Abstract
Ergosterol is an essential component for fungi, including entomopathogenic fungi like Cordyceps. Cordyceps has been used as a traditional medicine in Japan and China and possesses various unique ergostane-type sterols, those exhibit bioactivities. In this manuscript, we reported the isolation of 2 new ergostane-type sterols, 1 and 2 along with ten sterols (3-12) from 7 strains of Cordyceps related fungal strains, Cordyceps takaomontana NBRC 101754, Metarhizium owariense NBRC 33258, Polycephalomyces formosus NBRC 109994, Cordyceps tuberculata NBRC 106948, Cordyceps tenuipes NBRC 108997, Cordyceps sp. NBRC 106954, and Tolypocladium paradoxum NBRC 106958 collected fruiting bodies of Ophiocordyceps heteropoda. In addition, the antitrypanosomal activity and antimicrobial activity of isolates were tested to find 6 showed the antitrypanosomal activity, and the minimum inhibitory concentration (MIC) value was confirmed as 1.41 µg/mL. In the antimicrobial assay, the MIC value of 8 against methicillin-resistant Staphylococcus aureus was determined to be 3.1 µg/mL.
Keywords
Introduction
Entomopathogenic fungi are not only used as biological pesticides due to their interesting life cycle of infecting insects, 1 but also possess various secondary metabolites which have valuable biological activities. Among them, Cordyceps is used as traditional medicines in China, Japan, and other Asian countries and secondary metabolites of them have been studied as sources of medicinal seeds. For example, fingolimod, developed from myriocin which was found in the fruiting bodies of Isaria sinclairii was approved for the treatment of multiple sclerosis by US Food and Drug Administration in 2010. 2 We have studied the secondary metabolites of Cordyceps and related entomopathogenic fungi to find novel and bioactive compounds.3,4 Sterols are a class of compounds that are essential for fungal survival. Ergosterol is not only a basic component of the fungal cell membrane but also plays an important role in their morphogenesis. Ergostane-type sterols were reported to show antitumor, 5 anti-inflammation, 6 immunomosuppressive, 7 antimicrobial activity, 8 and other various activities.9,10
African trypanosomiasis (sleeping sickness) is a fatal disease in East and West Africa and is caused by 2 subspecies of parasite Trypanosoma brucei gambiense and Trypanosoma brucei rhodesiense. 11 The third subspecies, Trypanosoma brucei brucei does not infect humans but causes infections in cattle. Trypanosomes frequently mutate cell surface proteins, making it difficult to develop a vaccine to prevent human infection. In addition, many studies were conducted to develop new medicines, however, only 0.1% of the total was invested in neglected diseases including trypanosomiasis, malaria, and leishmaniasis. 12 As for bacterial infections, methicillin-resistant Staphylococcus aureus (MRSA) also causes serious problems in hospitals and community settings. 13 S aureus is a gram-positive bacterium found in the human nasal mucosa and causes a variety of serious infections ranging from skin infections to pneumonia, and septicemia. The infection control of drug-resistant bacteria always requires new antibiotics and their appropriate use. Due to these circumstances, the development of new medicines and therapies is urgently needed. In this report, the isolation and structural elucidation of 2 new ergostane-type sterols (1-2) and other known compounds (3-12) were described. In addition, the antitrypanosomal activity and antimicrobial activity of isolates against S aureus and MRSA were investigated.
Result and Discussion
Structure Elucidation of Isolated Compounds
Each strain was cultured and chromatographed with the following methods shown in the experimental section to isolate 1 to 3 and 8 to 12. The fruiting bodies of Ophiocordyceps heteropoda were extracted and fractionated to yield 3 to 7.
Compound 1 was isolated from the culture of Cordyceps takaomontana NBRC 101754 as white amorphous. The molecular formula of 1 was determined as C28H44O2 from molecular ion peak [M] + m/z 412.3321 (calculated for 412.3441, Δ −2.0 mmu) by HRCIMS analysis with 7° of unsaturation. 1H and 13C nuclear magnetic resonance (NMR) analysis suggested that 1 was an ergostane-type sterol with a ketone group. Three substructures were estimated by the COSY spectrum and their bindings were determined by HMBC analysis (Figure 2). In the HMBC spectrum of 1, a long-range correlation observed between H-5 (δH 2.70)/C-6 (δC 201.4) revealed the ketone group placed at C-6. From these analyses above, a 3-hydroxyergosta-7,22-diene-6-one structure was constructed. In the NOESY spectrum, correlations between H-5/H-9 (δH 2.29) and H-9/H-14 (δH 2.07) were observed, indicating α orientation of H-5 (Figure S7). The coupling constant of H-3 (δH 4.21, quin, J = 2.8 Hz) was observed, which suggests the β-equatorial position of H-3. In the comparison of the 13C NMR data of 1 and the reported values of (22E,24R)-3β-hydroxyergosta-7,22-diene-6-one, 14 upfield shifts of 2.3 to 5.4 ppm were observed in C-1 to C-5 which carbons around C-3 due to the γ-gauche effect of 3α-OH group. Additionally, NOE correlation observed between H-19/H-2β (δH 1.62) and H-4β (δH 1.67), H-3/H-2α, H-2β, H-4α, and H-4β indicated the presence of an α-axial hydroxy group at C-3 and the structure of 1 was determined as (22E,24R)-3α-hydroxyergosta-7,22-diene-6-one (Figure 1).
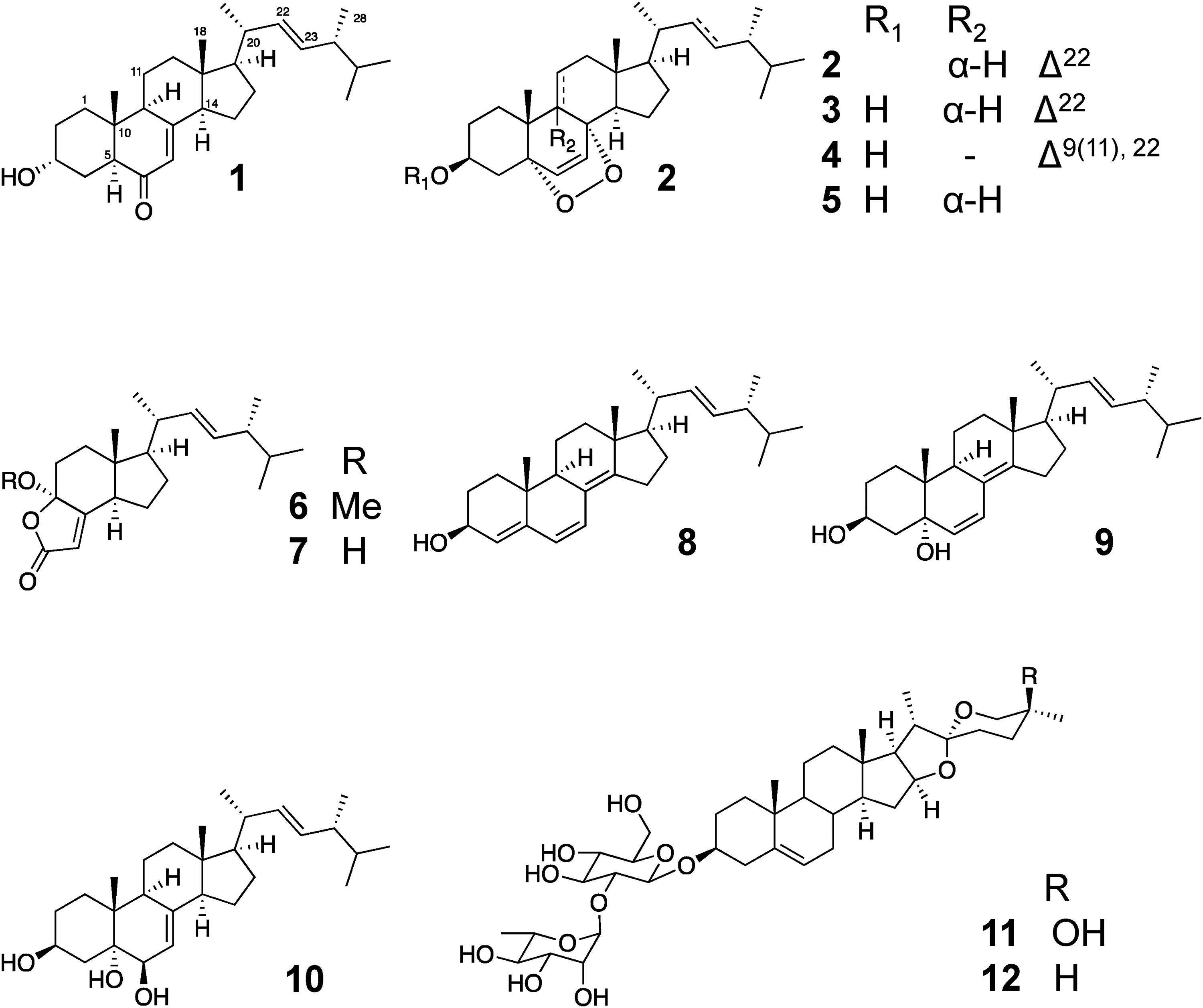
Structures of isolated compounds 1 to 12.

COSY and selected HMBC correlations of 1 and 2.
Compound 2 was isolated from the culture of Metarhizium owariense NBRC 33258. The molecular formula of 2 was proposed as C46H76O4 by HREIMS analysis with 9° of unsaturation. 1D and 2D NMR spectrum of 2 showed resemblance to those of ergosterol peroxide (3), indicating that 2 possess ergosterol peroxide structure as substructure. The remaining elements of 2, were deduced as C18H33O with 2° of unsaturation. The presence of unsaturated fatty acid moiety was suggested by the signals of terminal methyl (δH 0.88 ppm), methylene (δH 1.25-1.34 ppm), olefin protons (δH 5.33 and 5.35 ppm) in the 1H NMR spectrum. The position of an unsaturated bond of fatty acid moiety was analyzed by fragmentation patterns of the MS spectrum of 2. Fragment ions at m/z 567 and m/z 411, corresponding to [M–non-2-ene]+ and [M–octadec-11-enoic acid]+, which caused by the cleavage at the allyl position at C-9 and ester position at C-3, respectively (Supplemental Figure S16). The Z-configuration of C-11/C-12′ was determined from the coupling pattern of these corresponding proton signals which showed double triplets (J = 5.8, 11.2 Hz). The chemical shift value of an unsaturated olefin proton can be estimated by considering the effect of substituents. 15 The predicted chemical shift values of H-11′ and H-12′ also showed a more similar value of cis formation of H-11′ and H-12′ at δH 5.42 than these of trans formation of δH 5.48, supporting the Z-configuration of a double bond at C-11′/C-12′. Compare the 13C NMR of 2 and ergosterol peroxide (3), 2.7 ppm of upfield shift at C-3 (δC 69.2), and 3.8 and 3.7 ppm of downfield shifts at C-2 and C-4 of 2 were observed in 13C NMR, indicating fatty acid moiety connected to C-3 of ergosterol peroxide. Fragment ion of m/z 411, which corresponds to aglycon moiety, also indicated this connection. The coupling pattern of H-3 was dddd (J = 11.5, 11.4, 5.3, 5.3 Hz), indicating the presence of H-3α. NOESY correlations observed between H-19/H-2β and H-4β, and H-3/H-1α, H-2α, and H-4α also clarified the configuration at C-3. From these analyses, the structure of 2 was determined as ergosterol peroxide-3-O-(11Z)-octadecenoate (Figure 1).
Ten known sterols were isolated from entomopathogenic fungal strains. Ergosterol peroxide (3) 16 was isolated from Cordyceps tenuipes NBRC 108997, Cordyceps sp. NBRC 106954 and fruiting bodies of O. heteropoda. 9,11-dehydroergosterol peroxide (4), 17 (3β,5α,8α)-5,8-epidioxyergost-6-en-3-ol (5), 18 volemolide (6), 19 and demethylincisterol A3 (7) 20 were isolated from the fruiting bodies of O. heteropoda. 3β,22E, ergosta-46,8(14),22-tetrane-3-ol (8) 21 and 3β,5α,22E, ergosta-6,8(14),22-triene-3,5-diol (9) 22 were isolated from Polycephalomyces formosus NBRC 109994. From the culture of C. takaomontana NBRC 101754 and Cordyceps tuberculata NBRC 106948, (22E,24R)-ergosta-7,22-diene-3β,5α,6β-triol (10) 23 was isolated. Isonutageninn I (11) 24 and diosgenin 3-O-α-L-rhamnopyranosyl-(1→2)-β-D-glucopyranoside (12)25–27 were isolated from the EtOAc soluble fraction of fungal culture of Tolypocladium paradoxum NBRC 106958. The structures of these known compounds were determined by the comparison of reported values in each literature. Isolated compounds from each strain were summarized in Supplemental Table S1 (Table 1).
1H NMR (600 MHz) and 13C-NMR (150 MHz) Data of 1, (22E)-3β-Hydroxyergosta-7,22-diene-6-one and 2 in CDCl3.

Biological Activity of Compounds
As the evaluation of biological activities of isolates, antitrypanosomal activity, and antimicrobial activity were performed. 2 wasn’t applied for assays because of the amount of sample material. In our antitrypanosomal activity screening of natural products, a strain Trypanosoma brucei brucei strain GUTat 3.1, was applied. 28 Volemolide (6) showed strong activity of an IC50 of 1.41 µg/mL against T brucei, while 6 showed cytotoxicity against MRC-5 cells, human normal diploid fibroblasts with an IC50 value of 8.24 µg/mL. Antitrypanosomal activity of 6 in protozoa was comparable to those of the existing drug, suramin (IC50 of antitrypanosomal activity: 1.58 µg/mL, cytotoxicity against MRC-5 cells: >100 µg/mL), and bioactive compounds described in our previous report, 29 while the selectivity of 6 was lower than that of suramin. The IC50 value of ergosterol peroxide (3) against Trypanosoma cruzi, the cause of Chagas disease, was reported to be 6.74 µg/mL. 30 This activity was reported to be attributed to the peroxide moiety. Although the protozoan species were different, 6 showed comparable activity to 3. Detailed investigation of 6 may reveal a mechanism of action of ergosterols against Trypanosoma, not associated with the peroxide group.
In the antimicrobial assay, 8 and 3 showed growth inhibitory activity against gram-positive strains, S. aureus, and MRSA while others didn’t show significant activity. Especially, 8 showed growth inhibitory activity and the minimum inhibitory concentration (MIC) value was determined as 3.1 µg/mL, while the MIC value of oxacillin, a positive control, was 32 µg/mL. New compound 1 did not show significant bioactivity in this antimicrobial assay. (22E,24R)-3β-hydroxyergosta-7,22-diene-6-one, 3β-OH epimer of 1 was reported to inhibit antigen-induced degradation of RBL-2H3 cells 31 and human neutrophil elastase. 32 The hydroxy group at C-3 of ergosterol is reported not to affect the bioactivity such as anticancer activity and is used as a linkage position for preparing molecular proves.33,34 Considering these reports, 1 may show similar activity to its epimer. Isolation of ergosterol peroxide fatty acid ester was rarely reported from natural resources. 35 Recently, fatty acid ergosterol esters were synthesized by lipase-catalyzed reactions, and the antimicrobial activities of derivatives were described. 36 New ergosterol derivative 2 is expected to have similar activity.
Conclusion
In this report, 2 new ergostane-type sterols (1-2) and ten known sterols (3-12) were isolated from 7 entomopathogenic fungal strains and naturally occurring fruiting bodies of O. heteropoda. Ergosterol peroxides were isolated from some fungi, including Ganoderma lucidum and cordyceps. These compounds were reported to have biological activities, such as antitumor activity, anti-inflammatory activity, and antimicrobial activity. Compound 2, an ergosterol peroxide derivative with 11Z-octadecenoate moiety at C-3 was isolated from the culture of M. owariense NBRC 33258. In our screening studies, 8 showed strong growth inhibitory activity against MRSA, and 6 showed strong antitrypanosomal activity. To the best of our knowledge, there has been no research on the antimalarial activity of 6, indicating further analysis of those compounds and derivatives may contribute to the development of new medicines or laboratory tools.
Experimental
General Experimental Procedures
Optical rotations were recorded on a JASCO P-1030 polarimeter (Jasco). Infrared (IR) spectra were recorded on a Jasco FT/IR-410 spectrometer with an attenuated total reflectance (ATR) accessory, or Shimadzu FTIR-8400S instrument (Shimadzu). NMR spectra were recorded on JEOL ECA600 NMR (600 MHz, JEOL) or Bruker AVANCE III HD (500 MHz) spectrometers with deuterated solvent (CDCl3 or pyridine-d6, Kanto Chemical Co., Inc.), and the solvent chemical shifts were taken as the internal standard. MALDI-TOF MS was performed on a JMS-S3000 (JEOL). HREIMS and HRCIMS were obtained on a JEOL JMS-700 MS station. HRESI( + )MS was performed on a Q Exactive Focus (ThermoFisher Scientific). HPLC was performed using a JASCO LC-900 series instrument.
Fungal Material
Cordyceps strains, C. takaomontana NBRC 101754, M. owariense NBRC 33258, P. formosus NBRC 109994, C. tuberculata NBRC 106948, C. tenuipes NBRC 108997, Cordyceps sp. NBRC 106954 and T. paradoxum NBRC 106958 were purchased from the Biological Resource Center, National Institute of Technology and Evaluation (NBRC). Fruiting bodies of O. heteropoda were collected in Kyoto prefecture, Japan in 2007. The fruiting bodies of O. heteropoda were identified by morphological characteristics and a voucher specimen has been deposited in the Herbarium of Faculty of Pharmaceutical Sciences, Tokushima Bunri University.
Fungal Culture
Cordyceps strains were cultured in potato sucrose broth (PSB, 200 g potato, 20 g sucrose, and 1 L distilled water), on potato sucrose agar medium, added agar 15 g/L to PSB, Tre-P-Y medium (20 g trehalose, 3 g peptone, 1 g yeast extract, and 1 L distilled water) or oatmeal medium (50 g of oatmeal and 1 L of distilled water). Each strain was cultured at 25 °C, 120 rpm.
Extraction and Isolation
C. takaomontana NBRC 101754 was cultured in a PSB medium for 4 weeks (25 °C, 120 rpm). Mycelium was collected by filtration and extracted with MeOH twice and the mixture of CHCl3: MeOH (1:1) once. This extract was then partitioned with EtOAc and H2O to get 54.0 g of EtOAc soluble layer. EtOAc layer was then chromatographed over a silica gel column chromatography using an n-hexane: CHCl3: MeOH gradient system (1:1:0 to 0:1:9) to increase polarity to give 23 fractions (Fr.1A-1W). Fr.1I (401.2 mg) was then purified by Sephadex LH-20 column (ø36 × 470 mm) with CHCl3: MeOH (1:1) to yield 5 fractions (Fr.2A-2E). Fr.2E (201.2 mg) was purified by a silica gel column (SNAP Ultra, 10 g, Biotage) using an n-hexane: EtOAc gradient system to increase the polarity to get 8 fractions (Fr.3A-3H). Fr.3E (146.2 mg) was further purified by flash column chromatography (SNAP Ultra, 10 g) with CHCl3: MeOH gradient system to increase the polarity to isolate
M. owariense NBRC 33258 was cultured on PSA medium (500 mL × 20 flasks) for 4 weeks at room temperature. Mycelium was collected by filtration and extracted with MeOH twice and a mixture of CHCl3: MeOH (1:1) once. This extract was then partitioned with EtOAc and H2O to get 4.7 g of EtOAc soluble layer. The EtOAc layer was then chromatographed over a silica gel column chromatography using a CHCl3: MeOH: H2O gradient system (9:1:0 to 1:8.6:1.4) to increase polarity to give 9 fractions (Fr.1A-1I). Fr.1F (27.2 mg) was purified by silica gel column chromatography with n-hexane: CHCl3 (7:3) to get 2 fractions (2A-2B). Fr.2A (14.7 mg) was further purified by preparative TLC to isolate 9.7 mg of 2.
The fruiting bodies of O. heteropoda (31.0 g) was extracted with MeOH twice and CHCl3: MeOH (1:1) once to yield 4.7 g of extract. The extract was then partitioned with a silica gel column with n-hexane: CHCl3: 95% MeOH gradient system (1:1:0 to 0:1:9) and get 5 fractions (Fr. 1A-1E). Fr.1C (7.5 mg) was determined as volemolide (6). Fr.1D (385.0 mg) was chromatographed over a silica gel column with n-hexane: CHCl3: 95% MeOH gradient system (6:4:0 to 0:1:9) to give 10 fractions (Fr.2A-2J). Fr.2G (22.0 mg) was purified with a silica gel chromatography (eluent: n-hexane: EtOAc gradient system, 9:1 to 1:1) to yield 0.1 mg of demethylincisterol A3 (
C. tenuipes NBRC 108997 was cultured on PSA medium for 4 weeks at room temperature. The culture was extracted with MeOH twice and a mixture of CHCl3: MeOH (2:1) once. The extract was then partitioned with liquid-liquid separation to get 31.2 g of BuOH soluble layer. Then the BuOH layer was chromatographed on a silica gel column to give 17 fractions (Fr.1A-1Q). Fr.1F (360.0 mg) was purified with a silica gel column (30 g) to give 12.0 mg of ergosterol peroxide (3).
P. formosus NBRC 109994 was cultured on Tre-P-YA medium (500 mL × 32 flasks). Mycelium and medium were separated and extracted with 3 L of MeOH twice and 3 L of a mixture of CHCl3: MeOH (2:1) to get 21.5 g of extract. Mycelium extract was then suspended with H2O and partitioned with EtOAc to get EtOAc soluble layer (1.0 g). This layer was chromatographed over an amino silica gel column (SNAP NH 11 g) using n-hexane to give seven fractions (Fr.1A-1G). Fraction 1F (123.2 mg) was purified on Sephadex LH-20 column (ø36 × 500 mm) with CHCl3: MeOH (1:1) to yield 2 fractions (Fr.2A-2B). Fraction 2B (54.0 mg) was further purified with reverse phase flash column chromatography (SNAP Ultra C18, 12 g) to isolate 3.9 mg of 3β,5α,22E, ergosta-6,8(14),22-triene-3,5-diol (9). Fr.1E (88.5 mg) was then chromatographed with reverse phase flash column chromatography (SNAP Ultra C18, 12 g) to get Fr.3A-3C. Fr.3B was further purified with ODS HPLC (column: Cosmosil AR-II ø10 × 250 mm; UV wavelength: 205 nm; flow: 3.0 mL/min; solvent system 90%MeOH isocratic system; Rt: 24.1 min) to isolate 8.3 mg of 3β,22E, ergosta-46,8(14),22-tetrane-3-ol (8).
C. tuberculata NBRC 106948 was cultured with PSB medium and extracted to yield 19.8 g of MeOH extract. 9.0 g of extract was chromatographed over a silica gel column (900 g) to get ten fractions (1A-1J). Fr.1H (420.0 mg) was then purified by Sephadex LH-20 column (ø36 × 470 mm) using a mixture of CHCl3: MeOH 1:1 to get 4 fractions (2A-2D). In total, 18.0 mg of Fr.2C was further purified with ODS HPLC (column: Cosmosil AR-II ø20 × 250 mm; UV wavelength: 205 nm; flow: 3.0 mL/min; solvent system 0.1% AcOH MeOH/ 0.1% H2O gradient system; Rt: 23.0 min) to isolate 2.9 mg of cervesterol, (22E,24R)-ergosta-7,22-diene-3β,5α,6β-triol (
T. paradoxum NBRC 106958 was cultured in oatmeal medium (500 mL × 27 flasks) for 3 weeks at 25 °C (120 rpm). Mycelium was collected by centrifugation (5600 rpm, 20 min) and extracted with 6 L of MeOH twice and 6 L of a mixture of CHCl3 and MeOH (2:1) to get mycelial extract (42.8 g). Then, mycelial extraction was partitioned with liquid-liquid extraction using EtOAc, BuOH, and H2O to yield 20.0 g of EtOAc soluble layer. EtOAc layer was column chromatographed by a silica gel column (SNAP Ultra, 100 g) using an n-hexane: CHCl3: MeOH gradient system to increase the polarity to get 21 fractions (1A-1U). Fr.1Q (49.7 mg) was further purified with ODS flash column chromatography (SNAP Ultra C18, 12 g; H2O MeOH gradient system) and HPLC (Column: Cosmosil 5C18 PAQ ø20 × 250 mm; Flow: 4.0 mL/min, UV wavelength 205 nm; 50% to 60% MeOH gradient system) to isolate 13.2 mg of isonuatigenin I (11). Fr.1P (56.4 mg) was chromatographed over an ODS column (Cosmosil 75C18-OPN, ø15 × 80 mm; 10% to 100% MeOH gradient system) to get 5 fractions (Fr.2A-2E). Fr.2C (37.5 mg) was further purified by HPLC (Column: Cosmosil 5C18 PAQ ø20 × 250 mm; Flow: 4.0 mL/min, UV wavelength 205 nm; 50% to 60% MeOH gradient system) to isolate 3.8 mg of diosgenin 3-O-α-L-rhamnopyranosyl-(1→2)-β-D-glucopyranoside (12).
Spectral data of 1: white amorphous, 1H and 13C NMR: see Table 1; HRCIMS 412.3321 [M]+ (calculated for C28H44O2, 412.3341, Δ −2.0).;
Spectral data of 2: white amorphous, 1H and 13C NMR: see Table 1; HREIMS 692.5749 [M]+ (calculated for C46H76O4, 692.5744, Δ + 0.5).;
Supplemental Material
sj-docx-1-npx-10.1177_1934578X221105363 - Supplemental material for Ergostane-Type Sterols From Several Cordyceps Strains
Supplemental material, sj-docx-1-npx-10.1177_1934578X221105363 for Ergostane-Type Sterols From Several Cordyceps Strains by Tatsuro Yoneyama, Hiroki Takahashi, Aleksandra Grudniewska, Sayaka Ban, Akemi Umeyama and Masaaki Noji in Natural Product Communications
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online. NMR, MS, IR, and UV spectra of isolated compounds 1 and 2 are available.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
