Abstract
Five naphthoquinones including 4 compounds with new absolute configurations, (–)-2'S-trypethelone methyl ether (
Cultured lichen mycobionts have shown to be capable of producing diverse metabolites under stressed conditions that were not detectable in natural lichens, but are structurally related to fungal metabolites. 1 -3 Trypetheliaceae is a family of crustose lichens with approximately 15 genera and 418 species around the world recorded. 4,5 Thailand is located in the tropical region with diverse distribution of lichens. In 2017, the total Thai lichens were reported to be at least 1292 species and, currently, Trypetheliaceae crustose lichens were found in 44 species. 6 -8 The genus Marcelaria, 9 especially Marcelaria cumingii, possesses dominant morphological characteristics and is an abundant source of secondary metabolites, comprising anthraquinones and xanthones. 10 -12 Nevertheless, no chemical data on the cultured mycobiont of this species have been found.
1,2-Naphthoquinone derivatives were regarded as major components of mycobionts derived from some lichens of Trypetheliaceae family, such as Astrothelium sp and Trypethelium sp. 13 -15 Several mycobiont-derived o-naphthoquinones were isolated; nonetheless, the absolute stereochemistry of these metabolites was not completely assigned. 3,13,14,16 Using the ascospore discharge technique, 17 M. cumingii was successfully cultured. Herein we reported the separation of the methanol extract of this cultured mycobiont (Figure 1) resulting in the isolation of 5 naphthoquinones. The structural elucidation and their cytotoxicity against several cancer cell lines were conducted.
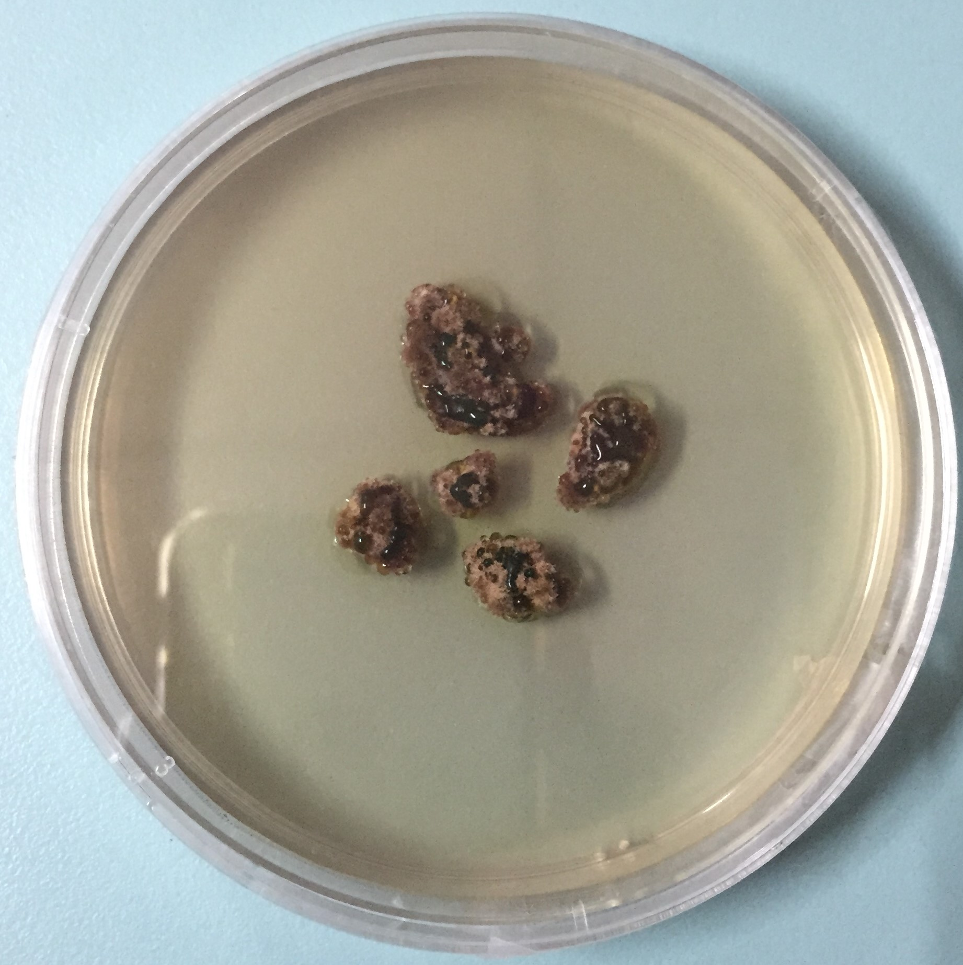
Cultured mycobiont of Marcelaria cumingii.
Results and Discussion
Compound
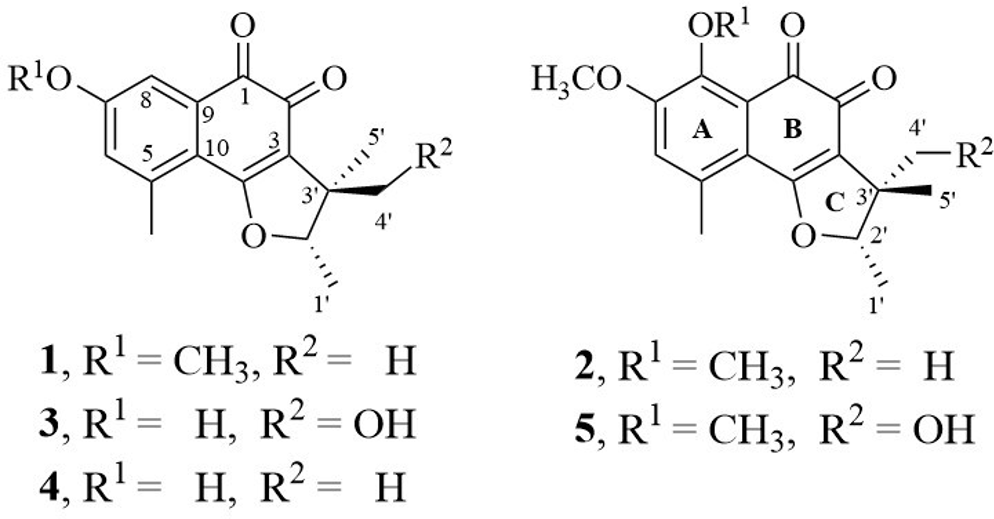
Chemical structures of 1 to
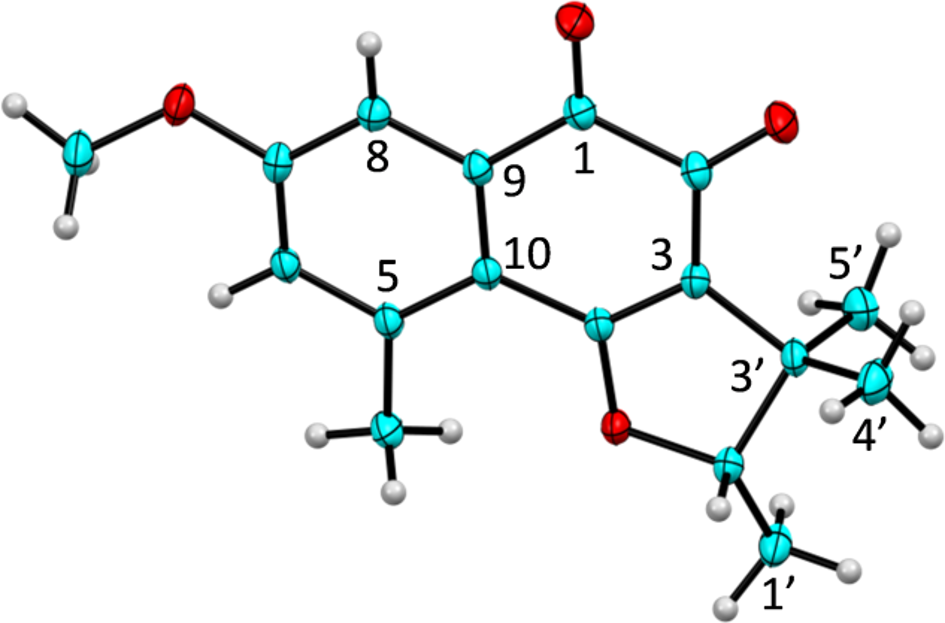
OOak Ridge Thermal Ellipsoid Plot of
All isolated compounds were preliminarily evaluated for cytotoxic activity against KB and HeLaS3 cell lines. (–)-2'S-trypethelone methyl ether (
In Vitro Cytotoxicity of

IC50, half-maximal inhibitory concentration.
aPositive control.
In Vitro Cytotoxicity of 1 Against Vero, MRC-5, HT29, HCT116, and A549 Cell Lines.

IC50, half-maximal inhibitory concentration.
Computational chemistry was applied to clarify the SAR and showed the comparison of molecular binding pose between doxorubicin and all inhibitors (supplemental Figure S1). All compounds bound at the same site and laid on the protein surface. The binding region was found to be similar to that reported by Kongkathip.
18
Doxorubicin was shown to be the most active compound against KB and HeLaS3. This corresponded with the docking result, which showed that it formed H-bond with Arg99 (2.05 Å), Met101 (2.18 and 2.25 Å), Val103 (1.75 Å), Gly105 (1.68 Å), Ile130 (2.83 Å), and Asp515 (1.86 Å). (–)-2'S-Trypethelone methyl ether (
(–)-2'S-Trypethelone methyl ether (
Conclusions
The methanol extract from cultured mycobiont of Marcelaria cumingii yielded 4 naphthoquinones with new absolute configurations: (–)-2'S-trypethelone methyl ether (
Experimental
General Experimental Procedures
The NMR spectra were measured on a Bruker Avance III (400 MHz for 1H NMR and 100 MHz for 13C NMR) and Varian Mercury-400 Plus NMR (400 MHz for 1H NMR and 100 MHz for 13C NMR) spectrometers. The HRESIMS were recorded on a HRESIMS Bruker microTOF. Thin-layer chromatography was carried out on precoated silica gel 60 F254 or Sephadex LH-20 and spots were visualized by UV254nm, UV365nm lamp. Column chromatography was performed with silica gel 60 (0.040-0.063 mm). Solvents used for isolation were n-hexane, dichloromethane, ethyl acetate, acetone, and methanol.
Fungal Isolation, Cultivation, and Identification
Specimen of M. cumingii was collected from tree bark at Pak Chong district, Nakhon Ratchasima, Thailand (700 m alt.). The mycobiont of M. cumingii was successfully isolated from perithecia of lichen thallus by the ascospore discharge technique 17 and was cultivated in 90 mm Petri dishes containing Malt-Yeast-Extract agar at room temperature (30-32°C) for 9 weeks. The voucher specimen was identified by Dr Theerapat Luangsuphabool and was deposited at the Lichen Herbarium, Ramkhamhaeng University (voucher No. RAMK027993). The fungal culture is maintained in the lichen research unit at Ramkhamhaeng University, Thailand. The molecular data were analyzed to confirm the species identification. The genomic DNA was extracted from the mycobiont culture using CTAB precipitation protocol. 19 The nuclear large subunit ribosomal DNA (nuLSU) and mitochondrial small subunit ribosomal DNA (mtSSU) loci were amplified using primer pairs LR0R/LR3 20 and mrSSU1/MSU7, 21,22 respectively. Polymerase chain reaction conditions and DNA sequencing followed protocols previously described. 23 The nuLSU and mtSSU sequences were deposited in DDBJ (accession number LC223104 and LC223105) and confirmed similarity sequences to M. cumingii (KM453789; 99%) and (LC034284; 99%), respectively.
Extraction and Isolation
The fungal biomass and agar were separately extracted. The mycobiont colonies were extracted with methanol at room temperature to yield the crude extract (0.45 g). This crude extract was applied to normal phase silica gel column, eluted with the solvent system of dichloromethane:methanol (25:1) to afford 5 fractions
(–)-Trypethelone methyl ether (1 )
Dark violet-red crystalline solid (4.9 mg);
(–)-8-Methoxytrypethelone methyl ether (2 )
Violet-red crystalline solid (2.9 mg);
(–)-4'-Hydroxytrypethelone (3 )
Violet crystalline solid (4.1 mg);
(–)-Trypethelone (4 )
Violet amorphous solid (3.0 mg);
(–)-4'-Hydroxy-8-methoxytrypethelone methyl ether (5 )
Orange amorphous solid (1.1 mg);
Single-Crystal X-Ray Crystallography
With the help of a Bruker X8 PROSPECTOR KAPPA CCD diffractometer equipped with an IµS X-ray microfocus source operated at 45 kV, 0.65 mA, producing an intense monochromatic CuKα radiation (λ = 1.54178 Å), the diffraction data of
Cytotoxicity Assay
Cytotoxic activities of isolated compounds were tested using the standard MTT colorimetric method previously described. 18
Molecular Docking Calculations
Protein and inhibitors preparation
The protein structure of topoisomerase II was retrieved from the Protein Data Bank (pdb code: 1AB4). 25 The crystallographic water molecules were deleted and the hydrogen atoms were added to protein by using AutoDockTools 1.5.6. 26 The inhibitors were constructed and the geometry was optimized with PM6 level of theory by using MOPAC2009. 27,28
Molecular docking calculations
The atomic potential grid box for molecular docking was constructed with the grid size of 126 126 126 point. The distance spacing between each point is 0.375 Å. Subsequently, doxorubicin and all 5 inhibitors were docked into the protein by using AutoDock 4.2.6 program. 26 The Lamarckian Genetic Algorithm was employed and the Gasteiger charge was employed to all inhibitors. The calculations of each inhibitor were set to 500 cycles while the other parameters were set as default. The most populous pose was selected for further molecular binding analyses.
Supplemental Material
Supplementary data - Supplemental material for Naphthoquinones From Cultured Mycobiont of Marcelaria cumingii (Mont.) and Their Cytotoxicity
Supplemental material, Supplementary data, for Naphthoquinones From Cultured Mycobiont of Marcelaria cumingii (Mont.) and Their Cytotoxicity by Suekanya Jarupinthusophon, Theerapat Luangsuphabool, Thammarat Aree, Thuc-Huy Duong, Kiattisak Lugsanangarm, Prayumat Onsrisawat, Pongpun Siripong, Ek Sangvichien and Warinthorn Chavasiri in Natural Product Communications
Footnotes
Acknowledgments
SJ would like to thank the 90th Anniversary of Chulalongkorn University Fund (Ratchadaphiseksomphot Endowment Fund, GCUGR1125603021D) and Phranakhon Rajabhat University Development Fund. The Thailand Research Fund via Directed Basic Research Grant (Grant no. DBG6180029) for financial support is also acknowledged.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
