Abstract
Shan-Dou-Gen (山豆根) is a crude drug mainly derived from the roots of Leguminosae plants, and it has antipyretic, antidotal, anti-inflammatory, and analgesic effects. In Japan, the root of Euchresta japonica has been used as a material of Shan-Dou-Gen. However, E. japonica is not used for medicinal purposes today, and commercial Shan-Dou-Gen products are imported from China. In China, several plant species have been used as Shan-Dou-Gen materials, but a crude drug derived from the root of Sophora tonkinensis is now officially used as Shan-Dou-Gen. However, it is difficult to morphologically identify the species used in Shan-Dou-Gen. In the present study, we showed that the Shan-Dou-Gen products commercially available in Japan are derived from S. tonkinensis using phylogenetic and sequencing analyses of the chloroplast trnH–psbA region. Furthermore, we performed species-specific polymerase chain reaction using conserved sequences of S. tonkinensis. Amplification was observed with Shan-Dou-Gen, whereas no amplification occurred with other crude drugs derived from the roots of S. flavescens and S. japonica. These results indicated that the genetic approach can be useful to authenticate Shan-Dou-Gen.
Keywords
Shan-Dou-Gen (山豆根) is a crude drug mainly derived from the roots of Leguminosae plants, and it has antipyretic, antidotal, anti-inflammatory, and analgesic effects. It was first listed in the book of Kai-baa-ben-cao (開宝本草) in the 10th century. Several Leguminosae plant species have been used as materials of Shan-Dou-Gen in China and Japan. 1,2 A survey by the Japan Kampo Medicines Manufacturer’s Association indicated that approximately 300 kg of the drug is produced in Japan annually. Shan-Dou-Gen is listed in the Japanese Standards for nonpharmacopoeial crude drugs, 2018, in which the origin of the crude drug is defined as Sophora subprostrata Chun et T. Chen. According to the Flora of China, S. subprostrata is a synonym for Sophora tonkinensis. 3,4 In the Edo era of Japan (AD 1603-1867), Ranzan Ono stated that Shan-Dou-Gen was derived from the root of Euchresta japonica. Although E. japonica grows naturally in the southwestern parts of Japan, the number of individuals is small. Therefore, E. japonica is not used for medicinal purposes in Japan today, and commercial Shan-Dou-Gen products are imported from China. In China, a crude drug derived from the root of S. tonkinensis, which is collected mainly from the Guangxi Province, is called Guang-Dou-Gen (広豆根), and it is now officially used as Shan-Dou-Gen. S. tonkinensis is the only species listed as the source of Shan-Dou-Gen in the Pharmacopoeia of the People's Republic of China, 2015, 10th edition. However, the plant species used as materials of Shan-Dou-Gen vary between areas in China. In northern and northeastern China, the rhizome of Menispermum dauricum, which belongs to the Menispermaceae family, has been used as a material of Shan-Dou-Gen. In Guangxi Guilin, the root of Cyclea hypoglauca, which also belongs to the Menispermaceae family, has been used. In the provinces of Hubei, Henan, Shanxi, Gansu, Shaanxi, and Jiangsu, the roots of plants from the genus Indigofera (I. amblyantha, I. fortune, and I. ichangensis), which belongs to the Leguminosae family, have been utilized. In Hunan, Guizhou, and Yunnan Provinces, the roots of plants from the genus Ardisia (A. crenata, A. hortorum, and A. mamillata) have been used. Because various species have been used as materials of Shan-Dou-Gen, it is difficult to identify them morphologically. Recently, genetic analysis based on DNA barcoding regions has been a powerful tool for identifying species in crude drugs. In the present study, we showed, using DNA analysis, that the Shan-Dou-Gen products commercially available in Japan were derived from S. tonkinensis.
To examine the phylogenetic relationship of the origin of Shan-Dou-Gen, sequences of the chloroplast trnH–psbA region were obtained from the DDBJ/EMBL/GenBank databases. The trnH–psbA region has been widely used for species identification and phylogenetic analysis. We used the sequences of S. tonkinensis (accession no. GQ434960, GU396707, KJ766122, KJ766123, and KC902513), M. dauricum (accession no. GQ434987), and A. crenata (accession no. HQ427113), which were registered in the DDBJ/EMBL/GenBank databases. Because the sequences of I. amblyantha, I. fortune, I. ichangensis, and C. hypoglauca were not registered, the sequences of I. tinctoria (accession no. GU396729) and C. polypetala (accession no. HG005018) were used instead. The sequences of 19 Sophora species were used as the in-group for the genus Sophora, and those of Glycyrrhiza uralensis were used as the out-group (accession no. GU396733). Using these sequences, phylogenetic relationship was examined (Figure 1). Five sequences of S. tonkinensis were classified into one group and other Sophora species were not included in the clustered group of S. tonkinensis. Dendrogram data showed that the trnH–psbA region was useful to discriminate S. tonkinensis from other species.

Molecular phylogenetic analysis by the neighbor-joining method. Dendrogram was constructed based on the 0.2 kb aligned nucleotide sequence of the chloroplast trnH–psbA region. Numbers at nodes indicate bootstrap values with 1,000 replicates. Branch length is proportional to the number of base substitutions per site.
Multiple alignment of the trnH–psbA regions was performed, and polymorphism sites are shown in Table 1. The differences between 5 sequences of S. tonkinensis (accession no. GQ434960, GU396707, KJ766122, KJ766123, and KC902513) were limited to the number of repeat sequences at nucleotide positions 105-117 and 159-170 (Table 2). Alignment data showed that positions 80 and 130 were sequences specific to S. tonkinensis. At position 80, there was a “T” in the S. tonkinensis sequence and a “C” in the sequences of other Sophora species. At position 130, there was a “T” in the S. tonkinensis sequence and a “G” in the sequences of other Sophora species. Therefore, S. tonkinensis can be theoretically discriminated from other species by DNA analysis. To identify the origin of Shan-Dou-Gen, 2 samples (samples A and B) obtained from different companies in Japan were analyzed. DNA was extracted from Shan-Dou-Gen and amplified by polymerase chain reaction (PCR) using universal primers of the trnH–psbA region. The results of the sequencing analysis showed that the trnH–psbA region sequence of Shan-Dou-Gen (sample A) was the same as that of S. tonkinensis without repeat sequences at positions 159-170, and that the nucleotides of Shan-Dou-Gen (sample A) at positions 80 and 130 were “T.” The trnH–psbA region of Shan-Dou-Gen (sample B), except for partial regions (positions 118-158), was sequenced successfully, and the sequence was the same as the S. tonkinensis sequence. The nucleotide at position 80 of Shan-Dou-Gen (sample B) was “T,” whereas the nucleotide at position 130 was not identified, probably because different haplotypes might have different numbers of repeat sequences at positions 105-117 and 159-170. These results showed that S. tonkinensis was the origin of the commercial Shan-Dou-Gen products available in Japan.
Alignment Data of Chloroplast trnH–psbA Regions.

Single-nucleotide polymorphism sites of genus Sophora and commercial Shan-Dou-Gen products are shown. Accession numbers of DDBJ/EMBL/GenBank Databases are shown in parentheses.
a N: unidentified.
Repeat Sequences of Sophora Tonkinensis and Shan-Dou-Gen.

We next performed species-specific PCR from crude drugs using conserved sequences of S. tonkinensis. Sophorae Radix (苦参; the root of Sophora flavescens), Huái gēn (塊根; the root of Sophora japonica), and Glycyrrhizae Radix (甘草; the root of G. uralensis) were used as negative controls. Because the trnH–psbA region of S. japonica was not registered in the DDBJ/EMBL/GenBank databases, we confirmed that the nucleotides at positions 80 and 130 of S. japonica were "C" and "G," respectively, through sequencing analysis. S. tonkinensis-specific primers were designed; the 3′-terminal nucleotide of the forward primer was position 80, and that of the reverse primer was position 130 (Figure 2a). The unique PCR products were obtained from Shan-Dou-Gen using S. tonkinensis-specific primers (Figure 2b). However, no amplification occurred in other species, including S. flavescens, which has only 2 single-nucleotide polymorphisms at positions 80 and 130 in its primer binding sites. Therefore, we concluded that the unique primers are highly specific to S. tonkinensis. DNA quality of all samples was confirmed by PCR using universal primers (Figure 2c).
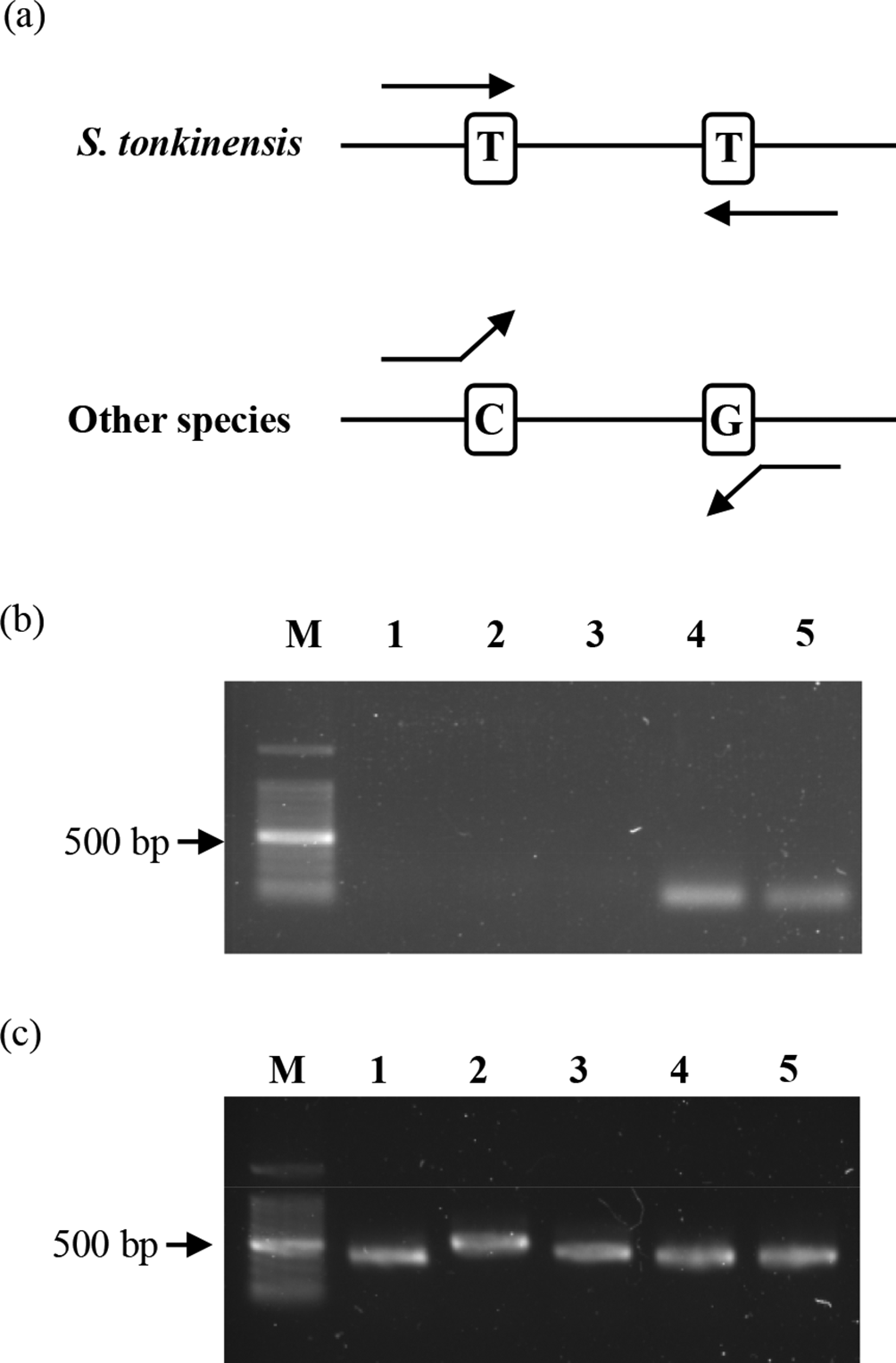
Binding sites of Sophora tonkinensis-specific primers (a). Polymerase chain reaction using S. tonkinensis-specific primers (b) and universal primers (c). 1, Sophorae Radix (苦参; the root of S. flavescens); 2, Huái gēn (塊根; the root of S. japonica); 3, Glycyrrhizae Radix (甘草; the root of G. uralensis); 4, Shan-Dou-Gen (山豆根, sample A); 5, Shan-Dou-Gen (山豆根, sample B). M: 100 bp ladder.
In conclusion, through DNA analysis, we showed S. tonkinensis as the material of the Shan-Dou-Gen products commercially available in Japan. Sequences of the trnH–psbA region were useful to distinguish S. tonkinensis from other species because species-specific sequences were observed at 2 nucleotide positions. Species-specific PCR using these sequences was successfully performed on crude drugs. These results showed that genetic analysis can be used to authenticate Shan-Dou-Gen.
Experimental
Sample collection and DNA extraction: Shan-Dou-Gen products (samples A and B) were purchased from Uchidawakanyaku, Ltd (Japan) and Tochimoto Tenkaido Co., Ltd (Japan), respectively. Sophorae Radix and Glycyrrhizae Radix were purchased from Uchidawakanyaku, Ltd. The root of S. japonica was obtained from the botanical garden of Josai University, Saitama, Japan. Total genomic DNA was extracted using NucleoSpin Plant II (Takara, Japan) and quantified using a NanoDrop 1000 instrument (ThermoFisher Scientific, Waltham, MA, USA).
PCR and sequencing: PCR targeting the trnH–psbA region was conducted at a final volume of 25 µL. The final volume contained the following components: 1× PCR buffer, 0.4 mM of each deoxyribonucleotide triphosphate, 0.5 µM each of universal forward (5′-3′, GTT ATG CAT GAA CGT AAT GCT C) and reverse (5′-3′, CGC GCA TGG TGG ATT CAC AAT CC) primers, 0.5 units of KOD FX Neo (Toyobo, Japan), and 10 ng of genomic plant DNA. 5,6 PCR products were purified and sequenced using BigDye Terminator v1.1 (Applied Biosystems, USA) on a 310 DNA genetic analyzer (Applied Biosystems) following the manufacturer’s protocol. Phylogenetic analysis based on the trnH–psbA region sequences was performed using Molecular Evolutionary Genetic Analysis version 6. 7 Molecular phylogenetic analysis was conducted using the neighbor-joining method. Evolutionary distances were computed using the Kimura 2-parameter method. 8 S. tonkinensis-specific forward (5′-3′, TAG AAA AAA CTC TAT TGC TCC TTT ACT TTA TAT) and reverse (5′-3′, CTT TTT GGA ATA CAT ATG ACT TCG A) primers were designed manually. PCR amplification was performed using the reaction mixture (containing 0.5 µM each of the forward and reverse primers, 0.5 units of KOD FX Neo, and 10 ng of genomic plant DNA) and the following conditions: a cycle of 2 minutes at 94°C; 30 cycles of 10 seconds at 98°C, 30 seconds at 63°C, 10 seconds at 68°C; and 1 cycle of 2 minutes at 68°C. PCR products were analyzed by 1.2% agarose gel electrophoresis.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) declared no financial support for the research, authorship, and/or publication of this article.
