Abstract
Background:
A core challenge in managing diabetes is predicting glycemic responses to meals. Prior work identified significant interindividual variation in responses and developed personalized forecasts. However, intraindividual variation is still not well understood, and the most accurate approaches require invasive microbiome data. We aimed to investigate (1) whether postprandial glycemic responses (PPGRs) can be predicted with limited data and (2) sources of intraindividual variation.
Methods:
We used data collected from 397 people with Type 1 Diabetes (T1DEXI) and 100 people with Type 2 Diabetes (ShanghaiT2DM) who wore continuous glucose monitors (CGMs) and logged meals. Using dietary, demographic, and temporal features, we predicted 2 hours PPGR, and peak 2 hours postprandial glucose rise (Glumax). We evaluated the contribution of food features (eg, macronutrients, food category) and use of personal training data and investigated intraindividual variability in responses.
Results:
We achieved comparable accuracy to prior work for PPGR (T1DEXI R = 0.61, ShanghaiT2DM R = 0.72) and Glumax (T1DEXI R = 0.64, ShanghaiT2DM R = 0.73), without using invasive data like microbiome. Including food category features led to higher accuracy than macronutrients alone. Analysis of glycemic responses to duplicate meals identified time of day (PPGR: T1DEXI P < .05 for lunch, ShanghaiT2DM P < .001 for lunch and dinner) and menstrual cycle (Glumax: P < .05 for perimenstrual) as sources of variability.
Conclusions:
We demonstrate that in individuals with T1D and T2D, glycemic responses to meals can be predicted without personalized training data or invasive physiological data.
Keywords
Introduction
Managing blood glucose (BG) around meals is critical for individuals with Type 1 and Type 2 diabetes (T1D, T2D), and there is growing understanding of the risks of elevated postprandial BG in the general population. 1 Dietary plans like DASH (Dietary Approaches to Stop Hypertension) have been promoted for managing diabetes, 2 but there is a need for personalized advice. 3 Zeevi et al 4 demonstrated that adults without diabetes can differ significantly in glycemic responses to the same foods, and that a personalized diet can lead to healthier BG patterns. Randomized controlled trials (RCTs) of personalized dietary interventions showed more improved HbA1c compared to general diet plans.5-8
Thus, many studies aimed to predict individuals’ glycemic responses to meals. Table 1 summarizes the state of the art for predicting postprandial glycemic response (PPGR), measured with the incremental area under the curve (iAUC) in the 2 hours following a meal; and postprandial glucose rise (Glumax), the difference between glucose at meal onset and maximum glucose in the following 2 hours. Evaluations used Pearson’s correlation coefficient (R) between predicted and measured response, and coefficient of determination (R2), indicating the proportion of variance explained by the model. The most accurate works used personal meal training data. For example, Tily et al 9 obtained an R of 0.80 for PPGR in individuals without diabetes, but this dropped to 0.64 without personal meal data in the training set. Another model trained on an Israeli population had lower accuracy when applied in a Midwestern American population, and improved with more data from the target population. 10 Even with personalized training data, predictions for individuals with T1D were less accurate than predictions for the general population, as shown in Table 1. The most accurate models used microbiome data,9,11 which requires stool samples and is not widely available.
Summary of Previous Studies on Predicting Glycemic Responses to Meals.

Source: Bold indicates best value of R for an outcome.
Macronutrient, micronutrient, caffeine (mg), and carb-to-fat ratio.
Macronutrient, micronutrient, caffeine (mg), carb-to-fat ratio, caloric sums, carb sums, and fiber sums.
Macronutrient, micronutrient, glycemic index, glycemic load, organic acids (g), and ash (g).
Macronutrient, micronutrient, meal total weight, carb-to-fat ratio, caffeine (mg), and cholesterol (mg).
While nutrition drives changes in glucose, models using only carbohydrate intake had the lowest accuracy. However, dietary intake has been represented using macronutrients and their ratios rather than food categories or other nutrients, which could better predict responses. Furthermore, while prior work highlighted interindividual variation,4,11 Hengist et al 16 identified significant intraindividual variation, with responses to the same meal a week apart being only moderately correlated (R = 0.43 and 0.47 depending on continuous glucose monitor [CGM]). Thus, there remain two key limitations in personalized nutrition: (1) PPGR prediction relies on invasive and expensive data and (2) reasons behind intraindividual variation are not yet understood.
To address this, we developed a model to predict glycemic responses (PPGR, Glumax) using limited data and investigated whether features such as time of day and menstrual cycle phase were related to intraindividual variation in responses. Our model relied on dietary (food type, macronutrients) and temporal (eg, meal time, time since last meal) features. We evaluated our approach on two data sets from individuals with T1D 17 and T2D. 18
Methods
Data Sets
We used T1DEXI, which included 497 adults with T1D (mean age 37 ± 14 years; 363 female, 134 male), 17 and ShanghaiT2DM, 18 which included 100 adults with T2D (mean age 60.17 ± 13.71 years; 44 female, 56 male). The variables in each data set are summarized in Table 2.
Data Available in T1DEXI and ShanghaiT2DM Data Sets.

Data Processing
First, we excluded meals if there was a CGM gap of ≧30 minutes during the 30 minutes before and 120 minutes after the meal start. Gaps under 30 minutes were filled using linear interpolation. 19 Base CGM was defined as the median CGM value during the 30 minutes before a meal as in prior work. 4
As ShanghaiT2DM included only food names, we mapped these to standardized food codes and nutrients. We verified the English translations, then unified food names (eg, 红萝卜, 胡萝卜, and 胡卜 all refer to carrot and were mapped to 胡萝卜). To obtain nutrition information, we used the FatSecret API 20 which provides macronutrients for Chinese foods. For complex dishes not in FatSecret (eg, pork noodle soup), we identified each component and combined macronutrients. Foods were matched to codes in the Food and Nutrient Database for Dietary Studies (FNDDS), 21 which provides nutrient values for over 5000 foods. The FNDDS has a hierarchical structure, where, for example, carrots (73100000) are a subcategory of orange vegetables (73000000). Foods were matched to codes using the Python fuzzywuzzy package. For complex dishes without an exact match, we coded each component (eg, two codes for “tomato and eggs”), and estimated the proportion of each component using ChatGPT, leveraging its knowledge of common recipes and typical ingredient proportions. For example, ChatGPT suggested a 1:1 ratio for tomatoes and eggs in “tomato and eggs” (See Supplemental Appendix A1 for details). Since ShanghaiT2DM participants did not categorize their eating occasions as in the T1DEXI data set, we determined time windows for breakfast, lunch, and dinner using the trend of average CGM values (see Supplemental Appendix A2 for details). Figure 1 shows the distribution of the nine top-level categories of FNDDS by time of day for both data sets, and Supplemental Appendix A3 shows the distribution of macronutrients.
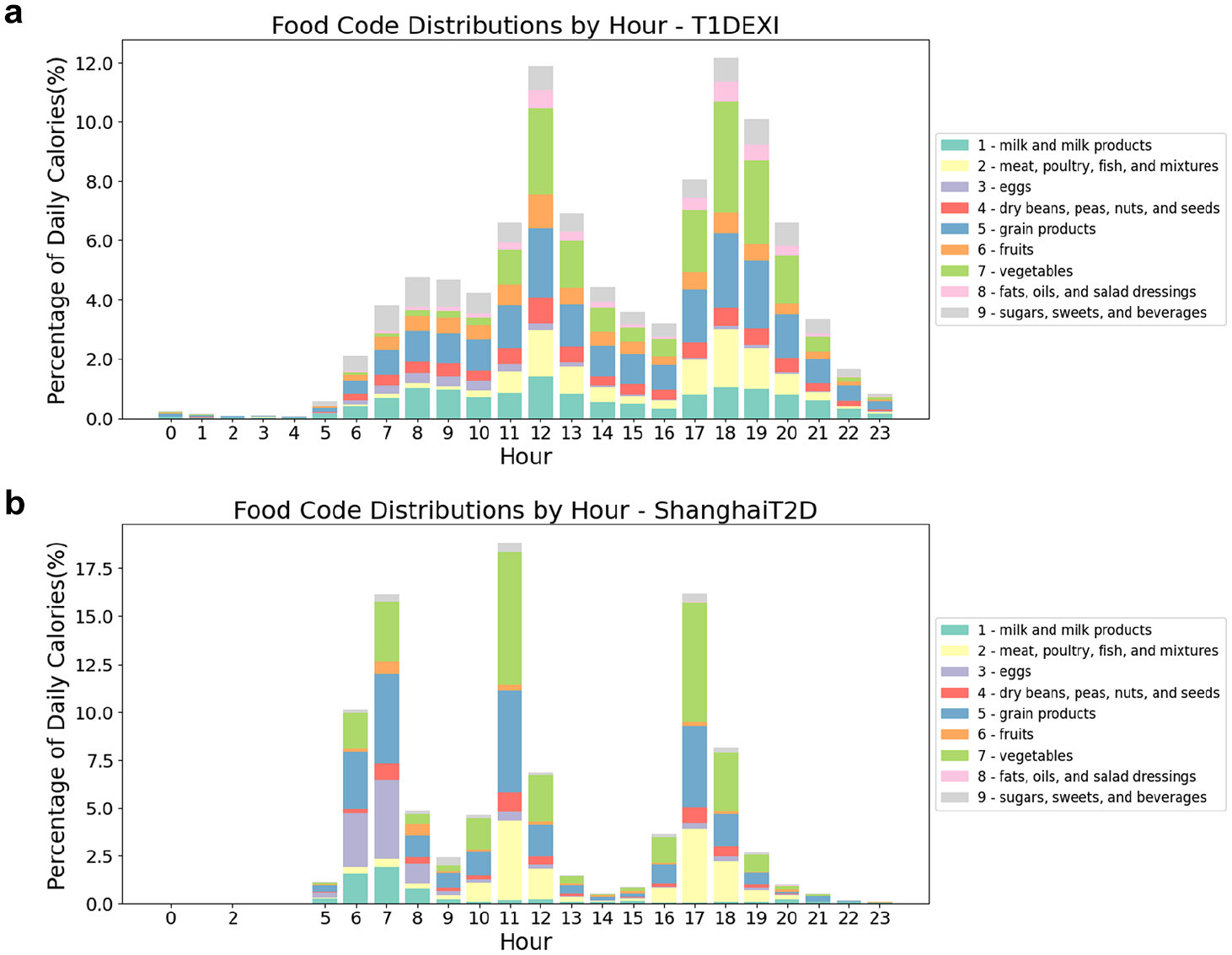
Distribution of top-level food categories by hour: (a) T1DEXI; (b) ShanghaiT2DM.
For both data sets, we calculated caloric ratios of macronutrients for each meal (carbohydrate-to-fat ratio, carbohydrate-to-protein ratio, and protein-to-fat ratio) as follows: fat (9 kcal/g), protein (4 kcal/g), and carbohydrate (4 kcal/g). 22
Hierarchical Information Criterion Feature Selection on Food
There are many unique foods in our data sets (T1DEXI 3,244 and ShanghaiT2DM 2,144), posing a challenge for feature selection. We leveraged that FNDDS codes form a hierarchy and used Hierarchical Information Criterion (HIC)
23
to determine the optimal level for each food item. The HIC is based on mutual information between the feature and outcome and takes into account the number of samples of a feature and its level, with a preference for more general (higher) levels. Equation (1) is the mutual information formula, where

HIC is defined in equation (2), where
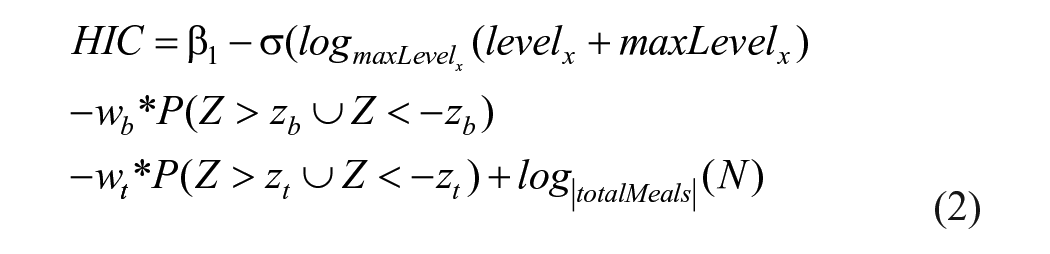
Postprandial Glycemic Response Prediction Models
We developed three base models using CatBoost in Python: (1) carbohydrates only, (2) energy only, and (3) full model. The variables in the full model for both data sets are: energy (kcal), macronutrients (grams) and their interaction ratios, base CGM (mg/dL), time since last meal (hours), sum of insulin bolus (units) within 1 hour of the meals, binary indicator of non-insulin hypoglycemic agents within an hour of the meal (only for ShanghaiT2DM), meal type (breakfast, lunch, dinner), and demographics (age, gender, body mass index [BMI] [kg/m2], duration of diabetes [years]). To evaluate the use of personalized data, we ran a 10-fold cross-validation split by meal and split by person (ie, ensuring that individuals appear only in training or test data). We conducted hyperparameter tuning using scikit-learn in Python, which selected 200 iterations and a learning rate of 0.05. L2 Leaf Regularization was used to prevent overfitting (l2_leaf_reg = 3). The top 5, 10, and 20 food features selected on each fold were added separately to the CatBoost models as one variable. CatBoost automatically processes these categorical data using its native encoding capabilities and learns relationships among the categories.
To assess feature importance, we used SHapley Additive exPlanation (SHAP) values, which indicate the average change in the model’s output when conditioned on a specific feature. 24 The overall workflow of data processing and prediction is shown in Figure 2.
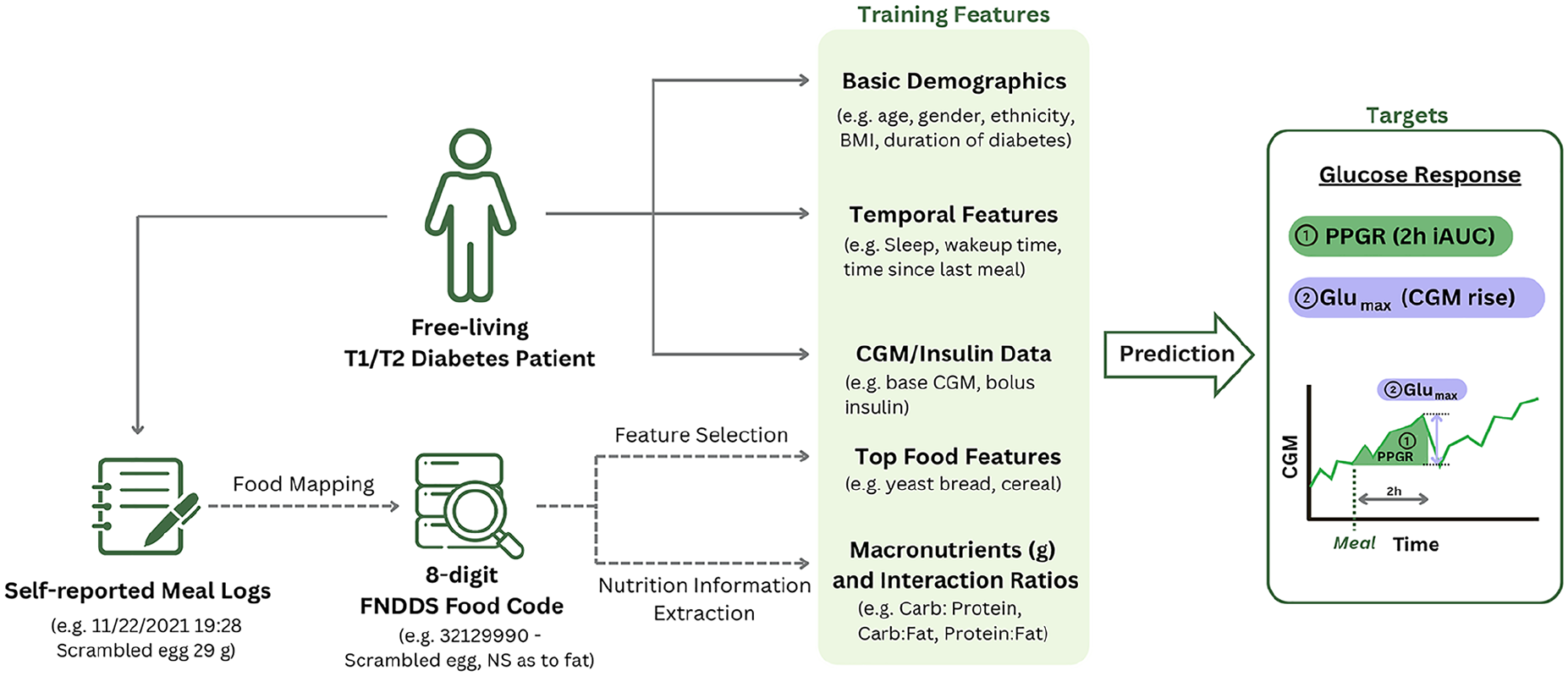
Overview of our method for predicting glycemic response. Food logs are used to obtain food names, and mapped to food codes, which are input to our feature selection method. The top selected food features, along with other personal features, are used to predict the two glycemic responses.
Modeling Intraindividual Variation
We investigated intraindividual variation in glycemic responses using data from individuals who consumed the same meal at least twice. Meals were considered duplicates when food items were identical, allowing for a 10% variation in quantity. Thus, 30 g of plain yogurt is a duplicate of 33 g of plain yogurt but not 30 g of strawberry yogurt. Meals were excluded if another meal over 200 kcal 16 was consumed within 2 hours after, to avoid confounding. For each subject, we extracted CGM data from 30 minutes before to 2 hours after each meal, and then calculated Pearson correlations between duplicate meals. We then performed hierarchical clustering on the correlation matrix of raw CGM time series data from duplicated meals using the seaborn.clustermap package with the following parameters: method = “ward” for linkage and distance metric = “euclidean.” We calculated the standard deviation (SD) of PPGR by meal type (breakfast, lunch, dinner) to test if responses varied by time of day:
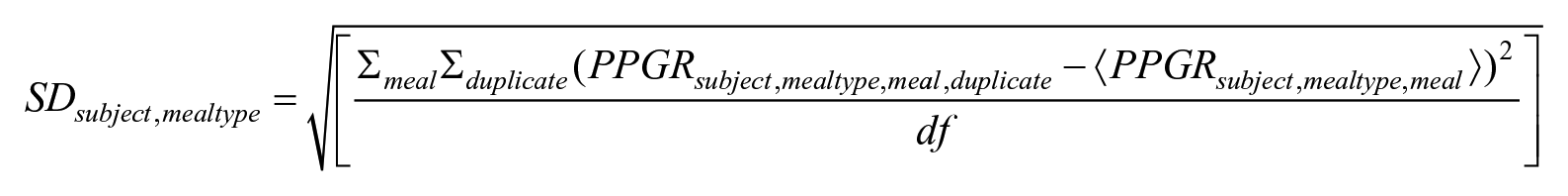
Above, subject refers to subject ID, PPGRsubject,mealtype,meal is the average PPGR for duplicate meals within a specific meal type, and df denotes the degree of freedom. We repeated this for Glumax. Individual SDs were compared using the Kruskal-Wallis H test.
Meal timing
To examine how timing of food consumption affects glycemic response, we constructed a linear mixed-effects model for each data set using the Python statsmodels library. Each response variable (PPGR, Glumax) was used as an outcome, with meal type, energy, and an interaction term between carbohydrate percentage and insulin included as explanatory variables. Random effects for subject ID and duplicated meal were added in all models to account for interindividual and meal variability and used random slopes and intercepts.
Menstrual cycle
Prior work suggests menstrual cycle phases play a role in glycemia,25-28 so we investigated this as a second source of intraindividual variation. We used the T1DEXI data set, which included the start date of menstrual periods. Since it did not include hormonal data or cycle length, we first calculated the duration between the reported start dates for each individual to determine their cycle length. For durations within 21 to 35 days, 29 that value was used as the cycle length. We excluded individuals with cycle lengths between 36 and 41 days (as these could be two shorter or one longer cycle) and assumed a 28-day cycle for others. We categorized each menstrual cycle into four phases—perimenstrual (days −3 to 2), mid-follicular (days 4 to 7), periovulatory (days −15 to −12), and mid-luteal (days −9 to −5)—based on prior work 30 with day 1 as menstrual onset. We computed one cycle before and after each reported date based on the determined cycle length. We analyzed duplicate meals consumed during any phase.
We constructed a univariate linear mixed-effects model for each outcome (daily median CGM, PPGR, and Glumax), with explanatory variables including menstrual cycle phases and meal factors (meal type, base CGM, energy, and the interaction between carbohydrate percentage and insulin). We added random effects for subject ID and duplicated meal to account for interindividual and meal variability and used random slopes and intercepts. Daily median CGM was calculated as the median of the 24-hour CGM data (12:00 am to 11:59 pm) recorded on the meal date and was used for comparison to prior work. 26
Results
Feature Selection
The top 20 food features selected by HIC (for all 10 folds) are shown in Table 3. Foods high in carbohydrates (eg, yeast bread, sweet potato, cereals) and alcoholic drinks had positive coefficients, while foods high in protein or fat (eg, chicken, nuts, avocado) and vegetables (eg, celery) generally had negative coefficients. Notably, the algorithm determined the optimal hierarchy level and selected “alcoholic beverages” rather than individual kinds (eg, beer, white wine).
Top 20 Food Features Selected for Each Data Set for Each Glycemic Response Metric.

Features with negative coefficients (ie, linked to lower responses) in our feature selection algorithm are shown in parentheses.
Postprandial Glycemic Response Prediction
Table 4 reports quantitative prediction results for both data sets, and Figures 3 and 4 show predicted versus actual responses for each model and training split. For both responses, models based solely on carbohydrates or energy had low accuracy (all R < 0.15, R2 < 0.03). Models with 20 food features had significantly higher accuracy than the full models without food features for both data sets (two-tailed t-test P < .05 for R and R2). These models had comparable accuracy to prior work reported in Table 1. For example, on the T1DEXI data set with split by person (no personal training data), our Glumax prediction had R = 0.64 and R2 = 0.38, closely matching R = 0.61 and R2 = 0.37 reported by Shilo et al 14 despite not using microbiome or personal meal data. However, for both responses and data sets, R and R2 values were significantly higher (all P < .05) in models with personalized training data (ie, split by meal). The highest SD across cross-validation folds was 0.040 for R and 0.017 for R2, highlighting the consistency of model performance across the folds.
Results of Prediction of PPGR and Glumax for Each Cross-Validation Split (By Meal, By Person).

Bold indicates the highest value for each column for each data set.
Significantly higher R/R2 compared to the full model are indicated by *P < .05, ** P < .01, *** P < .001 indicated by two-sample t-test.
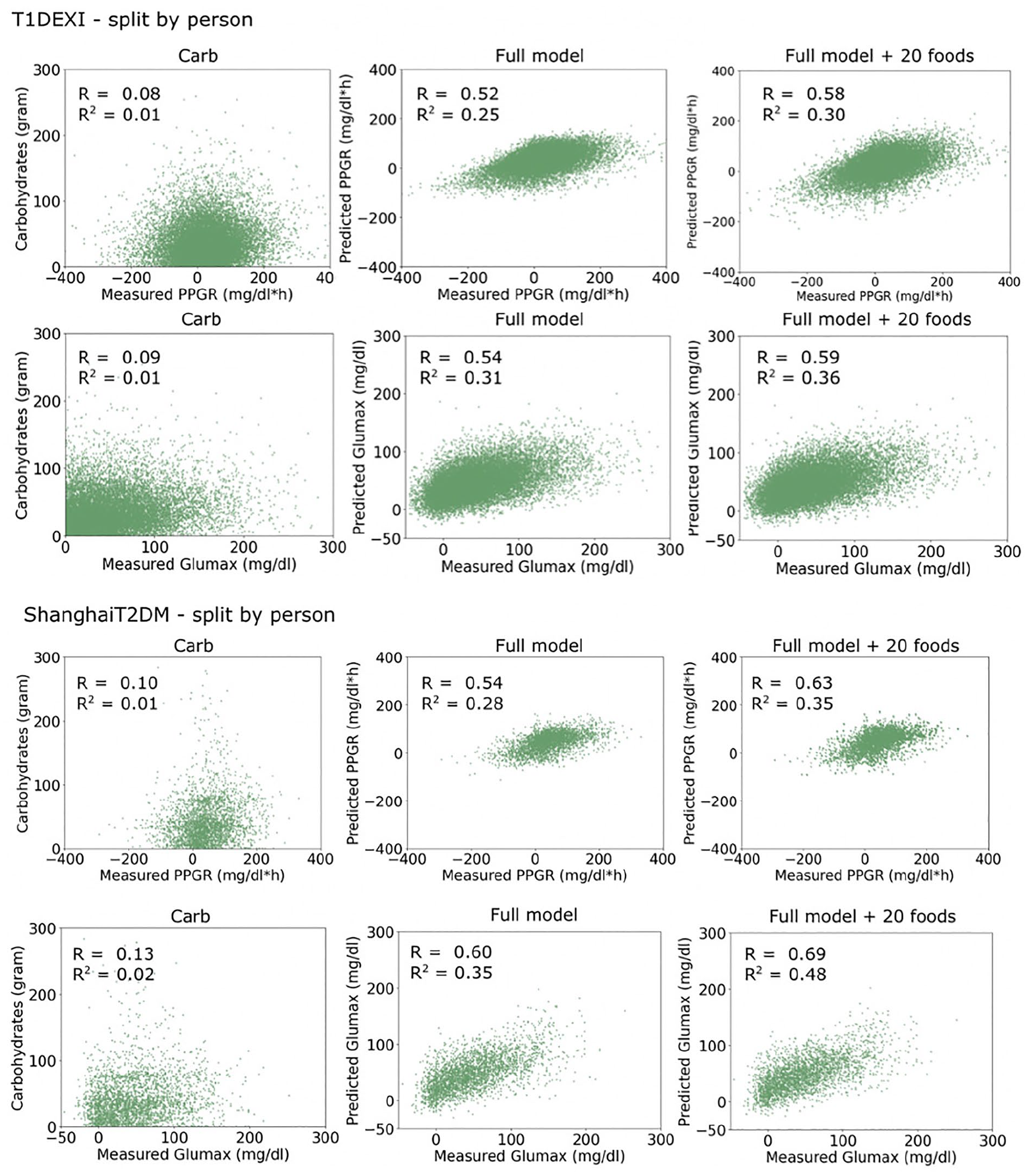
Predicted and measured PPGR and Glumax on T1DEXI and ShanghaiT2DM, split by person.
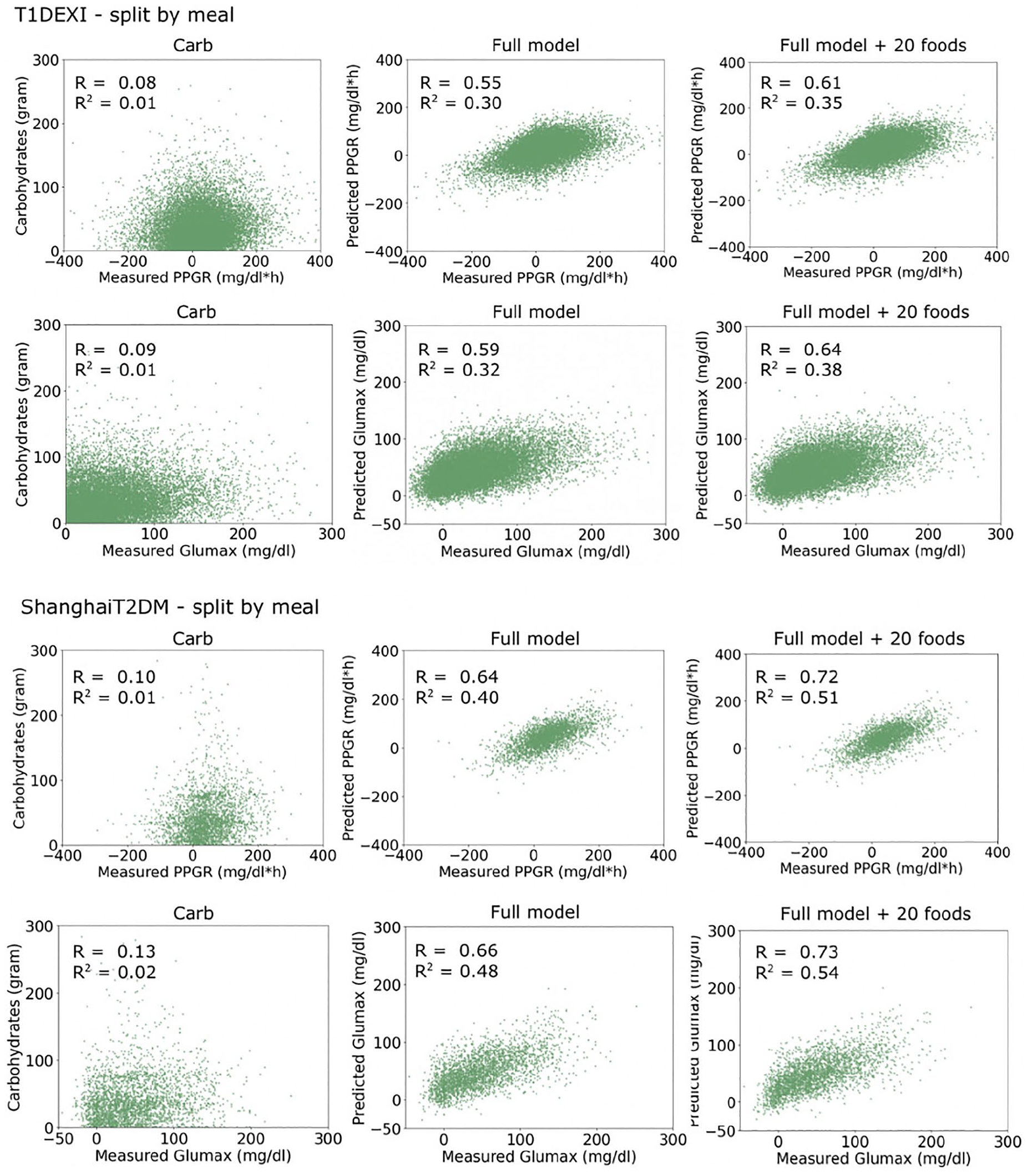
Predicted and measured PPGR and Glumax on T1DEXI and ShanghaiT2DM, split by meal.
Figure 5 shows feature importance for the full models with 20 food features and training split by person (see Supplemental Appendix B for split by meal). Base CGM was the top feature, and while macronutrients had expected effects, food features were still ranked among the top 10.
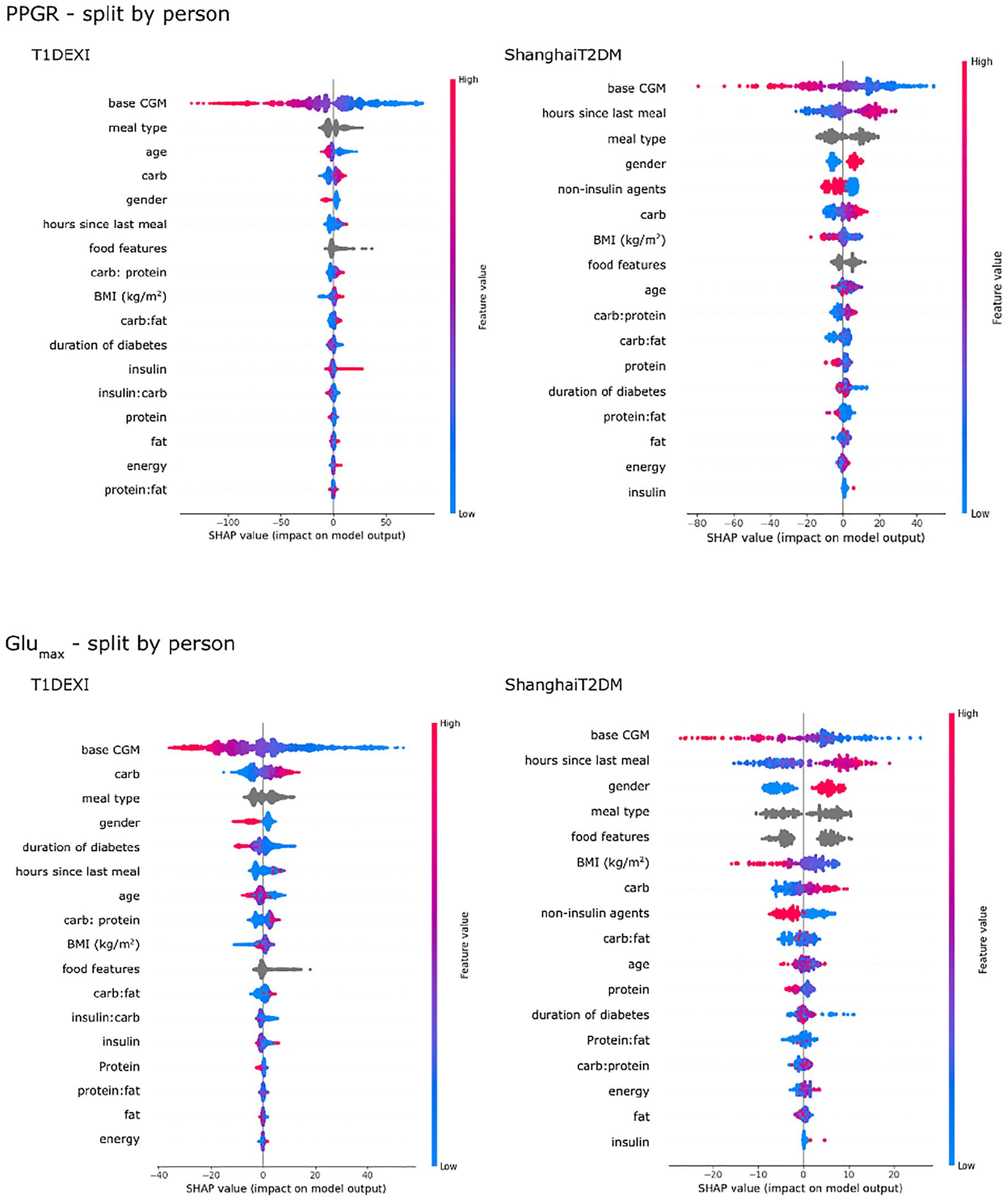
SHAP values of the features on the PPGR and Glumax prediction model, split by person.
Analysis of Intraindividual Variation
Our criteria for repeated meals yielded 116 individuals with 232 meals and 658 duplicates in T1DEXI (mean 1.54 ± 0.78 meals per person, mean 2.58 ± 1.65 duplicates per meal) and 48 individuals with 139 meals and 361 duplicates in ShanghaiT2DM (mean 2.90 ± 2.60 meals per person, mean 2.60 ± 1.68 duplicates per meal). Figure 6 shows the hierarchical clustering heatmaps of CGM data from two subjects in the ShanghaiT2DM data set, illustrating how meals clustered based on time of day rather than food type.
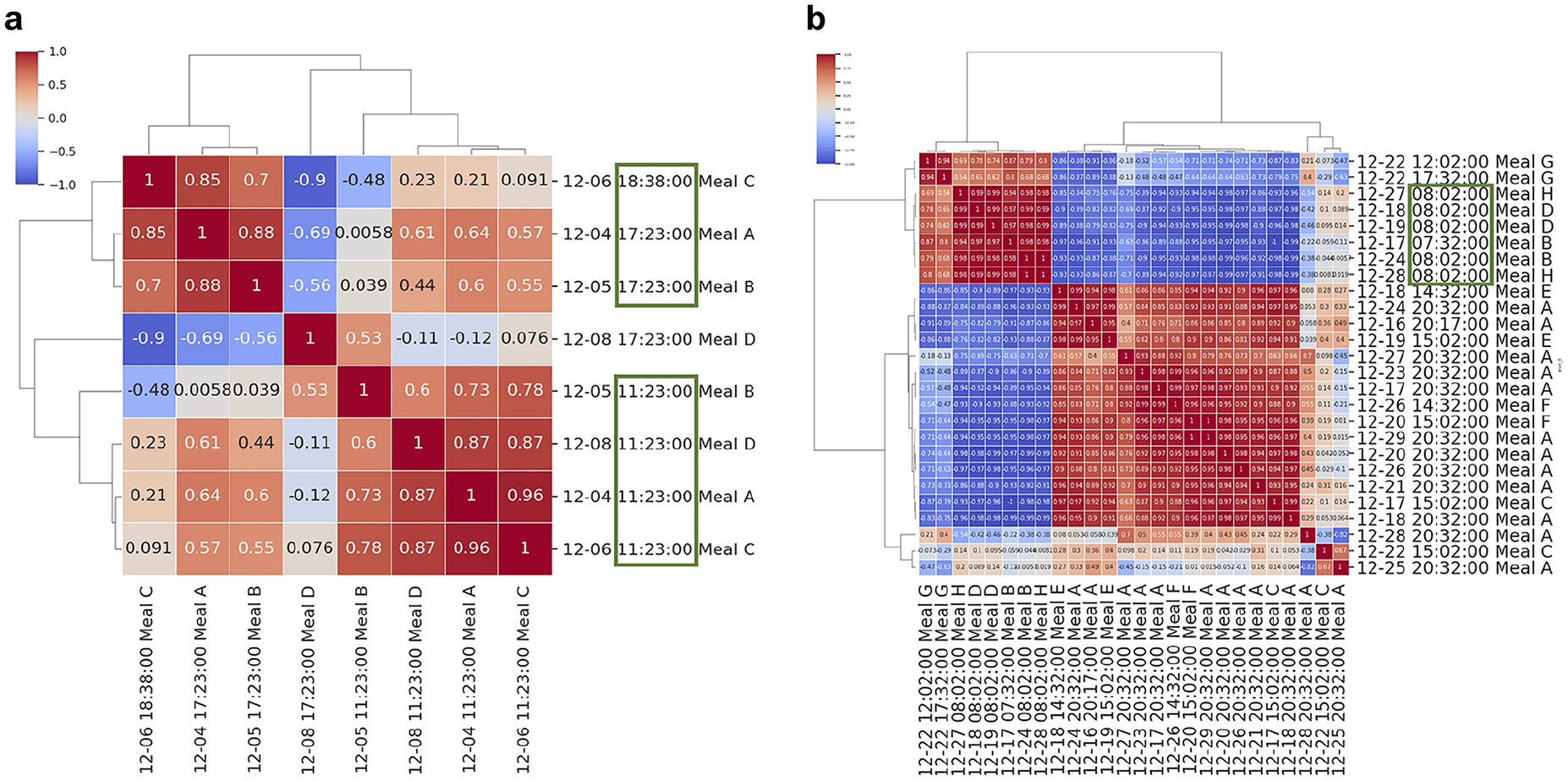
Clustering based on meal times from the ShanghaiT2DM data set. (a) Clusters formed around lunch and dinner times. (b) Two clusters formed, one around breakfast time, and another combining lunch and dinner. Green boxes highlight temporal patterns [lunch and dinner in (a) and breakfast in (b)].
Table 5 shows the mean SD of PPGR and Glumax by meal type. Breakfast was the most repeated meal. Figure 7 shows SD by person, highlighting intraindividual (glycemic responses vary even for the same meal at similar times) and interindividual (some individuals have higher SD) variance. While dinner had the highest SD for PPGR and Glumax in both data sets, differences between meal types were not statistically different for PPGR (T1DEXI: 1.75, P = .42; ShanghaiT2DM: 0.10, P = .95) or Glumax (T1DEXI: 0.73, P = .69; ShanghaiT2DM: 0.87, P = .64), using a Kruskal-Wallis H test. The mean SD of PPGR and Glumax did not differ by menstrual cycle phases (see Supplemental Appendix C).
Mean PPGR Standard Deviation and Glumax Standard Deviation of Repeated Meals by Meal Type.

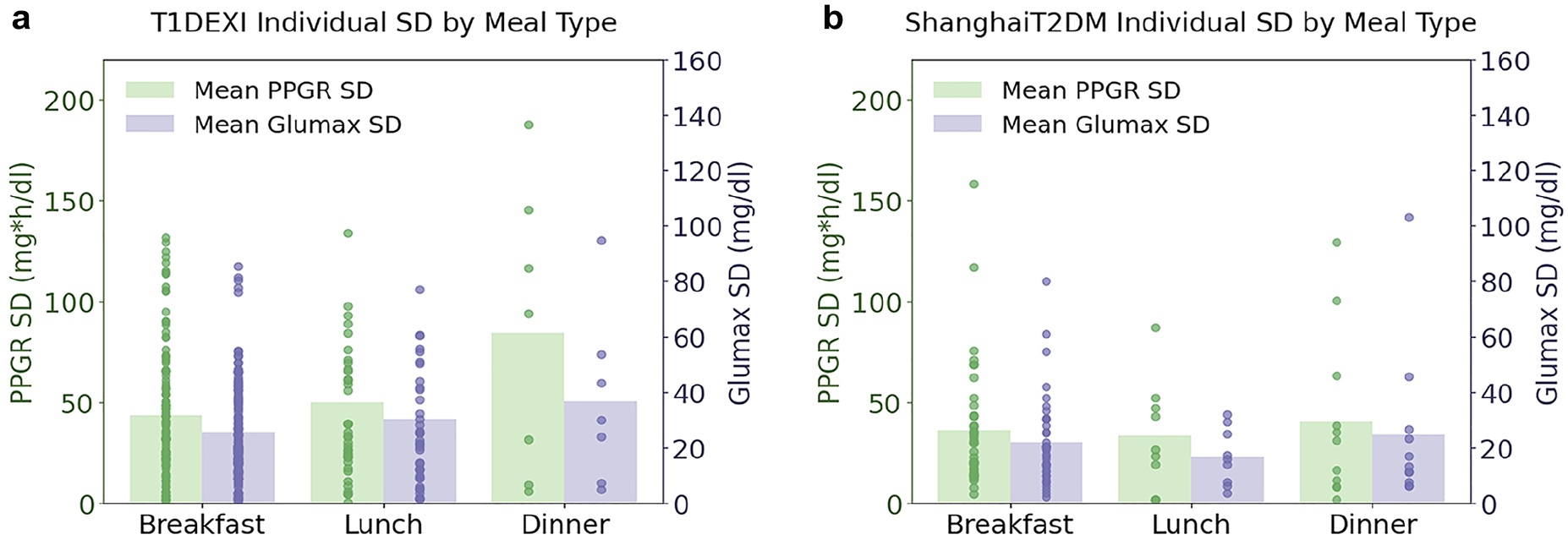
Average of individuals’ standard deviation by meal type: (a) T1DEXI; (b) ShanghaiT2DM.
Meal timing
Results for the mixed-effects model are shown in Table 6. For PPGR, meal timing was significant in both data sets (ShanghaiT2DM: P < .001 for lunch and dinner; T1DEXI: P < .05 for lunch with breakfast as the reference). We found the same effect for Glumax in ShanghaiT2DM (P < .05 for lunch and P < .001 for dinner), although not for T1DEXI. We additionally tested non-insulin hypoglycemic agents and activity as potential confounders, but neither was significant (see Supplemental Appendix D1). The results of adding base CGM as an explanatory variable are presented in Supplemental Appendix D2.
Univariate Linear Mixed-Effects Results With Meal Types.

P < .05, **P < .01, ***P < .001.
Menstrual cycle
Our method led to a total of 1280 cycles and 4836 cycle phases from 261 individuals (mean 4.90 ± 1.34 cycles and mean 18.53 ± 4.74 cycle phases per person) in T1DEXI. Applying our criteria for duplicate meals, we identified 120 duplicate meals from 37 individuals (mean 1.20 ± 0.51 meals per individual, mean 3.24 ± 2.13 duplicates per meal).
Table 7 shows the results of the mixed-effects models. First, the perimenstrual phase (day −3 to 2 days after menses onset, equivalent to day 26 to +2 for a 28-day cycle) had significantly higher daily median CGM compared to the periovulatory phase (days 14 to 17), which is consistent with prior findings in people without diabetes (higher daily median CGM during the luteal phase [day 24.5 ± 8.0] compared to the late-follicular phase [day 13.6 ± 3.4]). 26 The Glumax was significantly higher for duplicated meals during the perimenstrual phase (P < .05), suggesting that hormonal factors may contribute to intraindividual variation in responses.
Univariate Linear Mixed-Effects Results With Menstrual Cycle Phases.

P < .05, **P < .01, ***P < .001.
Discussion
We aimed to predict PPGRs from limited data and investigate the reason for high intraindividual variation. We found that (1) PPGR and Glumax can be accurately predicted without invasive data such as microbiome, (2) including food type in models led to higher accuracy than only including its macronutrients, and (3) intraindividual variation is partly explained by temporal and hormonal factors.
While prediction of glycemic responses to meals has been a highly active research area, the most accurate models have relied on extensive individual data including gut microbiome. We find that comparable accuracy to prior work can be achieved10,14 with much more limited data, namely demographics, meal features, and temporal features. Furthermore, our results of split-by-person validation show that high accuracy can be obtained with no person training data. Ultimately, this may make it easier and cheaper to scale personalized nutrition interventions.
Our work raises new questions on which aspects of food contribute to glycemic responses. While prior work focused on macronutrients, we find that food categories (selected with HIC) led to higher accuracy even with macronutrients in the model. However, future work is needed to understand what the mechanism may be, such as whether it is due to other nutrients (eg, fiber), features such as processing level, or factors correlated with food type (eg, eating speed which may be partly determined by physical properties of a food). There is some evidence, for example, suggesting ultra-processed foods may have higher glycemic impact. 31
Prior work raised questions about high degrees of intraindividual variation in glycemic responses. 16 Taking advantage of habitual eating behavior, we now show that this variation may be partly due to timing (eating the same food for lunch vs dinner) and hormones (menstrual cycle phase). However, there are many other factors (eg, stress, activity before and after meals) that may also be influential. Understanding the reasons behind intraindividual variation could significantly improve guidance to patients and reduce the frustration experienced when the same behavior (ie, same meal, same insulin dosing) yields different effects.
Our study has several limitations. First, since our data sets did not include microbiome, it is an open question as to whether that would further improve our models. While many individuals repeated meals, there were fewer duplicate meals at dinner, so future work may be needed in controlled settings to fully examine the role of time. Similarly, in both data sets, subjects self-reported their meals, which is another potential source of errors. In addition, the ShanghaiT2DM data set required additional post-processing to determine nutrients and food codes, which could have affected accuracy. Moreover, additional potential confounders were missing from the data, such as physical activity (eg, daily step count) in ShanghaiT2DM, which could have influenced the results. Low power could also be a limitation, especially for the T1DEXI data set. As part of future work, we aim to test on a more comprehensive data set and further explore the generalizability of models. Finally, without hormonal data, we cannot confirm the menstrual cycle phases identified for each participant.
Conclusions
We showed that PPGRs can be accurately predicted from limited data. Food categories identified using HIC led to improved accuracy compared to models using macronutrients. In addition, our analysis provided new insights into reasons behind intraindividual variation in glycemic responses. Our findings suggest that personalized nutrition could be used widely in diabetes management and may not require expensive or invasive data to obtain sufficient accuracy.
Supplemental Material
sj-docx-1-dst-10.1177_19322968251321508 – Supplemental material for Predicting Postprandial Glycemic Responses With Limited Data in Type 1 and Type 2 Diabetes
Supplemental material, sj-docx-1-dst-10.1177_19322968251321508 for Predicting Postprandial Glycemic Responses With Limited Data in Type 1 and Type 2 Diabetes by Yiheng Shen, Euiji Choi and Samantha Kleinberg in Journal of Diabetes Science and Technology
Footnotes
Acknowledgements
This publication is based on research using data from Jaeb Center for Health Research Foundation that has been made available through Vivli, Inc. Vivli has not contributed to or approved and is not in any way responsible for, the contents of this publication. During the preparation of this work, the authors used ChatGPT to assist in estimating ingredient portions for complex food items based on common recipes.
Abbreviations
BG, blood glucose; CGM, continuous glucose monitor; DASH, Dietary Approaches to Stop Hypertension; FNDDS, Food and Nutrient Database for Dietary Studies; HIC, Hierarchical Information Criterion; iAUC, incremental area under the curve; PPGR, postprandial glycemic response; RCT, randomized controlled trial; SD, standard deviation; SHAP, SHapley Additive exPlanation; T1D, type 1 diabetes; T1DEXI, type 1 diabetes and exercise initiative; T2D, type 2 diabetes.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The research was supported by NIH U54TR004279 and NSF 1915182.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
