Abstract
Due to the significantly negative impact of the coronavirus (CoV) disease (COVID-19) pandemic on the health of the community and the economy, it remains urgent and necessary to develop a safe and effective treatment method for COVID-19. Huang-Lian-Shang-Qing-Wan (HLSQW) is a herbal formula of traditional Chinese medicine (TCM) that has been applied extensively for treating “wind-heat-associated” symptoms in the upper parts of the body. The objective of the present in silico study was to investigate the potential effects of HLSQW in the context of severe acute respiratory syndrome (SARS)-CoV-2 infection. We analyzed the possible interactions between bioactive compounds within HLSQW on targets that may confer antiviral activity using network pharmacology and pharmacophore-based screening. HLSQW was found to potentially target a number of pathways and the expression of various genes to regulate cell physiology and, consequently, the anti-viral effects against SARS-CoV-2. Bioactive compounds contained within HLSQW may exert combined effects to reduce the production of proinflammatory factors, which may trigger the “cytokine storm” in patients with severe COVID-19. Results from molecular modeling suggested that the bioactive HLSQW components puerarin, baicalin, and daidzin exhibit high binding affinity to the active site of 3-chymotrypsin-like cysteine protease (3CLpro) to form stable ligand-protein complexes, thereby suppressing SARS-CoV-2 replication. In addition, our results also demonstrated protective effects of the HLSQW extract against cell injury induced by the proinflammatory cytokines tumor necrosis factor-α, interleukin (IL)-1β, and IL-6, against reactive oxygen species production and nuclear factor-κB activity in normal human lung cells in vitro. To conclude, HLSQW is a potential TCM remedy that warrants further study with the aim of developing an effective treatment for COVID-19 in the future.
Keywords
Introduction
Coronavirus (CoV) disease (COVID-19) was first identified towards the end of 2019 before rapidly developing into a global pandemic that severely affected human health worldwide. 1 At present, social interactions among economies have become severely hindered by the pandemic, with a severe adverse impact on the health of the community. Therefore, preventative measures and development of effective treatment strategies are required for this infectious disease. The most recent advancement in COVID-19 prevention is the advent of vaccines, which have been rolled out in a number of countries. However, concerns about a fresh outbreak remain due to the emergence of new virus variants, which convey additional uncertainties regarding vaccine efficacy and hinders the advancement of treatment strategies.2,3
Several repurposing agents, including remdesivir, chloroquine, and dexamethasone, have been urgently approved in a number of countries for treating COVID-19 due to their reported marginal benefit to patients. However, clinical results showed little or no improvements in the mortality rate.4-6 In addition, antiviral drugs have also been repurposed in a number of clinical trials for COVID-19, either as monotherapy or in combination.6-8 Since severe COVID-19 outbreaks remain in progress at present in a number of countries, finding a specific therapeutic option with high efficacy for treating this disease is urgently required. Investigations into applying traditional remedies to target COVID-19 have been encouraged by the governments of various countries. 9 Traditional medicine, including traditional Chinese (TCM), 10 Ayurveda, 11 and Unani, 12 has been attracting attention in the field of COVID-19 therapy. In particular, TCM is attracting the interest of researchers, as there have been initial promising reports of efficacy in the prevention, treatment, and recovery of patients with COVID-19.13,14 However, complementary studies on the pharmacological activities and the underlying mechanism of these remedies must be performed prior to further clinical trials.13,15
Huang-Lian-Shang-Qing-Wan (HLSQW) is a traditional concoction based on a formula that has been used extensively in TCM. It consists of the following: Rhizoma Coptidis (Huanglian), Huangqin, Huangbai, Gardenia jasminoides, Flos Chrysanthemi (Chrysanthemum), Angelica tail, Gegen, Mint, Figwort, Chuanxiong, Turmeric, Forsythia, Dahuang, and honey. 16 According to TCM, HLSQW possesses detoxifying activities and is particularly effective in alleviating adverse symptoms in the upper body, such as inflammatory conditions in the upper respiratory tract. In addition, this traditional formula has been standardized, quality-controlled, and modernized for safe and convenient use in TCM clinical practice. 16 Huanglian (Rhizoma Coptidis) 17 and Chrysanthemum (Flos Chrysanthemi) 18 are the main bioactive components of HLSQW that have been previously investigated for their effects on RNA viruses experimentally. In addition, components of HLSQW have also been reported to exert inhibitory effects on coronaviruses.19,20 These previous findings suggest that HLSQW may prove to be a potential remedy for COVID-19.
In the present study, the potential pharmacological effects of HLSQW were explored by deploying a number of integrated multidisciplinary methods, including ligand profiler, pharmacophore fitting, ingenuity pathway analysis (IPA), and molecular modeling using the Discovery Studio 2020 program. We focused on the potential regulatory roles of the bioactive compounds contained within HLSQW in the signaling pathways that can be manipulated by the viruses. Furthermore, we analyzed the possible effects of HLSQW on the “cytokine storm,” which is associated with the inflammatory symptoms exhibited by patients with severe COVID-19. The study design and schematics of the in silico target fishing for the components of HLSQW are presented in Figure 1. Data derived from this in silico assay may be useful for further studies on the therapeutic properties of HLSQW and its constituents on a molecular level, with the aim of developing a novel strategy for treating COVID-19.

Flowchart of the in silico target fishing protocol for the components of Huang-Lian-Shang-Qing-Wan.
Materials and Methods
In Silico Studies
Structure Preparation
The 3-chymotrypsin-like cysteine protease (3CLpro; PDB code: 6W63), papain-like protease (PLpro; PDB code: 6WX4), and RNA-dependent RNA polymerase (RDRP; PDB code: 7BV2) protein structures were retrieved from the Protein Data Bank. Each entity was prepared for fixing common structural problems by applying for the protonation states at pH 7.4 using the BIOVIA Discovery Studio 2020 software (Dassault Systèmes). The 29 components of HLSQW (aloe-emodin, berberine, baicalein, baicalin, berberine, chrysophanic acid, coptisine, curcumin, daidzin, daidzein, emodin, emodin-3-methyl ether, ferulic acid, ferulic acid, fructose, gardenin, glucose, glucoside, magnoflorine, iridoid, lignan, menthol, menthone, phenylethane, phellodendrine, palmatine, pectin, puerarin, and triterpenes) were sketched using BIOVIA Draw and prepared for generating the appropriate compound for protonated isomers and tautomers at pH 7.4.
In Silico Target Fishing by Pharmacophore Models
Interactive pharmacophore models were generated based on the complexities of the prepared protein and its corresponding ligand in situ. Each model was validated with their known inhibitors. The sensitivities of the pharmacophore models for 3CLpro, PLpro and RDRP were 0.73913 (true positive/total active: 17 of 23), 0.5 (true positive/total active: 1 of 2) and 0.75 (true positive/total active: 3 of 4), respectively. In total, 3 COVID-19-related target models and 16 035 pharmacophore models in PharmaDB were then used as screening targets for the 29 components of HLSQW. 21
Molecular Docking for 3-Chymotrypsin-Like Cysteine Protease (3CLpro)
The binding site on the 3CLpro (PDB code: 6W63) of SARS-CoV-2 was identified by using the “CDOCKER” protocol embedded into the Discovery Studio 2020 software. For each molecule, the top 10 docking poses were generated and ranked using the CDOCKER score (-CDOCKER interaction energy). In accordance with this, the top 4 compounds, namely baicalin, puerarin, daidzin, and iridoid, were selected for further binding interaction analysis alongside ritonavir. 21
Molecular Dynamics Simulation for the Ligand-Protein Complexes
The GROMACS package (version 2020.4) was used for all simulation courses. The ligand-protein complexes were prepared in UCSF Chimera (version 14; Regents of the University of California) based on a previous molecular docking result. 22 Topology was built for each complex using the CHARMM36 General Force Field (CGenFF version 4.4, updated July 2020) before the simulation box was created using water (TIP3P model) as solvent. Na+ and Cl− ions were used for charge neutralization. A simulation was prepared for each complex as described previously. 23 Briefly, the energy minimization step was conducted with 50 000 steps before the equilibrium stages (canonical ensemble-isothermal-isobaric ensemble, 50 000 steps) were assembled at a temperature of 300 K and pressure of 1 bar. Molecular dynamics (MD) production was performed using a computing system accelerated by a graphics processing unit with a 12-cores central processing unit run on Ubuntu 18.04. The output trajectory file was collected for root-mean-square deviation (RMSD) analysis and inter-molecule interactions.
Network Analysis Using Ingenuity Pathway Analysis (IPA)
A goodness-of-fit value of >0.6 was considered to be a potential compound target. To generate a corresponding molecular network from these targets, the 419 human target genes were set as focus molecules and analyzed using a core analysis tool in IPA (IPA 2020; Qiagen Sciences, Inc.). The enriched molecular networks and associated ontology groups in their respective canonical pathways, upstream regulators, diseases, functions, and rankings of network analysis were calculated based on statistical significance. All presented pathways and downstream functions were deemed to be statistically significant according to Fisher's exact t-test (P < .05). Specifically, the network was started from the focus molecule and from there each molecule was connected with others. Molecules with specific ontology associations in the QIAGEN knowledge base were then added to the network.
In Vitro Studies
Chemicals and Reagents
High-performance liquid chromatography (HPLC)-grade methanol, acetonitrile, and phosphoric acid were obtained from Sinopharm Chemical Reagent Co., Ltd, ultrapure water by a Milli-Q water purification system (EMD Millipore), and HLSQW from Taiji Group (No. 1915011). Nine standard compounds, liquiritin, baicalin, berberine, baicalein, aloe-emodin, rhein, isoimperatorin, chrysophanol, and physcion, were acquired from The National Institute for the Control of Pharmaceutical and Biological Products (Beijing, China). The purities of the standards were >98% through HPLC determination. 3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) and Hoechst 33342 were obtained from Sigma-Aldrich; Merck KGaA. CM-2′,7′-dichlorodihydrofluorescein diacetate (CM-H2DCFDA) was obtained from Thermo Fisher Scientific, Inc., Dulbecco's modified Eagle's medium (DMEM), fetal bovine serum (FBS), L-glutamine, penicillin G, and trypsin-EDTA from Life Technologies; Thermo Fisher Scientific, Inc., and nuclear factor κB (NF-κB) p65 Colorimetric Transcription Factor Assay Kit from Abcam.
HPLC Analysis
HLSQW (1.0 g) was extracted by methanol (25 mL) using ultrasound (100 W, 40 kHz) for 30 min. HPLC analysis was performed on a Waters 1525 system (Waters Corporation) equipped with an online vacuum degasser, automated injection value, and a diode-array detector. Chromatographic separation was performed on a WondaSil C18 Superb column (5 μm; 250 × 4.6 mm i.d.; Shimadzu Corporation). Water containing 0.3% phosphoric acid (solvent A) and acetonitrile (solvent B) were selected as the mobile phase in the following gradient elution protocol: 0 to 45 min, 10 to 35% solvent A; 45 to 70 min, 30 to 60% solvent A; 70 to 85 min, 60 to 90% solvent A; and 85 to 90 min, 90 to 5% solvent A. Column temperature was set at 25°C and the flow rate was 1.0 mL/min; the detection wavelength was 254 nm. 16
Ultra-Pure Liquid Chromatography (UPLC)-Q-Time of Flight/Mass Spectrometry (MS) Analysis
The HLSQW extract was prepared using the same protocol as that described in the “HPLC analysis” section. Chromatographic analysis was conducted using an ACQUITY UPLC I-Class/Xevo TQ-XS IVD System (Waters Corporation) equipped with a ZORBAX SB-C18 column (4.6 × 50 mm; Agilent Technologies, Inc.), which was operated following a procedure described previously. 24 The column temperature was set at 25°C; the mobile phase consisted of 0.1% formic acid in aqueous solution (solvent A) and acetonitrile (solvent B). The gradient elution was set as follows: 0 to 33 min (10%-30% solvent B), 33 to 51 min (30%-60% solvent B), 51 to 60 min (60%-90% solvent B), 60 to 63 min (90% solvent B), 63 to 64 min (90%-10% solvent B), and 64 to 67 min (10% solvent B). The injection volume was 2 μL with the flow rate set at 0.2 mL/min. Parameters set for MS with the ion source of electrospray ionization were as follows: (i) Both positive and negative ion modes were used for MS operation; (ii) ion source temperature, 120°C; (iii) scanning range m/z, 50 to 1200; (iv) capillary voltage, 3.0 kV (positive ion mode) and 2.5 kV (negative ion mode); and (v) cone voltage, 40 kV. The desolvation temperature was set at 350°C with the cone gas flow rate set at 400 L/h. The low- and high-energy function values of MSE were set correspondingly at 6 and 20 to 35 eV.
Cell Culture
The normal human embryonic lung fibroblast cell line HEL 299 was obtained from the Bioresources Collection and Research Center (cat. no. 60117), Food Industry Research and Development Institute, and maintained in culture in DMEM with 2 mM L-glutamine, 10% FBS, 100 U/mL penicillin, and 100 μg/mL streptomycin. 25 The cells were placed in 75-cm2 (75T) tissue culture flasks and cultured at 37°C under a humidified atmosphere with 5% CO2.
Cell Viability and Morphological Analysis
HEL 299 cells at 2.5 × 105 cells/mL/well were individually maintained in 24-well plates and then treated with tumor necrosis factor-α (TNF-α; 10 ng/mL), interleukin (IL)-1β (10 ng/mL), IL-6 (10 ng/mL), and 100 μg/mL HLSQW for 24 h. The cell morphology changes were examined using a phase-contrast light microscope (Leica Microsystems GmbH; magnification, 400 × ). Cell viability was assessed by performing an MTT assay, as previously described. 26
Reactive Oxygen species (ROS) Production Assay by Image Cytometry Analysis
HEL 299 cells at 2.5 × 105 cells/mL/well density were individually maintained in 24-well plates and then treated with TNF-α (10 ng/mL), IL-1β (10 ng/mL), IL-6 (10 ng/mL), and 100 μg/mL HLSQW for 12 h. The cells were harvested and re-suspended in 500 μL 10 μΜ CM-H2DCFDA and Hoechst 33342 (10 μg/mL) at 37°C for 15 min for ROS determination. All cells were incubated at 37°C for 30 min and analyzed using a NucleoCounter® NC-3000™ image cytometer (ChemoMetec A/S) according to the manufacturer's protocol and the NucleoView NC-3000 software (version 1.4; ChemoMetec A/S), as previously described. 27
NF-κB p65 Activity Assay
HEL 299 cells at 2.5 × 107 cells/mL/well density were individually maintained in a 75-T flask and then treated with TNF-α (10 ng/mL), IL-1β (10 ng/mL), IL-6 (10 ng/mL), and 100 μg/mL HLSQW for 12 h. The cells were harvested and nuclear proteins from the cells were extracted using the Nuclear Extraction kit from Abcam (cat. no. ab113474), according to the manufacturer's protocols. Total nuclear protein concentration was also measured using the Pierce™ Bicinchoninic Acid Protein Assay Kit (Thermo Fisher Scientific, Inc.). The obtained supernatants were used to quantify the level of NF-κB p65 activation using the NF-κB p65 Transcription Factor Assay Kit (Colorimetric) from Abcam (cat. no. ab210613), according to the manufacturer's protocols. The level of NF-κB p65 activation was derived by measuring the absorbance at 405 nm using an enzyme-linked immunosorbent assay reader. 28
Statistical Analysis
The results are presented as the mean ± standard deviation for the indicated numbers of independent experiments. Statistical comparison of data was performed using one-way ANOVA followed by Tukey's test. ***P < .001 was considered to indicate statistically significant differences between the control and TNF-α/IL-1β/IL-6 treated groups, while ###P < .001 was considered to indicate statistically significant differences between the TNF-α/IL-1β/IL-6 treated and HLSQW-treated groups.
Results
In Silico Target Analysis on the Major Compounds of HLSQW With 3CLpro, PLpro, and RdRp
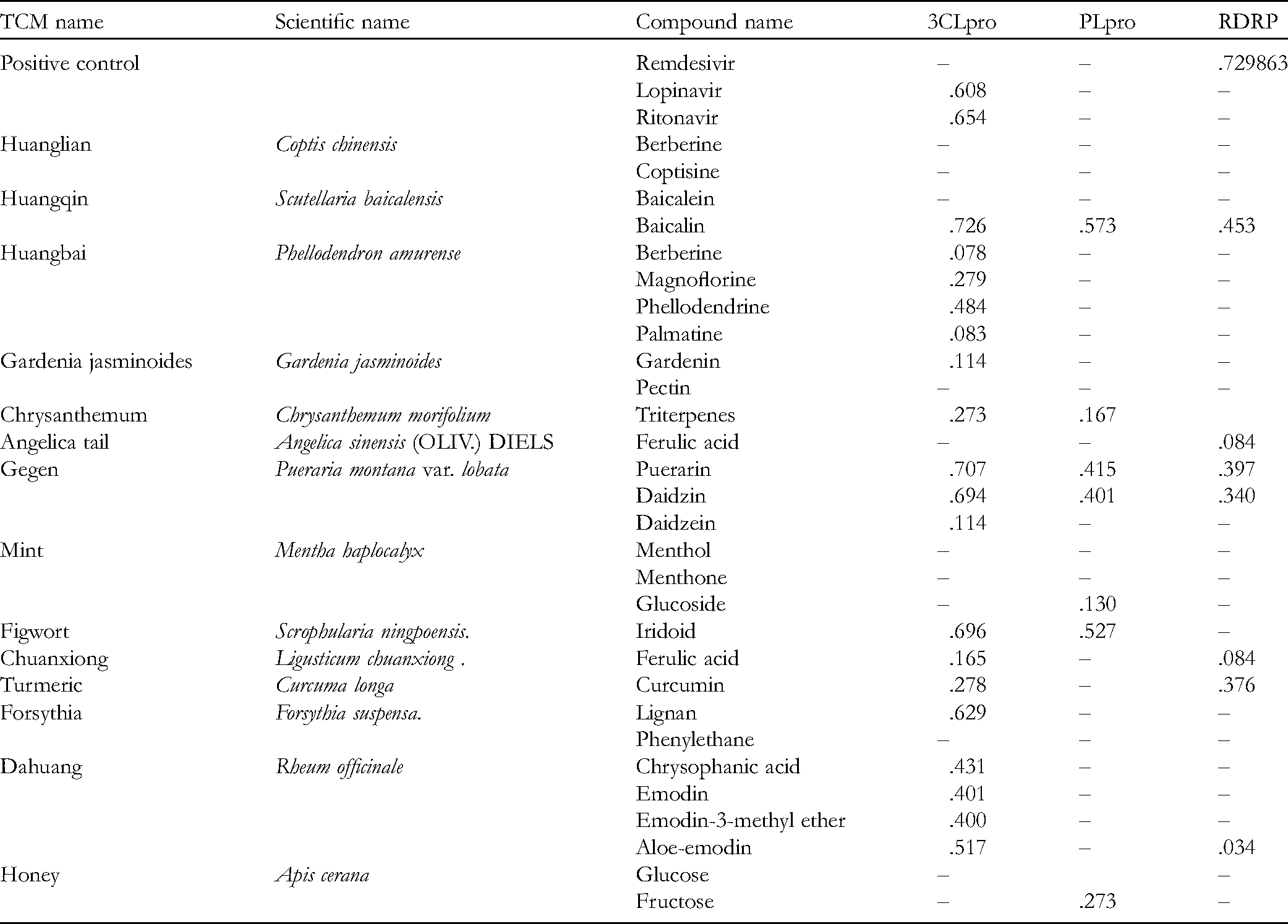
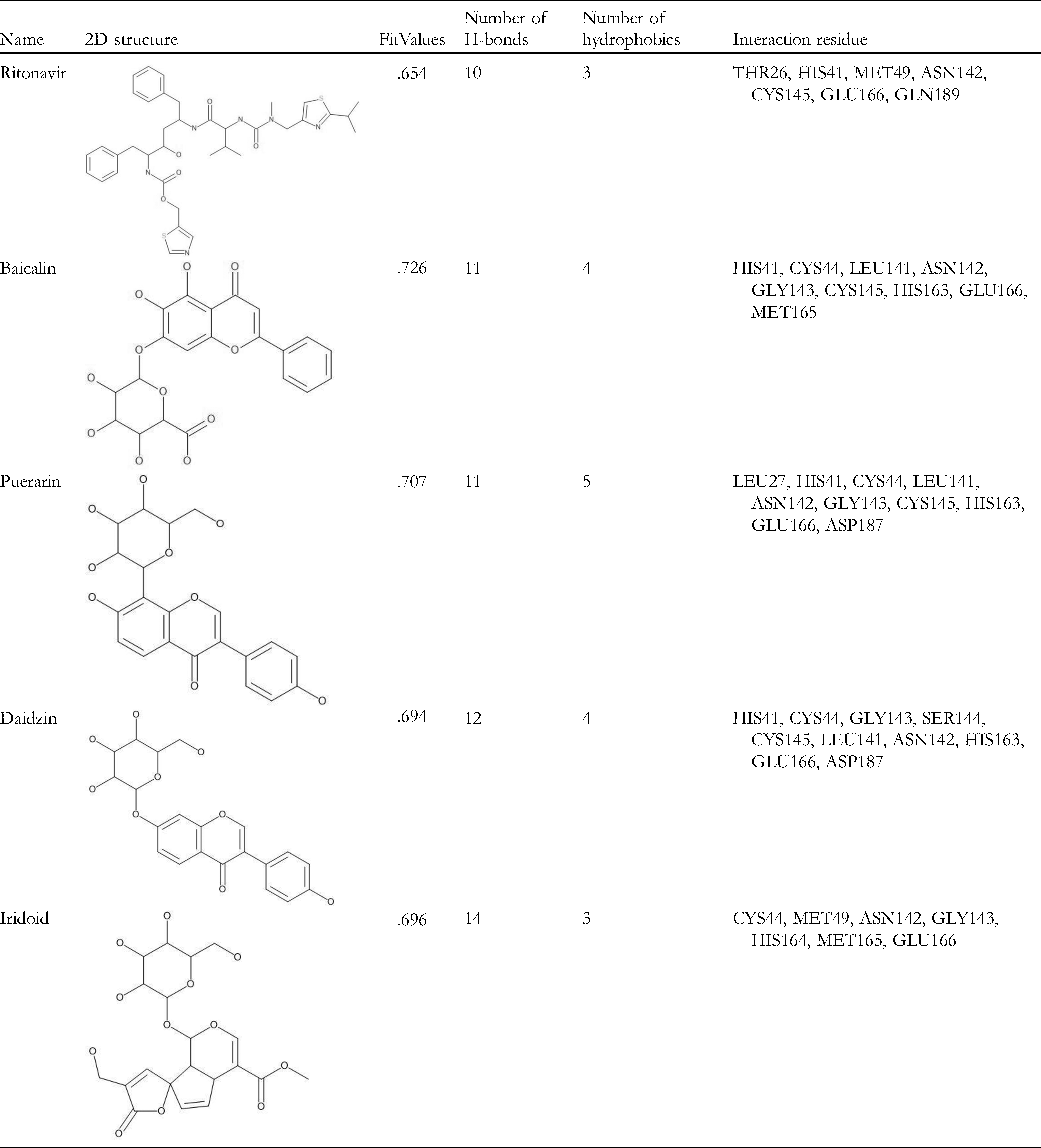
Since the HLSQW formula and its herbal compositions were standardized in TCM, we collected the major bioactive compounds contained within HLSQW from the relevant monographs of the Chinese Pharmacopoeia (2015 edition). Twenty-nine major bioactive compounds, which are chemical markers and most high-content herbal ingredients, were selected. At first, the binding affinities of the 29 components of HLSQW, namely berberine, coptisine, baicalein, baicalin, berberine, magnoflorine, phellodendrine, palmatine, gardenin, pectin, triterpenes, ferulic acid, puerarin, daidzin, daidzein, menthol, menthone, glucoside, iridoid, ferulic acid, curcumin, lignan, phenylethane, chrysophanic acid, emodin, emodin-3-methyl ether, aloe-emodin, glucose, and fructose, to 3CLpro, PLpro, and RdRp proteases of SARS-CoV-2, were calculated by using the pharmacophore models in PharmaDB. The results are shown in Table 1. Baicalin (CDOCKER score: .726), puerarin (CDOCKER score: .707), daidzin (CDOCKER score: .694), and iridoid (CDOCKER score: .696) exhibited high binding affinities onto 3CLpro, compared with those of the positive control ritonavir (CDOCKER score: .654). Baicalin (CDOCKER score: .573), iridoid (CDOCKER score: .527) puerarin (CDOCKER score: .415), and daidzin (CDOCKER score: .401) also showed high binding affinity to PLpro. However, no compound showed sufficient binding affinity to RdRp, unlike the positive control remdesivir (CDOCKER score: .729). The three- and two-dimensional ligand interaction analyses of ritonavir (Figure 2), baicalin (Figure 3), puerarin (Figure 4), daidzin (Figure 5), and iridoid (Figure 6) with 3CLpro are shown. Baicalin interacts with His41, Cys44, Leu141, Asn142, Gly143, Cys145, His163, Glu166, and Met165 of 3CLpro (Figure 3). As shown in Figure 4, puerarin can interact with residues Leu27, His41, Cys44, Leu141, Asn142, Gly143, Cys145, His163, Glu166, and Asp187 of 3CLpro. In Figure 5, daidzin interacts with His41, Cys44, Gly143, Ser144, Cys145, Leu141, Asn142, His163, Glu166, and Asp187 of 3CLpro. Puerarin interacts with residues Cys44, Met49, Asn142, Gly143, His164, Met165, and Glu166 of 3CLpro in Figure 6. As shown in Table 2, ritonavir, baicalin, puerarin, daidzin, and iridoid can form either hydrogen bonds or hydrophobic interactions with residues Asn142 and Glu166 (Figures 2 to 6). Baicalin exhibited superior binding affinity with 3CLpro according to CDOCKER (Figure 3), suggesting that baicalin has a more stable docked structure than ritonavir (Figure 2) and lopinavir (Table 1). To verify the stabilities of the ligand-protein complexes, MD simulation was performed for each ligand-protein complex, as obtained from molecular docking. As shown in Figure 7, puerarin and ritonavir, with RMSD values <0.5 nm, are the 2 most stable ligands in the binding pocket. The RMSD values of baicalin and daidzin were mostly maintained at <1 nm, but varied greatly. Finally, iridoid-3CLpro was demonstrated to be an unstable complex, since the ligand diffused out of the binding site 10 ns after simulation. To conclude, among the 4 tested compounds in simulation courses, the potential ligand with the strongest interaction with 3CLpro is puerarin, with the stability of its ligand-protein complex comparable to that of ritonavir.

Structure-based docking model of ritonavir selectively targeting the binding sites of 3-chymotrypsin-like cysteine protease (3CLpro) in SARS-CoV-2. (A) Three-dimensional and (B) two-dimensional ligand interaction diagrams.

Structure-based docking model of baicalin selectively targeting the binding sites of 3CLpro in SARS-CoV-2. (A) Three-dimensional and (B) two-dimensional ligand interaction diagram of baicalin with 3CLpro.

Structure-based docking model of puerarin selectively targeting the binding sites of 3CLpro in SARS-CoV-2. (A) Three-dimensional and (B) two-dimensional ligand interaction diagram of puerarin with 3CLpro.

Structure-based docking model of daidzin selectively targeting the binding sites of 3CLpro in SARS-CoV-2. (A) Three-dimensional and (B) two-dimensional ligand interaction diagram of daidzin with 3CLpro.

Structure-based docking model of iridoid selectively targeting the binding sites of 3CLpro in SARS-CoV-2. (A) Three-dimensional and (B) two-dimensional ligand interaction diagram of iridoid with 3CLpro.
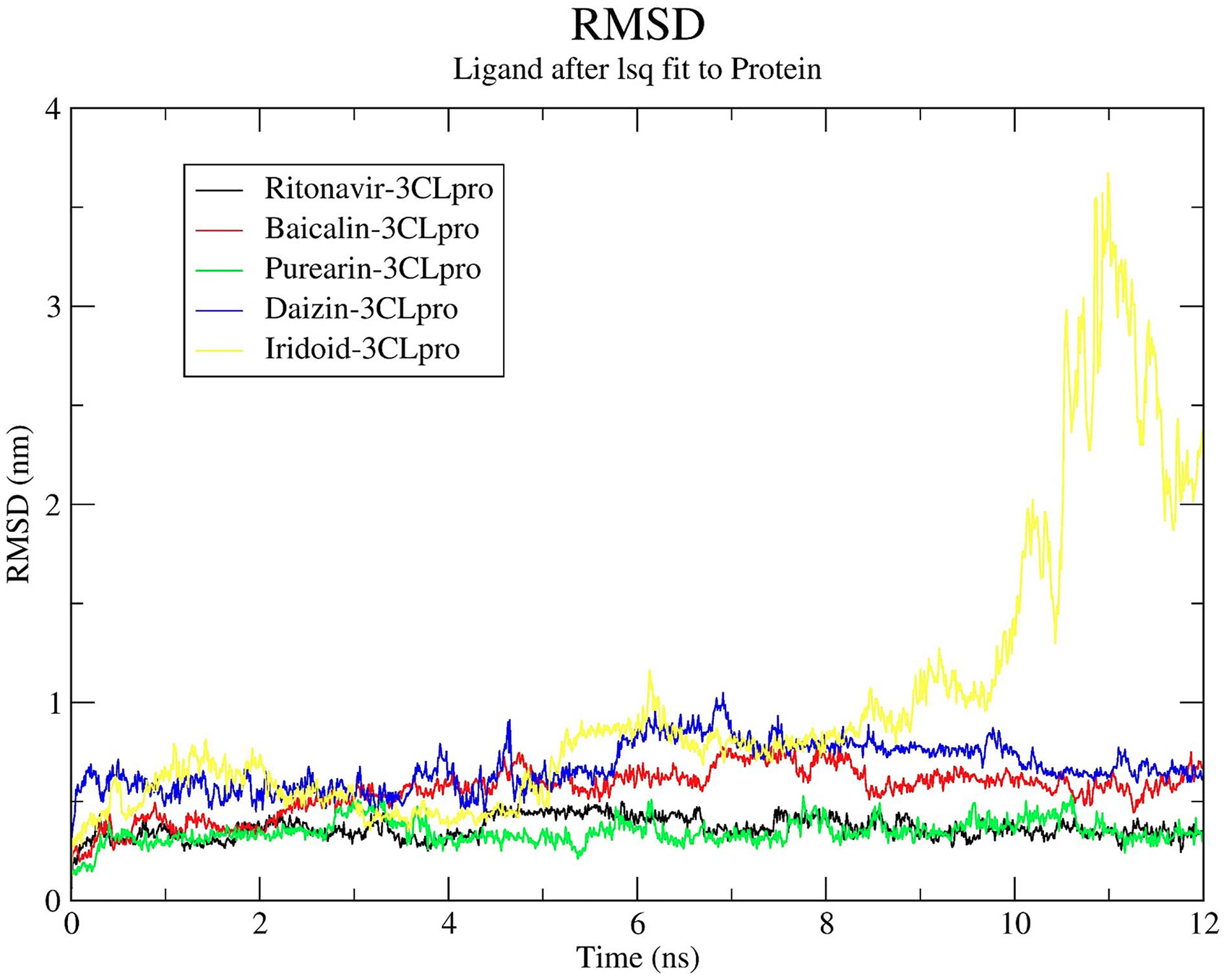
Root-mean-square deviation fitting of the major bioactive Huang-Lian-Shang-Qing-Wan compounds with the 3CLpro protein structure. PDB ID, 6W63. 3CLpro, 3-chymotrypsin-like cysteine protease; PDB, protein data bank.
The Pharmacophore Fitting Results for the Components of HLSQW With 3CLpro, PLpro, and RDRP of SARS-CoV-2.
The Interaction Statistics Between the major Components of HLSQW and 3CLpro.
Network Analysis on the Effects of HLSQW on Anti-Virus and Anti-Inflammatory Cytokines Using IPA
Since TCM is typically comprised of a multitude of components with various targets, we hypothesized that HLSQW has other biological targets and can regulate different signaling pathways. Therefore, the potential therapeutic anti-virus signaling network of HLSQW was analyzed using IPA (Figure 8). The potential anti-viral targets associated with each compound were found based on the HLSQW information contained in the TCM. We next performed viral infection pathway analysis on the target information. In Figure 9, HLSQW exerted therapeutic effects by interfering with viral intrusion into cellular processes. Our results suggested that the main effect of HLSQW was to interfere with pathways related to viral infection. We subsequently collected the targets corresponding to the molecular pathways involved in the cellular function of HLSQW in Figure 10 and Supplemental Table S1. The top 10 molecular pathways associated with HLSQW include “cell death and survival,” “cellular movement,” “organismal injury,” “lipid metabolism,” “cellular movement,” “gene expression,” “hematological system development,” “cell cycle,” “cell signaling and interaction,” and “cellular development.” A network of the associations of HLSQW with anti-inflammatory cytokines was also generated following IPA analysis. Figure 11 demonstrates the cytokines which are most regulated by HLSQW-targeted genes. Details of targeted genes are as shown in Table 3. Cytokines corresponding to these genes include TNF, IFN-γ, IL-4, IL-6, IL-1β, IL-2, IL-15, CSF-2, IL-13, and IL-1α. Finally, we focused on the effects of the target molecules on the IL-6 pathway. Our results revealed that all compounds of HLSQW could trigger IL-6 production through the phosphoinositide 3-kinase (PI3K)/protein kinase B (AKT), Janus kinase (JAK)2/signal transducer, and activator of transcription 3 (STAT3), Ras/Raf/mitogen-activated protein kinase (MAPK), and NF-κB signaling pathways, according to Figure 12.

Viral infection network constructed using ingenuity pathway analysis core analysis. Focus molecules, represented by grey color, are potential targets of Huang-Lian-Shang-Qing-Wan.

Virus entry pathways. The molecules, represented in grey color, are potential targets of Huang-Lian-Shang-Qing-Wan. Inhibition of c-ABL and phosphoinositide 3-kinases may block virus entry by actin cytoskeleton reorganization and polymerization. Inhibition of the SRC family may block viral penetration by membrane refilling.

Statistical results of the top 10 related ontology terms associated with the potential targets of Huang-Lian-Shang-Qing-Wan. These targets can be related to the downstream cell functions and phenotypes. The statistical results indicated that the top 3 functions are “cell death and survival,” “cellular movement,” and “organismal injury.”

Statistical results of the top 10 cytokines using upstream analysis by ingenuity pathway analysis for the potential targets of Huang-Lian-Shang-qing-Wan. The most counted genes associated with the potential regulatory cytokines are tumor necrosis factor, interferon-γ, and interleukin-4.

Canonical IL-6 signaling pathway. IL-1, tumor necrosis factor-α, and IL-6 induce signaling through the JNK, MAPK, signal transducer, and activator of transcription 3 and PI3K pathways. It causes the cell to generate factors during the acute-phase responses and lymphocyte stimulation. Several components of Huang-Lian-Shang-Qing-Wan were predicted to target JNK, p38 MAPK, mitogen-activated protein kinase, extracellular signal-related kinases 1/2, Janus kinase 2, and PI3K. JNK, c-Jun N-terminal kinases; MAPK, mitogen-activated protein kinase; PI3K, Phosphoinositide 3-kinases; IL, interleukin.
The top ten Inflammatory Cytokines Which Regulated by HLSQW-Targeted Genes.

HLSQW Extract Exerts Protective Effects Against Inflammatory Cytokines and Inflammation-Associated Factors in HEL 299 Cells
The obtained results suggested a potential suppressive effect of HLSQW on cytokine-regulated pathways and on cellular factors that activate inflammation in the host organism. To verify this, we prepared an extract of HLSQW and analyzed it by liquid chromatography. The results showed high concentrations of baicalin, berberine, baicalein, rhein, liquiritin, aloe-emodin, isoimperatorin, chrysophanol, and physcion (Figure 13). Further analysis by UPLC-Q-TOF/MS in both positive and negative ion modes also detected the aforementioned compounds, as presented in Figure 14, with retention times corresponding to those of reference standards.

High-performance liquid chromatographic analysis. (A) Detection of major bioactive compounds in Huang-Lian-Shang-Qing-Wan extract; (B) standard compounds.

Ultra-pure liquid chromatography-Q-time-of-flight/mass spectrometry analysis results for the detection of major bioactive compounds in Huang-Lian-Shang-Qing-Wan extract.
Guided by the in silico results, we investigated the protective effects of the HLSQW extract on cytokine-induced cell injury in normal human lung cells in vitro. In vitro experiments were performed, using TNF-α/IL-1β/IL-6 with and without treatment with HLSQW extract (Figure 15A). Treatment with the cytokines decreased the cell viability to <60% of the control group, induced changes in cellular morphology, and elevated NF-κB activity and ROS production. By contrast, HLSQW treatment preserved the cellular morphology while protecting cell viability, which was ∼80% of the control group, in the presence of the 3 inflammatory cytokines (Figure 15B and C). Furthermore, the effects of the individual components, including puerarin, daidzin, and baicalin, were also examined alongside the HLSQW extract at different concentrations. The obtained results (Figure 16) demonstrated that the HLSQW extract significantly preserved cell viability following cytokine treatment in HEL 299 cells in a concentration-dependent manner. Our results also revealed that puerarin and baicalin can significantly attenuate the cytotoxicity of cytokines in HEL 299 cells at a concentration of 100 µM, compared with that in the group treated with the cytokines alone. HLSQW treatment was also found to significantly reduce ROS production (Figure 17A) and NF-κB activity (Figure 17B) in HEL 299 cells. To conclude, these results suggest that HLSQW exerts cytoprotective effects against cytokine-induced injury on the HEL 299 cell line.

In vitro experimental results of the cytoprotective effects of HLSQW extract against cytokine-induced cell injury and cytotoxicity. (A) Effects of treatment with cytokine tumor necrosis factor-α/IL-1β/IL-6, and HLSQW extract on HEL 299 cells. (B) Cell morphology changes and (C) cell viability of HEL 299 cells treated with cytokines, with or without HLSQW extract. HLSQW, Huang-Lian-Shang-Qing-Wan; IL, interleukin.
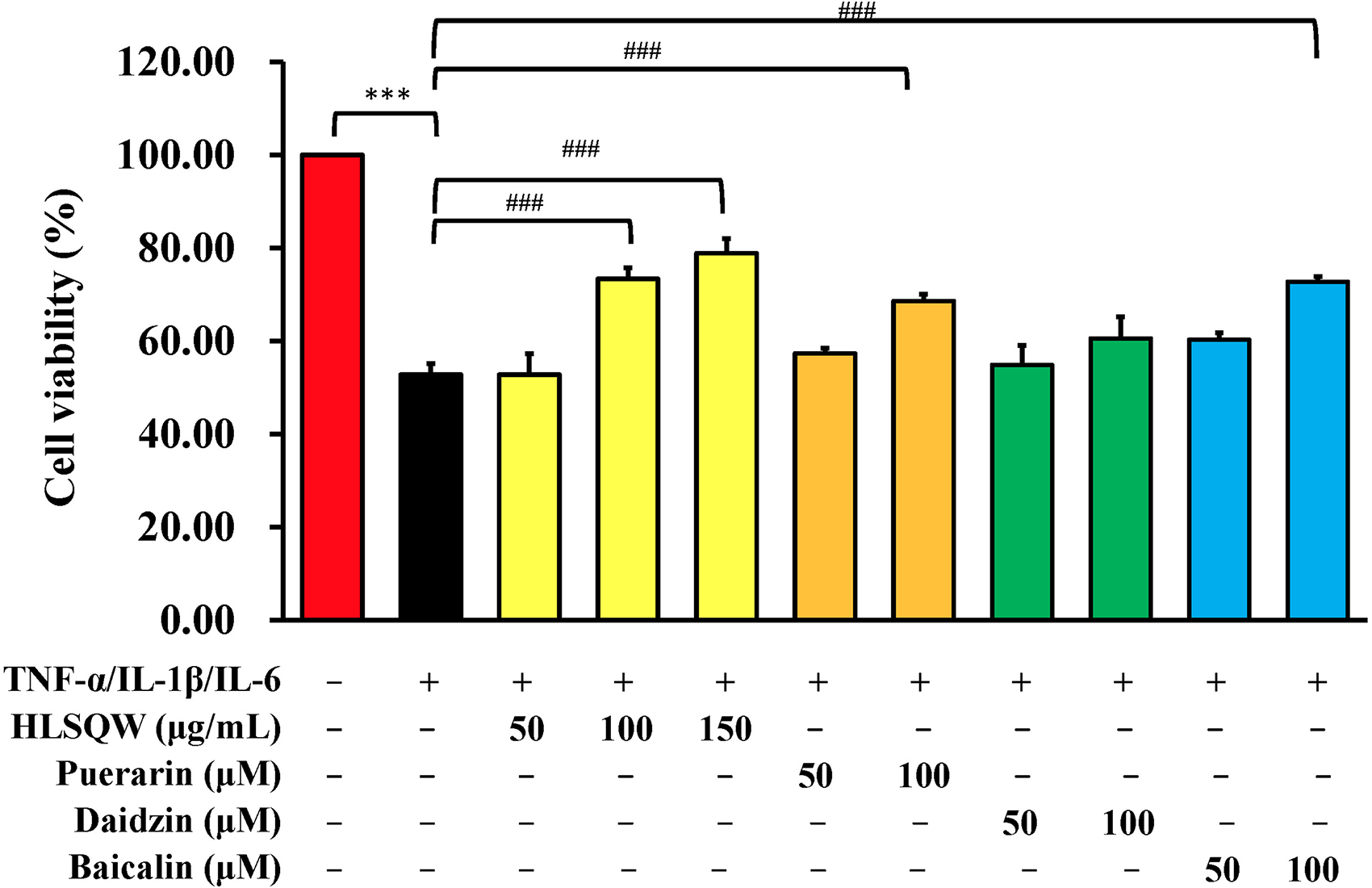
In vitro experimental results of the cytoprotective effects mediated by different concentrations of Huang-Lian-Shang-Qing-Wan extract, puerarin, daidzin, and baicalin against cytokine-induced cytotoxicity. Data were analyzed using one-way ANOVA followed by Tukey's post hoc test. ***P < .001 and ###P < .001.

Effects of HLSQW extract on cellular inflammation-associated factors in HEL 299 cells following treatment with cytokines. (A and B) Nuclear factor-κB p65 activity and (C) reactive oxygen species production in HEL 299 cells after treatment with cytokines, with or without HLSQW extract. Data were analyzed using one-way ANOVA followed by Tukey's post hoc test. ***P < .001 and ###P < .001. HLSQW, Huang-Lian-Shang-Qing-Wan.
Discussion
Understanding the replication cycle of the severe acute respiratory syndrome (SARS)-CoV-2 is important for identifying the molecular targets for the development of anti-viral treatment agents for COVID-19. The viral replication cycle consists of fusion, translation, proteolysis, RNA replication, and packaging before virion release.29,30 Initial studies mainly focused on molecular agents that can specifically target viral proteases31,32 and RdRp,33,34 because their relevant protein structures were among the first to be characterized.35,36
Bioactive compounds contained within HLSQW were screened in the present study on 3 major targets of SARS-CoV-2, specifically 3CLpro, PLpro, and RdRp, with particularly promising results found with 3CLpro. In addition, baicalin, puerarin, daidzin, and iridoid all exhibited high binding affinities to 3CLpro compared with those of remdesivir, the positive control (Table 1). Mechanistically, these compounds can form various types of intermolecular interactions with 3CLpro, with both the backbone and specific residues, including hydrogen bonds, π-π stacks, and hydrophobic interactions (Figures 2 to 6). According to previous studies, interactions with target proteins may hinder the flexibility of amino acids at the catalytic site.35,37 As presented in Table 2, baicalin, puerarin, and daidzin formed direct interactions with the residues Cys145 and His41, which were previously proposed to be responsible for the catalytic activity of 3CLpro. 37 At the binding site, the predicted affinity and intermolecular interactions of baicalin, puerarin, daidzin, and iridoid suggested that all 3 of these components can form inhibitory ligand-protein complexes with 3CLpro. Furthermore, MD results revealed that the puerarin-3CLpro complex exhibited superior stability in the simulation system compared with other complexes (Figure 7), suggesting that puerarin may act as an inhibitor of 3CLpro by stably binding to the active site. In addition, acceptable results were also observed for baicalin and daidzin, meaning that they can also be considered as potential compounds in the drug discovery process of targeting SARS-CoV-2 3CLpro. 38 Recently, several small molecules have been developed and/or repurposed for COVID-19 treatment through inhibiting 3CLpro to suppress viral replication. 39 Some of these compounds have yielded promising results both in vitro and in vivo.40-43 Notably, PF-00835231 from Pfizer has now entered the first phase of clinical trials. 40 All these compounds were assumed to interact directly with 3CLpro at the same binding pocket as observed in our docking and simulation model. Collectively, the results obtained in the present study and those from the recent studies suggest that HLSQW may interrupt the viral replication cycle of SARS-CoV-2 during the proteolysis process by specifically inhibiting 3CLpro. However, the nature of the interactions between the individual HLSQW bioactive components and 3CLpro during the inhibition of its activity should be systematically verified experimentally, such as by fluorescence resonance energy transfer assay.
The nature of the alterations in the host cell after infection with SARS-CoV-2 remains poorly understood. In general, the pathogen manipulates the machinery of the host by interfering with intracellular cell signaling pathways to facilitate pathogen replication. 44 As presented in our IPA network analysis, the proteins and cytokines appear to be triggered immediately after viral entry (Figures 8 and 9), which consequently involve a number of important pathways with a complex cross-talk network (Figure 8). These pathways are mainly responsible for cellular metabolism, behavior, and death/survival. Of note, cell death/survival-related pathways were found to be the main item among the mechanisms of action found for HLSQW (Figure 10). During the progression of acute pneumonia, the death/survival decision of the host cell is a crucial defense mechanism for preventing viral replication by eliminating infected cells. 45 Conversely, when a large number of cells die in the lung in a short space of time it can result in impaired lung function. 46 In addition, the immune system, inflammatory response, and other intercellular cell death/survival signals are crucial during the course of pneumonia. 8 Some bioactive compounds of HLSQW have been previously reported to exert anti-viral effects,17,18 particularly on coronaviruses. 19 The present study revealed that the mechanism of action underlying the effects of HLSQW mainly depended on the regulation of death-survival signaling pathways in the host cell. However, further investigation on the mechanism underlying the HLSQW-induced effects on these signaling pathways is required.
An association between cytokine-release syndrome (CRS) and poor outcome of COVID-19 has been previously identified.8,47 To control the “cytokine storm,” treatment options based on modern medicine and TCM have both been considered.8,43 Our analysis results demonstrated that HLSQW potentially affects a number of cytokine genes, including TNF, IFN-γ, interleukins (IL-1 and IL-6), and CSF2 (Figure 10 and Table 3), all of which were previously detected in patients with COVID-19. 48 In Figures 11 and 12, our IPA analysis results on the downstream signaling pathways of the CD14IL-1, tumor necrosis factor (TNFR), and glycoprotein 130 IL-6 receptors (GP130IL-6R) showed a complex relationship between the bioactive compounds of HLSQW and these pathways. The interactions were predicted to occur in both the cytoplasm and the nucleus. The important nodes in the network, as shown in Figure 11, were found to be the PI3K/AKT, JAK2/STAT3, Ras/Raf/MAPK, and NF-κB signaling pathways. Correspondingly, many important bioactive compounds of HLSQW also participate in this cross-talk to possibly regulate the cytokine signaling pathways. The anti-inflammatory effect of HLSQW may be mediated by synergistic effects among several bioactive constituents contained within this formula. Curcumin 49 and daidzin 50 have been reported to confer inhibitory effects on cytokine synthesis and secretion mediated by berberine 51 and puerarin.52-54 through the activation of p38 MAPK and JNK. In addition, previous studies demonstrated that baicalin and baicalein can attenuate cytokine-induced and chemokine-induced inflammation.55-58 The consistency of this finding with the present study suggests that HLSQW may be applied as a potential treatment option for ameliorating cytokine-release syndrome (CRS) to provide beneficial effects for patients with COVID-19.
The combination of multiple components with a variety of targets is a characteristic feature of TCM, such that the therapeutic outcome is likely due to synergistic or additive effects mediated by a multitude of components. This increases the difficulty for investigating the pharmacological mechanisms of the formulas used in TCM. In the present study, we screened the effects of the bioactive compounds in HLSQW on the major targets of SARS-CoV-2 and then conducted a compound-target relationship and network pharmacology analysis by IPA (Figure 1). The results suggested that several pharmacological effects of HLSQW were associated with the potential targets of SARS-CoV-2. According to the theoretical system of TCM, the herbal compositions of a remedy can be categorized into 4 groups: Monarch; minister; assistant; and envoy. 59 The combination of “monarch and minister” is the core power of the remedy and confers the main efficacy. Following this traditional analogy, in HLSQW, the “monarch” consists of Huanglian, Huangqin, Huangbai, Gardenia jasminoides, and Da Huang. The “minister” compositions are Forsythia, Chrysanthemum, Angelica tail, Mint, and Chuanxiong. Interestingly, our analysis using IPA and in silico assay also revealed that the main bioactive compounds contained within the “monarch” and “minister” groups are mainly responsible for the therapeutic effects against COVID-19. By targeting the pathways that result in the cytokine storm and 3CLpro of SARS-CoV-2, these compounds contained within HLSQW not only serve important roles, but can also provide potential starting points for developing anti-SARS-CoV-2 agents.
An excessive inflammatory response leading to severe illness in patients with COVID-19 was identified to be a consequence of cytokine release. 41 NF-κB activation and ROS production are known to be important mediators for activating inflammation.60,61 Our in vitro results not only support our in silico data, but also suggest a protective role of the formula against pneumonia. In the present study, we investigated the potential protective effects of HLSQW on cytokine-induced cell injury on the normal human lung HEL 299 cell line in vitro. The protective effects against cytokine-induced injury (Figure 15B), cytotoxicity (Figures 15C and 16), ROS production (Figure 17A), and NF-κB activity (Figure 17B) in the cells suggested that HLSQW exerts its effects on multiple targets. The cytokine-induced ROS production and consequent mediation of NF-κB p65 activity have been previously reported as an inflammatory response at the cell level.62,63 HEL299 cells were treated with cytokines and HLSQW extract for 12 h, therefore long enough for examining the responsive reactions of the treated cells, not only for the cytokine-induced effects, but also the biological activity induced by HLSQW extract. Previous studies demonstrated that blocking cytokine-mediated inflammation and cell injury may benefit patients with COVID-19 with cytokine storm syndromes.8,47 Therefore, HLSQW may attenuate CRS in patients with COVID-19 and possibly improve prognosis. However, our study has a number of limitations. All in silico approaches require further experimental verification. In addition, although most of the main bioactive components of HLSQW were selected for the present study, the list of compounds investigated was not representative of all the chemical components in the mixture. Due to insufficient data on the metabolic profile of the investigated compounds, this matter was beyond the scope of this study.
In conclusion, our results revealed the potential of HLSQW as a therapeutic agent based on TCM that may confer beneficial effects on patients with COVID-19 by targeting multiple mechanisms. In addition, the main bioactive compounds of HLSQW are worthy candidates for further anti-viral drug discovery studies, particularly for SARS-CoV-2. Indeed, the present study was based on bioinformatics and used an in silico approach, which requires confirmation by further experimental analysis.
Supplemental Material
sj-docx-2-npx-10.1177_1934578X211030818 - Supplemental material for In Silico Target Analysis of Treatment for COVID-19 Using Huang-Lian-Shang-Qing-Wan, a Traditional Chinese Medicine Formula
Supplemental material, sj-docx-2-npx-10.1177_1934578X211030818 for In Silico Target Analysis of Treatment for COVID-19 Using Huang-Lian-Shang-Qing-Wan, a Traditional Chinese Medicine Formula by Ching-Wen Huang, Hai-Anh Ha, Shih-Chang Tsai, Chi-Cheng Lu, Chao-Ying Lee, Yuh-Feng Tsai, Fuu-Jen Tsai, Yu-Jen Chiu, Guo-Kai Wang, Chung-Hua Hsu and Jai-Sing Yang in Natural Product Communications
Footnotes
Acknowledgments
The authors thank Kuan-Wen Chen and Tzu-Mao Hung in GGA Corporation, Molecular Science and digital innovation center (Taiwan) for performing the molecular docking to 3CLpro, PLpro, and RdRp.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the China Medical University Hospital (grant no. DMR‑110‑155), and the Shin-Kong Wu Ho-Su Memorial Hospital (grant no. 2021SKHADR030).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
