Abstract
Background:
Abnormal glucose variability (GV) is a risk factor for diabetes complications, and tens of indices for its quantification from continuous glucose monitoring (CGM) time series have been proposed. However, the information carried by these indices is redundant, and a parsimonious description of GV can be obtained through sparse principal component analysis (SPCA). We have recently shown that a set of 10 metrics selected by SPCA is able to describe more than 60% of the variance of 25 GV indicators in type 1 diabetes (T1D). Here, we want to extend the application of SPCA to type 2 diabetes (T2D).
Methods:
A data set of CGM time series collected in 13 T2D subjects was considered. The 25 GV indices considered for T1D were evaluated. SPCA was used to select a subset of indices able to describe the majority of the original variance.
Results:
A subset of 10 indicators was selected and allowed to describe 83% of the variance of the original pool of 25 indices. Four metrics sufficient to describe 67% of the original variance turned out to be shared by the parsimonious sets of indices in T1D and T2D.
Conclusions:
Starting from a pool of 25 indices assessed from CGM time series in T2D subjects, reduced subsets of metrics virtually providing the same information content can be determined by SPCA. The fact that these indices also appear in the parsimonious description of GV in T1D may indicate that they could be particularly informative of GV in diabetes, regardless of the specific type of disease.
The likely effect of glucose variability (GV) on the increased risk of complications in diabetes1-6 has pushed researchers to propose a number of indices that can be used to quantify GV from either self-monitoring blood glucose (SMBG) or continuous glucose monitoring (CGM) time series. CGM profiles, in particular, because of their capability to capture components of glucose dynamic invisible even in the full 7-point SMBG profiles, are powerful tools to be exploited for the retrospective assessment of GV. Indices for GV quantification proposed in the literature include metrics quantifying the dispersion of glucose readings, the amplitude of glucose fluctuations, the risk of extreme glucose conditions (ie, hypo- and hyperglycemia), and the overall quality of glycemic control. We refer the reader to other sources7-11 for recent reviews. In this pool of available GV indices, however, several metrics have similar formulations or measure almost the same physiological quantity, with highly redundant information in the characterization of GV. Thus, a method capable of extracting a reduced subset of metrics still effective for describing GV in a specific population would be desirable.
In a recent work, 12 we have proposed the use of sparse principal component analysis (SPCA)—a technique introduced by Zou et al 13 —as an approach to select a subset of GV indices from a wider pool of metrics and provide a parsimonious description of GV in type 1 diabetes (T1D). As detailed in Fabris et al, 12 SPCA extracts a combination of a reduced number of GV indices that together preserve the majority of the variance of the initial wider pool of metrics; in terms of variance, GV indicators selected by SPCA are able to convey and reconstruct almost the whole information expressed by the original set of indices. In our previous work, 12 SPCA was successfully applied to a data set of CGM time series collected in 33 T1D subjects, selecting a reduced number of up to 10 GV indices able to span more than 60% of the original variance. The aim of the present article is to extend the application of SPCA to type 2 diabetes (T2D).
Materials and Methods
Database
The data set consists of CGM time series collected from 13 T2D males for an average period of 6 days under normal life conditions using the Medtronic® Guardian REAL-Time® CGM System (Medtronic, Northridge, CA). Mean ± SD demographic information of the data set is age, 54.4 ± 10.0 years; duration of diabetes, 10.7 ± 7.0 years; glycosylated hemoglobin, 7.4 ± 1.4 %; and body mass index, 29.4 ± 3.6 kg/m2.
The Original Pool of GV Indices
Twenty-five well-established indices for GV quantification were considered similarly to Fabris et al. 12 The pool of metrics includes mean and sample standard deviation (SD) of all glucose readings, percentage coefficient of variation (%CV), mean of daily SDs (SDw), SD of daily means (SDdm), J-index (a combination of mean and SD), 14 percentages of values below, within and above the euglycemic target range (70-180 mg/dl), 50th percentile, interquartile range (IQR), range of glucose readings, and the mean amplitude of glycemic excursions (MAGE) index.15,16 Moreover, measures derived from nonlinear transformations of glucose values quantifying the risk associated to a glucose profile were also evaluated: we considered low and high blood glucose indices (LBGI, HBGI),17,18 average daily risk range (ADRR), 19 and blood glucose risk index (BGRI); 11 hyperglycemic index, hypoglycemic index, and index of glycemic control (IGC); 8 the glycemic risk assessment diabetes equation (GRADE) score and the 3 contributions due to the different glycemic states, that is, %GRADEhypo, %GRADEeu, %GRADEhyper; 20 and, finally, the M100 index. 21 For all these indices, hypo and hyperglycemic thresholds, when appropriate, were set at 70 and 180 mg/dl, respectively.
The listed GV indices were evaluated on the CGM time series of the data set under analysis (the correlation matrix of the 25 metrics in the considered T2D population is reported in the Appendix to this paper). Then, before entering SPCA, each GV index was mean centered and scaled (ie, divided by its sample SD) to avoid any bias in the SPCA results and to facilitate the comparison of results obtained from different data sets. 12
SPCA
SPCA is a 2-step data processing technique of general applicability introduced by Zou et al. 13 Details on the implementation for our purposes can be found in Fabris et al 12 ; here, we report only a brief description.
Let
Step (a): apply the principal component analysis (PCA) technique
22
to the matrix
Step (b): apply the least absolute shrinkage and selection operator (LASSO) constraint 23 to each selected PC to obtain sparse estimates of regression coefficients and maintain in the PC regressor a reduced number of GV indices. Since PCs are linear combinations (regressions) of all the original variables, this will reduce the number of explicitly considered GV indices for each PC, still preserving a high percentage of the total original variance. The number of nonzero coefficients for each PC was determined so that the selected subset was of the same size as the one obtained in our previous work, 12 that is, we selected 5 GV indices for each PC, to get a final subset of 10 metrics.
Calculation of GV indices and implementation of SPCA were performed with software developed by us in MATLAB® environment.
Results
The
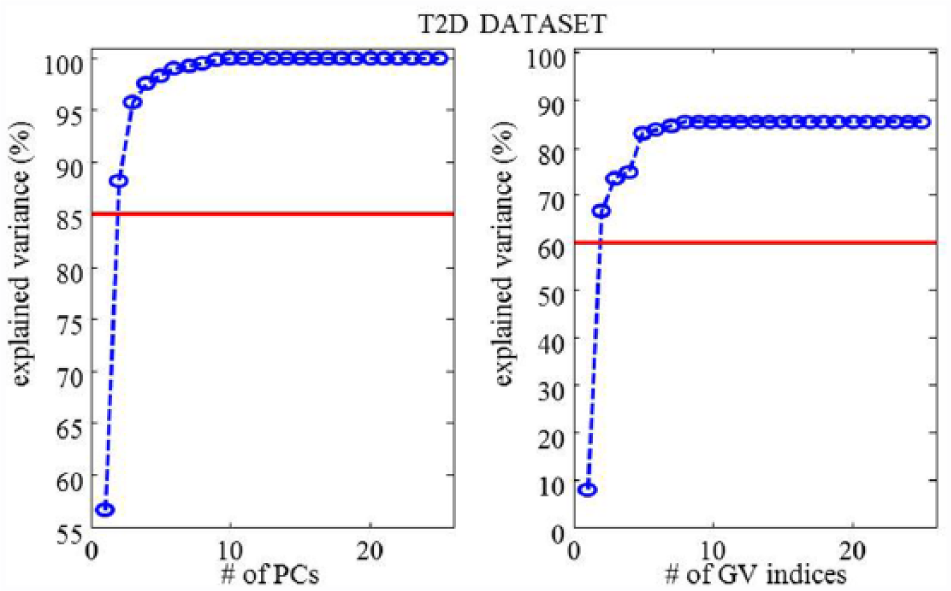
T2D data set—percentage explained variance as a function of selected PCs (left panel) and selected GV indices per PC (right panel).
T2D Data Set—GV Indices Selected by SPCA for Each PC.
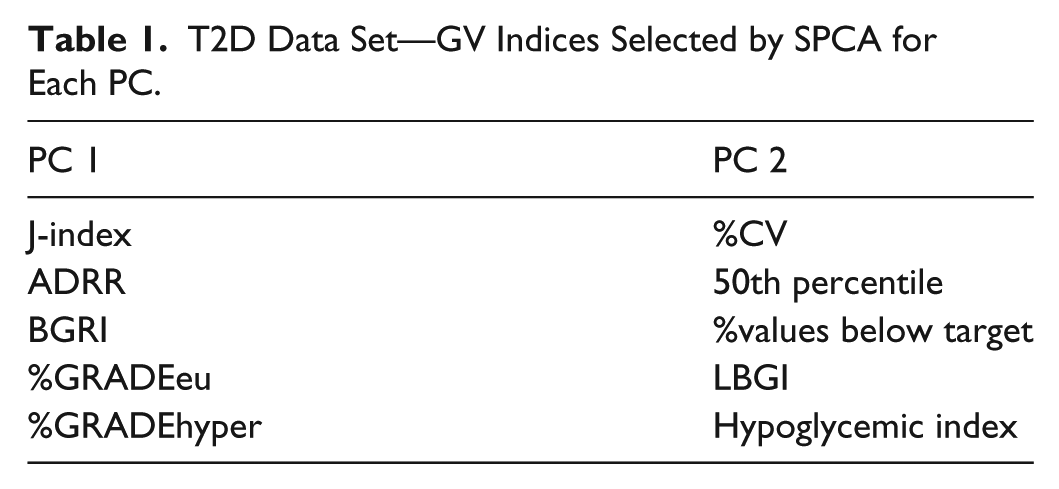

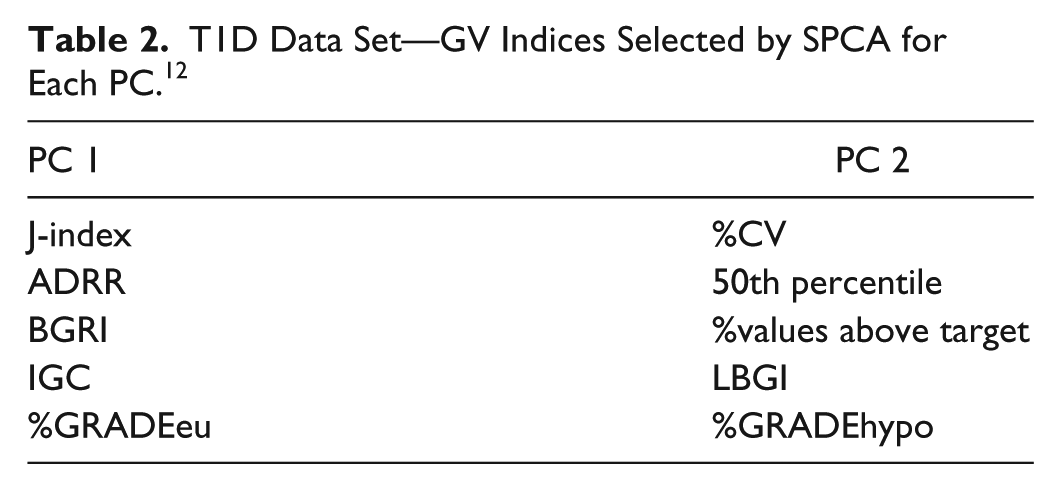
These results obtained in T2D are in line with those reported in Fabris et al, 12 where SPCA was applied to the same 25 GV indicators, evaluated on 33 CGM time series acquired from T1D subjects monitored for 4 days using the Dexcom® SEVEN® Plus CGM System (Dexcom, San Diego, CA, USA). To help the reader in the comparison, we report the plots used for the choice of SPCA parameters in Figure 2 and the set of indices finally selected in Table 2. Two out of 25 PCs were selected also for the T1D data set and 5 GV indices per PC were necessary to save more than 60% of the original variance (62% was finally saved). Interestingly, the 4 GV indices selected for the T2D data set are all included in the T1D parsimonious set, seeming to be particularly informative of GV in diabetes, regardless of the specific type of disease.
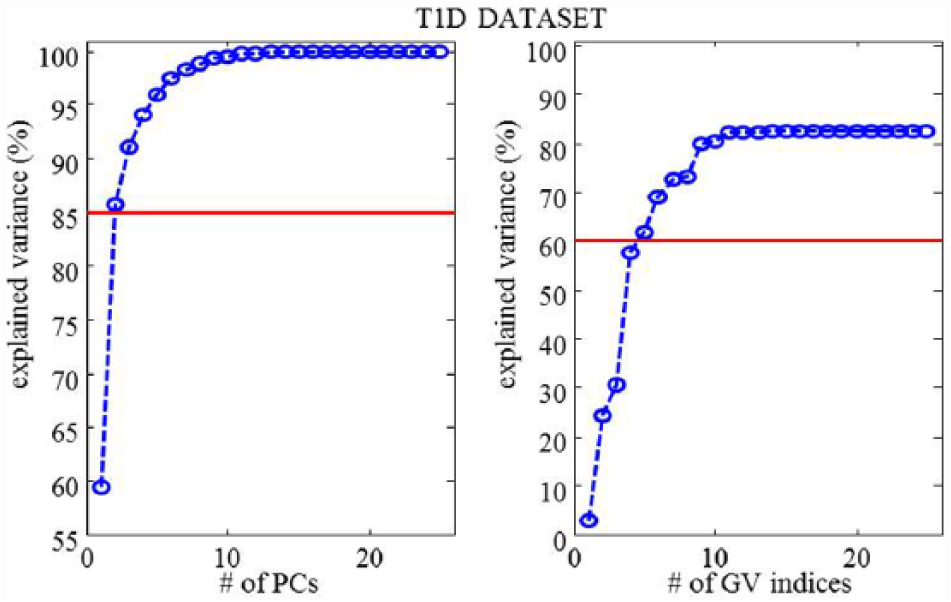
T1D data set—percentage explained variance as a function of selected PCs (left panels) and selected GV indices per PC (right panels).
T1D Data Set—GV Indices Selected by SPCA for Each PC. 12

Discussion
Robustness of SPCA Results
In the previous section, results obtained from the application of SPCA to 25 GV indicators evaluated on 13 T2D CGM time series have been documented. Nevertheless, no information has been provided about the robustness of what was achieved. This section will discuss this aspect, investigating if and how measurement noise and the low sample size data context that we have (less CGM signals than GV indices) influence the obtained results.
SPCA robustness to measurement noise can be theoretically proved by observing the intrinsic robustness to noise of PCA, which is indeed widely used in denoising applications. As detailed in Fabris et al 12 and mentioned in Section 2.3, PCs obtained from PCA are sorted by the amplitude of the covariance matrix eigenvalues. This means that the first PCs, as associated to the largest eigenvalues, are the most significant, while other PCs associated to small eigenvalues account mostly for noise. Selecting the first PCs allows to filter the noise affecting the data, and this is what we achieved in our application by selecting 2 out of 25 PCs. Beyond this intrinsic robustness to noise of PCA, to further prove that SPCA does not respond to noise, we performed 3 different perturbations of the original data set SPCA was computed on. Given that a direct perturbation of GV indices was not possible because of the correlation among them, we achieved variations of their values through a perturbation of the original (independent) CGM signals. Specifically, we (1) smoothed the CGM time series using a Savitzky–Golay filter, (2) added white noise with zero mean and 5% SD to the original CGMs, and (3) added white noise with zero mean and 10% SD to them. In all 3 scenarios, we got exactly the same results, with the same 10 GV indices selected and 83% of the original variance saved, confirming the robustness of SPCA to intrinsic or additional noise in our data.
The second technical aspect that deserves a discussion is the consistency of PCA, that is, the precision in estimating the PC coefficients, given the small number of available observations (
Clinical Interpretation of SPCA Results
SPCA results need also to be considered from a clinical viewpoint. Specifically, as already seen in Fabris et al, 12 we can observe from this analysis that GV can be reduced to a 2-dimensional phenomenon, well described by 2 combinations of indicators (either PCs or sparse PCs) identified through SPCA. A clinical interpretation of the meaning of each PC/sparse PC is not trivial, but we can observe that the first sparse PC, that is, the one describing the largest part of the variance of the original pool of GV indices, is mainly influenced by glucose control and hyperglycemia indices, while the second one, less significant in terms of variance than the first one, but still meaningful, accounts for hypoglycemia. This seems reasonable because hyperglycemic conditions may be more varying from 1 patient to the other than hypoglycemic ones, given the larger amplitude of the hyperglycemic range and the consequent much wider range of possible hyperglycemic scenarios. Of note is that the independence between the 2 PCs allows to capture and characterize the 2 extreme glycemic conditions separately.
Starting from the result that GV can be considered as a 2-dimensional phenomenon described by the first 2 PCs/sparse PCs, a clinically relevant usefulness of SPCA and its outputs can take shape. Specifically, the next step will consist in investigating if PCs and sparse PCs can be used for classification purposes, for example, to discriminate between groups of subjects affected by different metabolic disorders and between patients having different levels of quality of glycemic control. The development of a classifier based on PCs and sparse PCs is currently being conducted and preliminary results are promising.
Conclusions
GV is a risk factor for the development of complications from diabetes and tens of indicators for its quantification from CGM signals have been proposed in the literature. Despite the large number of available GV indices, a gold-standard metric to assess GV has not been identified yet, and a combination of indices is very likely to be needed to reliably characterize GV from glucose profiles.27,28 However, because some indices have similar mathematical formulations or measure almost the same physiological entity, considering all available GV metrics will provide highly redundant information and some indices could be of limited added value in the characterization of GV within a diabetic population. In this context, we have recently proposed a SPCA-based approach to select, from a pool of
Footnotes
Appendix
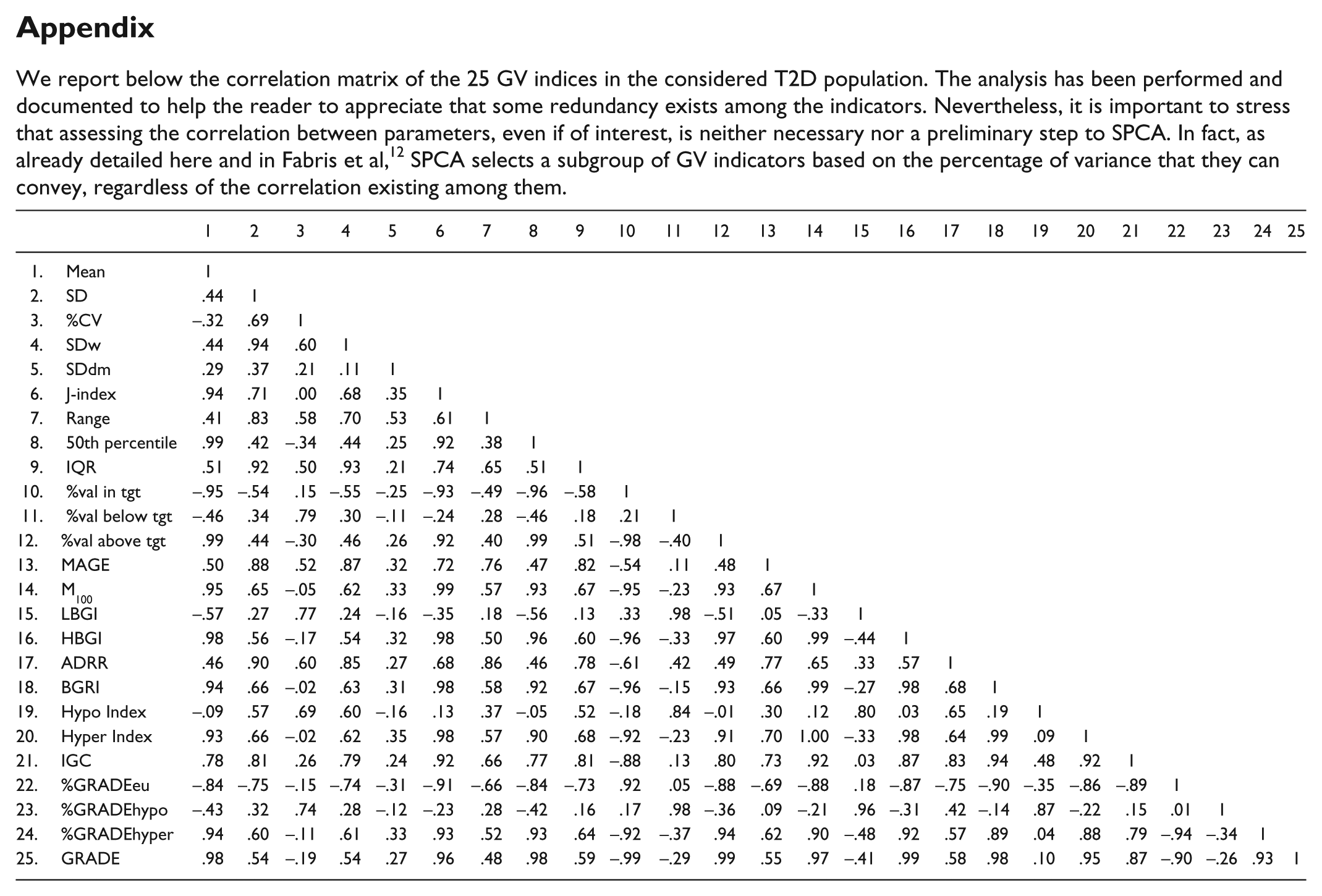
We report below the correlation matrix of the 25 GV indices in the considered T2D population. The analysis has been performed and documented to help the reader to appreciate that some redundancy exists among the indicators. Nevertheless, it is important to stress that assessing the correlation between parameters, even if of interest, is neither necessary nor a preliminary step to SPCA. In fact, as already detailed here and in Fabris et al, 12 SPCA selects a subgroup of GV indicators based on the percentage of variance that they can convey, regardless of the correlation existing among them.

| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 | 21 | 22 | 23 | 24 | 25 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1. Mean | 1 | ||||||||||||||||||||||||
| 2. SD | .44 | 1 | |||||||||||||||||||||||
| 3. %CV | –.32 | .69 | 1 | ||||||||||||||||||||||
| 4. SDw | .44 | .94 | .60 | 1 | |||||||||||||||||||||
| 5. SDdm | .29 | .37 | .21 | .11 | 1 | ||||||||||||||||||||
| 6. J-index | .94 | .71 | .00 | .68 | .35 | 1 | |||||||||||||||||||
| 7. Range | .41 | .83 | .58 | .70 | .53 | .61 | 1 | ||||||||||||||||||
| 8. 50th percentile | .99 | .42 | –.34 | .44 | .25 | .92 | .38 | 1 | |||||||||||||||||
| 9. IQR | .51 | .92 | .50 | .93 | .21 | .74 | .65 | .51 | 1 | ||||||||||||||||
| 10. %val in tgt | –.95 | –.54 | .15 | –.55 | –.25 | –.93 | –.49 | –.96 | –.58 | 1 | |||||||||||||||
| 11. %val below tgt | –.46 | .34 | .79 | .30 | –.11 | –.24 | .28 | –.46 | .18 | .21 | 1 | ||||||||||||||
| 12. %val above tgt | .99 | .44 | –.30 | .46 | .26 | .92 | .40 | .99 | .51 | –.98 | –.40 | 1 | |||||||||||||
| 13. MAGE | .50 | .88 | .52 | .87 | .32 | .72 | .76 | .47 | .82 | –.54 | .11 | .48 | 1 | ||||||||||||
| 14. M100 | .95 | .65 | –.05 | .62 | .33 | .99 | .57 | .93 | .67 | –.95 | –.23 | .93 | .67 | 1 | |||||||||||
| 15. LBGI | –.57 | .27 | .77 | .24 | –.16 | –.35 | .18 | –.56 | .13 | .33 | .98 | –.51 | .05 | –.33 | 1 | ||||||||||
| 16. HBGI | .98 | .56 | –.17 | .54 | .32 | .98 | .50 | .96 | .60 | –.96 | –.33 | .97 | .60 | .99 | –.44 | 1 | |||||||||
| 17. ADRR | .46 | .90 | .60 | .85 | .27 | .68 | .86 | .46 | .78 | –.61 | .42 | .49 | .77 | .65 | .33 | .57 | 1 | ||||||||
| 18. BGRI | .94 | .66 | –.02 | .63 | .31 | .98 | .58 | .92 | .67 | –.96 | –.15 | .93 | .66 | .99 | –.27 | .98 | .68 | 1 | |||||||
| 19. Hypo Index | –.09 | .57 | .69 | .60 | –.16 | .13 | .37 | –.05 | .52 | –.18 | .84 | –.01 | .30 | .12 | .80 | .03 | .65 | .19 | 1 | ||||||
| 20. Hyper Index | .93 | .66 | –.02 | .62 | .35 | .98 | .57 | .90 | .68 | –.92 | –.23 | .91 | .70 | 1.00 | –.33 | .98 | .64 | .99 | .09 | 1 | |||||
| 21. IGC | .78 | .81 | .26 | .79 | .24 | .92 | .66 | .77 | .81 | –.88 | .13 | .80 | .73 | .92 | .03 | .87 | .83 | .94 | .48 | .92 | 1 | ||||
| 22. %GRADEeu | –.84 | –.75 | –.15 | –.74 | –.31 | –.91 | –.66 | –.84 | –.73 | .92 | .05 | –.88 | –.69 | –.88 | .18 | –.87 | –.75 | –.90 | –.35 | –.86 | –.89 | 1 | |||
| 23. %GRADEhypo | –.43 | .32 | .74 | .28 | –.12 | –.23 | .28 | –.42 | .16 | .17 | .98 | –.36 | .09 | –.21 | .96 | –.31 | .42 | –.14 | .87 | –.22 | .15 | .01 | 1 | ||
| 24. %GRADEhyper | .94 | .60 | –.11 | .61 | .33 | .93 | .52 | .93 | .64 | –.92 | –.37 | .94 | .62 | .90 | –.48 | .92 | .57 | .89 | .04 | .88 | .79 | –.94 | –.34 | 1 | |
| 25. GRADE | .98 | .54 | –.19 | .54 | .27 | .96 | .48 | .98 | .59 | –.99 | –.29 | .99 | .55 | .97 | –.41 | .99 | .58 | .98 | .10 | .95 | .87 | –.90 | –.26 | .93 | 1 |
Acknowledgements
CGM data were kindly provided by Alberto Maran (University of Padova, Padova, Italy) for T1D and by Alejandra Guillén (Medtronic Iberica, Madrid, Spain), Giuseppe Fico, and Maria Teresa Arredondo for T2D, data collected within the framework of METABO, a research project funded by EU (grant agreement FP7-216270).
Abbreviations
ADRR, average daily risk range; BGRI, blood glucose risk index; CGM, continuous glucose monitoring; CV, coefficient of variation; GRADE, glycemic risk assessment diabetes equation; %GRADEhypo/eu/hyper, percentages of GRADE due to hypo/eu/hyperglycemia; GV, glucose variability; HBGI, high blood glucose index; IGC, index of glycemic control; IQR, interquartile range; LASSO, least absolute shrinkage and selection operator; LBGI, low blood glucose index; MAGE, mean amplitude of glycemic excursions; PC, principal component; PCA, principal component analysis; SD, standard deviation; SDdm, SD between daily means; SDw, SD within day; SPCA, sparse principal component analysis; T1D, type 1 diabetes; T2D, type 2 diabetes.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The MOSAIC project was funded by the EU within the 7th Framework Program (grant agreement FP7-600914)
