Abstract
Polyalthic acid (PA) is a diterpene present in several trees of the
The

Structure of (−) ent-polyalthic acid.
The diterpene PA was encapsulated in poly lactic-co-glycolic acid (PLGA) nanoparticles
using the nanoprecipitation method. Scanning electron microscopy (SEM) images showed
particles with semispherical morphology (Figure 2), some with sizes between 100 and 200 nm probably due
to the aggregation during sample processing.
5
The average size of particles in suspension
obtained using dynamic light scattering (DLS) was 98.64 ± 28 nm (Figure 3), with a polydispersity
index of 0.057. The surface charge of the particles was negative (

SEM images of the PA-containing nanoparticles. PA, polyalthic acid; SEM, scanning electron microscopy.
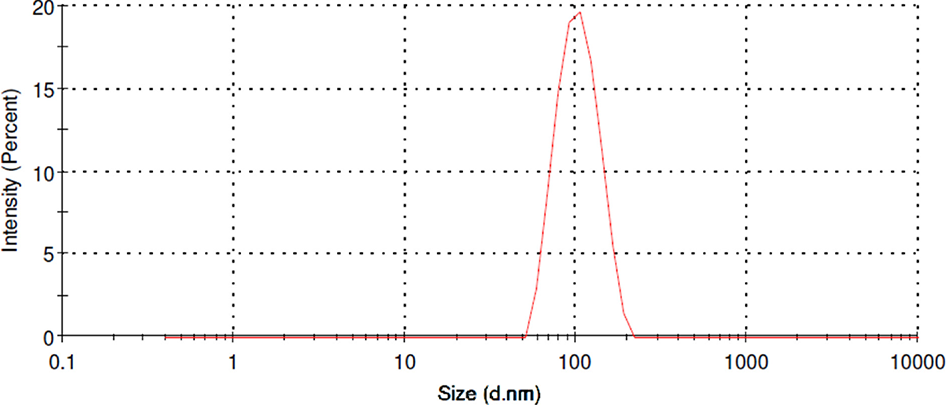
Representative size distribution of PLGA nanoparticles containing polyalthic
acid. PLGA, poly lactic-co-glycolic
acid.
A decrease in the release rate was observed until it reached an approximately constant value from 6 to 24 hours, with almost 64% of the PA released at the end of the experiment (Figure 4). The initial burst is probably due to the release of molecules adsorbed on nanoparticle surfaces, followed by a more slow diffusion from the polymer matrix. This release kinetic is consistent with the properties of PLGA 50:50 (Mw 24 000–38 000) used in this work, which is hydrolyzed faster than other polymers of the same family with higher proportions of lactic or glycolic acid. 6 Calculated IC50 values for the treatments showed that the nanoparticles containing PA have higher cytotoxicity for both cell lines (Table 1).

In vitro release kinetics of PLGA nanoparticles containing polyalthic acid
studied during 24 hours. PLGA, poly
lactic-co-glycolic acid.
IC50 Values and SI for the Different Treatments.
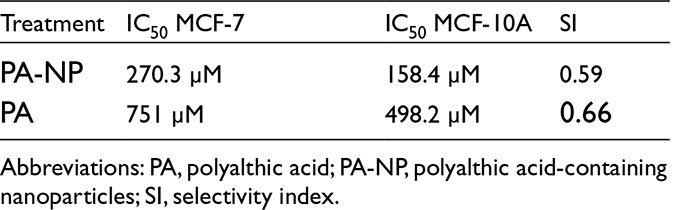
Abbreviations: PA, polyalthic acid; PA-NP, polyalthic acid-containing nanoparticles; SI, selectivity index.
The effects on MCF-7 cell proliferation were significant from 24.65 μM for PA incorporated in nanoparticles and from 197.52 for free PA. Although treatments also showed antiproliferative capacity in the MCF-10A lineage, the significant effects began to be evident at higher concentrations (98.76 μM for PA incorporated in nanoparticles and 395.03 μM for PA) than those required for MCF-7 cells. The described facts evidenced a selective inhibition of cell proliferation for the formulation of PA in nanoparticles on the MCF-7 tumor lineage. To compare the effect of the 2 treatments on each cell line, we performed a 2-way analysis of variance (ANOVA) and the Bonferroni post hoc test. For MCF-7 cells, when comparing the effects of PA incorporated in nanoparticles with those of the free molecule, significant differences were found in the range of concentrations from 24.65 to 395.03 μM, evidencing a higher antiproliferative activity for the nanoparticle formulation. No significant effects were found between the 2 treatments on MCF-10A cells in the range of concentrations assessed (Figure 5). As seen in Figure 6, treatments with PA incorporated in nanoparticles damaged selectively DNA of MCF-7 cells in a concentration-dependent manner, with significant damage scores for the 3 assayed concentrations, when compared to negative control. In MCF-10A cells, the treatment with PA incorporated in nanoparticles did not induce significant damage at any concentration. Free PA was tested as a control in the highest concentration assayed for nanoparticles and showed selective damage in MCF-7 cells but in less extension when compared with the last 2 concentrations of PA incorporated in nanoparticles. These results show that PA has an intrinsic capacity to induce selective DNA damage in MCF-7 cells, which is enhanced by its incorporation in PLGA nanoparticles. This study shows that the incorporation of PA in polymeric nanoparticles enhances the cytotoxicity of this molecule and its antiproliferative and genotoxic effects on MCF-7 cells. Analysis of cell viability revealed a decrease in the IC50 for treatment with PA incorporated in nanoparticles when compared to the free molecule. This effect has been reported for antitumor drugs incorporated in nanoparticles. Lozano et al 7 reported a decrease in the IC50 of docetaxel in MCF-7 and A549 cells when this molecule was incorporated in polymeric nanocapsules. In another study, methotrexate showed increased cytotoxicity when encapsulated in PLGA–β-cyclodextrin nanoparticles. 8 Characteristics such as the size, surface charge, and surfactant layer of the nanoparticles are factors that enhance the interaction with cells. It has been demonstrated that sizes below 300 nm and a negative surface charge increase the interaction with the cell surface mainly due to the high surface area caused by the small size and the interaction with the phospholipids of the outer surface of the cell membrane, mainly phosphatidylcholine, which has a positive charge. 9 Based on these facts, the higher cytotoxicity of PA incorporated in nanoparticles observed in this work could be explained by the small particle size and the negative surface charge, which could increase the interaction of the nanoparticles–PA system with the cells and enhance its cytotoxic effects. However, despite this increased cytotoxicity, treatments with these formulations did not appear to be selective on the tumor cells. This fact agrees with previous in vitro studies, where molecules such as docetaxel were not selective when incorporated into polymeric nanocapsules and tested in vitro. 7 In regard to this fact, it has been reported that the selectivity of nonfunctionalized nanoparticles is observed mainly in vivo, due to the enhanced permeability and retention (EPR) effect. This effect occurs with particles with sizes below 300 nm and is due to the high vascularization and low lymphatic drainage of solid tumors, in addition to the abnormal architecture of the vascular endothelium in tumor blood vessels, which causes selective accumulation of nanoparticles when compared to the normal tissues. 10 Thus, it is very likely that the nanoparticles obtained in this work can exhibit the aforementioned effect if tested in vivo, due to their small size. Because the 2,3-bis-(2-methoxy-4-nitro-5-sulfophenyl)-2H-tetrazolium-5-carboxanilide (XTT) test determines cell viability through the activity of cellular dehydrogenases, a cell can metabolize tetrazolium despite being under cell cycle arrest and unable to divide. The ability of a compound to inhibit cell proliferation can be tested using more specific techniques such as the 5-bromo-2′-deoxyuridine (BrdU) incorporation assay. This assay determines the rate of cell proliferation using BrdU, a synthetic thymidine analog. Using this method, the effect of the different treatments was evaluated. PA, either encapsulated in nanoparticles or free, showed a selective antiproliferative capacity on MCF-7 cells. This effect was more significant in the case of PA-incorporated nanoparticles, as was evidenced using 2-way ANOVA. These facts support that PA has an intrinsic and selective antiproliferative effect, which is evidenced at subcytotoxic concentrations and can be enhanced when incorporated into PLGA nanoparticles, probably due to the same factors discussed earlier for cell viability. These results are evidence of cytostatic rather than cytotoxic action for PA incorporated in PLGA nanoparticles. To investigate if the selective antiproliferative effect was due to DNA damage, we performed the alkaline version of comet assay, which can detect single- and double-strand breaks. 11 As is shown in our results, treatments with PA–nanoparticles system caused selective DNA damage on MCF-7 cells in a dose-dependent manner even at subcytotoxic concentrations, whereas no damage was found in MCF-10A cells.

Cell proliferation for MCF-7 and MCF-10A cell lines treated with PA-NP and free
PA compared using 2-way ANOVA and the Bonferroni post test. ANOVA, analysis of variance; NP, polyalthic
acid in nanoparticles; PA-NP, polyalthic acid-containing
nanoparticles.

Score of DNA damage determined by comet assay for treatments with PA incorporated
in nanoparticles on MCF-7 and MCF-10A cells. The results evidenced selective DNA
damage on the tumor line. NCs were not treated. DMSO, dimethyl sulfoxide; NC, negative control; NP, empty
nanoparticles; PA, polyalthic acid; PC, positive control.
Previous studies showed that MCF-7 cells have a tendency to accumulate more DNA lesions than MCF-10A cells after genotoxic treatments. Using comet assay, Ortiz et al 12 showed that sandalwood essential oil has selective genotoxic effects in MCF-7 cells compared with noncancerous MCF-10A cells when assayed at noncytotoxic concentrations. In another study, Francisco et al 13 detected increased DNA damage on MCF-7 cells when compared with MCF-10A cells, using comet assay immediately after low doses of γ-radiation. These authors also found that the total antioxidant capacity in MCF-7 cells was lower than in their nontumor counterparts, as well as the DNA repair capacity.
These studies provide evidence on the decreased susceptibility of MCF-10A cells to DNA damage. Based on these results, we can explain the selective genotoxicity on MCF-7 cells assuming that PA have the capacity to induce DNA lesions, which is enhanced when encapsulated in polymeric nanoparticles. The increased tendency to accumulate DNA lesions and the deficiency in DNA repair mechanisms observed naturally in MCF-7 cells may probably be the cause of the observed selectivity. This selective genotoxic effect is consistent with the selective inhibition of cell proliferation observed in the BrdU assay results. Consequently, it can be assumed that the selective DNA damage induced by PA was the cause of the decreased proliferation in MCF-7 cells when compared with MCF-10A cells. This work shows that PA has selective antiproliferative and genotoxic effects on MCF-7 cells, which are enhanced by its incorporation in PLGA nanoparticles. The development of a formulation of PA in nanoparticles, which not only preserve but also improve the biological activity of this molecule, opens a way to the possible elimination of toxic solvents in future studies. In summary, PA incorporated in polymeric nanoparticles is a promising selective cytostatic antitumor agent with possibilities of being used in breast cancer therapy.
Experimental
PA and Nanoparticles
PA was donated graciously by the Natural Products Research Group of the
University of Franca (UNIFRAN). Nanoparticles were obtained by the
nanoprecipitation method.
14
Briefly, 50 mg of 50:50, acid-terminated
poly(
To determine the drug loading by mass (DL), the amount of PA in the nanoparticles
was quantified indirectly and the weight of the nanoparticles was measured after
freeze-drying. DL was calculated as follows:
In all cases, PA was quantified by HPLC, using a C18 column with isocratic elution conditions and ACN-water (65:35, v/v) as mobile phase, with a flow rate of 1.0 mL/min and run time of 15 minutes. The wavelength of the detector was 201 nm. A standard curve made with PA standard solutions was used for the quantification. All the chromatographic conditions were validated before the study. For release studies, 2 mL of the nanoparticle suspension was placed into a dialysis sack (MWCO 12 kDa, Sigma) and immersed into a beaker with 25 mL of 2% Tween 80 aqueous solution under magnetic stirring. Aliquots of 1 mL were taken at different time intervals, with reposition of the dialysis solution.
Cell Lines and Culture Conditions
MCF-7 (human breast tumor) and MCF-10A (human breast normal) cell lines were obtained from the Rio de Janeiro Cell Bank. Dulbecco’s modified Eagle’s medium (DMEM) + HAM F10 (1:1, v/v, Sigma-Aldrich) supplemented with 10% fetal bovine serum (Sigma-Aldrich), 1% stabilized penicillin/streptomycin solution (Sigma-Aldrich), and kanamycin-stabilized solution (Sigma-Aldrich) was used as culture medium. Cells were cultured in T25 flasks (TPP, Switzerland) and incubated at 37°C and 5% CO2. In all cases, cells were used between the third and eighth passage.
Evaluation of Cell Viability by the XTT Assay
Cell viability was assessed using the XTT assay. For this experiment, 104 cells per well were seeded in 96-well plates. After 24 hours of incubation, the cells were treated with PA either incorporated in nanoparticles or dissolved in dimethyl sulfoxide (DMSO), at concentrations in the range of 24.65 to 3160.26 μM. DMSO (10%) was used as a positive control, whereas PLGA nanoparticles without PA and 1% DMSO were used as vehicle control. Untreated cells were used as negative controls. After 24 hours of treatment, the culture medium was removed, cells were washed with phosphate-buffered saline (1× PBS) and incubated with DMEM without phenol red, mixed with the XTT/electron coupling reagent solution (50:1, v/v, Roche Applied Science). After incubation for 4 hours, the absorbance of each well was measured with a spectrophotometer (Tecan A-5082, Salzburg, Austria) using a wavelength of 492 nm and 690 nm as reference. The experiments were performed in triplicate and the percentage of viable cells was expressed in relation to the negative control, considered as 100% cell viability. For the cell proliferation assay, the BrdU ELISA kit was purchased from Roche Applied Science. Treatments were similar to those applied in the XTT assay, except for the positive control, which was doxorubicin 0.5 μM. After 24 hours of treatment, the BrdU solution was added to each well at a final dilution of 1:100 and incubated at 37°C for 90 minutes. Cells were then fixed with the fixative solution from the kit according to recommendations provided by the manufacturer and the DNA was denatured for 30 minutes using the appropriate solution from the kit. This solution was removed and the plates were incubated 30 minutes with anti-BrdU-peroxidase (anti-BrdU-POD) Fab fragments in solution (1:100 dilution) for 1 hour. Finally, each well was washed 3 times with PBS and the plates were incubated for 30 minutes with peroxidase substrate. The absorbance was measured with a spectrophotometer (Tecan A-5082, Salzburg, Austria) using a wavelength of 370 and 492 nm as reference. Experiments were carried out in triplicate and the percentage of proliferation was expressed in relation to the negative control considered as 100% of cell proliferation.
Assessment of Genotoxicity Through Comet Assay
To perform this assay, 105 cells per well were seeded in 6-well plates and incubated for 24 hours. After incubation, PA, either in PLGA nanoparticles or dissolved in DMSO, was added at 98.8, 197.5, and 395 μM. Doxorubicin 0.5 μM was used as positive control, whereas nanoparticles without PA and 1% DMSO were used as vehicle controls. No treatment was applied in the case of the negative control. Immediately after treatment, the cells were incubated for another 4 hours. Cells were harvested after incubation and 20 μL of cell suspension was homogenized with low-melting-point agarose (0.75%, m/v) and placed onto a microscope slide precoated with standard agarose (1.5%, m/v). The slides were then covered with a coverslip and maintained at −20°C for 5 minutes. After that, the coverslip was removed and the slides were submerged overnight at 4°C in an alkaline lysis solution (pH 10). Slides were then washed with phosphate buffer and incubated in alkaline buffer solution (NaOH 0.3 M and Na2EDTA 1 mM, pH > 13) for 20 minutes. Electrophoresis was performed in the same alkaline buffer at 25 V and 300 mA for 25 minutes. After electrophoresis, the slides were neutralized in a buffer (0.4 M Tris-HCl, pH 7.5) for 15 minutes and fixed 10 minutes in a solution containing trichloroacetic acid 15%, zinc sulfate 5%, and glycerol 5%. For visualization, slides were stained for 15 minutes with a mixture of AgNO3 0.1%, NH4NO3 0.1%, tungstosilicic acid 0.25%, and formaldehyde 0.15%. The stained slides were placed in acetic acid 1%, dried, and visualized at 40× magnification in a light transmission microscope (Nikon, Japan). For each treatment, 2 slides were prepared and 100 comets were scored according to the level of DNA migration, with the absence of DNA migration scored as 0 and maximum DNA migration scored as 4. The score limits for 100 comets were 0 (100% undamaged cells) and 400 (100% damaged cells). 11 Each treatment was prepared in triplicate.
Statistical Analysis
In viability, proliferation, and comet assays, the means of treatments were compared using 1-way ANOVA coupled to Dunnett’s and Tukey’s post-tests, with 0.05 as level of significance. Two-way ANOVA and the Bonferroni’s post-test were applied to compare the antiproliferative activity of PA incorporated in nanoparticles and free PA on each cell line. All statistical tests and the IC50 calculations were performed using the software GraphPad Prism version 5.0.
Footnotes
Acknowledgments
The authors appreciate the funding institutions and also thank Dr Priscila Marcato (University of São Paulo) for the support in DLS measures.
Authors’ Note
Leniher Castan Chibas and Priscila Pavini Cintra contributed equally to this work.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES, PhD scholarship number 12467-13-8) and Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP Grants number 2011/13630-7 and 2014/12465-0) supported the study financially.
