Abstract
Continuous glucose monitoring (CGM) data stored in data warehouses often include duplicated or time-shifted uploads from the same patient, compromising data quality and accuracy of resulting CGM metrics. We developed a processing algorithm to detect and resolve these errors. We validated the algorithm using two weeks of CGM data from 2038 patients with diabetes. Duplication errors were identified in 528 patients, with 25.7% showing significant differences in at least one metric (Time in Range, Coefficient of Variation, Glycemic Management Indicator, or Glycemic Episode counts) between raw and processed data. Eleven patients crossed clinically meaningful thresholds in one or more metrics after processing. Our results underscore the importance of real-world CGM data processing to maintain accurate and reliable CGM metrics for research and clinical care.
Introduction
The widespread use of continuous glucose monitoring (CGM) technology has led to the integration of CGM metrics in clinical and research settings. 1 CGM data are increasingly stored in data warehouses, and the transition from patient-level data collection to clinical-level data analysis creates a complex storage architecture. 2 Unlike curated CGM data from clinical studies, CGM data stored in a data warehouse may contain duplicated or overlapping profile uploads from the same patient, compromising data quality (Figure 1, left). Potential sources of duplication include multiple device uploads, time-shifted data caused by user time zone changes, or repeated patient IDs. However, identifying the exact source is challenging using the data alone, and the issue typically becomes apparent only when glucose measurements exceed the expected time frequency. This duplication problem has received limited attention in the literature, leading to CGM metrics being computed on data “as is” or ad-hoc processing approaches, such as averaging measurements from the same timestamp or arbitrarily selecting one measurement per timestamp. These solutions are unreliable because parallel glucose profiles create ambiguity in determining glucose levels at a specific time. At the same time, individual-level manual data curation is not resource-feasible within large data warehouse settings. Automatic identification and correction of duplication issues are essential in ensuring accurate and reliable CGM metrics in data warehouse settings.
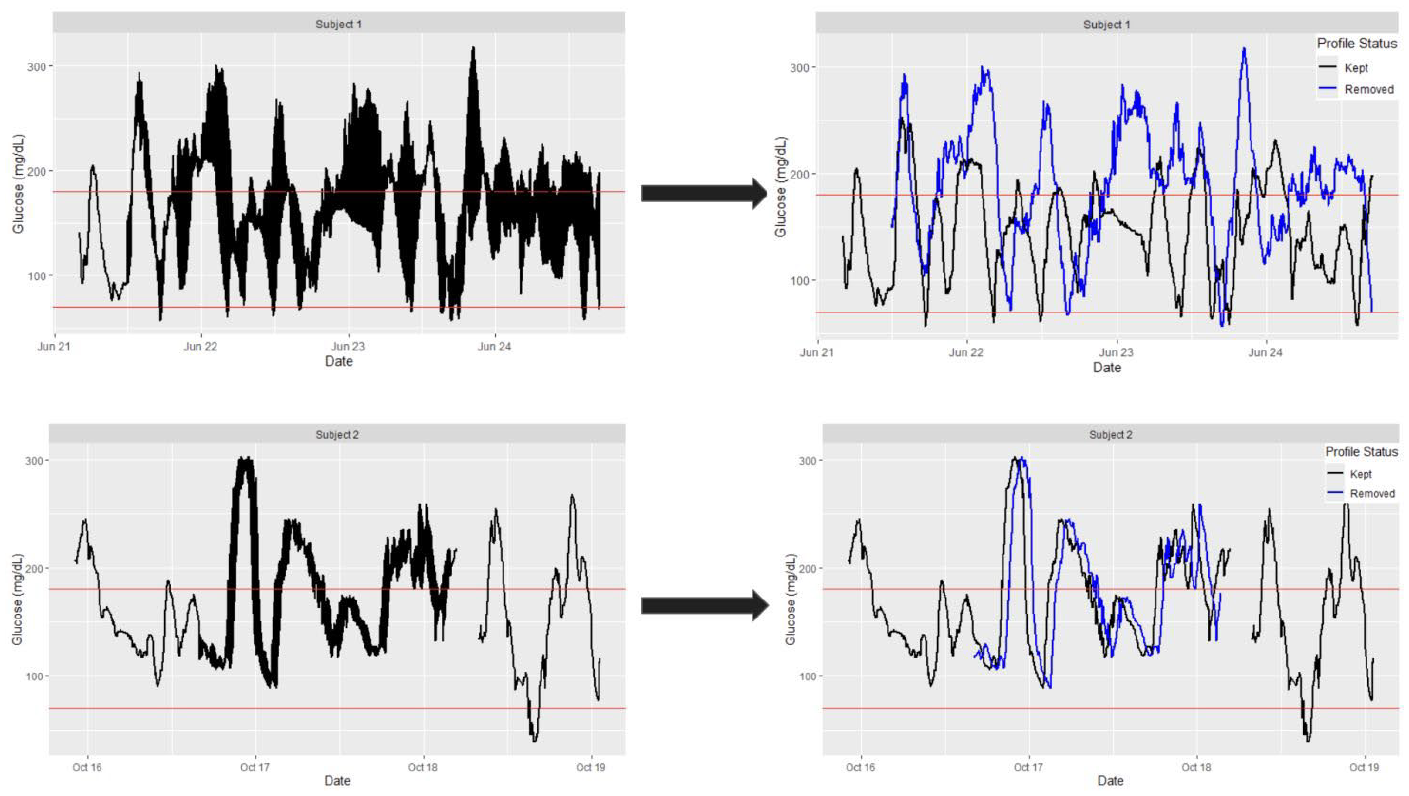
Example CGM data stored in the data warehouse from two subjects with more frequent than expected measurements (left) and after applying the proposed processing algorithm (black profiles, right). Blue profiles represent data removed by the algorithm, highlighting the presence of multiple parallel profiles for a single individual.
To address data quality issues due to duplication, we develop a data-cleaning algorithm that uses observation-level unique identifiers to distinguish between multiple CGM data streams. Our algorithm leverages preceding non-duplicated glucose measurements to filter out duplicates, and we validate its performance on retrospective observational CGM data from a single center. Our findings confirm that unprocessed duplication can lead to clinically significant differences in CGM metric values for some patients, highlighting the importance of standardized data processing for reliable and accurate CGM data integration into electronic health records (EHRs). 2
Methods
We retrospectively collected 2 weeks of CGM data from a cohort of pediatric and adult patients with diabetes receiving outpatient care at a single-center academic health system. Our data set, generated in September 2024, included an initial sample size of 2,348 unique individuals. Following international consensus guidelines, 1 we retained profiles with at least 70% data availability over the 14 most recent days of CGM readings, where data availability was calculated using R package iglu. 3 The final cohort comprised of n = 2,038 individuals. Each observation included patient ID, time stamp, glucose value (in mg/dL), Device ID (associated with a unique device measuring glucose levels), and Observation ID (a unique identifier for each measurement). This research was approved by the Michigan Medicine Institutional Review Board (HUM00249748).
We identified duplicated CGM observations by flagging data points recorded at intervals shorter than the expected 5-minute measurement frequency (top Figure 2). Although duplication may stem from parallel profiles with distinct Device IDs, resolving using only Device ID is problematic. Patients often switch devices, leading to multiple Device IDs within a profile. Furthermore, Device ID information may be missing, or duplication may occur within the same ID. To address this, our stepwise processing algorithm considers glucose values, time stamps, Device IDs, and Observation IDs. It progresses sequentially to identify a unique glucose profile, avoiding abrupt switching between parallel profiles. Specifically, the algorithm selects one value from a duplicated set following the logic illustrated in Figure 2. The algorithm begins at time zero for each CGM profile and progresses until it encounters the first duplicated set of measurements (red box surrounding g2 and g3). For exact duplicates (identical glucose values and time stamps), extra copies are removed. For non-exact duplicates (differences in glucose values or time stamps), the algorithm references the preceding non-duplicated entry (g1 in Figure 2) to identify a unique match based on Device ID or Observation ID. If no unique match is found, it matches by expected time frequency (5 min from g1). In the most complex scenario, where multiple distinct glucose values share the same time and ID information, the algorithm selects the glucose value closest to g1, assuming minimal glucose change over short intervals. The selected value becomes the new verified observation (g1) for resolving subsequent duplicate sets.
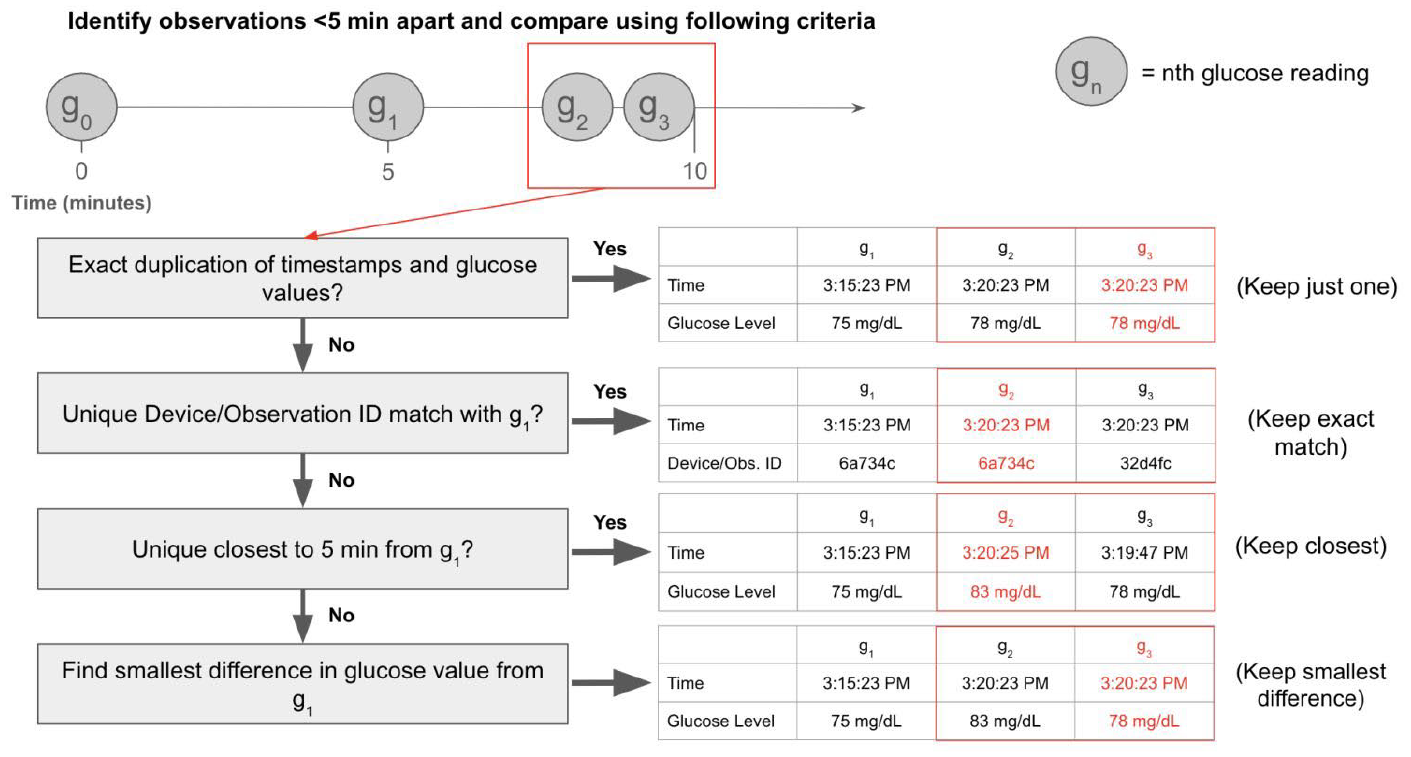
Illustration of the proposed stepwise processing algorithm for removing duplicate observations.
To assess the impact of our algorithm on the value of resulting CGM metrics, we calculated consensus CGM metrics of glycemic control for each subject before and after processing using the R package iglu.3,4 The consensus CGM metrics included time in range (TIR, % between 70 and 180 mg/dL), time hyperglycemic (% >180 mg/dL), time hypoglycemic (% <70 mg/dL), coefficient of variation (CV), Glycemic Management Indicator (GMI), 5 and Level 1 glycemic episode count. 1 A Level 1 hyperglycemia episode is defined as >=15 consecutive minutes above 180 mg/dL, and a Level 1 hypoglycemia episode is defined as >=15 consecutive minutes <70 mg/dL. We evaluated the absolute differences in each CGM metric between the raw and processed data and divided patients into lower and higher magnitudes of difference groups for each metric. We used difference thresholds of 2% for TIR, 2% for time hyperglycemic, and 1% for time hypoglycemic, following the previously established bounds on clinically acceptable similarity, 6 and a threshold of 0.1 for both CV and GMI, representing standard reporting accuracy for these metrics. For episodes, we calculated differences in the reported counts. We counted the number of patients crossing a clinical threshold as a result of processing, including CV levels of less than or equal to 36%, 7 target TIR (between 70 and 180 mg/dL) of ≤ or >70%, 8 target GMI of < or ≥ 7%. 5 Finally, we used data from 10 individuals with the highest number of duplications to verify algorithm performance. We visually inspected their profiles before and after processing to assess continuity and identify the characteristics of removed observations.
Results
Duplication detection procedure within the algorithm revealed that 25.9% (n = 528) of the 2,038 individuals in our study cohort had some form of duplication errors. In this affected subgroup, between 1 and 6446 observations were removed, with an average of 129 observations per profile, representing 10.75 hours of data over the two-week period. We identified three types of duplication errors: exact duplication of both timestamp and glucose values, glucose value duplication with a predictable time shift, and glucose value duplication with an unclear time shift pattern.
Table 1 categorizes differences in CGM metrics before and after processing profiles of the affected 528 patients. A higher magnitude of differences in at least one metric is observed in 136 patients (25.7%), with the highest count being change in CV values, followed by a count of differences in Level 1 hyperglycemia episodes.
Classification of the Absolute Differences in Consensus Metrics Before and After Data Processing for a Subgroup of 528 Patients Containing Duplicate Measurements.

In total, 11 individuals shifted across clinically meaningful thresholds in at least one CGM metric due to data processing: six crossed the 70% TIR threshold, one crossed the 7% GMI threshold, and four crossed the 36% CV threshold.
Discussion
We developed a CGM data-processing algorithm to remove duplicated measurements. Comparing consensus glycemic control metrics between raw and processed data, we found duplication can affect metric interpretation for some individuals, with a minor impact on clinical interpretation. Furthermore, as research studies increasingly pull CGM data from data warehouses, even minor clinical differences in CGM metric values may have a significant effect on the research conclusions. Thus, our findings highlight the need for systematic data processing before the analysis of CGM data from data warehouses and its integration with EHR, emphasizing the value of an informed data-processing approach over “as-is” metric computation.
Our algorithm, while complex, improves data reliability, and its accessibility can be enhanced by integration within existing open-source CGM software. 9 Although we applied it to 5-minute frequency data in our study, the algorithm easily adapts to other frequencies without altering the core logic.
However, the algorithm has limitations. It is specifically designed to address duplication errors, meaning other data quality issues, if present, remain unaddressed. In the most ambiguous cases, the algorithm uses glucose values to resolve duplications, which may occasionally result in alternating profile selection. Despite these challenges, the algorithm’s automation enables efficient processing of large-scale, high-volume data sets; this is particularly valuable in data warehouse settings where manual review of each profile is infeasible.
Future efforts to quantify the sources of time shifts and duplication errors and establish standardized CGM data-processing guidelines will help ensure consistent and accurate CGM data handling in large data warehouses. Our results highlight opportunities to improve CGM data accuracy and reliability, ultimately benefiting both clinical care and research studies.
Footnotes
Acknowledgements
The authors thank Ashley Garrity and Emily Dhadphale for their technical support and guidance in this project.
Abbreviations
CGM, continuous glucose monitor; EHR, electronic health records; TIR, time in range; CV, coefficient of variation; GMI, glycemic management indicator.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: WW and IG declare no competing interests. JML has served as a consultant to Tandem Diabetes Care, has participated on the Sanofi Digital Advisory Board, and is on the Medical Advisory Board for GoodRx.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: WW received support from the Institute for Healthcare Policy and Innovation Summer Student Fellowship. IG and WW received research support through NIH grant R01HL172785. Additional support for this work was provided by the National Institute of Diabetes and Digestive and Kidney Diseases through grants P30DK089503 (MNORC), P30DK020572 (MDRC), and P30DK092926 (MCDTR), as well as by the Elizabeth Weiser Caswell Diabetes Institute at the University of Michigan.
