Abstract
In the present study, the synergistic effects of BASk, a combination of betulinic acid (B), apigenin (A), and skimmianine (Sk) in the ratio of 1:1:1, were studied to construct a novel drug mixture against inflammation via the TLR4-nuclear factor Kappa light chain enhancer of activated B cells (NFκB) signaling pathway.
Keywords
Modern drugs usually target a single specific pathway, whereas plant products can act in an orchestral way with various bioactive compounds acting synergistically on their targets in complex cellular pathways and often being less toxic than synthetic drugs.
1
In order to reduce the rate of failure during drug development,
Inflammation is body’s defense mechanism either to remove or repair damaged tissues or to eradicate harmful pathogens thereby helps to attain a healthy state. However, anti-inflammatory condition can deteriorate due to excess and persistent pro-inflammatory mediators leading to chronic inflammatory diseases such as rheumatoid arthritis, atherosclerosis, diabetes, pulmonary diseases, and cancer 3 ; hence, it should be controlled in a proper way.
TLR4 is a transmembrane pattern recognition receptor which has the ability to bind with a spectrum of ligands for initiating an immune response. It is the only receptor that needs the accessory protein MD2 4 for its activation. Thus, the heterodimer, TLR4/MD-2 complex, initiates dimerization of intracellular domains which recruit the adapter protein molecules for the activation of the TLR4 signaling cascade, resulting in an immune response. This protein-protein interaction mediates the activation of the transcription factor NFκB resulting in the release of pro-inflammatory mediators responsible for the progression of inflammation. 5
Our previous laboratory studies reported the beneficial role of betulinic acid (B),
6
apigenin (A),
7
and skimmianine (Sk)
8
against inflammation, and thus, we selected them for conducting experiments on oxidized low density lipoprotein (ox-LDL) induced inflammation. In the present study, we combined 3 compounds together, named BASk, for the first time to develop a new drug formulation against inflammation. Betulinic acid is a triterpenoid, Apigenin is a flavonoid, and skimmianine (Sk) an alkaloid. We designed the work in such a way to find the effect exerted by 3 compounds when used in a combined form instead of using individually. Hence, to investigate the effect of BASk on inflammation, we selected the TLR4-Nuclear factor Kappa light chain enhancer of activated B cells (NFκB) signaling pathway since it plays a key role in the generation of inflammatory responses during an antigenic stimulus. As a preliminary step, we conducted
The drug likeness properties of 3 bioactive compounds (Figure 1) were predicted according to Lipinski’s RO5 and confirmed by Molinspiration property engine v2016.10 (Table 1). The molecular weights (MWs) of 3 compounds were below 500 which indicate their better absorption, diffusion, and transportation in body fluids. Similarly, topological polar surface area and number of rotatable bonds (nrotb) were below 160 Å and 10, respectively, which indicate the better efficacy of 3 compounds in intestinal absorption, bioavailability, and crossing the blood brain barrier. Thus, all 3 compounds fulfilled the limitations of both RO5 and Molinspiration. Similarly, their bioactivity score was identified using Molinspiration bioactivity score v2016.03 (Table 2) and it ranges from moderately active to active. Thus, the results reveal the possibility that the multiple physiological functions exhibited by the compounds were due to their interaction with drug targets such as G protein-coupled receptor (GPCR) ligands, ion channel modulators, kinase inhibitors, nuclear receptors, protease inhibitors, and enzyme inhibitors.

Structure of (a) betulinic acid, (b) apigenin, and (c) skimmianine generated in Molinspiration cheminformatics software using simplified molecular input line entry system (SMILE) notation.
Drug Likeness Score of 3 Bioactive Compounds Predicted by Lipinski’s Rule of Five and Molinspiration Property Engine v2016.10.

GPCR, G protein-coupled receptor.
Bioactivity Score of Betulinic Acid, Apigenin, and Skimmianine Predicted by Molinspiration Bioactivity Score v2016.03.

MW, molecular weight; Natom, number of atoms; nOHNH, number of hydrogen bond donor; nON, number of hydrogen bond acceptor; nrotb, number of rotatable bonds; TPSA, topological polar surface area.
Molecular interaction of ligand molecules toward the active site of the receptor (MD2) helps to determine their strong affinities toward the target protein and it was examined via docking interaction studies. From the docking study (Table 3 and Figure 2), it is clear that 3 compounds show strong binding interactions with the amino acid residues present in the active site of the MD2 receptor. These 3 compounds share some common amino acid interacting residues such as Val82 and Met85 with MD2 via hydrophobic bonds. Similarly, Sk, through hydrogen bonds, and B, through hydrophobic bonds, interact with a common residue Leu87 on MD2, and A showed a strong salt bridge interaction with the residue Lys128. Thus, B, A, and Sk all exhibit strong binding interactions with the active site of MD2 via hydrogen and hydrophobic bonds and a salt bridge, respectively.
Interacting Amino Acid Residues and Type of Binding Interactions With the Active Site of the MD2 Receptor Shown by Betulinic Acid, Apigenin, and Skimmianine.

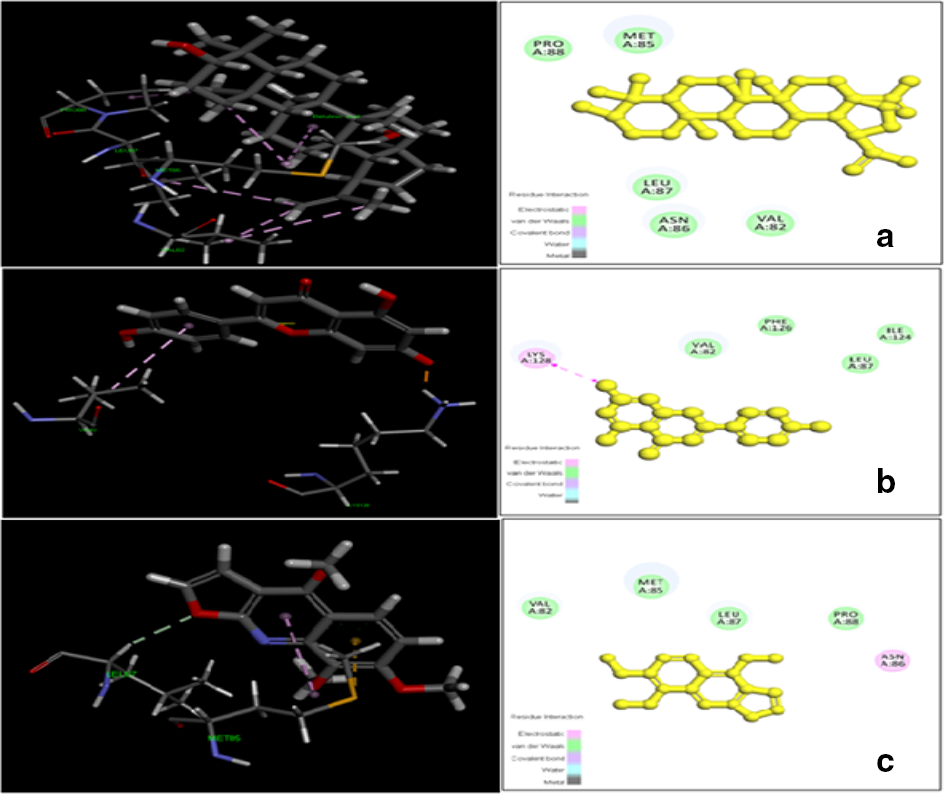
Molecular docking interactions of (a) betulinic acid, (b) apigenin, and (c) skimmianine in the active site of the human receptor protein MD2.
The synergistic effect of BASk on MD2 and further activation of TLR4 were examined in ox-LDL induced hPBMCs. In the present study, the cultured hPBMCs were pretreated with the bioactive compounds prior to ox-LDL induction. TLR4 (Figure 3(a)) and MD2 (Figure 3(b)) reached higher levels in the ox-LDL group compared with normal hPBMCs. However, its level was depleted during pretreatment with BASk and showed a significant depletion of TLR4 and MD2 in comparison with the individual compounds. Thus, BASk inhibited TLR4 activation by preventing its interaction with MD2 and showed a better effect compared with its individual compounds. This significant reduction of MD2 and TLR4 by BASk highlighted its inhibiting role in the activation of the TLR4 signaling pathway during inflammation.

Inhibitory effect of BASk on (a) TLR4 and (b) adapter protein MD2. Human peripheral blood mononuclear cells isolated from human blood were pretreated with a specific concentration of BASk (25 µM) 1 h prior to ox-LDL induction. Cell lysates were collected and the protein levels of the transmembrane receptor TLR4 and the adapter protein MD2 were determined by enzyme linked immunosorbent assay. Values are expressed as average of 6 samples ± SEM. aStatistical difference with group I at
The cytosolic and nuclear fraction of ox-LDL induced pretreated hPBMCs was collected to determine the nuclear translocation of NFκB during TLR4 activation. ox-LDL induced hPBMCs showed an increased nuclear translocation compared with the control group (Figure 4). However, the nuclear translocation of NFκB produced by BASk was significantly greater than that of the individual compounds. The result indicated the inhibiting role of BASk in NFκB activation.

Effect of BASk on NFκB activation and its nuclear translocation in human peripheral blood mononuclear cells. Isolated human peripheral blood mononuclear cells were pretreated with betulinic acid, apigenin, skimmianine, and BASk at a concentration of 25 µM 1 h before induction with ox-LDL. After 24 h, nuclear and cytosolic fractions were prepared and the level of NFκB was determined by enzyme-linked immunosorbent assay. Values are expressed as average of 6 samples ± SEM. aStatistical difference with group I at
The cyclooxygenase called prostaglandin-endoperoxide synthase is the key enzyme involved in arachidonic acid metabolism responsible for the formation of lipid mediators during inflammation. Cyclooxygenase - 2 (COX-2), its inducible isoform expressed in various cells types, is primarily responsible for the production of prostaglandins. Pretreatment of hPBMCs with BASk significantly reduced the expression of COX-2 (Figure 5(a)) during ox-LDL induction and in turn reduced the release of pain initiating prostaglandin, PGE2 (Figure 5(b)), in the culture supernatant.

Effect of BASk on mRNA expression of (a) COX-2 and (b) PGE2 in ox-LDL induced human peripheral blood mononuclear cells. mRNA was isolated from pretreated human peripheral blood mononuclear cells with the help of an RNA isolation kit. For quantification, mRNA expression data via reverse transcriptase polymerase chain reaction of bioactive compounds were normalized to glyceraldehyde-3-phosphate dehydrogenase. The inhibitory effect of BASk on the release of PGE2 was estimated by enzyme linked immunosorbent assay. Values are expressed as average of 6 samples ± SEM. Human peripheral blood mononuclear cells were grouped as follows: N, normal; O, ox-LDL; B, betulinic acid; A, apigenin; Sk, skimmianine; BASk, betulinic acid + apigenin + skimmianine. aStatistical difference with group I at
Cytokines belong to a diverse array of small protein molecules which are broadly grouped into pro- and anti-inflammatory cytokines based on whether they either promote or suppress the formation of atherosclerotic plaques. The supernatants from hPBMCs induced with ox-LDL showed a higher level of pro-inflammatory cytokines, tumor necrosis factor alpha (TNF-α) and interleukin 1 beta (IL-1β) (Figure 6(a)), and a depleted level of anti-inflammatory cytokine, IL-10 (Figure 6(c)). However, pretreatment of hPBMCs with BASk prior to ox-LDL induction decreased the release of TNF-α and IL-1β, and increased the production of IL-10 in the medium. Hence, it was confirmed that BASk attained this regulating potential of balancing the release of both pro- and anti-inflammatory cytokines since it inhibited the expression of pro-inflammatory (Figure 6(b)) and enhanced the expression of anti-inflammatory genes (Figure 6(d)).

Inhibitory effect of BASk on pro- and anti-inflammatory cytokines. (a) TNF-α, IL-1β and (b) IL-10; mRNA expression of (c) proinflammatory cytokines TNF-α, IL-1β and (d) anti-inflammatory cytokine IL-10 in ox-LDL induced human peripheral blood mononuclear cells. Cell lysates were used for the isolation of total RNA and the mRNA expression of pro-inflammatory cytokines (TNF-α and IL-1β) and anti-inflammatory cytokine (IL-10), was determined by reverse transcriptase polymerase chain reaction. Glyceraldehyde-3-phosphate dehydrogenase was used as control. Inhibiting potential of BASk on inflammatory cytokines was estimated by measuring the protein levels of TNF-α , IL-1β, and IL-10 by enzyme linked immunosorbent assay. Values are expressed as average of 6 samples ± SEM. Human peripheral blood mononuclear cells were grouped as follows: N, normal; O, ox-LDL; B, betulinic acid; A, apigenin; Sk, skimmianine; BASk, betulinic acid + apigenin + skimmianine. aStatistical difference with group I at
Bioactive compounds play a major role as lead compounds during drug design and development. Computer-based molecular modeling studies play a major role as a preliminary step for the selection of suitable drug candidates during drug development strategies since they are extremely fast and cost efficient. 9 Several reports demonstrated the drug likeness and bioactivity score of various ligand molecules via Lipinski’s RO5 and Molinspiration cheminformatics. 10,11 In the present study, the results obtained from RO5 and Molinspiration software proved that B, A, and Sk exhibited a better drug likeness and bioactivity score without violating RO5 and fulfilled the pharmacological properties essential for an orally active drug candidate. Since these 3 compounds exhibit drug likeness and bioactivity scores, they can be used as suitable drug candidates.
The essential role has been previously reported of hydrophobic amino acid residues present in the active site of the MD2 receptor to interact with TLR4 in order to undergo activation by endotoxins.
12
-14
Here, molecular docking studies revealed that B and Sk exert a strong binding interaction with the amino acid residue Leu 87 via hydrogen and hydrophobic bonds, and A through a salt bridge with Lys 128. Thus, 3 ligand molecules showed hydrophobic interaction with the receptor MD2 by sharing common amino acid intermediates (Met 85 and Val 82). Thus, on the basis of common interacting residues on the active site of the receptor MD2 from earlier reports, it has been stated that 3 ligand molecules play an antagonistic role toward MD2 from its dimerization with TLR4. MD2 is the major adapter protein involved in the activation of TLR4 because it is the only protein that complexes with TLR4, resulting in its activation.
TLR4 activation during ox-LDL induction creates a feedback loop by secreting inflammatory mediators in order to enhance inflammatory responses. An antagonist that can bind specifically to the active site of TLR4 is essential to break the feedback loop to reduce the severity of inflammation. 15 Here, BASk exerted this mechanism as an antagonist and blocked the TLR4 signaling cascade which triggers inflammation. Thus, BASk plays a major role as an inhibitor by preventing TLR4 signaling by significantly lowering the elevated level of TLR4 in ox-LDL induced hPBMCs. This result proves that the reduction of MD2 is actually responsible for blocking the TLR4 signaling cascade by preventing its dimerization with TLR4.
NFκB resides in the cytoplasm by interacting with the nuclear factor of kappa light polypeptide gene enhancer in B cells, inhibitor (IκB) protein complex and is inactive under normal conditions. 16 Phosphorylation and degradation of IκB is essential for the activation and translocation of NFκB toward the nucleus to elicit an immune response. 17,18 TLR4 has the ability to trigger the IκB phosphorylation essential for nuclear translocation of NFκB during an antigenic stimulus. 19 Here, BASk inhibited TLR4-mediated activation of NFκB from its nuclear translocation by preventing the rapid phosphorylation and degradation of IκB during ox-LDL induction and thereby reducing an inflammatory response. The result revealed that BASk exerted its anti-inflammatory effect much more than the individual compounds due to its synergistic action mediated by 3 bioactive compounds—B, A, and Sk. Finally, BASk attained a curative role against inflammation by exerting a synergistic action on ox-LDL induced hPBMCs and the present data support the studies that highlight the synergistic effects of bioactive compounds as therapeutic agents. 20 -24
Lipid mediators act as pro-inflammatory agents which play a major role in the activation of the NFκB signaling pathway. 25 Overexpression of COX-2 in activated inflammatory cells has been related to the time dependent and localized production of various pro-inflammatory mediators (PGE2) and pro-inflammatory cytokines (TNF-α and IL-1β) during chronic 26 -28 inflammatory conditions in response to an antigenic stimulus. Pro-inflammatory cytokines can make a disease itself or its symptoms worse by causing fever, inflammation, tissue destruction, and even shock and death, 29 whereas the anti-inflammatory cytokine, IL-10, inhibits the progression of inflammation and exerts a protective effect on various cell types. 30,31 Since BASk blocked the activation of the transcription factor, NFκB, by inhibiting the TLR4 signaling cascade antagonistically, COX-2 expression, and the release of pro-inflammatory mediators (TNF-α, IL-1β, and PGE2) in the medium reduced during ox-LDL induction. Similarly, BASk at the same time elevated the production of the anti-inflammatory cytokine, IL-10. Thus, BASk exerted its anti-inflammatory effect by balancing the release of pro- and anti-inflammatory cytokines, which play a crucial role in sound health.
The synergistic potential of BASk (betulinic acid + apigenin + skimmianine) in the ratio of 1:1:1 fulfilled the drug likeness property and stable binding interaction with the adapter protein MD2 by preventing its dimerization with TLR4. It is due to the inhibitory effect of BASk on MD2’s further activation of the TLR4 signaling pathway that reduces the progression of inflammation. Similarly, BASk inhibited the nuclear translocation of NFκB and in turn reduced the expression of COX-2. These inhibitions exerted by BASk reduced the inflammatory responses during ox-LDL induction by depleting the release of pro-inflammatory mediators and elevating the production of anti-inflammatory cytokine. This indicates the protective role of BASk in controlling the progression of disease. Thus, the present study concludes that BASk exhibits therapeutic efficacy much higher than the compounds when used individually. The therapeutic agents used commonly to treat inflammatory disease specifically block COX-2 activation from releasing pro-inflammatory mediators that trigger inflammatory responses. Here, BASk not only blocked COX-2 expression from releasing its pro-inflammatory mediators but also inhibited the TLR4-NFκB signaling cascade specifically. Hence, BASk can be preferred as a suitable drug candidate against inflammation.
Experimental
Lipinski’s RO5 32
According to this rule, the drug likeness of a compound can be predicted 33 on the basis of properties such as (i) LogP, octanol-water partition coefficient ≤5, (ii) MW ≤ 500, (iii) number of hydrogen bond donors ≤5, and (iv) number of hydrogen bond acceptors ≤10, with not more than 1 violation.
Molinspiration
This is a molecular processing and property calculation toolkit to compare the structures of representative ligands active on a particular target. 34 The molecular structures of corresponding ligand molecules generated using simplified molecular input line entry system (SMILE) notation (Table 4) were used for calculating the drug likeness and molecular properties such as molecular polar surface area (<160 Å), molecular volume, nrotb (<10), 35 and number of atoms (<30). Bioactivity scores were also calculated to find the biological activity of ligands toward the GPCR ligands, ion channel modulators, kinase inhibitors, nuclear receptors, protease inhibitors, and enzyme inhibitors. Thus, both drug likeness and bioactivity scores of the respective ligands were checked using the online software Molinspiration (www.molinspiration.com) on the basis of the following criteria. If the estimated bioactive score is >0, the ligand can be considered as an active compound and said to be inactive if the score is <−5.0; if the score value range is between −5.0 and 0.0, the ligand is said to be moderately active.
SMILE Notations of Betulinic Acid, Apigenin, and Skimmianine.

SMILE, simplified molecular input line entry system.
Protein Preparation for Molecular Docking
Human lymphocyte antigen 96 or MD2 having PDB ID: 2E56 with an X-ray diffraction resolution of 2 Å was selected as the target protein or receptor for docking studies.
36
The protein contains a single chain (A) having 144 amino acids (GLU17-ASN 160) and the lipopolysaccharide binding region is between 119 and 123 amino acids. The ligands present in them are 2 molecules of
Ligand Preparation and Molecular Docking of Bioactive Compounds
The 3D conformations of B, A, and Sk were downloaded from Pubchem (http://pubchem.ncbi.nlm.nih.gov). Each ligand was prepared and the energy minimized to obtain its correct conformation with least energy. The selected ligand and target protein molecule were subjected to molecular docking using CDOCKER in Biovia Discovery Studio 4.0.
Isolation of hPBMCs
Human blood was collected from healthy volunteers and the isolation of hPBMCs was carried out according to a published method. 37 The isolated hPBMCs were seeded onto Type I collagen coated plates. The cells were cultured in Roswell Park Memorial Institute Medium-1640 medium supplemented with 10% heat-inactivated autologous serum, 100 U/mL of penicillin/streptomycin solution overnight in an incubator supplied with 5% CO2 at 37°C at a cell density of 5 × 106 cells/mL.
In Vitro Study
All in vitro experiments were conducted on blood samples (12 mL) from healthy human donors taken with their consent and in agreement with institutional guidelines.
Oxidation of LDL
Isolation of LDL from human serum was carried out according to the method described by Nguyen-Khoa et al. 38 The isolated LDL was subjected to oxidation by incubating it at 37°C for 24 hours with 10 µM CuSO4 and then dialyzed overnight in phosphate buffered saline (PBS) containing 0.5 mM ethylene diamine tetraacetate (EDTA). 39 The degree of oxidation was measured on the basis of the amount of thiobarbituric acid reactive substances produced.
Experimental Design
All in vitro experiments were conducted with hPBMCs. The isolated hPBMCs were divided into 6 groups: Group I—normal, Group II—ox-LDL (50 µg/mL), Group III—ox-LDL + B (25 µM), Group IV—ox-LDL + A (25 µM), Group V—ox-LDL + Sk (25 µM), and Group VI—ox-LDL + BASk (25 µM).
An effective concentration of 25 µM was selected for conducting experiments according to our previous report. 11 BASk, which is a combination of B, A, and Sk, was prepared in the ratio of 1:1:1 (BASk) at a final concentration of 25 µM. Finally, the combination and synergistic effects of BASk were compared with each of its individual compounds. After 24 hours incubation, both the cell culture supernatants and cell lysates were collected and used for the determination of the role of various adapter proteins involved in the TLR4-NFκB signaling pathway by enzyme-linked immunosorbent assay (ELISA); the result was confirmed with reverse transcriptase polymerase chain reaction (RT-PCR).
Enzyme Linked Immunosorbent Assay
Isolated hPBMCs were pretreated with B, A, Sk, and BASk at a concentration of 25 µM, then induced with ox-LDL (50 µg/mL) and incubated for 24 hours. Then, the protein levels of PGE2 and cytokines (IL-1β, TNF-α, and IL-10) in the cell culture medium and the adapter proteins (TLR4 and MD2) in cell lysates were measured by ELISA. The colored end product formed due to the formation of enzyme substrate complex was read spectrophotometrically at 450 nm.
Determination of NFκB
The activation of transcription factor NFκB was determined according to a previous method.
40
Pretreated hPBMCs prior to ox-LDL induction were scraped into ice cold PBS after 24 hours incubation and centrifuged at 500
RNA Preparation and RT-PCR
Total cellular RNA was isolated from hPBMCs (5-10 × 106 cells) using trizol (TRI) reagent (Sigma Aldrich) and quantified at 260 nm. Reverse transcriptase PCR was performed with the reverse transcription system (Eppendorf Mastercycler), using a 2-step RT-PCR kit where cDNA synthesis and PCR reactions were conducted separately according to the manufacturer’s protocol. Total RNA (2 µg) was used as the template in the first step, which includes dNTPs, RNase inhibitor, and reverse transcriptase enzyme and the cDNA obtained was used for PCR amplification. The second step included amplification of cDNA using appropriate primers (Table 5). The PCR products were electrophoresed on 1.5% agarose gel and the bands obtained were visualized by staining with ethidium bromide. The image of the gel was captured in digital format and the intensities of corresponding bands were measured; glyceraldehyde-3-phosphate dehydrogenase served as control.
Details of Primer Sequences of Specific Genes.

GAPDH, glyceraldehyde-3-phosphate dehydrogenase.
Estimation of Protein
Protein was estimated by the method of Lowry et al. 41
Statistical Analysis
The results were examined using statistical software SPSS/PC+, version 11.0 (SPSS Inc., Chicago, IL, United States). One-way analysis of variance and pair-fed comparisons between the groups were made by Duncan’s multiple range tests and
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Kerala State Council for Science, Technology and Environment (KSCSTE) (019/SRSLS/2008/KSCSTE), Science Research Scheme, Thiruvananthapuram.
