Abstract
The chemical composition of 3 samples of Ligularia dictyoneura was studied. Nine furanoeremophilanes
We are studying the chemical and genic diversity in Ligularia (Asteraceae) in the Hengduan Mountains area of China using root chemicals and evolutionarily neutral base sequences as the indices. 1,2 Eremophilane-type sesquiterpenoids have been isolated from most of the major Ligularia species and many Ligularia species have been found to harbor intraspecific diversity in their root chemicals. On the basis of the isolated chemicals and our observation on the abundance of the plants in the field, we have proposed a hypothesis that furanoeremophilane-producing species or intraspecific groups are ecologically advantageous over those producing eremophilan-8-ones. 1
Ligularia dictyoneura (Franch.) Hand.-Mazz. grows in grassy slopes and scrubs in northwestern Yunnan and southwestern Sichuan Provinces. 3 Some eremophilane sesquiterpenes have been isolated from this species. 4 -7 We isolated eremophilan-8-one derivatives from samples collected in Shangrila (formerly Zhongdian) area, Yunnan, and furanoeremophilanes from samples collected in its surrounding areas (Figure 1). 8 On the basis of the TLC patterns of their root extracts, furanoeremophilane-producing L. dictyoneura samples were subgrouped into 6 chemotypes (types 1-6). 8 Because furanoeremophilanes are believed to be generated from eremophilan-8-ones 9 (Scheme 1), furanoeremophilane-producing populations of L. dictyoneura may have evolved from those producing eremophilan-8-ones.

Biosynthesis of furanoeremophilanes.
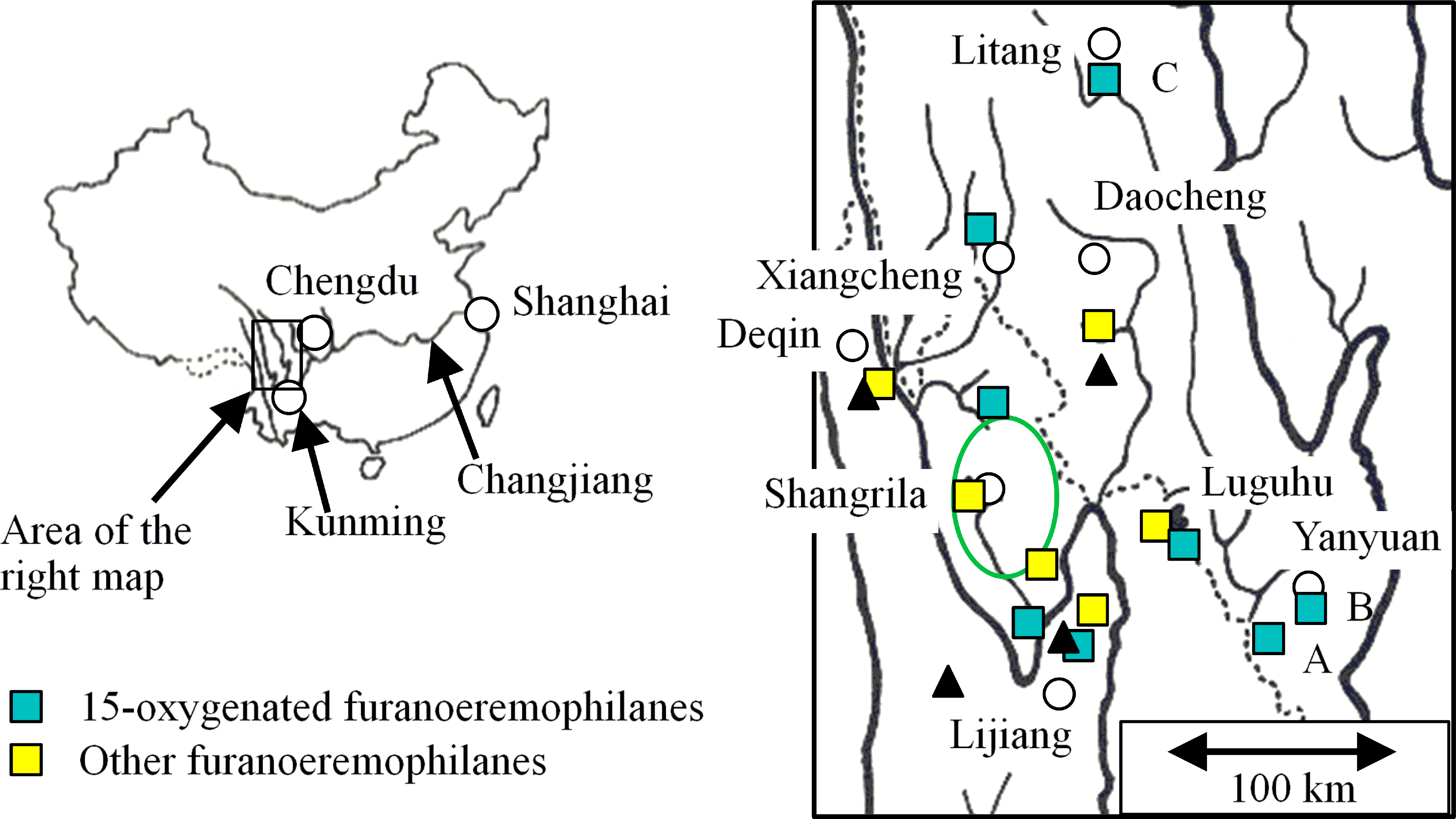
Collection locations of L. dictyoneura (squares). Circles and triangles indicate major cities and peaks, respectively. Solid and dotted lines indicate rivers and province boundaries, respectively. The green circle indicates the area of 8 samples with eremophilan-8-one. 8
Here we report the results of analyses on 3 samples of L. dictyoneura. They were collected further afield from our previous L. dictyoneura samples. Samples A to C were collected in Yanbian, Yanyuan, and Litang counties, Sichuan, respectively (Table 1 and Figure 1). One of the compounds isolated from sample A was new. The compounds from samples B and C were only listed previously 1 and here we describe the details of their analyses.
Collection Localities and Isolated Compounds of Ligularia dictyoneura Samples.

aCounty in parentheses. All in Sichuan Province.
The EtOH extract of each root sample was subjected to Ehrlich’s test, which is a facile method to detect furanoeremophilanes. 10,11 The 3 samples were positive to the test. The TLC pattern of sample A was similar to that of the previous type-2 samples and the chemotype of this sample was judged to be type 2 (Table 1). 8 Sample B showed 2 large pink spots at R f = 0.59 and 0.65 (hexane-EtOAc 7:3). In addition, 3 small pink spots (R f = 0.43, 0.76, and 0.88) and an orange spot of ligularone (R f = 0.80) were observed. Sample C showed the same 2 large spots; however, of the 4 minor spots, the orange spot and the least polar spot were absent. These TLC patterns were different from that of any of the previously analyzed samples of types 1 to 6, and, therefore, new chemotypes 7 and 8 were assigned to samples B and C, respectively (Table 1). 1

Compounds in each sample were isolated by use of silica gel column chromatography (CC) and HPLC. From sample A, compounds
The molecular formula of
1H and 13C NMR Data of New Compound 6 (C6D6).

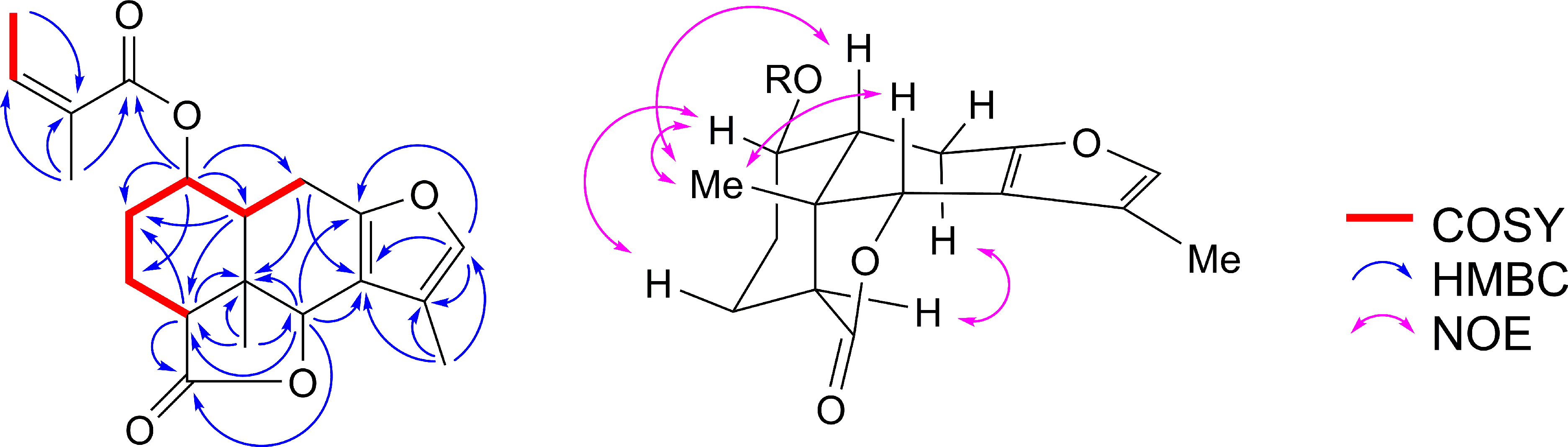
Selected COSY, HMBC, and NOESY correlations of compound 6.
The base sequence of the ITS1-5.8S-ITS2 region was determined for samples A to C. Samples 3 to 14 and 16 to 19 from our previous study on L. dictyoneura 8 were also reanalyzed. The adoption of a new set of primers 24 and a new sequencing reaction kit has improved the quality of data and allowed us to determine the sequences that had remained unknown in these samples. The results, as well as those previously determined for samples 1, 2, 15, and 20, are shown in Table 3. Subjecting the data to a phylogenetic analysis program MEGA X 25 returned no phylogenetic tree with a significant bootstrapping value, implying that the samples have not differentiated very much genetically. (See below for discussion on sample 9.)
aK = G + T; R = A + G; S = C + G; W = A + T; Y = C + T; - = deletion.
bThe base numbering based on the sequence of sample 15, which had been deposited in the database (AB299047).
cSamples A to C, the present work; samples 1 to 20, our previous work. 8
dA sequence with TTTT was also present.
eA sequence with AAA in place of AA at 252 to 253 of ITS1 was also present.
fA sequence without AC at 163 to 164 of 5.8S was also present.
gA sequence with T in place of GGA at 170 to 172 of ITS2 was also present.
Nine furanoeremophilanes were isolated from the 3 L. dictyoneura samples. Of them, 4 were 15-oxygenated derivatives (
The eremophilan-8-one type of L. dictyoneura is found in the Shangrila area, where the plant is most abundant (Figure 1, green circle); the furanoeremophilane type, in the surrounding areas. The only exception is sample 9 in our previous study: Although it was surrounded by eremophilan-8-one producers, furanoeremophilanes
The characteristic components in 4 of the 8 furanoeremophilane-producing types (types 2, 4, 7, and 8) are 15-oxygenated derivatives. 15-Oxygenated furanoeremophilanes have been isolated from L. tongolensis, 26 L. cymbulifera, 26 L. vellerea (Shangrila-type), 19 and L. hookeri. 27 Because the distribution areas of these plants and L. dictyoneura almost overlap (Shangrila County and the surrounding areas, see Figure 1), the production of similar compounds in these species may not be a coincidence.
Experimental
General
NMR, JEOL ECX-400 (400 MHz for 1H; 100 MHz for 13C) or JEOL AL 400 spectrometer; IR, JASCO FT/IR-230 spectrometer; MS, JEOL JMS-700 MStation or CMATE II. CC was performed on silica gel (Wakosil C-200 or C-300 or Kanto silica gel 60 N [spherical neutral]). Analytical TLC was carried out on Merck Kieselgel 60 F254, 0.2 mm thickness, using either Ehrlich’s reagent (p-dimethylaminobenzaldehyde and HCl) or p-anisaldehyde/AcOH/H2SO4 as the visualizing agent. HPLC was carried out using either a Shimadzu LC-20AT pump with a SPD-20A Prominence UV/VIS detector or a GL Sciences GL-7410 pump with a GL-7450 UV detector, and a Hitachi D-2500 Chromato-Integrator or a Shimadzu C-R8A Chromatopac, with either a GL Sciences Inertsil PREP-ODS column (20 × 250 mm) or a Kanto Mightysil Si60 (10 × 250 mm) column. DNA was purified from dried leaves using DNeasy Plant Mini Kit (Qiagen). A 30-cycle amplification of the ITS1-5.8S-ITS2 region by polymerase chain reaction was carried out with LC5 and LC6 primers 24 and HotStarTaq plus Master Mix Kit (Qiagen). The amplified DNA was separated by agarose gel electrophoresis and purified with High Pure PCR Product Purification Kit (Roche Diagnostics). Sequencing reactions were carried out with LC1 to LC4 primers 24 and BigDye Terminator Cycle Sequencing Kit Ver. 3.1 (Applied Biosystems). Sequence determination was carried out on a 3130xl Genetic Analyzer (Applied Biosystems).
Plant Materials
Samples were collected at locations shown in Table 1 and Figure 1 in August 2012 (sample A) and 2007 (samples B and C) and identified by X. G. (author). Voucher specimen numbers are 2012-52, 2007-25, and 2007-103, respectively (Kunming Institute of Botany).
Isolation of Compounds
Extraction of dried root of sample A (22.4 g) with EtOH afforded an extract (2.08 g). Silica-gel CC (n-hexane-EtOAc, gradient) of the extract afforded 5 fractions. CC (n-hexane-EtOAc) of Fr. 1 (eluted with n-hexane) afforded 7 fractions (Fr. 1-1 to 1-7). From Frs. 1-2 and 1-3,
CC (SiO2, n-hexane-EtOAc, gradient) of a part (182.8 mg) of the EtOH extract of dried root of sample B (7.7 g) afforded 4 fractions. From Fr. 1 (eluted with n-hexane),
CC (SiO2, n-hexane-EtOAc, 20:1) of a part (166.7 mg) of the EtOH extract of dried root of sample C (7.7 g) afforded 4 fractions. From Fr. 1,
Compound
An oil.
[α]D 25 +67.2 (c, 0.361, MeOH).
IR (neat/NaCl): 1780, 1711, 1645, 1234, 1157, 945, 756 cm-1.
1H and 13C NMR (C6D6): see Table 2.
MS (CI, CH4): m/z (%) = 345 [M+H+] (95), 245 [M+H+-AngOH] (100), 199 (51).
HRMS-CI m/z [M+H+] calcd for C20H25O5: 345.1702; found: 345.1697.
Footnotes
Acknowledgements
The authors thank Dr Takayuki Kawahara, Forestry and Forest Products Research Institute, and Dr Yoshinori Saito, Nagasaki University, for assistance in sample collection and discussion.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Grants-in-Aid for Scientific Research from JSPS (Nos. 16404008, 21404009, and 25303010) and Strategic Research Foundation Grant-Aided Projects for Private Universities from the Ministry of Education, Culture, Sports, Science, and Technology, Japan.

