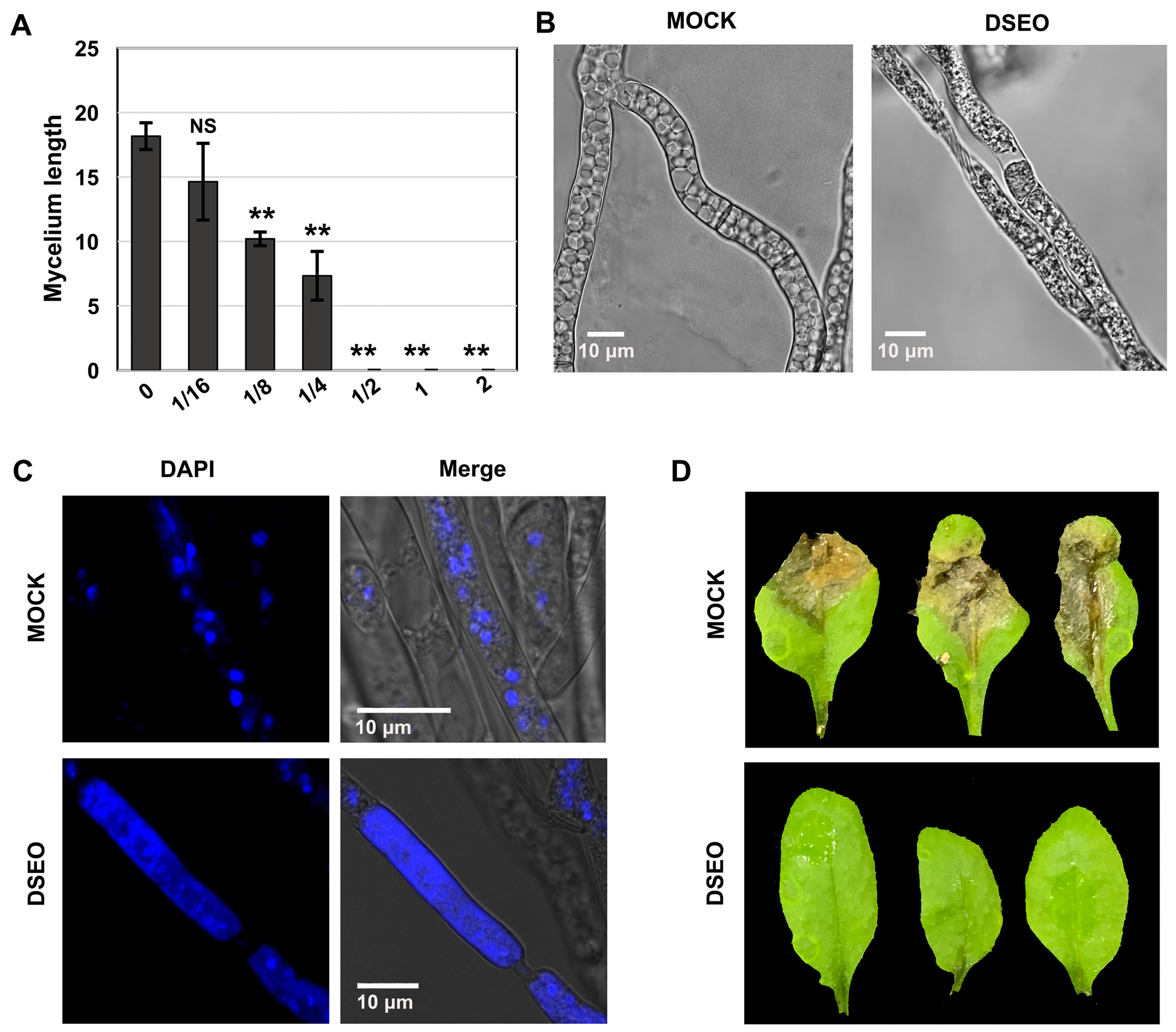
Antifungal effect of dill seed essential oil (DSEO) against Sclerotinia sclerotiorum. (A) Effects of different concentrations of DSEO (μL/mL) on hyphal growth in S sclerotiorum. (B) Effect of DSEO on hyphal morphology in S sclerotiorum. (C) Effect of DSEO on nuclear condensation in S sclerotiorum. Fluorescent microscopic pictures of the cells were recorded after nuclear staining with 4',6'-diamidino-2-phenylindole hydrochloride (DAPI). (D) Efficacy of DSEO against S sclerotiorum infection on Arabidopsis leaves. Statistical analysis was conducted by ANOVA Dunnett’s multiple comparisons test.
Overall Characteristics of RNA-seq Results
0.5 μL/mL of DSEO was used for treatment to profile the transcriptional response to DSEO in S sclerotiorum. RNA-Seq technique was performed on mycelia samples collected at 0.5, 1, and 2 h after DSEO treatment. The mycelia without DSEO treatment (0 h) were used as the control. Control and treatment groups performed triplicates, and 12 samples were collected for RNA-Seq (Table 2). After transcriptome sequencing, 79.46 GB of clean data was obtained. An average of 44,144,443 reads was obtained in each sample, and the Q30 value of each sample was higher than 92%. The clean reads were mapped to the S sclerotiorum reference genome, resulting in an average mapping percentage of 97.99% (Table 1). The correlation between any 2 triplicates for each treatment was > 95% (Supplementary Figure S1). The transcriptome sequencing quality and depth qualify for the following analysis.
Alignment Results of Sample Reads With the Reference Genome.

Statistical analysis of the RNA-Seq reads for Sclerotinia sclerotiorum before and after DSEO treatment. Mapping rate represented the percentage of mapped reads of the clean reads in each sample. SS_0, SS_1, SS_2, and SS_3 represented S sclerotiorum treated with DSEO for 0, 0.5, 1, and 2 h, respectively. Each treatment had 3 biological replicates.
DEG Profiles
We systematically compared the relative abundance of genes under DSEO treatment conditions (0.5, 1, and 2 h) with their abundance under normal conditions (0 h) to identify the dynamic expression patterns of DSEO-responsive DEGs. We found that thousands of genes were significantly and differentially expressed between DSEO treatment and normal conditions, whereas FPKM was used to define the expression level of each gene (Figure 2A). The results of differential expression analysis showed that most genes were regulated as early as 0.5 h (Figure 2A). Remarkably, these DEGs were further grouped into 2 clusters based on their expression patterns and constantly upregulated or downregulated during DSEO treatment, revealing that they were expressed in a specific regulation manner (Figure 2B). Comparing gene expression between samples treated for 0.5 h and 0 h (SS_1 vs SS_0), 2470 DEGs were found, comprising 1410 DEGs upregulated and 1060 DEGs downregulated (Figure 2C). A total of 3218 DEGs were found from the comparison between samples treated for 1 h and 0 h (SS_2 vs SS_0), of which 1912 were upregulated, and 1306 were downregulated (Figure 2C). Of all comparisons, 3793 DEGs were identified from the comparison between samples treated for 2 h and 0 h (SS_3 vs SS_0). Of these, 2334 were upregulated, and 1459 were downregulated (Figure 2C), suggesting that more genes were regulated at a more extended treatment period. The Venn diagram highlighted the shared and unique DEGs in each comparison (Figure 2D). The highest number of DEGs (2038) was shared by all comparisons. In particular, many DEGs (891) were specifically regulated in SS_3 versus SS_0 (Figure 2D). In contrast, the number of specific DEGs in the other 2 adjacent treatment periods (SS_2 vs SS_0, SS_1 vs SS_0) was almost identical, with 163 and 179 DEGs, respectively (Figure 2D). These data suggested that the expression profile of DEGs displayed significant differences, which may reflect specific cell changes in different treatment periods.

Overview of differentially expressed genes (DEGs) in Sclerotinia sclerotiorum response to dill seed essential oil (DSEO) treatment. (A) Volcano diagrams showed DEGs in S sclerotiorum treated with DSEO for 0.5, 1, and 2 h compared with 0-hour treatment. The red and green dots represented DEGs whose expression was upregulated and downregulated, respectively. The blue dots indicated the differences in gene expression, which were insignificant. The absolute value of log2 (fold change) ≥ 1 and P value <.05 indicated that the differences in gene expression were significant. (B) A cluster heat map illustrated the expression pattern of DEGs of RNA-Seq results in S sclerotiorum treated with DSEO for 0, 0.5, 1, and 2 h. (C) Number of DEGs in (B). All: Number of all DEGs; Up: Number of upregulated genes; Down: Number of downregulated genes. (D) Venn diagram depicted the number of DEGs and the overlapping DEGs between each time point.
Function Enrichment Analysis at Different Treatment Periods
GO enrichment annotations were classified into biological processes (BPs), cellular components (CCs), and molecular functions (MFs). Most of the DEGs in the 3 different treatment periods were enriched in similar MF terms, which were annotated as binding activity, indicating that binding-associated functions were regulated by DSEO treatment (Figure 3A). Guanyl nucleotide binding, heme binding, and tetrapyrrole binding were enriched in 3 treatment periods (Figure 3A). Two early adjacent treatment periods (SS_2 vs SS_0 and SS_1 vs SS_0) were enriched in similar GO terms, such as GO terms related to membrane; endoplasmic reticulum (ER) function in CC terms; and GO terms related to DNA integration, metabolic process, transportation, and protein glycosylation in BP terms (Figure 3B and C). Compared with SS_2 versus SS_0 and SS_1 versus SS_0, the DEGs of SS_3 versus SS_0 were enriched in the specific CC and BP terms, such as GO terms related to cytoskeleton, proteasome complex, and ribosome biogenesis (Figure 3C).

Scatter plot of Gene Ontology (GO) enrichment statistics for differentially expressed genes (DEGs) in Sclerotinia sclerotiorum treated with dill seed essential oil (DSEO) for 0.5, 1, and 2 h compared with that 0 h treatment. We displayed pathways that were enriched at (A) molecular function (MF), (B) cellular component (CC), and (C) biological process (BP).
Directed Acyclic Graph Representation of the Overrepresented GO Terms of CCs
The directed acyclic graph (DAG) was a graphical display method of the GO enrichment analysis results of DEGs. The branch represented the inclusion relationship, and the function range defined as the lower node function was subordinate to the upper node function. The top 5 most significant GO terms were selected for the DAG. The depth of the color represented the degree of enrichment. We made a DAG of enriched GO terms of CC at different treatment periods (Figure 4). Functional enrichment analysis of 2470 DEGs between samples treated for 0.5 and 0 h (SS_1 vs SS_0) revealed a significant enrichment (P = .01985) for genes with annotated membrane part (GO: 0044425) (Figure 4A). Other significantly over-represented GO terms, including endomembrane system (GO: 0012505), nuclear-ER membrane network (GO: 0042175), and ER membrane (GO: 0005789), were also enriched (Figure 4A). These results indicated that essential oil treatment primarily affected the membrane part of the fungal cell, particularly the ER membrane, as early as 0.5 h. Functional enrichment analysis of 3218 DEGs between samples treated for 1 and 0 h (SS_2 vs SS_0) revealed significantly enriched GO terms similar to SS_1 versus SS_0, and more GO terms related to peptidase complex (GO: 1905368) were enriched (Figure 4B). Furthermore, functional enrichment analysis of 3793 DEGs between samples treated for 2 and 0 h (SS_3 vs SS_0) revealed a significantly different enrichment for genes annotated with GO terms when compared with SS_1 versus SS_0 (Figure 4C). GO terms, including peptidase complex (GO: 1905368), proteasome core complex (GO: 0005839), and chromatin (GO: 0000785), were enriched (Figure 4C), indicating that more proteasome-related genes and chromatin genes were regulated at a more extended treatment period (2 h).

Directed acyclic graph (DAG) in the cellular component (CC) ontology for differentially expressed genes (DEGs) expressed in Sclerotinia sclerotiorum. (A) S sclerotiorum treated with dill seed essential oil (DSEO) for 0.5 h compared with 0-hour treatment, (B) S sclerotiorum treated with DSEO for 1 h compared with 0-hour treatment, and (C) S sclerotiorum treated with DSEO for 2 h compared with 0-hour treatment.

Differentially expressed genes (DEGs) involved in the endoplasmic reticulum (ER) function (A) and ribosome biogenesis (B). In (A) and (B), colors in the map represented the level of expression in the sample based on the RNA-Seq results. In (C), the relative expression levels of the 4 selected genes were measured by quantitative RT (qRT)-PCR.
DEGs Involved in the ER Function and Ribosome Biogenesis
DEGs in all 3 treatment periods were enriched in 1 of the 2 categories: ER membrane and ribosome biogenesis, which were considered critical components of cell protein translation (Figure 3). The relative expression levels of DEGs in different treatment periods were analyzed to identify DEGs associated with ER membrane in S sclerotiorum in response to DSEO treatment (Figure 5A). Only 1 gene, sscle_03g027530, encoded signal recognition SRP19/SEC65, was upregulated during DSEO treatment. Several genes related to signal peptidase complex, glucosyl transferases, ER lumen protein-retaining receptor, and Sec62/Sec66 domain-containing proteins, which exhibited the same expression pattern, were significantly downregulated (Figure 5A). Apart from ER function, several DEGs involved in ribosome biogenesis were differentially expressed in all 3 treatment periods (Figure 5B). DEGs involved in ribosomal proteins, rRNA methyltransferases, proteins related to ribosome and rRNA biogenesis, and some snoRNA-associated proteins were significantly downregulated in the 3 treatment periods (Figure 5B). These results indicated that protein synthesis, processing, and transport in the ER membrane were inhibited by DSEO treatment. In confirming the accuracy and predictability of the RNA-Seq results, qRT-PCR was employed to quantify the transcriptional levels of selected genes. Four DEGs, including sscle_03g027530, sscle_16g109460, sscle_01g00668, and sscle_03g030240, were randomly selected and evaluated. Data showed that each gene was significantly differential expression after DSEO treatment at different periods (Figure 5C). The differential expression pattern of selected DEGs between transcriptome data and qRT-PCR was consistent, indicating that RNA-Seq data were reliable.
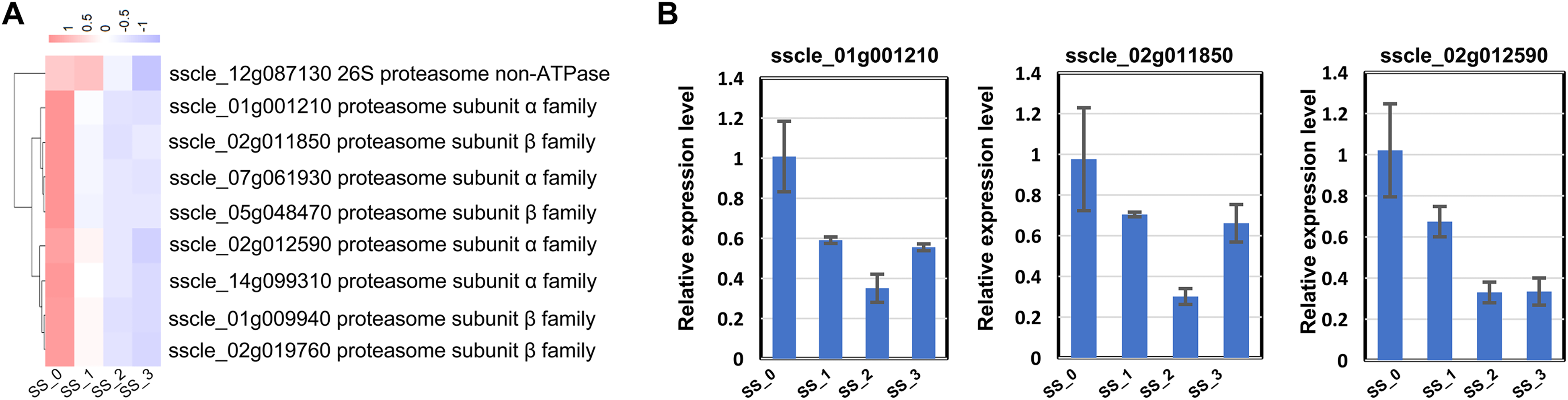
Differentially expressed genes (DEGs) involved in the proteasome complex. (A) Colors in the map represented the level of expression in the sample based on the RNA-Seq results. (B) The relative expression levels of the 3 selected genes were measured by quantitative RT (qRT)-PCR.
DEGs Involved in the Proteasome Complex
Proteasomes were protein complexes that degraded unneeded or damaged proteins by proteolysis and played a critical role in maintaining the homeostasis of cellular proteome. 21 In the present study, 1 26S proteasome ono-ATPase and 8 subunits were identified in the GO term proteasome complex (Figure 6A). Interestingly, all DEGs were downregulated in the proteasome complex (Figure 6A). We also performed qRT-PCR analysis of 3 randomly selected DEGs, including sscle_01g001210, sscle_02g011850, and sscle_02g012590 (Figure 6B), showing the same expression pattern of selected DEGs between qRT-PCR and transcriptome data, which further supported the reliability of the RNA-Seq data.
Discussion
In the present study, we characterized and investigated the antifungal activity of DSEO and the mechanisms underlying S sclerotiorum response to DSEO by transcriptome analysis. The MIC value of DSEO against S sclerotiorum on mycelial growth was 0.5 μL/mL at 24 h posttreatment. A previous study reported a higher MIC value (1.0 μL/mL) and a more extended growth period (4 days) required for thoroughly inhibiting the mycelial growth and sclerotial germination of S sclerotiorum. 13 We showed spraying 0.5 μL/mL DSEO on Arabidopsis leaves could significantly suppress lesion development, suggesting that DSEO has the potential to be a novel fungicide in crops for controlling Sclerotinia diseases. Although the previous study announced that inhibition of ergosterol synthesis, succinate dehydrogenase activities, malate dehydrogenase, and external medium acidification could explain the antifungal mechanism of essential oil, 13 the molecular mechanism by which DSEO inhibited fungal cells remained unknown. Nevertheless, transcriptional regulation of gene expression in fungal cells played a crucial role in the response of fungi to various stresses.22‐24 However, no report has been found regarding fungal cells’ global gene expression profile in response to DSEO treatment.
In this study, transcriptional profiling of S sclerotiorum showed that DSEO affected the expression of genes related to membrane part, transmembrane transport, ER part, DNA integration, cytoskeleton, proteasome complex, and ribosome biogenesis. Disruption and alteration of genetic material, inhibition of cell wall/cell membrane formation, and dysfunction of mitochondria have been proposed to the mechanism of the antifungal activity of essential oils.8,25‐27 The cell wall is a characteristic fungi structure that maintains cell morphology and protects the cell from osmotic and mechanical stresses.28,29 The activity of essential oil caused the cell wall to rupture and finally transform into an empty hyphal tip, which branches into a bud-like structure.30,31 However, DEGs related to cell wall integrity and biosynthesis/degradation have not been enriched in S sclerotiorum exposed to DSEO. This result may be due to essential oils’ non-transcriptional regulation of cell wall formation. GO-enriched terms such as membrane part, transmembrane transport, ER part, and DNA integration supported the previous conclusion that DSEO disrupted cell membranes and genetic material. Our results further demonstrated several novel cytological components, including ER part, ribosome biogenesis, cytoskeleton, and proteasome complex, which were regulated or targeted by DSEO. The rough ER on its ribosomes could produce proteins for the rest of the cell to function.32,33 Additionally, ER stress response could trigger PCD or apoptosis in mammals, plants, and fungus.34‐36 A strong correlation between the actin cytoskeleton and PCD or apoptosis has been reported in studies from plant and animal systems and yeast .37,38 The ubiquitin-proteasome system was the central executioner of apoptosis by triggering key cell death proteins, such as caspases.39,40 Accumulating polyubiquitinated proteins and declining proteasome activity activated the PCD program and demonstrated that proteasome dysfunction could lead to PCD. 41 Thus, DSEO treatment may induce ER stress, cytoskeleton, and proteasome-induced apoptosis or PCD in S sclerotiorum, finally inhibiting its growth and pathogenicity. These discoveries can be considered a potential antifungal mechanism by essential oils (Figure 7). Further study will determine the mechanism of DSEO in systemically and selectively affecting the transcription of S sclerotiorum genes, and searching key transcription factor targets would be the focus of the research. We also hypothesized that the new generation of essential oil-based fungicides would be applied in the future to control Sclerotinia diseases caused by fungus in crop production.

The molecular network recapitulates how Sclerotinia sclerotiorum cell responds to dill seed essential oil (DSEO) treatment.
Conclusion
Essential oils, natural products potentially for fungal inhibition, have been widely used in combating diseases and food preservation. However, the mechanism of essential oils in antifungal activity remained largely unknown. We identified DEGs in S sclerotiorum following inoculation with DSEO by RNA-Seq. A total of 2470, 3218, and 3793 DEGs have been identified in response to essential oil treatment at 0.5, 1, and 2 h, respectively. In addition, GO terms associated with membrane part, transmembrane transport, ER part, DNA integration, cytoskeleton, proteasome complex, and ribosome biogenesis were enriched in DEGs, which could provide references for further studies associated with the mechanism of action of essential oils’ antifungal activity. Furthermore, developing new generations of essential oil-based fungicides has significant potential for crop production in the future.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X221119910 - Supplemental material for Transcriptome Analysis Reveals the Mechanism of dill Seed Essential oil Against Sclerotinia sclerotiorum
Supplemental material, sj-docx-1-npx-10.1177_1934578X221119910 for Transcriptome Analysis Reveals the Mechanism of dill Seed Essential oil Against Sclerotinia sclerotiorum by Yu-Xin Chen, Wei Li, Hong Zeng, Gao Zhou and Qiang Cai in Natural Product Communications
Footnotes
Author’s Contribution
Y.X.C. and Q.C. conceived and designed the research. Y.X.C. performed most experiments and analyzed the data. W.L. performed bioinformatics analysis of deep RNA sequencing data. Y.X.C., H.Z., G.Z., and Q.C. analyzed the data. Y.X.C. and Q.C. drafted and revised the manuscript. Q.C. supervised the project. All authors approved the final version of the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Collaborative Grant-in-Aid of the HBUT National “111” Center for Cellular Regulation and Molecular Pharmaceutics, Doctoral Start-up Foundation of Hubei University of Technology, and Open Grant from Xinjiang Production & Construction Corps Key Laboratory of Protection and Utilization of Biological Resources in Tarim Basin, Hubei Provincial Natural Science Foundation of China (grant numbers XBTK-2020003, BSQD2020034, BRZD2002, 2020CFB197).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
