Abstract
Background:
Estimating body composition is relevant in diabetes disease management, such as drug administration and risk assessment of morbidity/mortality. It is unclear how machine learning algorithms could improve easily obtainable body muscle and fat estimates. The objective was to develop and validate machine learning algorithms (neural networks) for precise prediction of body composition based on anthropometric and demographic data.
Methods:
Cross-sectional cohort study of 18 430 adults and children from the US population. Participants were examined with whole-body dual X-ray absorptiometry (DXA) scans, anthropometric assessment, and answered a demographic questionnaire. The primary outcomes were predicted total lean body mass (predLBM), total body fat mass (predFM), and trunk fat mass (predTFM) compared with reference values from DXA scans.
Results:
Participants were randomly partitioned into 70% training (12 901) data and 30% validation (5529) data. The prediction model for predLBM compared with lean body mass measured by DXA (DXALBM) had a Pearson’s correlation coefficient of
Conclusions:
In this study, neural network models based on anthropometric and demographic data could precisely predict body muscle and fat composition. Precise body estimations are relevant in a broad range of clinical diabetes applications, prevention, and epidemiological research.
Introduction
Muscle and fat are important compartments of the human body composition, as both a shortage and an excess of these can impact health and physical function.1,2 Body mass index (BMI) is a method to distinguish those with an increased risk of morbidity such as diabetes mellitus and cardiovascular disease, as the World Health Organization defines overweight as a BMI above 25 and obesity as BMI above 30.3-5 A BMI above 30 is a substantial risk factor for increased mortality and morbidity.6,7 It is a common measurement of obesity and widely used in clinics and research purposes. It reflects the total body mass, which consists of fat mass (FM) and lean body mass (LBM). The FM can further be divided into subcutaneous fat and visceral adipose tissue (VAT), the latter being categorized as the dangerous fat due to an association with an elevated risk of morbidity.8-10 Epidemiological studies have shown that the distribution of fat and the amount of visceral fat is related to increased disease risk in both children and adolescents. 11 The LBM is identified to have a significant influence on various physiological pathways, and people with reduced LBM may have negative health consequences.12-14 These parameters are dynamic and change due to aging, illness, and physical performance. Therefore, BMI does not accurately define the relationship between FM, LBM, and increased morbidity incidence.15,16 Moreover, the body composition is an important factor in drug dose calculations and administration, especially in obese patients with diabetes. 6
Even though several advanced methods are available to get accurate estimations of the FM and LBM, such as dual X-ray absorptiometry (DXA) or imaging techniques (ie, magnetic resonance imaging), 17 these approaches are expensive and often unfitting for daily medical diabetes management and analysis of broad populations. Therefore, several studies have been proposing estimation models based on anthropometric measurements as an alternative to BMI.18,19,20-29 However, these models are focused on simple statistical models usability such as estimations calculated by hand in clinical settings rather than optimizing the performance of the estimates. 15 Also, many studies have been targeting specific subpopulation such as young athletes, nonobese subjects, or elderly patients, and many of these studies included a low number of participants. 30 These circumstances limit the transferability.
With the increased availability of computers in the clinical setting, tasks as calculating estimates can easily be delegated to a computer or a smartphone. Moreover, methods from machine learning have shown the potential to make accurate predictions of clinical events.31-35 It is of high value to investigate how a state-of-art machine learning algorithm could perform in estimating body composition using simple patients’ anthropometric measurements.
This research had the aim of exploring the potential of developing a machine learning model to predict accurate estimates of LBM and FM from simple anthropometric measurements in a wide sample of the general population which could be useful for patients with both type 1 and type 2 diabetes, people at risk, and young adults.
Methods and Materials
Data Source
The cohort was derived from individuals who were enrolled in the cross-sectional study: National Health and Nutrition Examination Survey 1999-2006 (NHANES). The study includes a stratified multistage probability sample of the noninstitutionalized US population. The survey comprised of home interviews of participants followed by physical examinations and laboratory measurements. The Centers for Disease Control and Prevention National Center for Health Statistics institutional review board approved the original survey protocol, and all participants provided written informed consent. Further information about the NHANES can be found online.
36
This research meets the recommendations standards described by the “
Study Sample
The study sample included participants aged 8-69 years who had a whole-body DXA scan examination, had a body measurement assessment (anthropometric assessment), and answered the demographic questionnaire. Exclusion from DXA assessment was pregnancy, self-reported history of radiographic contrast material in recent past, and reported weight over 300 pounds or height over 65″ (DXA table limitations). The DXA examinations were assessed with a Hologic QDR-4500A fan-beam densitometer (Hologic Inc., Massachusetts), and the body measures were obtained by trained specialists during the physical examinations. Only participants with complete data on the DXA scan, body measurements, and demographics were included in the cohort. A total of 18 430 individuals were qualified to enroll in the cohort.
Predictive Features
The study dataset included two classes of predictors (ie, features) that are easily obtainable in a clinical setting: (1) demographic predictors were age, sex, and ethnicity; (2) anthropometric measurements were height, weight, BMI, upper leg length, maximal calf circumference, upper arm length, arm circumference, waist circumference, thigh circumference, triceps skinfold, and subscapular skinfold. A total of 14 features were investigated for their potential to predict the study outcome.
Outcome
The primary outcomes were LBM in kg, body FM (FM) in kg, and TFM in kg (as an indicator of VAT11,38). The predicted predLBM, predFM, and predTFM were based on machine learning with a combination of the features described. The predicted estimates were compared with the DXA scan results of DXALBM, DXAFM, and DXATFM in order to assess the accuracy of the predicted values.
Prediction Approach
The cohort was randomly divided into the training dataset, of which the prediction models were developed, and the validation dataset, to obtain an unbiased estimate of the model’s performance. The dataset used for training included 70% of the study sample, and the dataset used for validation consisted of the remaining 30%. We split our study sample randomly so that participants could not be in both datasets. The procedure is illustrated in Figure 1. We derived three machine learning models from the training data: all based on artificial neural network regression algorithms (ANN) using Levenberg-Marquardt as a training function with ten hidden nodes; one model for predicting predLBM, one for prediction of predTFM, and another model for predicting predFM. We chose ANN for deriving the prediction models as they have several advantages, that is, flexible input layer, capacity to indirectly identify dynamic nonlinear interactions between dependent and independent variables, capacity to identify all potential interactions between predictor variables, and have proven good performance in the medical field of prediction.39,40
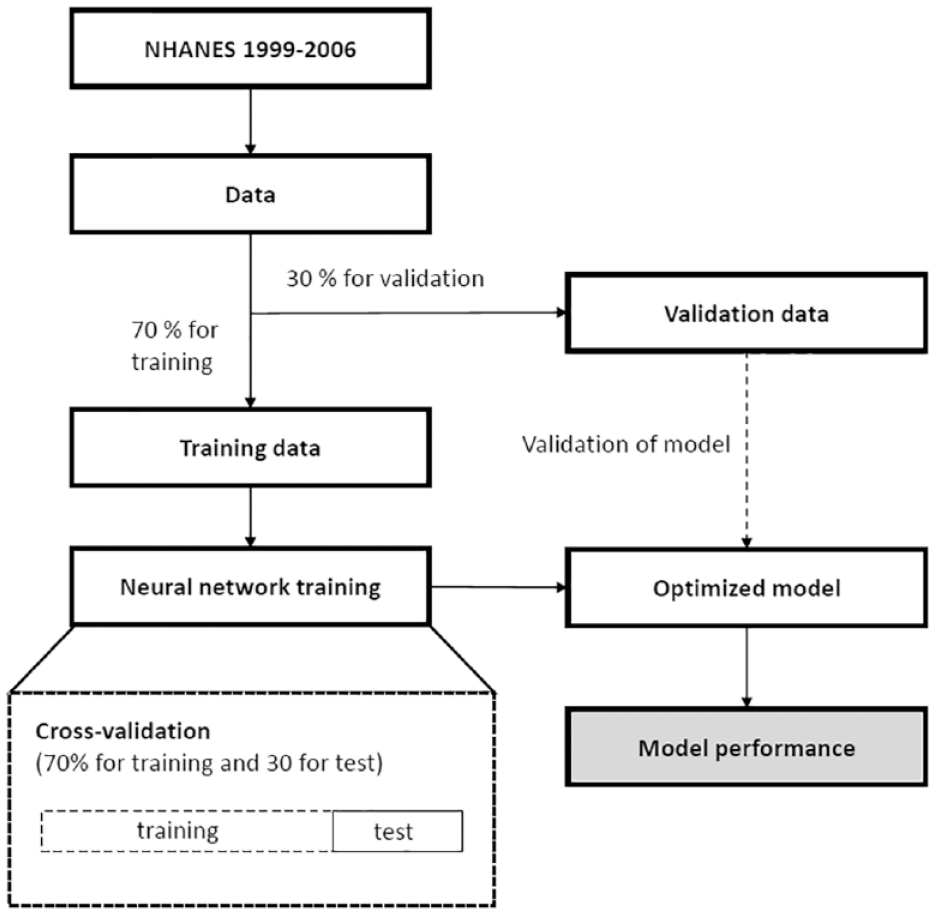
Illustrating the procedure for selecting the training dataset, validation dataset, and training the neural network prediction models using the training dataset. The training dataset is further divided into an actual training dataset and a test dataset to minimize the potential of overfitting the models.
To avoid the potential of overfitting the model to the training data, which would lead to decreased performance in the validation dataset (decreased generalizability), the training dataset was further split into 70%/30% for training and testing during the optimization of the models. This procedure minimizes the potential of overfitting the models. For transparency of potential overfitting, both results from the training dataset and the validation dataset are presented in the results. The ANN was implemented in Matlab R2016b (The Mathworks Inc., Natick, Massachusetts) using the Levenberg-Marquardt algorithm and with a hidden layer size of ten. The layer size was optimized using the training dataset. To investigate the performance of the features, the features were added to the model using forward selection and evaluated using the impact on total performance in the training dataset. Our study did not recalibrate the ANN models post the validation procedure.
Statistical Assessment
Statistical assessments were used to compare the predicted predLBM, predTFM, and predFM to the values obtained by the whole-body DXA scans. We calculated the Pearson correlation coefficient R for relations between predLBM/DXALBM, predTFM/DXATFM, and predFM/DXAFM, as well as
We also constructed a regression plot of the model predictions and DXA scan measurements. All analyses were implemented using the Matlab (vR2016b) (The MathWorks Inc., Natick, Massachusetts) software.
Results
Cohort and Study Population Characteristics
There were a total of 18 430 participants included in this study. The training and validation dataset contained 12 901 (70%) and 5529 (30%) participants.
The characteristics of the study population for the training and validation dataset are presented in Table 1. No substantial differences were seen between the training and validation dataset. A slight overrepresentation of males was included in the study cohort (57%). 62% of the cohort were adults (age above 18 years), and the composition of ethnicity was 40% white (non-Hispanic), 29% Mexican-American, 23%, black (non-Hispanic), and 8% other (including multiracial).
Characteristics of the Participants in the Training and Validation Dataset.
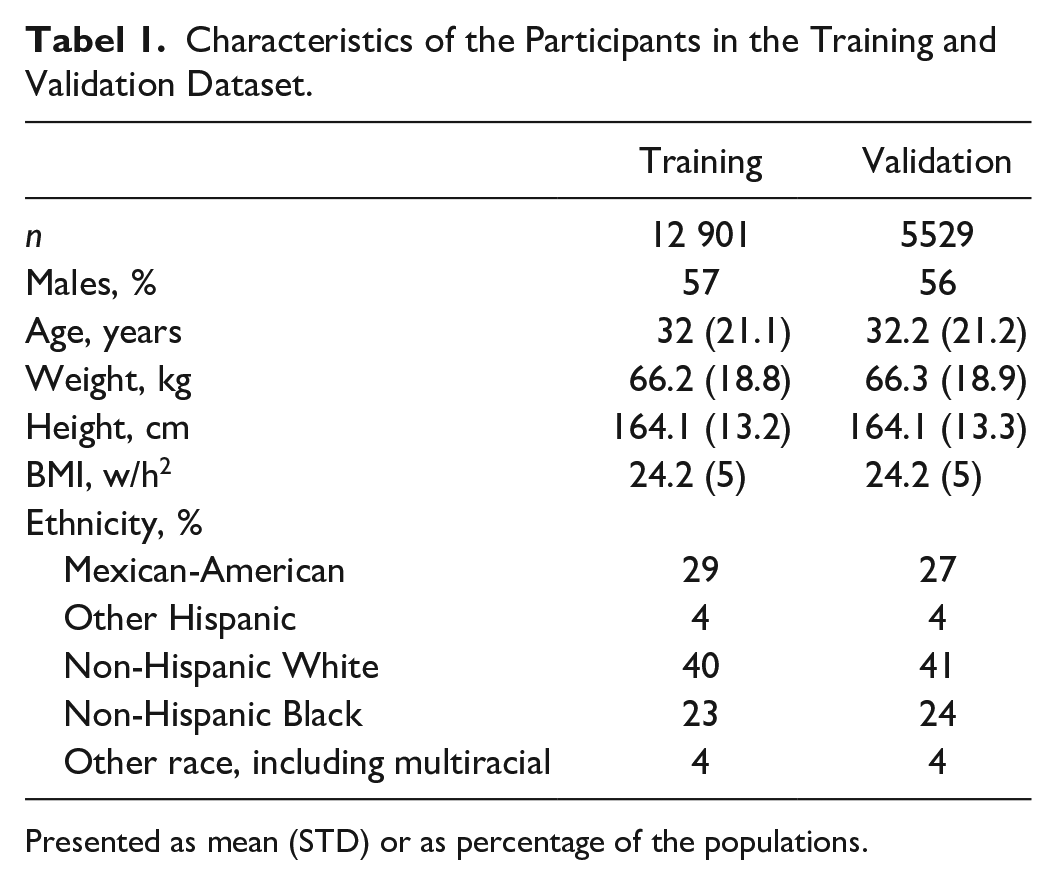
Presented as mean (STD) or as percentage of the populations.
Model Performance
In the validation dataset, the prediction model for lean body mass (predLBM) compared with LBM measured by DXA (DXALBM) had a Pearson’s correlation coefficient of
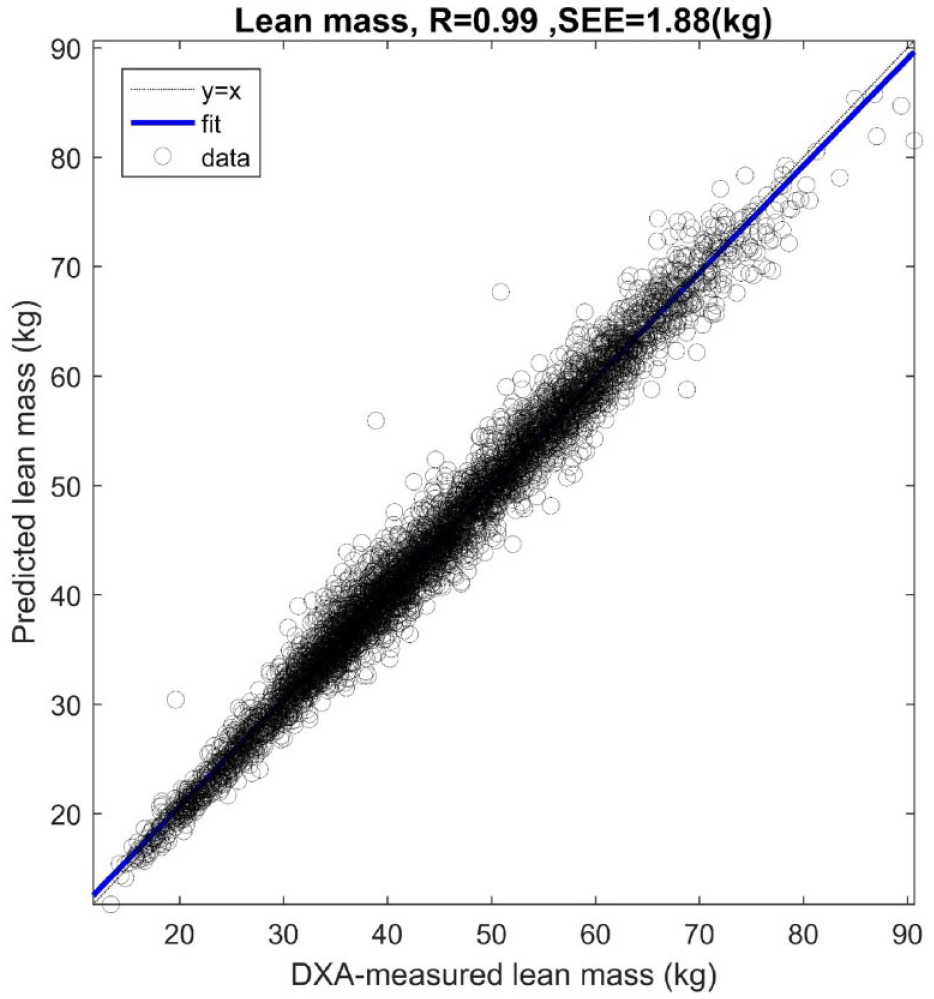
Regression plot of dual X-ray absorptiometry measured lean body mass and predicted lean body mass.
Performance of the Models in the Validation Dataset Among Subgroups in the Dataset.
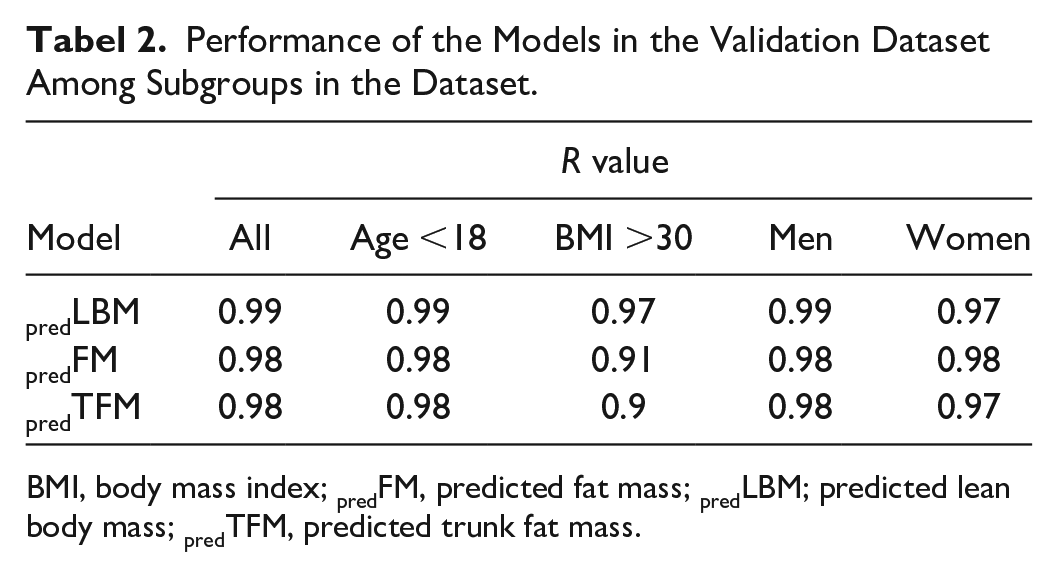
BMI, body mass index; predFM, predicted fat mass; predLBM; predicted lean body mass; predTFM, predicted trunk fat mass.
In the validation dataset, the predFM compared with FM measured by DXA (DXAFM) had a Pearson’s correlation coefficient of
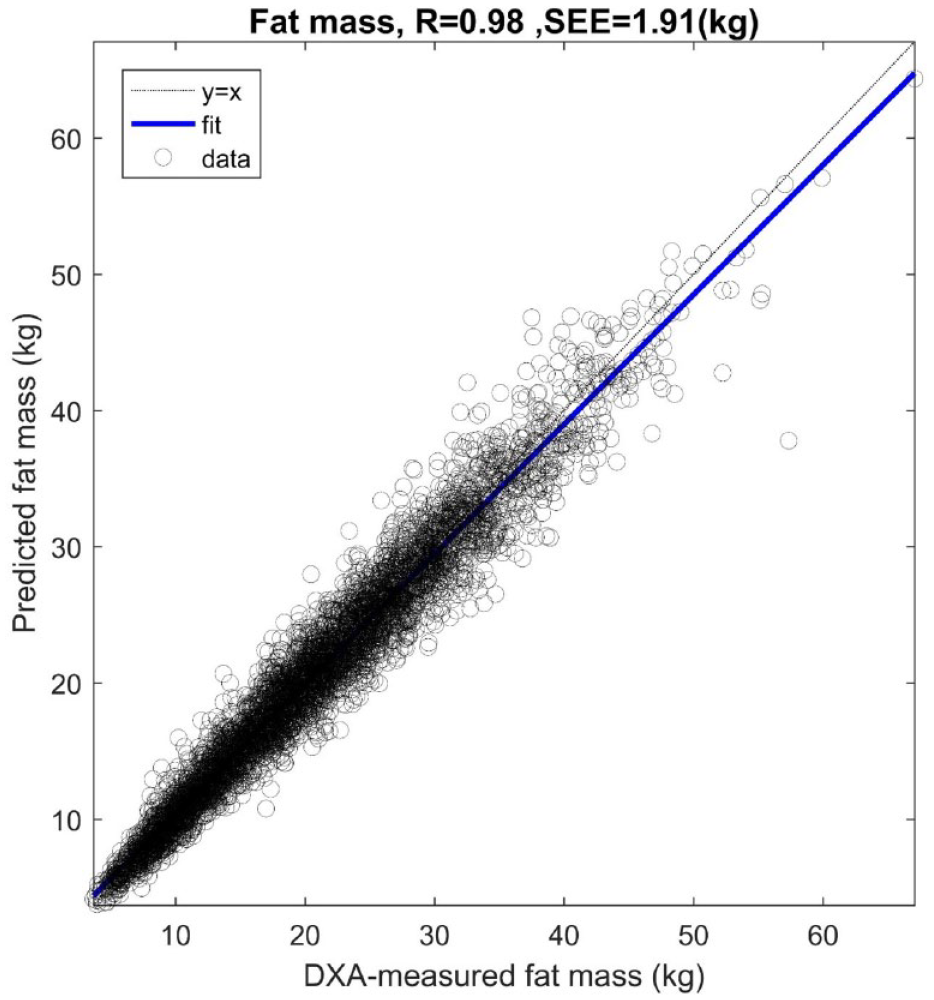
Regression plot of dual X-ray absorptiometry measured fat mass and predicted fat mass.
Moreover, in the validation dataset, the predTFM compared with TFM measured by DXA (DXAFM) had a Pearson’s correlation coefficient of
Algorithm Variable Importance
The importance of adding each feature into the model are presented in the supplement (eTable 2 in the Supplement) for predicting predLBM, in the supplement (eTable 3 in the Supplement) for predicting predFM, and in the supplement (eTable 4 in the Supplement) for predTFM.
For the model predicting predLBM, the features with the most influence on the SEE are: body weight, triceps skinfold, waist circumference, and gender—moreover, age, height, subscapular skinfold, BMI, arm circumference, ethnicity, and leg length also add to improve the prediction in a lower scale.
For the model predicting predFM, the features with the most influence on the SEE are BMI, triceps, waist circumference, weight, gender, age, and ethnicity. Adding calf circumference, thigh circumference, subscapular skinfold, arm length, arm circumference, and height further adds little predictive information.
Discussion
The present study was designed to develop whole-body LBM, FM, and TFM prediction models based on anthropometric dimensions and demographics information. With the use of whole-body DXA scanning as the reference, two prediction models were developed and then validated in an extensive and heterogeneous subject sample. Furthermore, the model’s performance was tested in subgroups of young people, men, women, and obese individuals. In general, the developed models had accurate predictive capability to assess LBM and FM. In obese participants, the prediction of FM was considerably reduced; this might be due to an increased interpersonal variation of fat distribution and difficulties in the accuracy of obtaining anthropometric measurements in obese people. 41
Several research studies have proposed anthropometric prediction models for the assessment of LBM and FM.18-29 However, most of these studies have been focusing on developing oversimplified prediction equations, have been targeting specific subpopulations, or have low numbers of participants. Kulkarni et al.
29
investigated a wide sample of Indian adult individuals (
A new study by Lee et al.
15
had the aim to develop simple anthropometric measurements to use calculation for prediction of LBM and FM in adults based on anthropometric from the NHANES 1999-2006. The proposed calculations included age, ethnicity, height, weight, and waist circumference. The results for predLBM were for men:
Study Strengths and Limitations
We demonstrated how a machine learning approach could be applied in predicting body composition using measures, which could easily be obtained in a clinical setting for the management of diabetes or an epidemiological study. The models were developed and validated from a wide representative sample of children, teens, and adults; multiple ethnic groups; and participants with large anthropometric diversity. Our separation of the data material into a training and validation dataset ensured that the potential of overfitting was minimized—this was further confirmed in the performance analysis between the training and validation dataset, which showed similar results.
However, there are several limitations to this analysis.
First, our data material was produced with a precise protocol for measuring participants’ anthropometrics. However, the bias in obtaining such measures is unavoidable even in a strictly and protocolized environment as the NHANES. Therefore, it is unexplored how this could influence the performance of the models when used in other settings.
Second, the reference used for LBM, FM, and TFM were measured using one type of DXA device—when developing our machine learning models, this was used as the ground truth. This means the models developed were produced with the aim of increasing accuracy compared with this ground truth. Interunit variability among DXA devices exists due to different causes, including the calibration procedure, calculation software, and variations in photon source intensities. 43 This could potentially influence the generalizability of our results (and others) to different settings and needs to be investigated in the future.
Third, our sample included multiple ethnic groups. However, some groups such as Asians and multiethnic groups were not well represented in our cohort.
Conclusion
This study describes the development and subsequent validation of three models for predicting the LBM, FM, and TFM from anthropometric and demographics information using a large representative sample. The approach was based on machine learning and with a focus on optimizing prediction accuracy. The results demonstrate that this approach could provide accurate estimates of the whole-body composition of FM and LBM in a heterogeneous population. This could potentially be a useful tool for the management of patients with both type 1 and type 2 diabetes as well as people at risk.
Supplemental Material
Supplement – Supplemental material for Precise Prediction of Total Body Lean and Fat Mass From Anthropometric and Demographic Data: Development and Validation of Neural Network Models
Supplemental material, Supplement for Precise Prediction of Total Body Lean and Fat Mass From Anthropometric and Demographic Data: Development and Validation of Neural Network Models by Simon Lebech Cichosz, Nicklas Højgaard Rasmussen, Peter Vestergaard and Ole Hejlesen in Journal of Diabetes Science and Technology
Footnotes
Author Contributions
Professor OH and SLC had access to all of the data analyzed in this study. OH and SLC take responsibility for the integrity and the accuracy of the study data analysis and results. SLC, NHR, PV, and OH were involved in the study design, concept, analysis, and interpretation of data. SLC drafted the manuscript and performed the statistical analysis. NHR, PV, and OH were involved in critical revision of the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
