Abstract
Background:
Pragmatic and easy-to-use alternatives to estimating body composition, such as lean body mass and fat mass, could be valuable tools for assessing the risk of diabetes or other metabolic diseases. Previous work has shown how demographic and anthropometric data could be used in a neural network to estimate body composition with high precision. However, there is still a need for a publicly available and user-friendly format before these results can have clinical impact.
Methods:
We used data from 18 430 NHANES participants and stepwise linear regression with inclusion of linear, interactions, and quadratic terms to model lean body and fat mass. HTML and Javascript was used to develop a webapp as a frontend of the model.
Results:
The models had a correlation cofficent R = 0.99-0.98 (P < .001) withstandard error of estimate [SEE] = 2.07-2.05.
Conclusions:
The results indicate that it is possible to develop a “white-box” model with high precision.
The proof of concept webapp is available as open source under the MIT license.
Introduction
Body mass index (BMI), a commonly and widely used measure of adiposity in clinical practice and research, has been criticized for its lack of ability to differentiate body composition.1,2 In individuals with the same BMI, fat mass (FM) and lean body mass (LBM) may be considerably different. Body composition with excessive FM is associated with an increased risk of morbidity, such as type 2 diabetes and cardiovascular disease.3-6 Estimating body composition is relevant in prediabetes and diabetes disease management, such as drug administration and risk assessment of morbidity/mortality. Several methods are available to obtain accurate estimations of the FM and LBM, such as DXA or imaging techniques.
7
Nevertheless, it is generally infeasible to use direct measures such as computed tomography (CT) and magnetic resonance imaging (MRI) due to cost, exposure to ionizing radiation, and availability, especially in larger research studies. Also, in places where a DXA is not available (eg, remote communities, developing countries) or for persons where it is not appropriate (eg, pregnancy, children), a need for an easy to access and inexpensive alternative is needed. To evade these limitations, an effort has been made to develop and test practical prediction models that can estimate body composition based on easily available data such as demographic and anthropometric data.
8
Recently, we published data, based on 18 430 adults and children from the US population, showing that it is possible to predict body composition with high precision using machine learning (neural networks).
9
In shorts, the results showed that there was a correlation of
However, the proposed model still needs further work before it can be used in clinical practice. Questions were raised regarding the use of a “black-box” model (artificial neural network), the potential for optimization in relation to practical usage, and the lack of a publicly available and user-friendly format. 10 We therefore sought to investigate the potential for further development of the concept such that a transparent, simple, and ready-to-use model could be released to the public domain. An easy-to-use algorithm using readily available and cheap measures may save both time, money, exposure to ionizing radiation, and increase availability in remote areas compared with say whole-body DXA.
Methods
The cohort used for modeling was derived from individuals who were enrolled in the cross-sectional study National Health and Nutrition Examination Survey 1999-2006 (NHANES). The study sample included participants aged 8 to 69 years who underwent a whole-body DXA scan examination, underwent a body measurement assessment (anthropometric assessment), and answered the demographic questionnaire. The cohort characteristics can be seen in Table 1 and are described in detail in a previous publication. 9
Characteristics of the Participants in the Training and Validation Data Set.
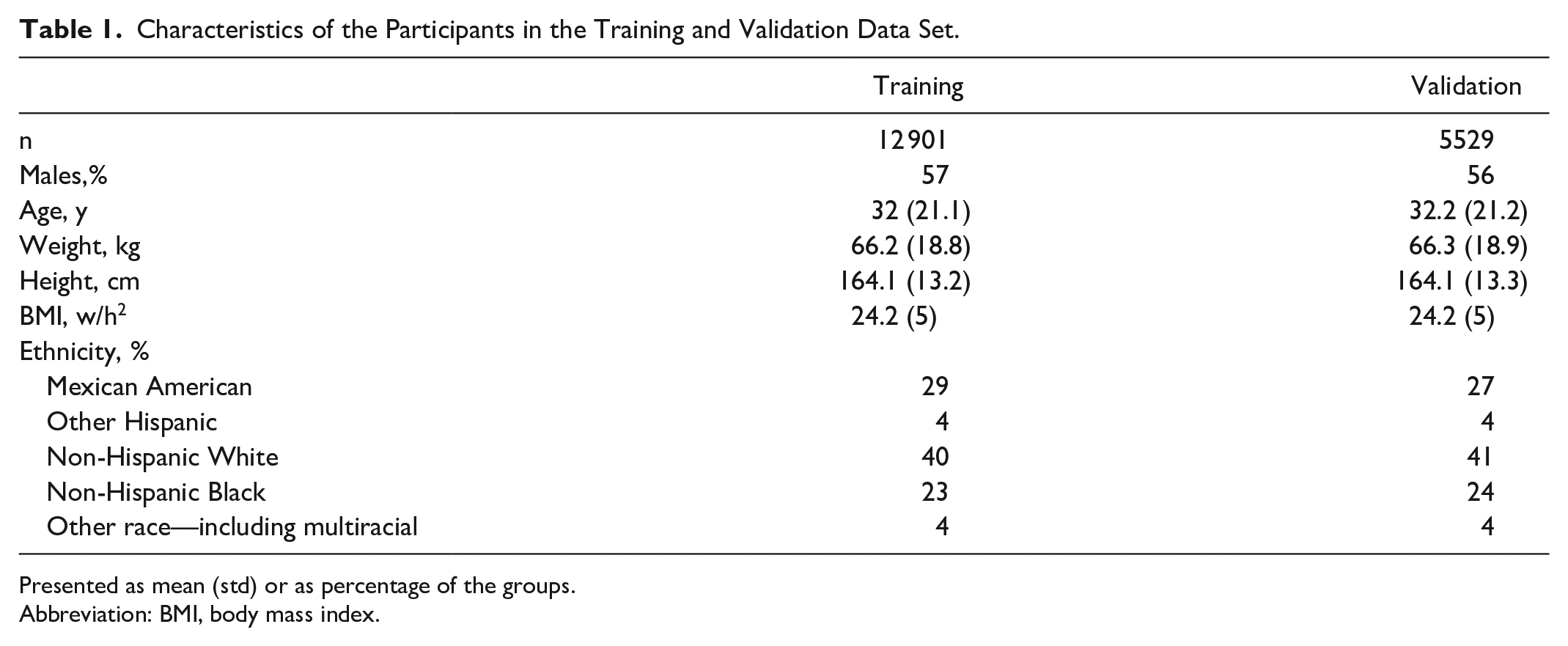
Presented as mean (std) or as percentage of the groups.
Abbreviation: BMI, body mass index.
The original study comprised 14 features for predicting body composition: age, sex, ethnicity, height, weight, BMI, upper leg length, maximal calf circumference, upper arm length, arm circumference, waist circumference, thigh circumference, triceps skinfold, and subscapular skinfold. To support the need for an applicable clinical prediction model with as few inputs as possible, for practical reasons, we included features that were shown to substantially improve the SEE. 9 The seven included features were sex, age, weight, height, BMI, waist circumference, and triceps skinfold.
The primary outcomes were predicted LBM in kilograms and body FM in kilograms compared with DXA-measured LBM and FM. The similarity between the predicted and measured LBM/FM was assessed using statistical measures: Pearson correlation coefficient (
Two models were trained on 70% of the cohort and tested on the remaining 30%: one model for predicting LBM and one model for predicting FM. To accommodate the request of a transparent model that could easily be implemented in practice, we choose to explore whether stepwise linear regression with inclusion of linear, interactions, and quadratic terms could solve the problem and keep some of the complexity of the artificial neural network, which leads to a precise prediction. For inclusion of terms, we choose a low
Proof of Concept Implementation
Furthermore, to make the prediction of body composition operable, we implemented the models in a simple and easy-to-use web app (MIT licensed). The code is published on GitHub, and a demo can be accessed, https://git.io/JRw3g. 12 To make the application easier to use, the input from BMI is calculated automatically from weight and height (weight [kg]/(height [cm]/100)2), and age is also automatically converted from years to months (mth = age [years]*12). This means that only six manual inputs are needed, and four of those require simple anthropometric measurements. A screenshot of the web app can be seen in Figure 1.
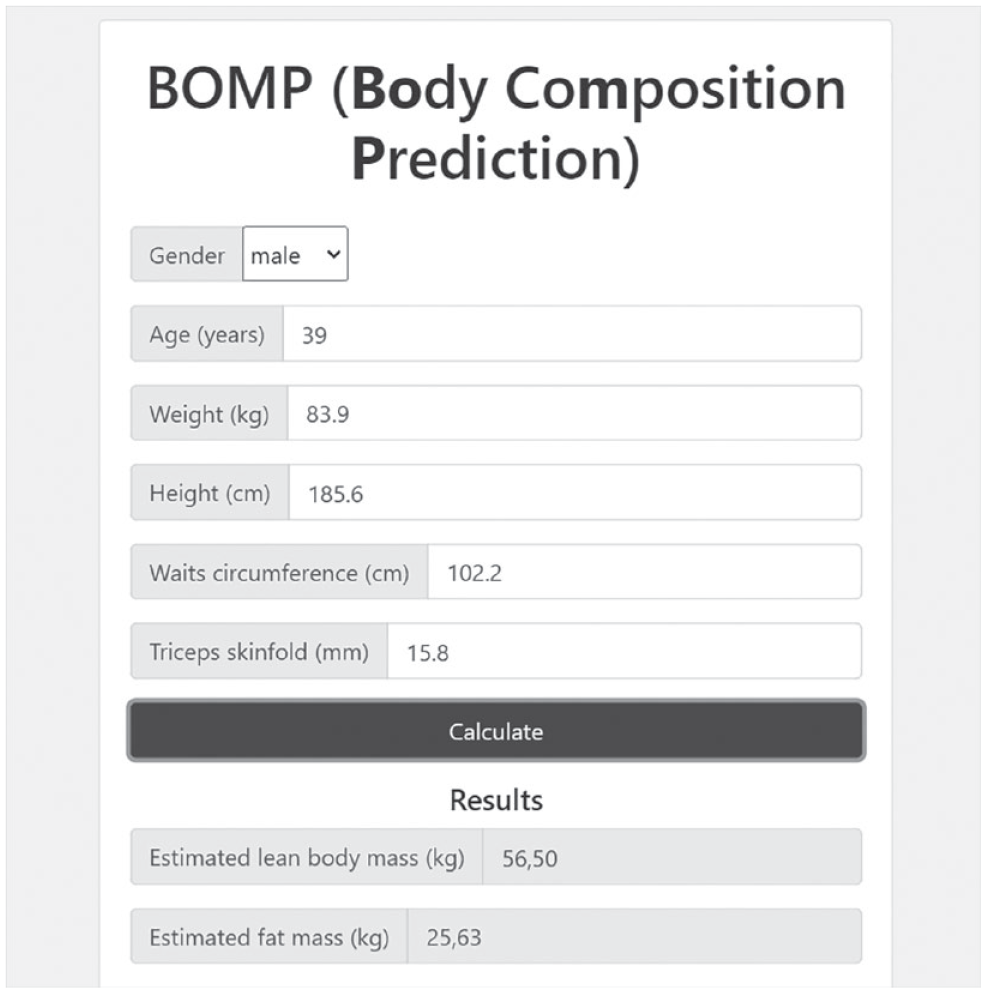
Screenshot from the implemented proof-of-concept web application.
Results
Participants were randomly partitioned on an individualized level into 70% training (n = 12 901) data and 30% validation (n = 5529) data. All seven features were included as linear terms; in addition, several interactions between the features and quadratic terms were also included in the finalized models. The model for LBM prediction had a correlation coefficient
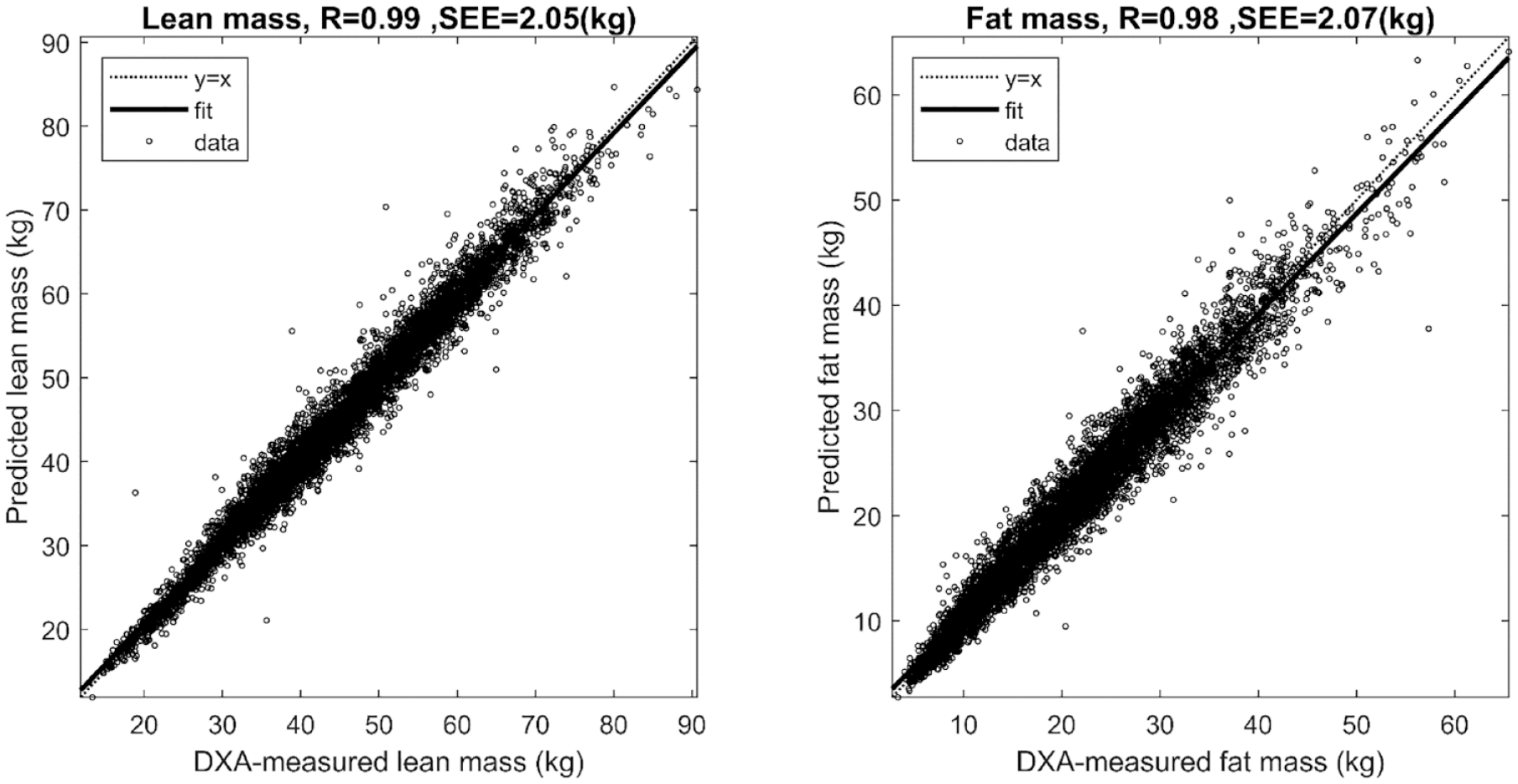
Regression plot for predicted lean mass and fat mass compared with DXA-measured lean and fat mass. Abbreviation: DXA, dual-energy x-ray absorptiometry.
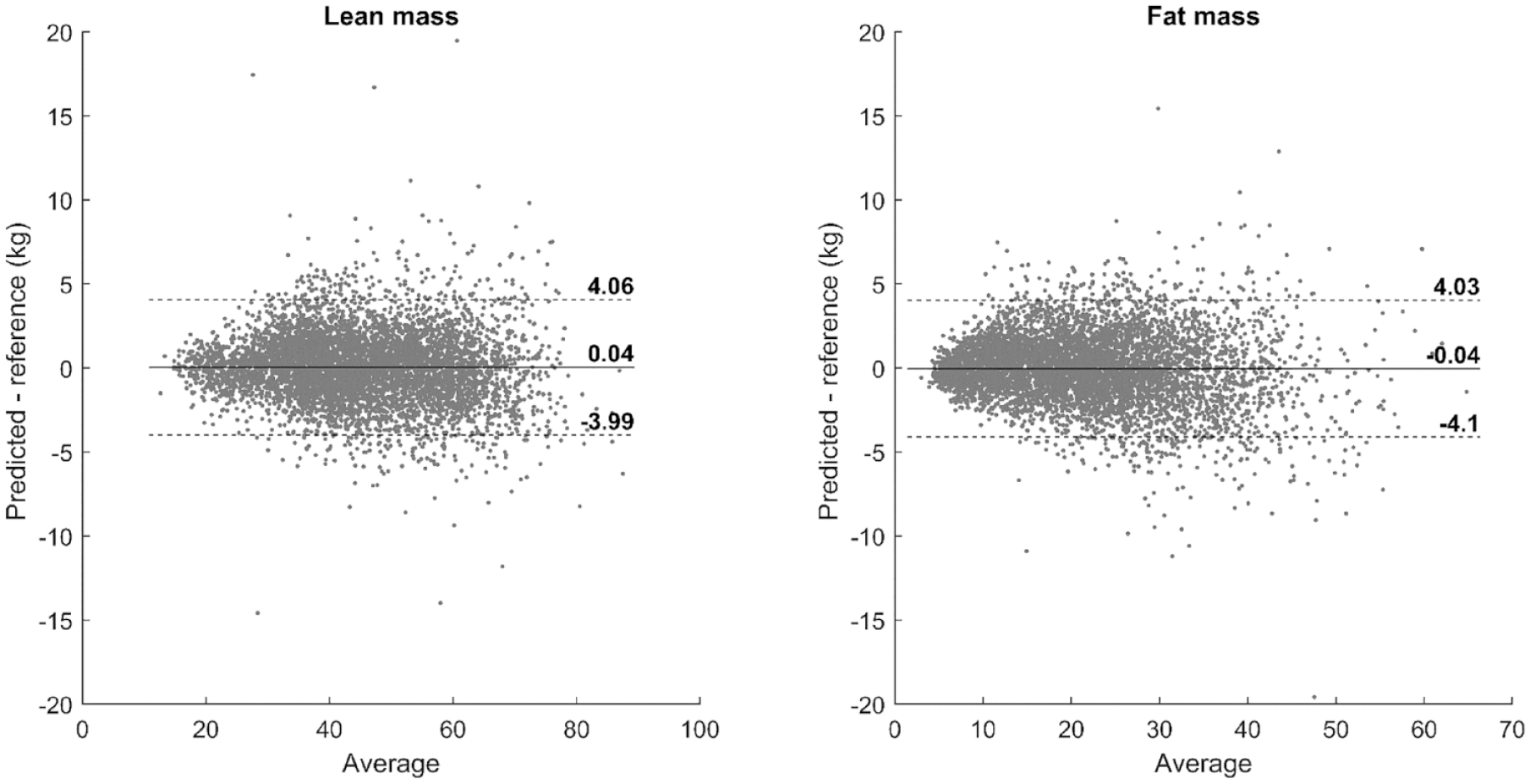
Bland-Altman plot for predicted lean mass and fat mass compared with DXA-measured lean and fat mass. The limits are mean ± 1.96SD. Abbreviation: DXA, dual-energy x-ray absorptiometry.
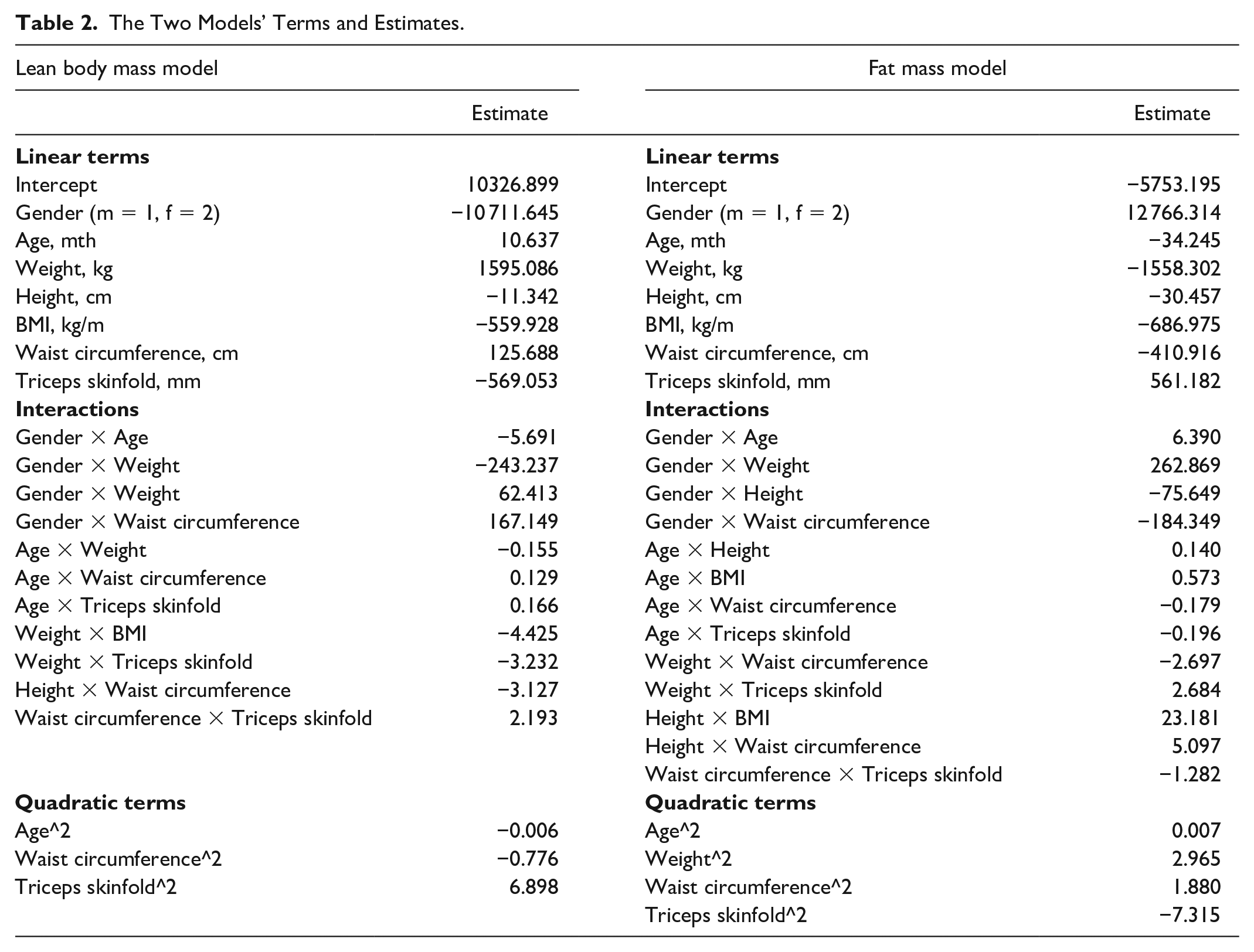
As an example, we can calculate the estimated LBM and FM from a male subject (39 years, weight of 83.9 kg, height of 185.6 cm, BMI 24.4 kg/m2, waist circumference(WC) of 102.2 cm and triceps skinfold measure of 15.8) by combining the terms in Table 2. This would yield an estimated LBM of 56.5 kg and FM of 25.6 kg. In comparison, the DXA-measured LBM is 57.3 kg and FM is 25.0 kg.
The Two Models’ Terms and Estimates.
Discussion
We developed a transparent, simple, and ready-to-use model with an additional web interface that was released to the public domain. The results show that it was possible to transformer the complexity of the previously used neural network 8 for the prediction of body composition into a transparent format using stepwise linear regression with linear, interactions, and quadratic terms. This transformation makes it straightforward to calculate the predicted LBM or FM from summarizing the terms in the equations. We made further steps toward the implementation of the prediction models in clinical practice by releasing a web interface 12 that can be accessed by a computer or mobile browser. It is now open to everyone to test and use the proposed models. The sample used to train and test the models was extracted from the NHANES and contains a representative and multiethnic cohort of people from the United States. However, using the predictions in another cohort outside the limits of the NHANES is of course encumbered with some uncertainty. Therefore, the models could benefit from additional validation in other countries. The results indicate that the predictions are robust in the obesity range. This is important as this is a major risk factor for developing type 2 diabetes mellitus. However, it could be valuable to test the models in estimating changes in body composition in longitudinal and interventional studies. With this work conducted, we may be able to estimate an individual’s risks of developing diabetes and other metabolic diseases using easy-to-conduct anthropometric measurements when advanced body composition methods, such as DXA scanning, are not available.
A limitation is that these are cross-sectional data. Prediction of body composition could potentially direct therapy in say type 2 diabetes toward reducing insulin resistance using say biguanides and physical activity in the case of high FM. In case of low FM, insulin deficiency and perhaps decreased muscle utilization of glucose may be a cause of the type 2 diabetes. Change in body composition over time could also aid in the monitoring of the effects of say physical activity and diet.
Footnotes
Abbreviations
MIT, Massachusetts Institute of Technology; FM, fat mass; LBM, lean body mass; BMI, body mass index; CT, computed tomography; MRI, resonance imaging; DXA, dual-energy x-ray absorptiometry; US, United States; NHANES, National Health and Nutrition Examination Survey; SEE, standard error of estimate.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
