Abstract
Background and Objective:
The aim of this study was to develop and validate explainable prediction models based on continuous glucose monitoring (CGM) and baseline data to identify a week-to-week risk of CGM key metrics (hyperglycemia, hypoglycemia, glycemic variability). By having a weekly prediction of CGM key metrics, it is possible for the patient or health care personnel to take immediate preemptive action.
Methods:
We analyzed, trained, and internally tested three prediction models (Logistic regression, XGBoost, and TabNet) using CGM data from 187 type 1 diabetes patients with long-term CGM monitoring. A binary classification approach combined with feature engineering deployed on the CGM signals was used to predict hyperglycemia, hypoglycemia, and glycemic variability based on consensus targets (time above range ≥5%, time below range ≥4%, coefficient of variation ≥36%). The models were validated in two independent cohorts with a total of 223 additional patients of varying ages.
Results:
A total of 46 593 weeks of CGM data were included in the analysis. For the best model (XGBoost), the area under the receiver operating characteristic curve (ROC-AUC) was 0.9 [95% confidence interval (CI) = 0.89-0.91], 0.89 [95% CI = 0.88-0.9], and 0.8 [95% CI = 0.79-0.81] for predicting hyperglycemia, hypoglycemia, and glycemic variability in the interval validation, respectively. The validation test showed good generalizability of the models with ROC-AUC of 0.88 to 0.95, 0.84 to 0.89, and 0.80 to 0.82 for predicting the glycemic outcomes.
Conclusion:
Prediction models based on real-world CGM data can be used to predict the risk of unstable glycemic control in the forthcoming week. The models showed good performance in both internal and external validation cohorts.
Introduction
Diabetes, a chronic metabolic disorder, affects millions of individuals worldwide and is classified into several subtypes, including type 1 diabetes (T1D) which is a distinct autoimmune diabetes subtype. The T1D is characterized by the autoimmune destruction of insulin-producing pancreatic beta cells, leading to an absolute insulin deficiency. 1 Consequently, people with T1D require life-long insulin treatment, and maintaining glycemic control is the cornerstone of managing T1D and preventing associated complications. 2 However, achieving stable glycemic control is often challenging due to the dynamic nature of the disease. Uncontrolled diabetes can lead to devastating short-term and long-term complications, such as diabetic ketoacidosis (DKA), hypoglycemia, cardiovascular disease, neuropathy, nephropathy, and retinopathy.2-4
Unstable glycemic control encompasses a spectrum of challenges, including hyperglycemia (elevated blood glucose levels), hypoglycemia (low blood glucose levels), and glycemic variability (fluctuations in blood glucose levels over time).5,6 These components of glycemic control are all intertwined and play significant roles in the management of T1D. Patients exhibit varying degrees of glycemic control, and the identification of those who experience unstable glycemic control is crucial for treatment personalization and tailoring of the most effective treatment strategy to the individual patient. 7 Understanding and managing individual components of glycemic control, namely hyperglycemia, hypoglycemia, and glycemic variability, are therefore of paramount importance.8-12
Continuous glucose monitoring (CGM) technology has revolutionized the monitoring and management of T1D.13,14 The CGM systems provide real-time data on glucose levels, enabling patients and health care providers to make informed decisions about insulin dosing, meal planning, and lifestyle modifications. The ability to capture data continuously allows for a more comprehensive assessment of glycemic control, including the detection of hyperglycemia, hypoglycemia, and glycemic variability. 15
Management of glycemic control has typically been reactive or based on historical data for a “naively” extrapolation to future glycemic control. 5 Nonetheless, many studies have been published in recent years, showing that hyperglycemia,16-19 hypoglycemia,20-27 and glycemic variability 28 are possible to predict in advance within minutes or hours using machine-learning approaches. However, a longer prediction horizon could enable a more clinically feasible timeframe for intervention.
In this study, we explored the application of explainable machine-learning models to predict the weekly risk of hyperglycemia, hypoglycemia, and glycemic variability in patients with T1D, using data obtained through CGM. By employing machine-learning models that provide transparent insights into their decision-making processes, we aim to contribute to the development of personalized treatment strategies that can effectively mitigate the risk of diabetes-related complications. Through this research, we hope to highlight the potential of CGM-based predictive modeling as a tool for identifying patients with unstable glycemic control, ultimately improving the quality of care, and enhancing the lives of individuals living with T1D.
Materials and Methods
Discovery Cohort
An overview of the study methods is provided in Figure 1. We trained and internally tested several prediction models using data from the WISDM study, 29 which includes CGM and baseline information. These models were designed to forecast the likelihood of hyperglycemia, hypoglycemia, or glycemic fluctuations in the upcoming week. The WISDM study aims to determine the effectiveness of CGM in reducing hypoglycemia in older adults with T1D in comparison to traditional blood glucose monitoring.
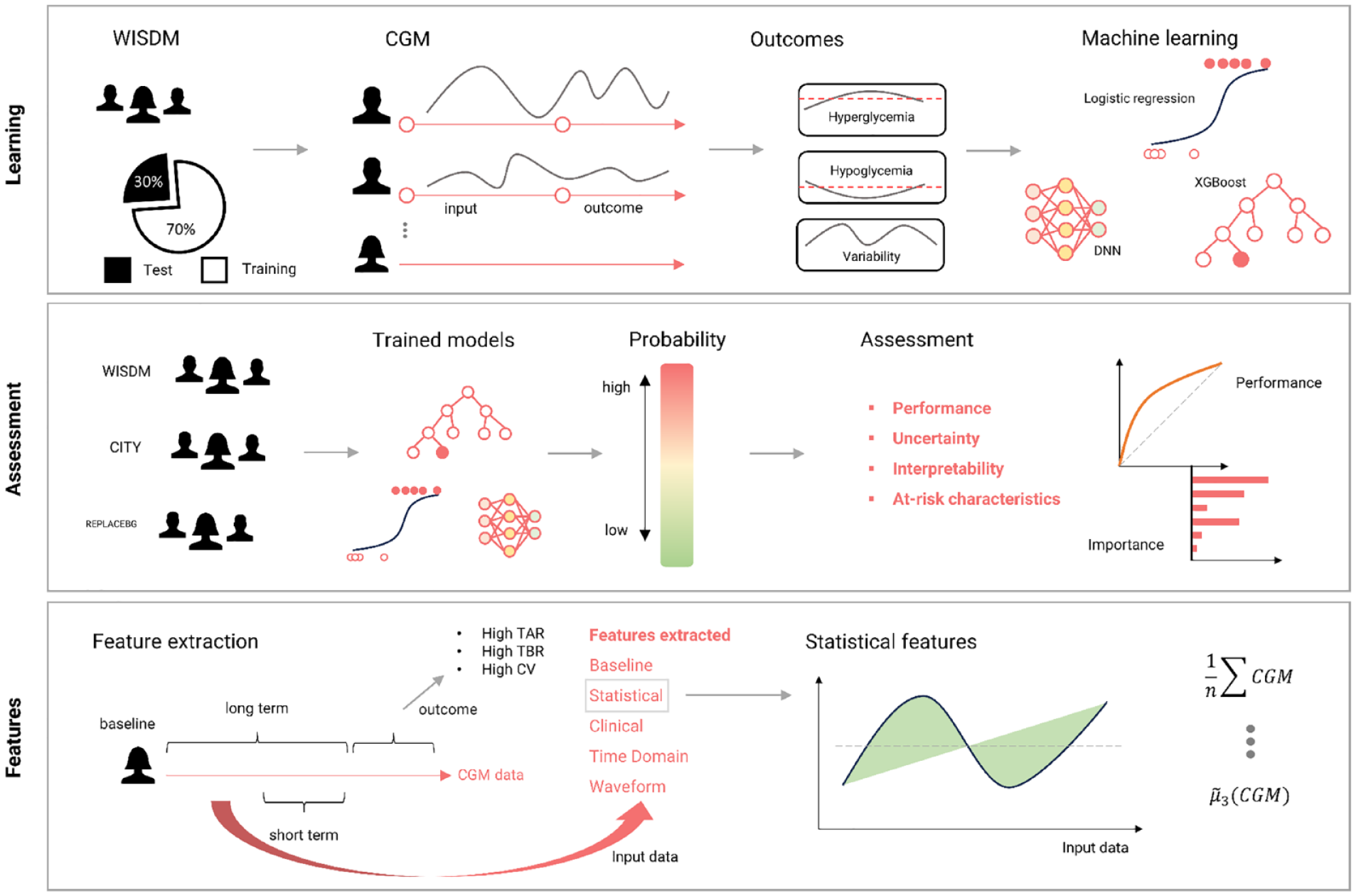
Training and validation of prediction models for prediction of hyperglycemia, hypoglycemia, and glycemic variability based on weekly CGM and baseline patient characteristics. The general ML workflow starts with partitioning the data into a training set (Train) and a test set (Test). The trajectories for training input are generated by sampling continuous subsequences of weekly CGM with adjacent weekly outcomes. A range of features are extracted from the weekly input CGM, and the binary outcome is defined in the following week in accordance with established CGM criteria. The training data sets are used for training to minimize the model’s prediction error. A model’s ability to predict a high risk of glycemic instability (hyperglycemia, hypoglycemia, and excessive glycemic variability) using the test data set and the two validation data sets is assessed by diagnostic performance characteristics, uncertainty estimates, interpretability, and at-risk characteristics.
A total of 206 individuals with T1D participated in the WISDM study and used CGM (Dexcom G5) for up to six months. We selected prediction cases where CGM was used for at least 70% of the time over five weeks. The first four weeks were used to extract features for forecasting the outcomes of the last week. Any participants without suitable CGM data for analysis were excluded.
External Validation Cohorts
We conducted external validation of our final prediction models using two additional independent data sets, namely the REPLACE-BG trial 30 and the CITY trial. 31 The rationale behind incorporating these two study populations alongside the WISDM population was to create a diverse and extensive data set that could represent the variations found in the broader T1D population, spanning from young to old individuals.
Prediction Model Targets
The targets of the prediction models were excessive hyperglycemia, hypoglycemia, and glycemic variability.
Hypoglycemia was measured using CGM and based on a time below range (TBR; <70 mg/dL), hyperglycemia was measured as time above range (level 2) (TAR; >250 mg/dL), and glycemic variability was measured as coefficient of variation (CV).15,32,33
We defined targets by binary thresholds that have all been recommended as treatment targets in recent CGM consensus guidelines:
Predictors and Feature Engineering
Predictor variables encompassed baseline characteristics including age, insulin pump usage, insulin type, and the number of insulin types. In addition, CGM features were categorized into clinical, statistical, time-domain, and waveform aspects to capture the dynamics of glucose variations. The calculation of CGM features was derived from short-term (1 week) and long-term (4 weeks) measurements before the target week. A total of 50 features were evaluated as potential predictors, as outlined in Table 1. Several of the features merit a more detailed explanation and mathematical definition, which is presented in Supplementary Material S1.
Features for Predicting Glycemic Targets for the Forthcoming Week.

The features were categorized into baseline, clinical, statistical, time-domain, and waveform features.
Model Development
For a comprehensive investigation, we employed tree classification approaches (General Linear Model, GLM; tree-based ensemble model; and deep neural network, DNN) to train, test, and validate each prediction model. These model types have fundamentally different approaches to modeling data, each with its own strengths and weaknesses.36-38
To maintain the independence of training and testing data, we partitioned the cases into a training data set (70%) and a test data set (30%) at the individual level (Figure 1). This methodology ensures that data from the same individual do not appear in both the training and test data sets. The final models were tested on the test data set and the validation data sets. We did not recalibrate the model after the training process. To attain good generalizable performance, careful tuning of the models is often necessary. In this study, grid search, 39 in combination with K-fold cross-validation 40 (5-fold), was employed for parameter optimization. For the GLM approach, we employed forward feature selection to reduce noisy features and improve generalization. 41
Logistic regression (general linear model)
Logistic regression, a classical statistical method in the GLM family, continues to be a cornerstone in predictive modeling and statistical analysis in clinical research.42,43 Widely used for binary classification tasks, it offers simplicity, interpretability, and computational efficiency. Its strengths include the ability to model probability directly, providing insight into the likelihood of outcomes. Logistic regression is robust in scenarios with a moderate number of features and a clear linear relationship with the log odds of the response variable. However, its linearity assumption can be a limitation when handling complex, nonlinear relationships.
XGBoost (tree-based ensemble model)
XGBoost, or Extreme Gradient Boosting, 44 is a machine-learning algorithm recognized for handling complex data sets by combining weak prediction models (decision trees) into a robust ensemble. It excels in capturing nonlinear relationships, managing missing/imbalanced data, and preventing overfitting, which generally leads to high predictive performance. This has been showcased in clinical prediction models across various medical applications.45-47
TabNet (deep neural network)
TabNet is a newly proposed DNN machine-learning model gaining prominence for its efficacy in handling tabular data. 48 TabNet supposedly excels at capturing complex relationships within structured data sets, making it particularly suitable for tasks such as classification and regression. Its strengths lie in interpretability, feature importance analysis, and robust performance across diverse data sets.
Model Assessment
The discrimination ability of the model was evaluated using the area under the receiver operating characteristic curve (ROC-AUC) and the area under the precision-recall curve (PR-AUC). 49 Confidence intervals (CIs) were estimated using bootstrap replicas. The determination of the cut-off between the binary targets (excessive hyperglycemia, hypoglycemia, and glycemic variability) is not deterministic. Hence, we calculated false positive predictions and true negative predictions for decision boundaries spanning from 1% to 100% of the cases for TAR, TBR, and CV (three CGM metrics widely used clinically) to provide a nuanced understanding of the model’s performance and clinical implications. To address the interpretability of the models, SHAP (SHapley Additive exPlanations) values 50 were used for explanatory purposes. All analyses were performed using MATLAB (R2021b), Python (v3), the Scikit-learn package (v0.23.2) for machine-learning utilities, the TabNet package (v0.1.6), SHAP package (v0.43.0), and the XGBoost package (v1.7.5).
Results
In this study, a cohort comprising 187 of 206 patients from the WISDM study was meticulously selected, contributing a total of 19 250 case weeks for the purpose of training and testing the prediction models. A supplementary validation data set was assembled, comprising 223 patients from the CITY and REPLACE-BG studies, with a cumulative case week count of 27 343 case weeks. The demographic and CGM characteristics of the enrolled patients are detailed in Supplementary Material S2 sTable 1, revealing statistically significant differences in age (mean = 18-68 years), diabetes duration (mean = 10-36 years), time in range (TIR) (median = 19%-63%), and TAR (median = 32%-56%) between the discovery cohort and the validation cohorts.
Model Performance
The models’ diagnostic performance characteristics are illustrated in Figure 2. Table 2 presents the ROC-AUC and PR-AUC values for each classifier and data set.
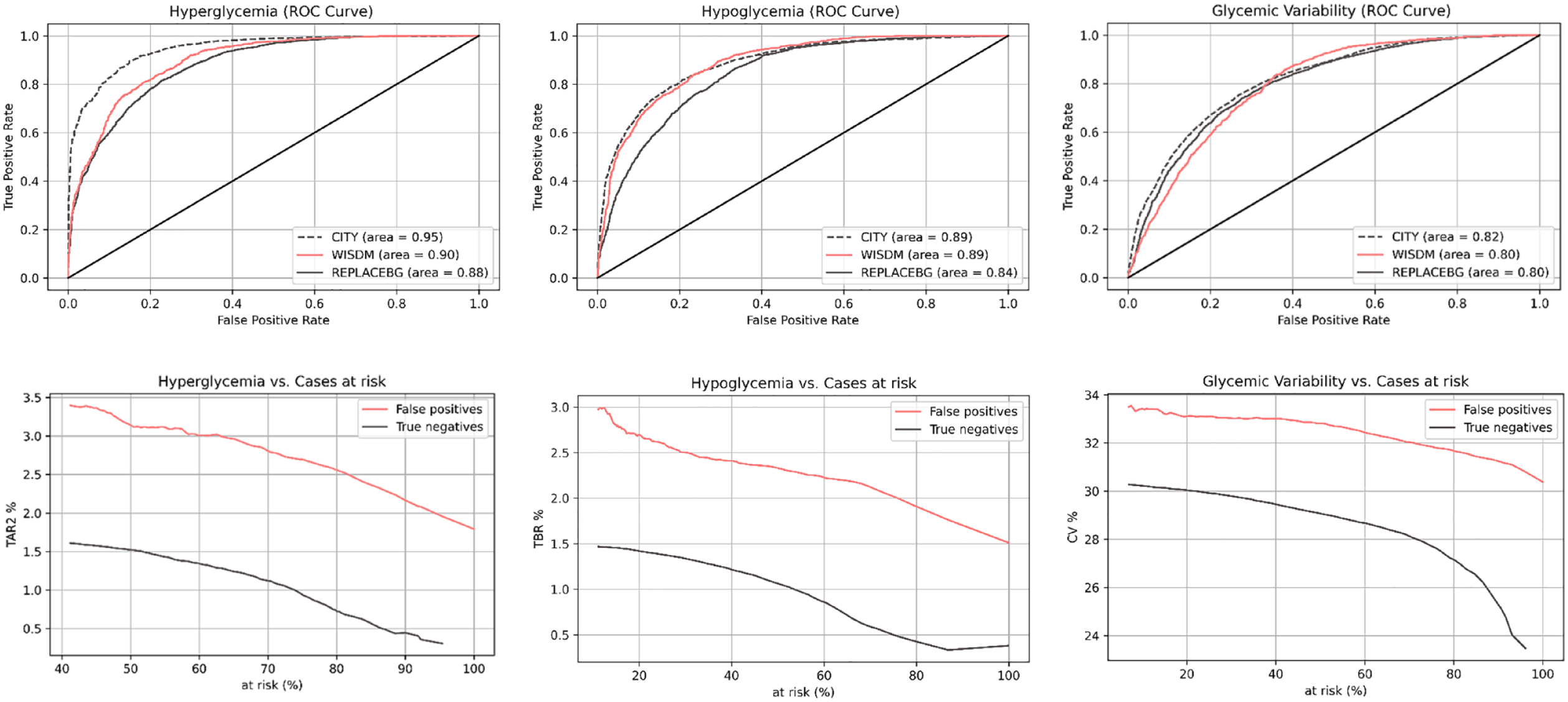
Receiver Operating Characteristic (ROC) curves for the prediction of excessive hyperglycemia, hypoglycemia, and glycemic variability (XGBoost models) (top panel). Data are shown for the three different data sets (CITY, WISDM, and REPLACE-BG). The corresponding at-risk models for each glycemic outcome are shown in the lower panels.
Diagnostic Performance Characteristics (ROC-AUC and PR-AUC) for Each Glycemic Target by Classifier Model.

Estimates are presented with 95% confidence intervals (CIs).
The XGBoost model exhibited ROC-AUC values of 0.9 [0.89-0.91], 0.89 [0.88-0.9], and 0.8 [0.79-0.81] for predicting hyperglycemia, hypoglycemia, and glycemic variability, respectively. Corresponding PR-AUC values were 0.9 [0.89-0.91], 0.72 [0.7-0.75], and 0.61 [0.58-0.63]. Notably, the utilization of nonlinear methods, such as XGBoost and TabNet, significantly improved performance in predicting future weeks with excessive hyperglycemia and hypoglycemia, with a more modest performance enhancement observed in the prediction of glycemic variability (0.8 vs 0.78 ROC-AUC; XGBoost vs logistic regression).
Moreover, the at-risk analysis (Figure 2) revealed that false-positive cases consistently exhibited poorer glycemic control (as evidenced by differences in TAR, TBR, and CV) compared with true-negative cases, irrespective of the chosen decision boundary and percentages of cases at risk. This suggests that false-positive cases could also be of clinical interest.
External validation
External validation of the model on the CITY and REPLACE-BG data sets confirmed the superiority of the XGBoost model performance compared with the other two models (P < .01).
For the CITY data set, the ROC-AUC was 0.95 [0.94-0.95], 0.89 [0.88-0.89], and 0.82 [0.81-0.82] for predicting hyperglycemia, hypoglycemia, and glycemic variability, respectively. The PR-AUC was 0.99 [0.99-1.0], 0.68 [0.66-0.69], and 0.83 [0.82-0.83] for predicting hyperglycemia, hypoglycemia, and glycemic variability, respectively.
For the REPLACE-BG data set, the ROC-AUC was 0.88 [0.87-0.89], 00.84 [0.84-0.85], and 0.8 [0.79-0.81] for predicting hyperglycemia, hypoglycemia, and glycemic variability, respectively. The PR-AUC was 0.91 [0.91-0.92], 0.73 [0.72-0.75], and 0.77 [0.75-0.78] for predicting hyperglycemia, hypoglycemia, and glycemic variability, respectively.
Feature Importance Analysis
The examination of feature importance, as depicted in Figure 3, revealed notable distinctions in the impact of individual features across the three prediction model targets investigated in this study.
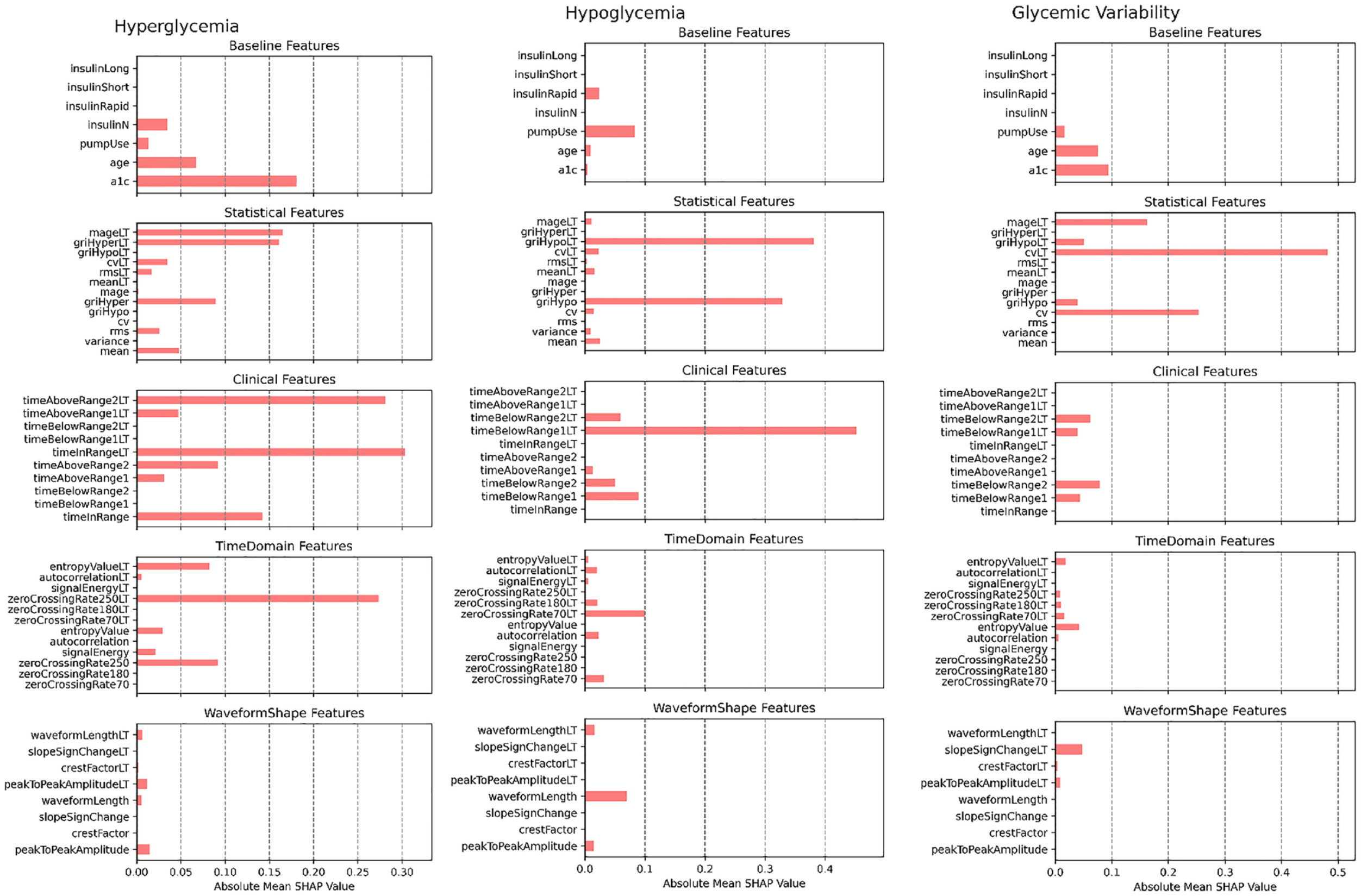
Feature importance for prediction of excessive hyperglycemia, hypoglycemia, and glycemic variability.
Hyperglycemia
In the context of hyperglycemia prediction, a diverse set of features spanning baseline, clinical, and time-domain categories played a substantial role. Specifically, previously documented hyperglycemia events, HbA1c levels, and zero crossing rate, serving as a measure of glycemic variability at the higher end of the glucose spectrum, emerged as pivotal predictors with significant impact on the model’s predictive performance.
Hypoglycemia
For hypoglycemia prediction, clinical features associated with both short-term and long-term hypoglycemia were of primary importance. Notably, the Glycemia Risk Index (GRI) hypo component, zero crossing rate at 70 mg/dL, and pump usage also contributed significantly to the predictive capacity of the model in this context.
Glycemic variability
The prediction of glycemic variability was predominantly influenced by statistical and clinical features. Previously documented glycemic variability emerged as the most crucial feature in this category. In addition, mean amplitude of glycemic excursion (MAGE), age, HbA1c levels, and occurrences of hypoglycemia were identified as contributing factors that enhanced the model’s predictive performance for glycemic variability.
Discussion
In this study, conducted within a cohort of individuals aged 60 years or older diagnosed with T1D, we trained and internally validated machine-learning models aimed at predicting the likelihood of experiencing excessive hyperglycemia, hypoglycemia, and glycemic variability in the ensuing week. Subsequently, external validation of these models was carried out using two independent cohorts of T1D patients, encompassing both adolescents and adults. The models demonstrated good generalizability across different age groups and settings.
The predictive models were constructed based on CGM dynamics observed in the preceding weeks, with the objective of forecasting the glycemic target of interest in the upcoming week. Notably, the final models exhibited robust discriminative capabilities, as indicated by AUC-ROC values ranging from 0.8 to 0.90. These findings were consistently validated in the external data sets, yielding AUC-ROC values ranging from 0.8 to 0.95.
The features influencing the predictive models varied depending on the glycemic target under consideration. Both clinically relevant and statistically significant CGM features demonstrated substantial discriminative power. It is noteworthy that baseline features, such as insulin type and HbA1c level, contributed only modestly to the additional predictive capabilities of the specified targets. In general, features related to the target had high predictive power, eg, hyperglycemia from the past 4 weeks had high predictive power for excessive hypoglycemia in the forthcoming week.
Moreover, our analysis revealed that beyond the binary discriminative ability of the models, false positives within the identified at-risk groups exhibited a more adverse metric for glycemic control compared with true negatives. This underscores the models’ capacity to effectively identify individuals at a clinically significant risk of experiencing either excessive hyperglycemia, excessive hypoglycemia risk, or increased glycemic variability.
Comparison to the Literature
We used a machine-learning–based modeling approach for predicting the risk of excessive hyperglycemia, hypoglycemia, and glycemic variability on a week-to-week basis using predictors derived from CGM and baseline patient characteristics, which, to our knowledge, has not been investigated before. Cichosz et al 51 recently published a study on the week-to-week prediction of only hypoglycemia, using only CGM data as input. This study expands the previous findings, showing that, in addition to the prediction of hypoglycemia risk, it is possible to predict week-to-week hyperglycemic excursions and variability. Also, increasing the number of weeks to include in the extraction of features significantly improves the predictability of the targets. The length of the sampling period has been shown to be associated with the predictive power of long-term glycemic control measured from CGM. 52
This study conducted a comparative analysis of three distinct model types to determine their predictive capabilities. The “simple” linear approach, as represented by logistic regression models, demonstrated commendable performance. Nevertheless, the adoption of nonlinear methods notably enhanced the overall performance of the model, aligning with findings reported in prior studies across diverse medical situations.53,54
Interestingly, the performance of the TabNet, a DNN model, consistently lagged behind that of the XGBoost tree-based model. This observation is in accordance with recent research by Shwartz-Ziv and Armon et al, 55 where a comparative analysis between XGBoost and TabNet similarly indicated a superior performance of the XGBoost model. These findings underscore the efficacy of nonlinear approaches, particularly tree-based models like XGBoost, in achieving enhanced predictive performance compared with certain DNN architectures such as TabNet or “simple” linear models such as logistic regression.
The approach we propose, centered on weekly risk prediction primarily derived from CGM data, holds the potential to empower patients, in collaboration with their health care providers, to implement proactive measures for the optimization of glycemic control. By providing advanced insights into the likelihood of encountering elevated risks of glycemic deterioration, our approach offers a valuable opportunity for patients to engage in preventative actions aimed at fostering more stable blood glucose regulation. The proactive involvement of both patients and clinicians in response to these predictions could contribute to a reduction in the frequency of hypoglycemic episodes, ultimately enhancing overall blood glucose management. One potential implementation involves integrating such predictive model into existing CGM user dashboards. Within these platforms, patients could be ranked based on either their current glycemic control status or the predicted control for the following week. This ranking system would enable health care providers, such as diabetes nurses, to efficiently identify patients at high risk of experiencing very poor glycemic control in the upcoming week.
A key consideration in the clinical utility of our predictive tool is how to effectively apply these models in practice, particularly when managing conflicting glycemic outcomes such as simultaneous risks of hyperglycemia and hypoglycemia. Therefore, we emphasize that this tool should not function as an automated decision-making system but rather as a component of a broader decision support framework. The models’ outputs should be interpreted in conjunction with other clinical data by health care providers, who can then tailor treatment decisions to the patient’s unique situation. This approach reinforces the need for clinician involvement in balancing complex glycemic targets and ensuring safe and personalized adjustments to therapies, such as insulin dosing, timing, and lifestyle interventions.
Strengths and Limitations
One notable strength of our study lies in the robustness achieved through the external validation of the model across expansive and diverse diabetes cohorts. The inclusion of large and heterogeneous cohorts mirrors the variations present in the broader T1D patient population, encompassing individuals from young to elderly age groups with varying risks of glycemic instability. In addition, a key strength is the high level of model explainability, facilitating a link between predictor variables and the predicted outcomes.
However, it is essential to acknowledge several limitations of our study. First, the generalizability of our findings is still constrained, particularly in the context of patients using closed-loop systems. The dynamics and variables associated with closed-loop systems introduce complexities that our model may not fully capture. Our observation of both an increase and a decrease in model performance during validation across the two independent data sets needs attention. This variability in performance may stem from divergent underlying glucose dynamics present in the distinct data sets. Consequently, a consideration for model calibration tailored to the specific characteristics of patient groups intended for use is advisable. This highlights the need for a nuanced approach to ensure the model’s applicability across diverse populations with varying glucose dynamics.
A notable limitation of this study is that the data were derived from clinical trial populations, which may not fully represent the broader population of individuals with T1D. These clinical trials apply strict inclusion and exclusion criteria, such as specific HbA1c cutoffs, which can limit the generalizability of the findings to real-world settings. For instance, patients with comorbidities such as liver or kidney failure, which are often exclusion criteria in trials, may not have been adequately represented in our sample. While we sought to address this by validating our results in a heterogeneous sample that included varying ages and A1c levels, this still does not fully capture the diversity of real-world populations. Therefore, future studies should focus on validating these findings in broader, real-world contexts to ensure the tool’s applicability across a wide range of clinical scenarios.
Even though the presented results seem promising, future prospective studies are needed to validate the finding in newer sensor technology, such as the Dexcom 7G/ Abbotts Libre 3 and to determine the potential effect and feasibility of implementing a clinical system for identifying patients at an excessive risk of reduced glycemic control.
Conclusion
We developed and externally validated machine-learning models to predict the weekly risk of hyperglycemia, hypoglycemia, and glycemic variability among people with T1D. The models were based on predictors derived from feature engineering on CGM data and baseline patient characteristics. We demonstrated that it is possible to use these data to forecast ensuing periods of reduced glycemic control. As such, our models could provide a potentially clinically useful tool to guide individual treatment.
Supplemental Material
sj-docx-1-dst-10.1177_19322968241286907 – Supplemental material for Explainable Machine-Learning Models to Predict Weekly Risk of Hyperglycemia, Hypoglycemia, and Glycemic Variability in Patients With Type 1 Diabetes Based on Continuous Glucose Monitoring
Supplemental material, sj-docx-1-dst-10.1177_19322968241286907 for Explainable Machine-Learning Models to Predict Weekly Risk of Hyperglycemia, Hypoglycemia, and Glycemic Variability in Patients With Type 1 Diabetes Based on Continuous Glucose Monitoring by Simon Lebech Cichosz, Søren Schou Olesen and Morten Hasselstrøm Jensen in Journal of Diabetes Science and Technology
Supplemental Material
sj-docx-2-dst-10.1177_19322968241286907 – Supplemental material for Explainable Machine-Learning Models to Predict Weekly Risk of Hyperglycemia, Hypoglycemia, and Glycemic Variability in Patients With Type 1 Diabetes Based on Continuous Glucose Monitoring
Supplemental material, sj-docx-2-dst-10.1177_19322968241286907 for Explainable Machine-Learning Models to Predict Weekly Risk of Hyperglycemia, Hypoglycemia, and Glycemic Variability in Patients With Type 1 Diabetes Based on Continuous Glucose Monitoring by Simon Lebech Cichosz, Søren Schou Olesen and Morten Hasselstrøm Jensen in Journal of Diabetes Science and Technology
Footnotes
Abbreviations
CGM, continuous glucose monitoring; CITY, CGM intervention in teens and young adults with T1D; CV, coefficient of variation; DKA, diabetic ketoacidosis; MAGE, mean amplitude of glycemic excursion; PR-AUC, area under the precision-recall curve; ROC, receiver operating characteristic; ROC-AUC, area under the receiver operating characteristic curve; SD, standard deviation; SHAP, SHapley Additive exPlanations; T1D, type 1 diabetes; TAR, time above range; TIR, time in range; WISDM, seniors with diabetes mellitus.
Authors’ Note
The source of the data is from the WISDM/REPLACE-BG/CITY trial, but the analyses, content, and conclusions presented herein are solely the responsibility of the authors and have not been reviewed or approved by trial group(s). During the preparation of this work, the author(s) used ![]() ’s “Curie” to improve grammar and language. After using this tool/service, the authors reviewed and edited the content as needed and take full responsibility for the content of the publication.
’s “Curie” to improve grammar and language. After using this tool/service, the authors reviewed and edited the content as needed and take full responsibility for the content of the publication.
Author Contributions
SLC had access to all data analyzed in this study. SLC takes responsibility for the integrity and accuracy of the study data analysis and results. SLC, MHJ, and SSO were involved in the study design, concept, analysis, and interpretation of the data. SLC drafted the manuscript and performed the statistical analysis. SSO and MHJ were involved in critical revision and preparation of the manuscript.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: The research was funded by i-SENS, Inc (Seoul, South Korea) and SLC’s involvement with the company did not influence the design, implementation, or interpretation of the study. SLC have received research funding from i-SENS, Inc (Seoul, South Korea), which manufactures some of the product types discussed in this paper. However, the study was conducted independently, and the authors declare that their involvement with i-SENS, Inc (Seoul, South Korea) did not influence the findings or conclusions of the study.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study was funded by i-SENS, Inc (Seoul, South Korea).
Ethics Approval
The presented study is a reanalysis of existing and anonymized data from the WISDM/REPLACE-BG/CITY clinical trials. The presented study in this paper did not need any approval from institutional and/or licensing committee, cf. Danish law on “Bekendtgørelse af lov om videnskabsetisk behandling af sundhedsvidenskabelige forskningsprojekter og sundhedsdatavidenskabelige forskningsprojekter” (Komitéloven, kap. 4, § 14, stk. 3). We confirm that all methods were carried out in accordance with relevant guidelines and regulations. The original WISDM/REPLACE-BG/CITY protocols and informed consent forms were approved by institutional review boards. Written informed consent was obtained from each participant prior to enrollment. An independent data and safety monitoring board provided trial oversight reviewing unmasked safety data during the conduct of the study.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
