Abstract
Two new natural products, mitrafungidione (1) elucidated as prototrop-isomers of (R-3-acetyl-5-ethyl-4-hydroxy-5H-furan-2-one, and maristachone F (2a), elucidated as 5-(1-hydroxyethyl)-4-(hydroxymethyl)-3-methoxy-2-methylphenol, together with 5 known compounds have been isolated from the solid cultures of an endophytic fungus associated with Mitragyna inermis (Rubiaceae) and identified as Cercophora samala. The structures of these compounds were elucidated by detailed spectroscopic analysis and by comparison of their spectroscopic data with those reported in the literature. The absolute configuration of 1 and 2a were determined by extensive DFT calculations.
Mitragyna inermis (Willd.) O. Kuntze (Rubiaceae) is a small tree growing on low alluvial plains and swampy savanna in West Africa 1,2 In traditional medicine, this plant is used to treat diseases such as diabetes, fever, diarrhea, dysentery, cholera, malaria, rheumatic infections, and osteoarthritis diseases. 3 -5 We have previously reported the isolation of a number of bioactive compounds from M. inermis. 6 Studies of ethyl acetate extracts from the mycelia of the culture media of the endophytic fungus Cercophora samala Udagawa & T. Muroi (Lasiosphaeriaceae) associated with M. inermis afforded 2 new natural products, mitrafungidione (1*) and maristachone F (2a), together with 5 known compounds. The structures of the new compounds were elucidated by detailed spectroscopic analysis, by comparison of their spectroscopic data with those reported in AntiBase, 7 and by DFT calculations. We report herein the isolation and structural elucidation of 1 and 2a.
For the taxonomic identification, the endophytic fungus associated with M. inermis (Rubiaceae) was grown on potato-dextrose-agar (PDA) at 25 °C under near-ultraviolet light with 12 hours photoperiods. Genomic DNA was extracted from 6-day-old colonies, using Quiagen Plant Mini Kit (QUIAGEN) following the manufacturer’s protocol. The fungal isolate ED-I was identified as C. samala, based on the ITS sequence, as described in the experimental part.
After fermentation on solid rice medium at room temperature (28 °C) for 30 days under static conditions, the culture was extracted with ethyl acetate, and the resulting material was subjected to column chromatography using a CH2Cl2/MeOH solvent system with gradient elution. Further purification of subfractions on Sephadex LH-20 afforded 2 new natural products, mitrafungidione (1) and maristachone F (2a), together with 5 known compounds namely: versiconol, 8 cerebroside B, 9 and the trivial metabolites ergosterol, thymine, and uracil.
Compound 1 (Figure 1) was obtained as reddish oil. The molecular formula C8H10O4 was deduced from ESI HRMS with 4 double bond equivalents (DBE). The 1H NMR spectrum (Table 1) showed resonances of an A3BCX-type spin system attributed to an oxymethine proton at δ X 4.03 and 2 methylene protons at δ H 1.72 (HB-6a) and 1.45 (HC-6b). The 1H,1H COSY spectrum (Figure 2) combined these fragments with a methyl triplet at δ HA 0.84 (H3-7) to an oxypropyl sequence CH3-CH2-CH-O-. The spectra showed additionally a methyl singlet at δ H 2.16 (H-9) and a low-field 13C signal at 191.0, suggesting a carbonyl group (C-8). HMBC correlations further revealed the moiety as a Cq-acetyl fragment. Additionally the D2O-exchangeable signal of a hydroxyl group at δ H 3.30 was seen.
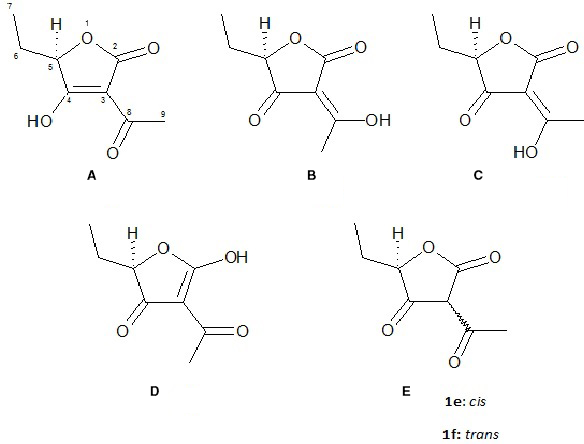
Classical structure of mitrafungidione (1a) and tautomers 1b – 1f.

Selected 1H-13C HMBC and 1H,1H COSY correlations of mitrafungidione tautomer 1a.
1H (300 MHz), 13C (125 MHz), HMBC and COSY Data of Mitrafungidione (1) in DMSO-D6 (δ in ppm).

The 13C NMR, APT, COSY, HMBC and HSQC spectra confirmed these assignments. HMBC correlations indicated additionally the connection of the oxypropyl fragment at C-5 with 2 carbonyl groups, forming a ketone (C-6, δ C 195.0) and, via the oxygen, of an ester carbonyl (C-2, δ C 174.7). The remaining C-acetyl unit must bridge these carbonyl groups, forming a furanedione derivative. The fact that no other methine proton was visible, suggested that one of the carbonyl groups is present in the enol form, resulting in 3-acetyl-5-ethyl-4-hydroxy-5H-furan-2-one (1a), or another prototrop-isomeric form thereof (Figure 1).
Compound 1, trivially named mitrafungidione, is a further natural member of the tetronic acid group and had been obtained previously by synthesis, but was published without NMR data. 10 Although 6 isomers may co-exist in equilibrium, in the literature the hydroxy group of tetronic acids is usually drawn as enol at C-4 (type 1a). However, our density-functional theory (DFT) calculations on the ωB97XV/6‐311 + G(2df,2p) level 11 predicted that this isomer contributes only 7% to the equilibrium, which is dominated by 1b (67 %); for 1c, a concentration of 26% was calculated, while the other isomers are neglectable. Nevertheless, for better comparison with the literature, we continued using structure 1a here. The shape of the experimental ECD spectrum was in agreement with DFT-data calculated for the Boltzmann-weighted (R)-1a-1c mixture (Figure 3). Mitrafungidione is therefore elucidated as the γ-butenolide (R)-1a, existing in methanol mainly as the isomers 1b and 1c (Figure 1 and Supplemental Figure S3).

Experimental (green, MeOH) and DFT-calculated ECD-spectrum (upper blue) of (
Compound 2a (Figure 4) was obtained as yellowish oil. The molecular formula C11H16O4, implying 4 DBEs, was deduced from the ESI HRMS spectrum. The 1H NMR spectrum of 2a in DMSO-d 6 showed 2 broad D2O-exchangeable proton signals at δH 4.53, 4.88, and one at 9.17, suggesting the presence of a phenol. All further spectra were measured in deuteriomethanol. With the singlet of only one aromatic proton at δH 6.80 (δC 109.2) and 5 additional 13C signals of fully substituted aromatic carbons at δC 117.5, 122.5, 145.5, 157.5 and 159.4, a pentasubstituted benzene was expected, which is in agreement with the 4 DBEs. From the 1D and 2D NMR spectra, a 1-oxyethyl sequence [CH3-CH(O-)-], an oxymethylene singlet at δH 4.67, a methoxy signal at δH 3.72, and a methyl singlet at δH 2.10 were derived.

Structure of (
The oxyethyl group was connected with C-5 (δC 145.6), as the HMBC correlation of this atom with Me-11 indicated. In both o-positions of C-5, the ring atoms C4- and CH-6 were assigned by correlations with CH-10. The oxymethylene group -O-CH2-9 correlated with C-5, C-4, and the methoxy-carbon C-3, and was therefore connected with C-4. The methyl singlet Me-7 at δΗ 2.11 showed a cross signal with C-3 and is therefore a direct neighbor of the latter in position 2, so that the remaining phenolic hydroxy group should be placed at C-1 (Figure 5). The other hydroxy groups were attached to C-9 and 10 to agree with the empirical formula, resulting in 5-(1-hydroxyethyl)-4-(hydroxymethyl)−3-methoxy-2-methylphenol (2a) (Figure 4). Surprisingly, this rare phenol type is closely related to maristachone A (2b), a metabolite isolated from a sponge-derived Stachylidium sp.
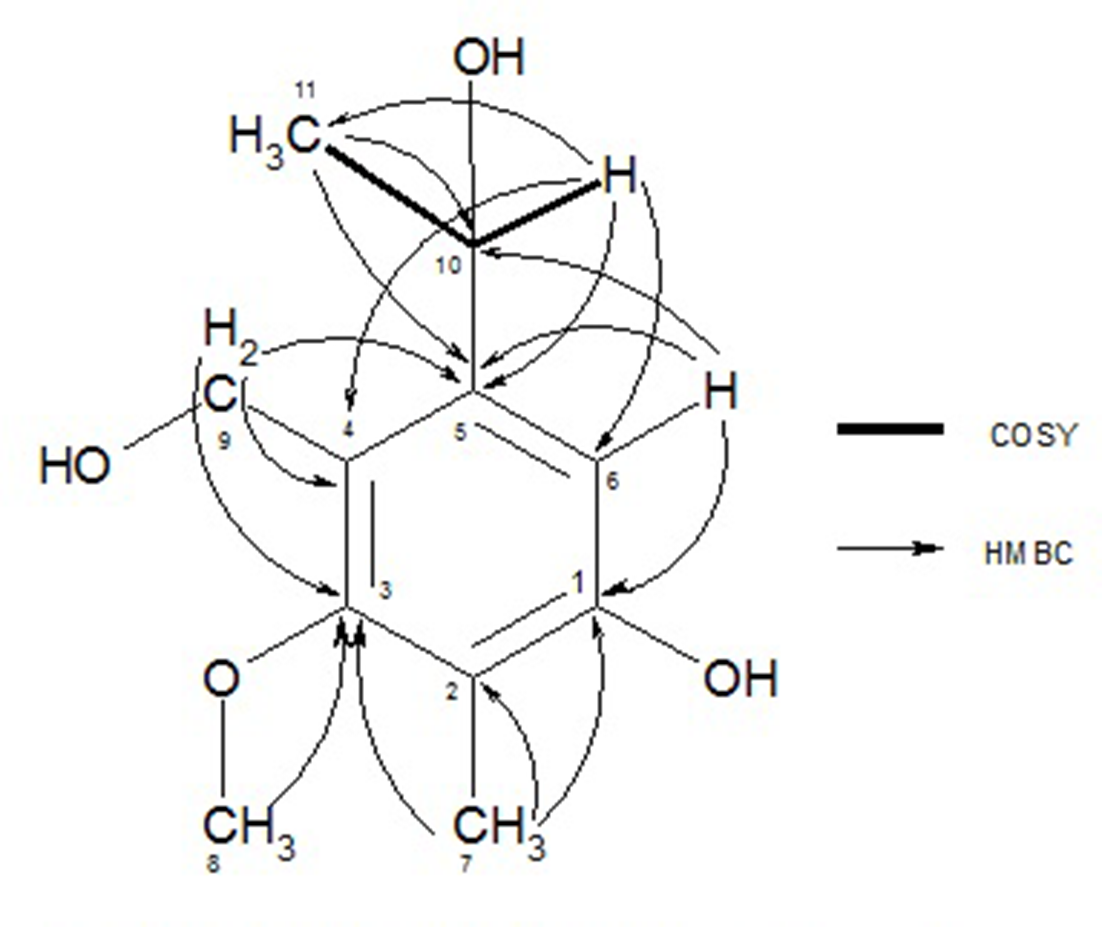
1H-13C HMBC and 1H-1H COSY correlations of 2a.
Experimental
General Experimental Procedures
Optical rotations: Polarimeter (Perkin-Elmer, model 343). ECD spectra were recorded on a JASCO J-810 spectrometer equipped with a JASCO etc.,-505S/PTC-423S temperature controller. The 1H NMR spectra were recorded on Varian Mercury-300 (300.141 MHz) and Varian VNMRS-300 (300.536 MHz) spectrometers equipped with 3 mm probes; 13C NMR spectra were measured at 125.707 MHz relatively to TMS as internal standard; shifts are reported as δ values. Electrospray-ionization mass spectrometry (ESIMS) and high-resolution mass spectra (ESIHRMS) were recorded on a micrOTOF time-of-flight mass spectrometer (Bruker Daltonics, Bremen, Germany), as well as on an Apex IV 7 Tesla Fourier-transform ion cyclotron resonance mass spectrometer (Bruker Daltonics, Billerica, MA). Column chromatography (CC): silica gel (230, 400 mesh; Macherey-Nagel, Düren, Germany). Size exclusion chromatography was done on Sephadex LH-20 (Lipophilic Sephadex, Amersham Biosciences Ltd; purchased from Sigma-Aldrich Chemie, Steinheim, Germany). TLC was performed on pre-coated silica gel sheets of Polygram SIL G/UV254 (Macherey-Nagel and Co., Düren, Germany), with mixtures of dichloromethane and methanol as eluents; spots were detected using UV light (254 and 365 nm) or by spraying with anisaldehyde/sulfuric acid staining reagent prepared by mixing of anisaldehyde (1 ml) and methanol (85 ml) with concentrated sulfuric acid (1 ml) and acetic acid (14 ml).
Plant and Fungal Material
The endophytic fungus C. samala was obtained from the branches of M. inermis collected in December 2011 at Nkoteng in the Centre Region of the Republic of Cameroon. The plant was identified by Mr. Victor Nana, botanist at the National Herbarium (Yaoundé, Cameroon), where a voucher specimen (N° 8886/SRF/Cam.), has been deposited.
The endophytic fungus was isolated from a branch sample using techniques as described previously. 13 A voucher specimen of the fungus is deposited in the culture collection at the Institute of Organic and Biomolecular Chemistry, Göttingen, Germany under the internal code ED-I.
Taxonomy of the Fungus
The ITS rDNA of the fungal strain ED-I was amplified by PCR reaction, with the primer pairs ITS1f 14 and ITS4. 15 PCR products were checked on 1% agarose electrophoresis gel stained with ROTI®GelStain (Roth, Germany). Purified amplicons were sequenced in both directions with the PCR primers by GATC (actually Eurofins Genomics Germany GmbH). Raw nucleotide sequences were edited in MEGA version 7 16 and deposited in NCBI GenBank database (Accession number MW177563).
ITS sequence of isolate ED-I was used for BLAST searches in NCBI GenBank (accessed 12.08.2020). Higher scoring hits were sequences of Cercophora samala (AY999134, NG708364, MH861345) and Podospora pauciseta (MH864359, MH858484, MH858208) belonging to the family Lasiosphaeriaceae. Additional sequences of Cercophora and Podospora were retrieved from the GenBank, including those of ex-type isolates and aligned in MEGA 7. Phylogenetic analysis based on PhyML 3.1 was conducted with the GTR model using the web service Phylogeny.fr (http://www.phylogeny.fr/one_task.cgi?task_type=phyml). In the phylogenetic tree (Supplemental Material, Figure S1), the ITS sequence of ED-I clustered to C. samala 2HD71-5. The topology of this tree reflects the polyphyletic character of both genera Cercophora and Podospora. 17
Fermentation, Extraction, and Isolation of Metabolites
The endophytic fungus strain ED-I was cultured on slants of potato dextrose agar (PDA: cooking water from 200 g potatoes/L, enriched with glucose 40 g/L and agar-agar 20 g/L) at 28 °C for 6 days. Pieces of a well-grown agar subculture were used to inoculate 10 P-flasks, each containing sterilized rice medium (200 g of rice suspended in 200 ml of tap water and sterilized) and incubated at room temperature under static conditions for 30 days. The culture was extracted 4 times with ethyl acetate (EtOAc) and the filtrate was concentrated to dryness in vacuo to afford 63 g of a yellow-brown extract. A portion of 60 g of the crude extract was fractionated by silica gel column chromatography using a CH2Cl2/MeOH gradient of increasing polarity (100:0‐80:20). The resulting 106 subfractions of each 200 ml were combined on the basis of TLC analysis to yield 7 main fractions: F1 (6.6 g), F2 (3.8 g), F3 (2.4 g), F4 (3.2 g), F5 (3.7 g), F6 (3.1 g) and F7 (7.7 g). Fractions F1 (composed of subfractions 1‐6) and F7 (composed of sub-fractions 78‐104) were complex mixtures containing mostly oils and polar compounds, respectively, and were not further studied. Fraction F2 (composed of subfractions 7‐21) was subjected to column chromatography over silica gel with CH2Cl2-MeOH (98:02), to yield ergosterol (76.7 mg). Fraction F3 (composed of subfractions 22‐34) was further separated by column chromatography over silica gel with CH2Cl2-MeOH (95:05); Sephadex LH-20 eluted with MeOH afforded finally maristachone F (2a, 8.6 mg). Fraction F4 (composed of subfractions 35‐47) was further separated by column chromatography over silica gel/CH2Cl2-MeOH (95:05) and on Sephadex LH-20/MeOH, affording mitrafungidione (1, 4 mg). Fraction F5 (composed of subfractions 48‐58), and fraction F6 (composed of sub-fractions 59‐77) were further separated by column chromatography over silica gel/CH2Cl2-MeOH (90:10) and yielded versiconol (6.3 mg), uracil (18.7 mg), cerebroside B (7.8 mg), and thymine (5-methyluracil, 16.3 mg). The remaining fractions F1 and F7 were not studied.
Mitrafungidione (1)
Reddish oil; ECD spectrum s. Figure 3.
(-)-ESI HRMS: m/z 169.0512 [M-H]- (calc. 169.0506 for C8H9O4) and m/z 361.0900 [2M-2H + Na]- (calc. 361.0904 for C16H18O8Na).
1H NMR, 13C NMR, 1H,13C HMBC and 1H,1H COSY data: Table 1 and Figure 2; see also Supplemental Material).
Maristachone F (2a)
Yellowish oil;
OR:
(+)-ESI HRMS: m/z 235.0946 [M + Na]+ (calc. 235.0941 for C11H16O4Na) and m/z 447.1988 [2M + Na]+.
1H (300 MHz, DMSO-d6 ): δ 9.17 (1H, s br, OH-1), 6.80 (1H, s, H-6), 2.01 (3H, s, Me-7), 3.65 (3H, s, Me-8), 4.44, 4.48 (2H, AB, 11.3 Hz, H2-9), 4.53, 4.88 (2H, 2 s br, 9,10-OH), 5.15 (1H, q, 6.3, H-10), 1.44 (3H, d, 6.3, H-11). 13C, 1H and 2D NMR in CD3OD: see Table 2 and Figure 4 and also the Supporting Information).
1H (300 MHz) and 13C (125 MHz) NMR Data of Maristachone F (2a) and 1H-13C and 1H-1H Correlations Exhibited in the 2D Spectra (125 MHz) in CD3OD (δ in ppm).

Supplemental Material
Online Supplementary file 1 - Supplemental material for New Metabolites From the Endophytic Fungus Cercophora samala Associated With Mitragyna inermis
Supplemental material, Online Supplementary file 1, for New Metabolites From the Endophytic Fungus Cercophora samala Associated With Mitragyna inermis by Erik Donfack Vouffo, Clovis Douanla-Meli, Angelbert Fusi Awantu, Bruno Ndjakou Lenta, Sylvère Ngouela, Etienne Tsamo and Hartmut Laatsch in Natural Product Communications
Footnotes
Acknowledgments
The authors wish to acknowledge the German Academic Exchange Service (DAAD) for awarding a grant to E. Donfack Vouffo (grant no. A/10/90160). They also acknowledge Mr. R. Machinek and Dr. H. Frauendorf for NMR and mass spectra, respectively, Mrs. F. Lissy and A. Kohl for technical assistance.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
