Abstract
We are engaged in the ongoing investigation into the chemical constituents of liverworts in our search for novel compounds and biologically active substances. In the present study, two new rosane diterpenoids were isolated from the Argentine liverwort Anastrophyllum species, together with known aromadendrane sesqui- and rosane diterpenoids. Two new ent-kaurene and 4 ent-kaurene diterpenoids were isolated from the Malaysian liverwort Jungermannia species. Their structures were determined using nuclear magnetic resonance spectroscopy techniques, circular dichroism spectroscopy, and chemical transformation.
In our work on the isolation of novel natural organic compounds and evaluation of their biological activity, we are continuing studies on the phytochemicals of liverworts. 1 -3 We have reported the distribution of sesqui- and diterpenoids from the genus Jungermannia 4 and diterpenoids from the Anastrophyllum species. 5 We reinvestigated phytochemicals of the unknown Argentine and Malaysian Anastrophyllum and Jungermannia species. Here we wish to report the newly isolated lipophilic terpenoids and their structural elucidation and biological activity.
ent-5,15-Rosadien-3-one (

Isolated compounds from the Anastrophyllum species and their derivatives.
The stereostructure of compound

Δδ values for MTPA ester of 3.
The stereostructure of compound
Compound
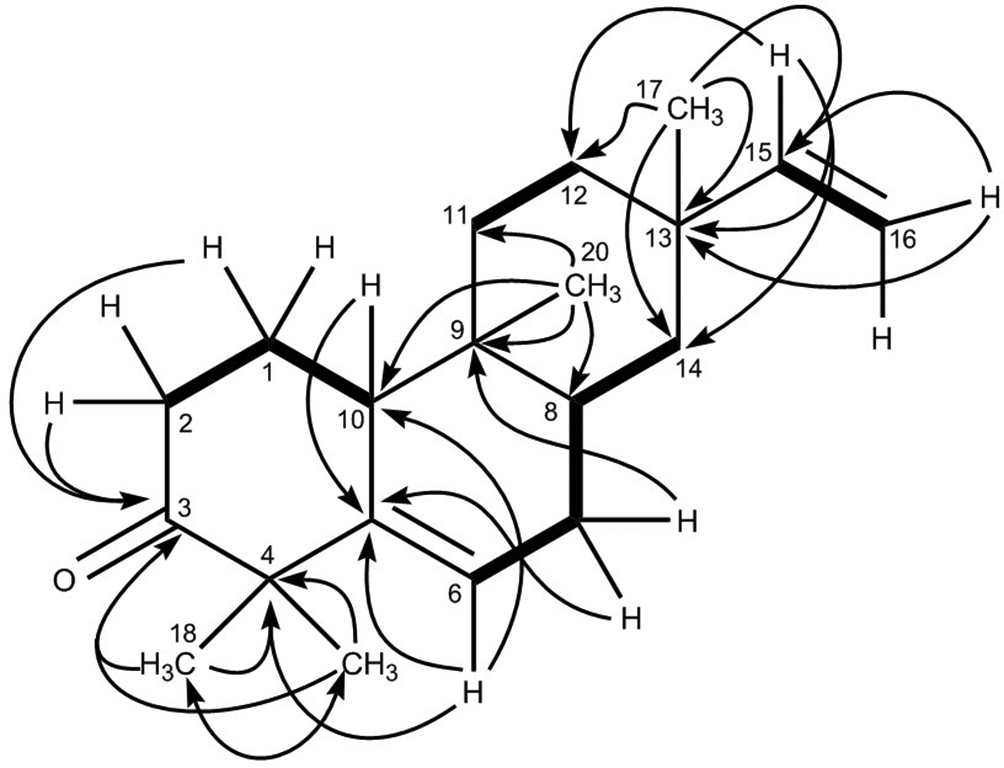
1H-1H COSY (bold lines) and HMBC correlations (arrows) for 1.
In the nuclear Overhauser effect spectroscopy (NOESY) spectrum of

(a) Important NOE correlations for 1. (b) Application of back octant rule to 1.
The 1H NMR spectrum of compound
The unknown Malaysian Jungermannia species produced ent-16-kauren-15-one (

Isolated compounds from Jungermannia species.
The EIMS spectrum of unstable compound
The 1H-1H COSY spectrum of

(a) 1H-1H COSY (bold lines) and HMBC correlations (arrows) for 13. (b) Important NOE correlations for 13.
The IR spectrum of

(a) 1H-1H COSY (bold lines) and HMBC correlations (arrows) for 14. (b) Important NOE correlations for 14.
The CD spectrum of
The absolute configuration of
So far, many isolations of ent-kaurane, labdane, rosane, and verticillane diterpenoids have been reported from the genus Jungermannia. 4 ent-Kaurane diterpenoids were isolated from the present unknown Malaysian Jungermannia species produce. Particularly, it is suggested that ent-kaurane and labdane diterpenoids are characteristic chemical markers of species in the genus Jungermannia.
Compounds
Experimental
General
1H and 13C NMR: 400 and 600 MHz (1H NMR), and 100 and 150 MHz (13C NMR). Chemical shift values are expressed in δ 7.26 (ppm) from CDCl3 as a standard (1H NMR) and δ 77.03 (ppm) from CDCl3 as a standard (13C NMR). Thin layer chromatography: visualized under UV (254 nm) light and by spraying with 15% H2SO4 and Godin’s reagent
16
followed by heating at 120°C-130°C. MeOH-CH2Cl2 (1:1) was used for Sephadex LH-20.
Plant Materials
Anastrophyllum and Jungermannia were collected in Argentina and Malaysia, respectively. These species were identified by Professor S. R. Gradstein and Dr T. Furuki. The voucher specimens were deposited at the Laboratory of Natural Products Chemistry, Daiichi University of Pharmacy.
Extraction and Isolation
The dried and ground material of the unidentified Anastrophyllum species and Jungermannia species was each extracted with Et2O for 1 month. The crude Et2O extract (1.1 g) of an unidentified Anastrophyllum species collected in Tucuman, Argentina, in 2005, was chromatographed on silica gel (n-hexane-EtOAc gradient) and divided into 8 fractions. Fraction (Fr.) 3 was rechromatographed on Sephadex LH-20, silica gel, and preparative high pressure liquid chromatography (prep. HPLC; Unison UK-silica, n-hexane-Et2O 19:1) to give ent-5,15-rosadien-3-one (
The crude Et2O extract (1.72 g) of the Jungermannia species collected in Cameron Highland, Malaysia, in 2005, was chromatographed on Sephadex LH-20 to give 3 fractions. Frs. 1-2 was rechromatographed on Sephadex LH-20 and silica gel to give ent-16-kauren-15-one (
Frs. 1-3 was chromatographed by silica gel and divided into 7 fractions. Frs. 1-3-4 and 1-3-5 were rechromatographed on silica gel and MPLC (Ultra Pack Si-40A, n-Hexane-EtOAc 9:1) to give ent-kauren-16β-ol (
e nt-5,15-Rosadien-3-One (1)
FTIR νmax cm−1: 1713;
1H NMR (600 MHz, CDCl3): δ 1.59 (1H, m, H-1α), 1.99 (1H, dddd, J = 13.5, 5.5, 5.5, 5.5 Hz, H-1β), 2.40 (1H, ddd, J = 15.4, 5.5, 5.5 Hz, H-2α), 2.53 (1H, ddd, J = 15.4, 13.5, 5.5 Hz, H-2β), 5.64 (1H, dd, J = 5.6, 2.2 Hz, H-6), 1.69 (1H, m, H-7α), 1.83 (1H, dddd, J = 17.7, 5.1, 5.1, 2.5 Hz, H-7β), 1.62 (1H, m, H-8), 2.24 (1H, m, H-10), 1.71 (1H, m, H-11α), 1.35 (1H, ddd, J = 13.5, 13.5, 3.8 Hz, H-11β), 1.52 (1H, ddd, J = 13.5, 13.5, 3.8 Hz, H-12α), 1.27-1.31 (2H, m, H-12β and H-14α), 1.16 (1H, ddd, J = 13.5, 3.8, 2.5 Hz, H-14β), 5.82 (1H, dd, J = 17.6, 10.8 Hz, H-15), 4.87 (1H, dd, J = 10.8, 1.4 Hz, H-16), 4.94 (1H, dd, J = 17.6, 1.4 Hz, H-16), 1.04 (3H, s, H-17), 1.24 (3H, s, H-19), 1.26 (3H, s, H-18), 0.69 (3H, s, H-20); 13C NMR: Table 1.
13C NMR Chemical Shifts for the 20 Carbon Atoms in Compounds 1, 2, 8, 13, and 14 (CDCl3, 100 MHz Except Where Indicated).

aMeasured at 150 MHz.
bValues in vertical columns may be interchanged.
EIMS m/z (rel. int.): 286 [M]+ (53), 271 (12), 243 (5), 229 (21), 229 (6), 173 (4), 149 (12), 138 (100), 136 (15), 133 (10), 121 (15), 109 (9), 107 (27), 93 (25), 81 (14), 79 (14), 69 (8), 67 (10), 55 (13), 41 (10).
HR-EIMS: calcd for C20H30O: 286.2297; found: 286.2299.
CD (MeOH): Δε296nm +2.07, Δε209nm −17.10 (c = 3.36 × 10−4)
(3S)-e nt-5,15-Rosadien-3-Ol (2)
FTIR νmax cm−1: 3319.
1H NMR (600 MHz, CDCl3): δ 1.12 (1H, m, H-1α), 1.78 (1H, m, H-1β), 1.81 (1H, m, H-2α), 1.54 (1H, m, H-2β), 3.25 (1H, dd, J = 11.4, 4.3 Hz, H-3), 5.60 (1H, m, H-6α), 1.69 (1H, m, H-7), 1.71 (1H, m, H-7), 1.46 (1H, m, H-8), 1.91 (1H, br d, J = 13.2 Hz, H-10), 1.26 (1H, m, H-11α), 1.65 (1H, m, H-11β), 1.48 (1H, m, H-12α), 1.25 (1H, m, H-12β), 1.28 (1H, m, H-14α), 1.13 (1H, m, H-14β), 5.81 (1H, dd, J = 17.6, 10.7 Hz, H-15), 4.92 (1H, dd, J = 17.6, 1.5 Hz, H-16), 4.85 (1H, dd, J = 10.7, 1.5 Hz, H-16), 1.02 (3H, s, H-17), 1.14 (3H, s, H-19), 0.97 (3H, s, H-18), 0.65 (3H, s, H-20).
13C NMR: Table 1.
EIMS m/z (rel. int.): 270 [M−18]+ (51), 255 (100), 241 (14), 227 (24), 201 (6), 199 (7), 187 (8), 173 (11), 161 (7), 147 (10), 133 (8), 119 (15), 105 (12), 93 (11), 81 (10), 67 (6), 55 (6), 43 (9).
Chemical ionization mass spectrometry (CIMS(CH4)) m/z: 287 [M− H]+.
FAB-MS m/z: 287 [M− H]+.
HR-FAB-MS: calcd for C20H31O: 287.2375; found: 287.2376.
Preparation of (R)- and (S)-MTPA Esters of 3
(R)-MTPA (5 mg), dicyclohexylcarbodiimide (5 mg), and 4-(dimethylamino)pyridine (8 mg) were added to compound
(R)-MTPA Ester 6
1H NMR (600 MHz, CDCl3): δ 1.109 (1H, ddd, J = 13.0, 13.0, 13.0, 3.7 Hz, H-1α), 2.048 (1H, dq, J = 13.0, 3.7 Hz, H-1β), 1.984 (1H, m, H-2α), 1.871 (1H, m, H-2β), 4.6965 (1H, dd, J = 11.5, 4.5 Hz, H-3α), 5.582 (1H, m, H-6), 1.991 (1H, m, H-7α), 1.880 (1H, m, H-7β), 1.766 (1H, m, H-8), 2.753 (1H, m, H-10), 2.725 (1H, d, J = 13.0 Hz, H-12a), 1.9555 (1H, dd, J = 13.0, 2.3 Hz), 1.757 (1H, m, H-14α), 1.359 (1H, br dd, J = 9.9, 2.3 Hz, H-14β), 5.8265 (1H, dd, J = 17.4, 10.7 Hz, H-15), 4.9435 (1H, dd, J = 10.7, 0.8 Hz, H-16), 4.961 (1H, dd, J = 17.4, 0.8 Hz, H-16), 0.972 (3H, s, H-17), 0.968 (3H, s, H-19), 1.024 (3H, s, H-18), 0.996 (3H, s, H-20).
(S)-MTPA Ester 7
1H NMR (600 MHz, CDCl3): δ 1.096 (1H, ddd, J = 13.0, 13.0, 13.0, 3.7 Hz, H-1α), 2.0195 (1H, dq, J = 13.0, 3.7 Hz, H-1β), 1.912 (1H, m, H-2α), 1.752 (1H, m, H-2β), 4.6665 (1H, dd, J = 11.6, 4.5, H-3α), 5.595 (1H, m, H-6), 1.989 (1H, m, H-7α), 1.883 (1H, m, H-7β), 1.763 (1H, m, H-8), 2.7335 (1H, m, H-10), 2.7215 (1H, d, J = 12.5 Hz, H-12a), 1.9495 (1H, dd, J = 12.5, 2.3 Hz), 1.757 (1H, m, H-14α), 1.3575 (1H, br dd, J = 9.9, 2.3 Hz, H-14β), 5.8245 (1H, dd, J = 17.4, 10.7 Hz, H-15), 4.9415 (1H, dd, J = 10.7, 0.8 Hz, H-16), 4.959 (1H, dd, J = 17.4, 0.8 Hz, H-16), 0.963 (3H, s, H-17), 1.067 (3H, s, H-19), 1.024 (3H, s, H-18), 0.994 (3H, s, H-20).
Oxidation of 2
Compound
Reduction of 4
Compound
e nt-5,15-Rosadiene-3β,11β-Diol (8)
FTIR νmax cm−1: 3431.
1H NMR (600 MHz, CDCl3): δ 1.14 (1H, dddd, J = 13.3, 13.3, 13.3, 4.0 Hz, H-1α), 1.73 (1H, m, H-1β), 1.80 (1H, m, H-2α), 1.59 (1H, m, H-2β), 3.25 (1H, dd, J = 11.5, 4.0 Hz, H-3), 5.58 (1H, m, H-6), 1.71 (1H, m, H-7), 1.85 (1H, m, H-7), 1.84 (1H, m, H-8), 2.68 (1H, br d, J = 13.2 Hz, H-10), 3.80 (1H, t, J = 3.2 Hz, H-11), 1.72 (1H, m, H-12α), 1.50 (1H, dt, J = 14.8, 3.2 Hz, H-12β), 1.32 (1H, t, J = 13.6 Hz, H-14α), 1.21 (1H, m, H-14β), 5.75 (1H, dd, J = 17.6, 10.8 Hz, H-15), 4.84 (1H, dd, J = 10.8, 1.2 Hz, H-16), 4.89 (1H, dd, J = 17.4, 1.2 Hz, H-16), 1.22 (3H, s, H-17), 1.15 (3H, s, H-18), 0.99 (3H, s, H-19), 0.64 (3H, s, H-20).
13C NMR: Table 1.
EIMS m/z (rel. int.): 286 [M −18]+ (100), 268 (88), 253 (86), 225 (92), 211 (22), 189 (85), 185 (50), 171 (73), 157 (47), 143 (37), 131 (42), 119 (44), 105 (45), 91 (34), 79 (18), 69 (16), 69 (8), 55 (23), 41 (20).
FAB-MS m/z: 327 [M+Na]+, 343 [M+K]+.
HR-FAB-MS: calcd for C20H32O2Na: 327.2300; found: 327.2284.
e nt-15-Oxo-16-Kauren-20-Al (13)
FTIR ν max cm−1: 1726, 1707.
1H NMR (400 MHz, CDCl3): δ 0.62 (1H, m, H-1β), 0.76 (3H, s, H-19), 0.95 (3H, s, H-18), 1.20 (1H, m, H-3β), 1.24–1.59 (m, overlapped signals H-2, 3, 6, 7, 11, 12 and 14), 1.35 (1H, m, H-5), 1.60 (1H, m, H-9), 1.80-1.91 (m, overlapped signals H-6, 11 and 14), 2.07 (1H, ddd, J = 13.5, 13.5, 4.2 Hz, H-12), 2.60 (1H, br d, J = 13.0 Hz, H-1α), 3.01 (1H, br s, H-13), 5.27 (1H, s, H-17), 5.95 (1H, s, H-17), 10.43 (1H, s, H-20).
13C NMR: Table 1.
EIMS m/z (rel. int.): 300 [M]+ (22), 288 (66), 271 (70), 255 (21), 245 (36), 215 (12), 201 (16), 188 (15), 173 (24), 162 (90), 160 (50), 150 (18), 147 (37), 135 (48), 126 (42), 111 (100), 105 (53), 91 (77), 79 (48), 69 (36), 55 (37), 43 (62).
HR-EIMS: calcd for C20H28O2: 300.2089; found: 300.2086.
UV (MeOH): λmax(log ε): 228 nm (4.46) (c 1.23 × 10−4).
CD (MeOH): Δε221nm −13.7, Δε237nm −5.1, Δε349nm −2.16 (c 1.23 × 10−4).
ent-20-Hydroxy-16-Kauren-15-One (14)
FTIR νmax cm−1: 3514, 1726.
1H NMR (600 MHz, CDCl3): δ 2.16 (1H, m, H-1α), 0.65 (1H, ddd, J = 13.5, 13.5, 3.8 Hz, H-1β), 1.50-1.57 (2H, m, H-2α and 12β), 1.42-1.48 (3H, m, H-2β, 3α and 14β), 1.20 (1H, ddd, J = 13.0, 13.0, 4.4 Hz, H-3β), 1.08 (1H, dd, J = 12.6, 2.0 Hz, H-5), 1.33-1.41 (2H, m, H-6α, 7α), 1.58 (1H, m, H-6β), 2.03 (1H, ddd, J = 13.9, 13.9, 4.9 Hz, H-7β), 1.32 (1H, m, H-9), 1.41 (1H, m, H-11α), 1.81 (1H, dd like, J = 15.2, 5.9 Hz, H-11β), 2.19 (1H, ddd, J = 12.8, 5.9, 2.7 Hz, H-12α), 3.03 (1H, br s, H-13), 2.56 (1H, d, J = 11.8 Hz, H-14α), 5.24 (1H, t, J = 1.2 Hz, H-17), 5.92 (1H, t, J = 1.2 Hz, H-17), 0.90 (3H, s, H-18), 0.86 (3H, s, H-19), 4.06 (1H, d, J = 12.6 Hz, H-20), 4.11 (1H, d, J = 12.6 Hz, H-20).
13C NMR: Table 1.
EIMS m/z (rel. int.): 302 [M]+ (86), 284 (24), 271 (100), 253 (51), 245 (79), 243 (81), 215 (19), 201 (48), 189 (22), 175 (40), 161 (21), 157 (26), 147 (28), 131 (27), 119 (29), 105 (45), 91 (67), 79 (43), 69 (32), 55 (25), 41 (27).
HR-EIMS: calcd for C20H30O2: 302.2246; found: 302.2248.
UV (MeOH): λmax (log ε): 236 nm (3.84) (c 4.44 × 10−4).
CD (MeOH): Δε217nm −8.31, Δε349nm −0.77 (c 4.44 × 10−4).
Bioassay of the Isolated Compounds
Capase-1 inhibitory and cytotoxic activity tests against HL-20 cells of the isolated compounds were conducted by Professor Dr T. Ito, Miss S. Nakamatsu, and Dr Y. Yagi (Tokushima Bunri University), respectively.
Footnotes
Acknowledgments
The authors thank Professor Dr S. R. Gradstein (Museo de Histria Natural de Paris, France) and Dr T. Furuki (Natural History Museum and Institute, Chiba, Japan) for their identification of the liverwort species. Thanks are also due to Dr M. Tanaka and Dr Y. Okamoto (Tokushima Bunri University) for conducting 600 MHz NMR and mass spectrometry measurements. We thank Professor T. Ito and Miss S. Nakamatsu for the capase-1 inhibitory activity, and Dr Y. Yagi for the cytotoxic activity.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
