Abstract
Parkinson's disease (PD) involves degeneration of dopaminergic neurons and dysfunction across multiple neurotransmitter systems, contributing to both motor and cognitive impairments. Aerobic exercise improves clinical outcomes; however, its underlying neural mechanisms remain unclear. Using conventional resting-state fMRI combined with Receptor-Enriched Analysis of functional Connectivity by Targets (REACT), we examined molecular-enriched motor network changes following six months of supervised aerobic training in PD. Exercise-related connectivity changes were inversely correlated with baseline PD–healthy control differences, reflecting a partial normalization of PD-altered motor networks. Molecular-enriched analyses revealed selective effects on dopaminergic (FDOPA-enriched) and cholinergic (VAChT-enriched) related networks, with no changes observed in networks associated with serotonergic or noradrenergic systems. These findings provide supporting evidence for potential mechanistic links between aerobic exercise and network reorganization in PD, highlight multisystem effects, and illustrate the utility of molecular-enriched fMRI for probing neurotransmitter-specific interventions.
Plain language summary
Parkinson's disease (PD) affects multiple aspects of brain function including movement. Exercise can improve symptoms, but it is unclear how it changes the brain. In this study, we combined brain imaging (fMRI) with maps of key neurotransmitters to see how exercise affects brain networks in PD.
We studied 22 people with PD, including a subgroup who underwent six months of aerobic exercise, and compared them to healthy controls. Using a method called REACT, we looked at brain circuits linked to dopamine, acetylcholine, serotonin, and noradrenaline.
Our results suggest that exercise may help partially restore normal function in circuits disrupted by PD, especially those related to dopamine and acetylcholine. This study provides supporting evidence that exercise changes brain networks in a way that could improve symptoms and supports using molecular-informed imaging to understand these effects.
Introduction
Parkinson's disease (PD) is a progressive neurodegenerative disorder characterized by motor deficits and a broad range of non-motor symptoms. 1 The hallmark pathology involves degeneration of nigrostriatal dopaminergic neurons, but converging evidence points to widespread dysfunction in multiple neurotransmitter systems, including serotonergic, cholinergic, and adrenergic pathways. This widespread neurotransmitter involvement highlights the need for approaches that can probe multiple transmitter systems simultaneously, rather than focusing exclusively on dopaminergic dysfunction.2–4
Exercise has emerged as a promising intervention for PD. Large randomized controlled trials, such as Park-in-Shape and SPARX, demonstrate that structured aerobic training can improve motor outcomes and potentially slow functional decline. However, the underlying neurobiological mechanisms remain poorly characterized. Elucidating how exercise influences neurotransmitter-related brain networks could provide insight into therapeutic targets and support the development of imaging biomarkers to guide future interventions.5–7
Resting-state functional MRI (rs-fMRI) is a powerful, non-invasive tool for investigating large-scale brain network organization and has provided important insights into both healthy brain function and alterations associated with disease. However, while conventional rs-fMRI can detect functional connectivity changes, it cannot specify the underlying neurobiological systems driving these alterations—a key limitation when studying potential multi-system changes such as those seen in PD or following exercise.8,9
Multimodal approaches such as Receptor-Enriched Analysis of functional Connectivity by Targets (REACT) extend conventional rs-fMRI by integrating PET-derived molecular target maps to estimate functional networks weighted by neurotransmitter distributions. 10 REACT has been used to characterize receptor-specific network modulation during pharmacological challenges10–14 and in clinical populations,15,16 demonstrating its ability to detect alterations not visible with standard fMRI analyses. Applying this framework to PD provides a novel opportunity to probe potential biological networks involved in disease and/or exercise, thereby linking molecular distributions with functional network reorganization induced by aerobic training.
We hypothesize that aerobic exercise promotes modulation of motor-related functional networks in PD, shifting connectivity toward normalization, particularly in networks associated with dopaminergic, serotonergic, cholinergic, and adrenergic systems, which are all affected by disease and sensitive to physical activity in healthy individuals. 17 To test this, we applied complementary approaches: conventional seed-based connectivity analyses to assess striato-cortical circuits, and molecular-enriched REACT analyses to probe neurotransmitter-specific network modulation in a longitudinal cohort of PD patients undergoing six months of supervised aerobic exercise, compared with PD control patients and healthy controls (HCs).
Methods
Participants
Twenty-two individuals with PD and twenty-one age-matched HCs were recruited as part of a PET/MR study investigating brain energetics, approved by the UBC Clinical Research Ethics Boards (Application Nos. H19-01839 and H19-02892) and conducted in accordance with the Declaration of Helsinki. Written informed consent was obtained from all participants.
Participants were excluded if they had any chronic diseases (other than idiopathic PD in the patient group) or any conditions preventing MRI scanning at 3 T. PD participants were also required to have mild-to-moderate disease severity, defined as Hoehn & Yahr 18 stage 1–3. Additionally, participants were excluded if cognitive impairment was suspected. Individuals with Montreal Cognitive Assessment 19 scores below the conventional cutoff of 26 were retained when lower scores were attributed to non-pathological factors (e.g., age or language proficiency), consistent with evidence that optimal MoCA cutoffs vary across demographic and sociocultural contexts.20–23 No other explicit motor or non-motor score cutoffs (e.g., Unified Parkinson's Disease Rating Scale Part III 24 ) were imposed beyond these criteria. Eligibility for participation in the aerobic exercise intervention was subsequently confirmed using the PAR-Q + questionnaire 25 to ensure safe participation.
All participants underwent a suite of assessments and provided demographic information before scanning, which included:
Age Disease duration (calculated from the reported date of symptom onset) Levodopa equivalent daily dose (LEDD)
26
Hoehn & Yahr (H&Y) stage
18
Unified Parkinson's Disease Rating Scale Part III (UPDRS-III)
24
Montreal Cognitive Assessment (MoCA)
19
Beck Depression Inventory (BDI)
27
Rapid eye movement sleep behavior disorder screening questionnaire (RBDSQ)
28
Maximal oxygen uptake (VO2max)
29
VO2max was computed from oxygen respiratory data collected using the COSMED system 30 during a ramp protocol on a stationary cycle ergometer.
A subset of PD patients (n = 7, randomly selected from eligible participants) participated in a longitudinal six-month supervised aerobic exercise intervention, consisting of thrice-weekly, 60-min stationary cycling sessions. Exercise followed a standardized structure comprising a 10-min warm-up, 40 min of steady-state moderate-intensity cycling, and a 10-min cool-down. Sessions were held in a group setting led by a certified personal trainer over Zoom due to the COVID-19 pandemic. Exercise intensity was monitored using heart-rate data, which participants viewed in real time on their wearable devices, and which were reviewed after each session by the study team. Moderate intensity was further ensured through continuous conversational engagement during sessions, consistent with established guidelines for maintaining exertion within the moderate range.
An additional PD subgroup (n = 9, randomly selected from eligible participants) served as a disease control group and were instructed to maintain existing low-level exercise habits. Six participants were not followed longitudinally because they were already engaged in independent exercise programs at a level expected to attenuate or obscure exercise-related (or lack-of-exercise-related) effects. As such, they were not eligible for inclusion in the longitudinal intervention component of the study. However, because disease-related differences were expected to be larger in magnitude than exercise-related changes, these participants could still be included in the baseline PD vs. HC comparisons to increase statistical power without compromising interpretability.
All PD participants were scanned in the OFF-medication state, having withheld PD medication for at least 12 h prior to imaging. A full summary of the demographic and clinical characteristics of the participants is provided in Table 1. No formal a priori power analysis was performed, as the study was designed as a pilot investigation.
Demographic and clinical characteristics of study participants.
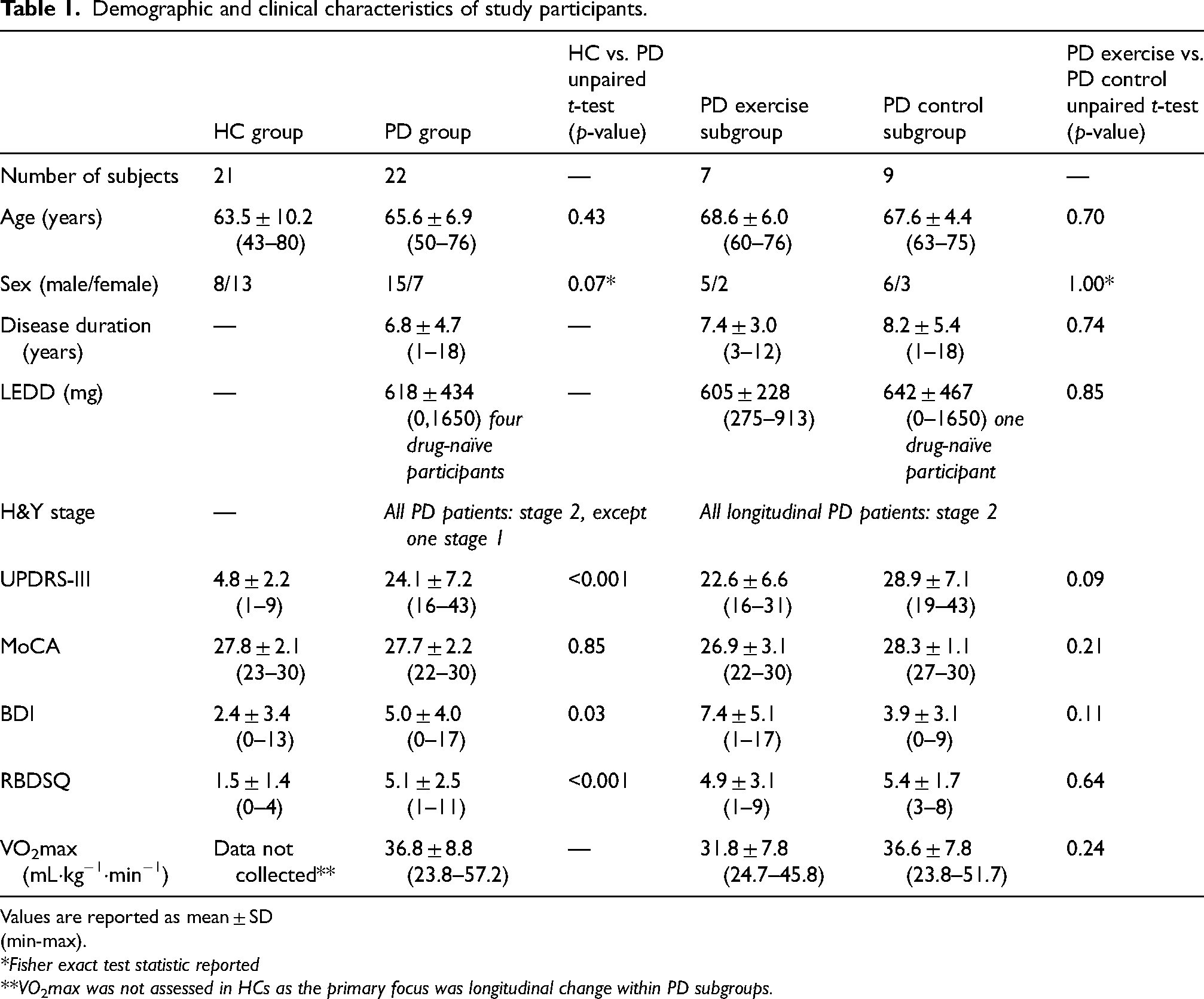
Values are reported as mean ± SD (min-max).
*Fisher exact test statistic reported
**VO 2 max was not assessed in HCs as the primary focus was longitudinal change within PD subgroups.
Consistent with prior studies of six-month exercise interventions in Parkinson's disease (e.g., Schenkman et al. 6 ), changes in clinical measures over this relatively brief intervention were generally small and often did not reach thresholds for meaningful improvement. Given the short duration, modest subgroup sample sizes, and the limited sensitivity of these clinical measures over six months, detailed tabulation of pre- vs. post-intervention changes were not included, as such reporting could risk overinterpretation of measurement noise rather than true intervention-related effects.
MRI acquisition
All imaging was performed on a GE SIGNA 3 T PET/MR scanner equipped with a 32-channel head coil. Participants in the longitudinal exercise and control subgroups underwent two MRI sessions (baseline and after the six-month period), while all other participants were scanned once. Resting-state fMRI data were acquired using a gradient-echo EPI sequence (TR = 3000 ms, TE = 20 ms, flip angle = 77°, voxel size = 2.5 × 2.5 × 2.5 mm3, FOV = 240 × 240 × 155 mm, 164 volumes). High-resolution structural images were acquired with a T1-weighted MPRAGE sequence (TR = 2700 ms, TE = 3.1 ms, TI = 1000 ms, flip angle = 8°, voxel size = 1 × 1 × 1 mm3, FOV = 256 × 256 × 202 mm) for use as an anatomical reference. All T1-weighted anatomical images were reviewed for major structural abnormalities, and no participants showed gross cerebrovascular lesions.
Preprocessing
Data were processed using FSL, 31 the CONN toolbox, 32 and MATLAB. 33 Preprocessing steps included realignment with correction for susceptibility interactions, slice timing correction, removal of the initial four volumes, outlier detection, direct coregistration to structural images, segmentation, normalization to MNI space, 34 and spatial smoothing using a 6 mm FWHM Gaussian kernel. Functional data were denoised by regressing out confounding effects of white matter, CSF, motion parameters and their derivatives, and linear trends, followed by temporal bandpass filtering (0.008–0.09 Hz).
Region-of-interest (ROI) parcellations were obtained using FreeSurfer's Desikan–Killiany atlas.35,36 FreeSurfer was run on each participant's structural image to generate subject-specific ROIs in native space, and separately on the MNI standard brain to define ROIs in MNI space. Due to the known progressive dopaminergic dysfunction in PD along the anterior–posterior axis, 24 the putamen was further subdivided into anterior, middle, and posterior portions of equal size using a k-means clustering approach applied to voxel coordinates. In total, 93 cortical and subcortical ROIs were defined. This set included only cortical and subcortical gray matter, excluding white matter regions. Mean time series were extracted from all subject-specific ROIs prior to any spatial smoothing, ensuring that signals were anatomically precise, whereas voxelwise MNI-space data were smoothed for group-level analyses. All preprocessing steps (including extraction of ROI time series) were performed using the CONN toolbox.
To account for asymmetry in motor symptom onset, PD patient scans were left–right flipped as necessary (performed using FSL's fslswapdim tool) such that the left hemisphere consistently corresponded to the more affected side, contralateral to worse motor symptoms. This was determined using lateralized motor UPDRS-III sub-scores.
Seed-based voxelwise connectivity analysis
Seed-based analysis was performed to assess connectivity between the left (more affected) posterior putamen, used as the seed region, and voxelwise targets in the cortical and cerebellar motor regions. The posterior putamen was selected given its strong involvement in sensorimotor processing and its well-established vulnerability to PD-related functional alterations, making it the striatal territory most relevant to motor impairment and recovery following exercise. These target regions were chosen for their central role in motor control, ensuring that the analysis focused on circuits most likely to exhibit disease- and exercise-related connectivity changes.
For each subject, the mean time series was extracted from their native-space posterior putamen ROI, providing a subject-specific definition of this small region. This time series was then Pearson correlated with voxelwise targets applied to the subject's functional data in MNI space, and the resulting correlation coefficients were Fisher transformed for group-level comparisons. Voxelwise targets were defined using an MNI-space mask comprising the primary and secondary motor cortices, supplementary motor area, primary sensory cortex, and cerebellum. The mask was expanded by one voxel in all directions to ensure complete coverage of the target regions across subjects, accounting for inter-subject anatomical variability and minor registration differences. Subject-level seed-to-voxel connectivity maps were computed using custom MATLAB code.
Group-level contrasts consisted of: (1) Effect of PD: baseline connectivity differences between PD and HC groups (PD–HC), (2) Effect of exercise: longitudinal connectivity changes in the PD exercise group (post–pre), and (3) Control group: longitudinal connectivity changes in the PD control group (post–pre). Conventional analyses were performed using FSL's randomise tool with 5000 permutations and threshold-free cluster enhancement (TFCE) for multiple-comparison correction. Statistical significance was considered at p < 0.05, family-wise error (FWE) corrected following TFCE.
To assess whether exercise-related connectivity changes aligned with baseline PD–HC differences, we applied a voxelwise regression-based pattern analysis. Unlike conventional voxelwise contrasts, which test each voxel independently and may be underpowered for distributed effects, this approach evaluates the full spatial pattern of PD-related alterations across the targeted motor system. This allows us to determine whether exercise-induced changes are specifically aligned with networks affected by disease, rather than being limited to isolated clusters.
For each subject, group-level post–pre changes in the PD exercise group were regressed against baseline PD–HC differences using orthogonal linear regression, which accounts for measurement error in both maps and provides a balanced estimate of the spatial relationship. To avoid regression-to-the-mean bias, data from PD participants contributing to each post–pre comparison were excluded from the corresponding baseline PD–HC map, under the assumption that the remaining baseline contrast is representative of a wider PD population.
To ensure that the observed pattern-level relationship between exercise-related connectivity changes and baseline PD–HC differences was robust and not driven by chance, permutation testing was performed by randomly shuffling group labels (PD exercisers and PD controls) across the longitudinal PD cohort. All 11,440 possible label assignments were exhaustively tested, generating a null distribution of correlation coefficients. Given our a priori hypothesis that exercise would normalize PD-related alterations—corresponding to a negative correlation—the observed effect was evaluated against the lower tail of this distribution to estimate its likelihood under the null hypothesis.
ROI-to-ROI-Based connectivity analysis
Seed-to-voxel and ROI-to-ROI analyses were used to provide complementary perspectives on motor network connectivity. The seed-to-voxel approach enabled voxelwise assessment of connectivity patterns originating from the posterior putamen, allowing visualization of distributed patterns of connectivity across motor-related circuits. In contrast, the ROI-to-ROI analysis summarized connectivity between predefined striatal and cortical regions, providing network-level quantification of pairwise functional connectivity strengths.
For the ROI-to-ROI analysis, striatal seeds consisted of the caudate and the anterior, middle, and posterior putamen, while targets consisted of the primary and secondary motor cortices, supplementary motor area, primary sensory cortex, and cerebellar motor regions. Mean time series for both seeds and targets were extracted in each subject's native space prior to any spatial smoothing to minimize partial volume effects, thus preserving anatomical specificity for small subcortical regions. Pairwise connectivity was quantified using Pearson correlation, and correlation coefficients were Fisher transformed for group-level comparisons.
Connectivity strengths were compared across groups using the same contrasts as in the voxelwise analyses: unpaired t-tests for the baseline PD–HC comparison, and paired t-tests for longitudinal changes in the exercise and control groups, with Bonferroni correction applied across the multiple connections. In addition, the regression-based pattern analysis framework was applied to the ROI-to-ROI connectivity matrices to test whether exercise-related changes aligned with baseline PD–HC differences at the network level, with specificity confirmed via permutation testing. ROI-to-ROI correlation matrices and significance testing were generated using custom MATLAB code.
REACT analysis
Molecular-enriched fMRI analysis was performed using the REACT method with six PET-derived molecular target maps representing the dopaminergic, serotonergic, cholinergic, and adrenergic systems: D2, DAT, FDOPA, 5-HTT, VAChT, and NAT. These population-based atlases were generated from independent PET datasets in healthy volunteers and normalized to MNI space.37–43 To standardize spatial resolution across maps and facilitate ROI-based analysis, all maps were regridded to 2 mm isotropic, which also aligned with the MNI-space FreeSurfer parcellation. The cerebellum was excluded from all PET maps, as it served as a reference region in the kinetic modeling of several tracers. All remaining 91 cortical and subcortical ROIs were included in the ROI-level REACT analysis, ensuring coverage of the whole brain outside the cerebellum. A custom script was implemented in MATLAB to perform the REACT analysis.
To address potential redundancy across molecular maps, variance inflation factors (VIFs) were calculated for all six PET-derived molecular regressors. Maps with VIF > 5 were iteratively removed, and VIFs were recalculated until all remaining regressors met this threshold. This procedure resulted in four molecular maps—one per neurotransmitter system—ensuring stable and interpretable model estimates. Sensitivity analyses including all six maps produced qualitatively similar results, which are reported in the Supplementary Materials.
The REACT framework was implemented using custom MATLAB code in two stages (see Dipasquale et al. for additional methodological details).
10
Extraction of molecular-enriched time series: Each molecular map was entered as a spatial regressor in a subject-level GLM of resting-state fMRI, after demeaning and z-scoring. fMRI signals were extracted as ROI averages in native space, while PET map values were averaged within the corresponding ROIs in MNI space. This ROI-level implementation improved stability for small subcortical regions and ensured equal weighting across ROIs regardless of their anatomical size. Estimation of molecular-enriched connectivity maps: The resulting molecular time courses were then entered as temporal regressors in a second GLM of the rs-fMRI data to generate subject-specific molecular-enriched functional connectivity maps. The molecular time series and fMRI ROI time series were demeaned and z-scored prior to regression, ensuring comparable scaling across regressors and improving model stability.
The REACT pipeline is illustrated in Figure 1.
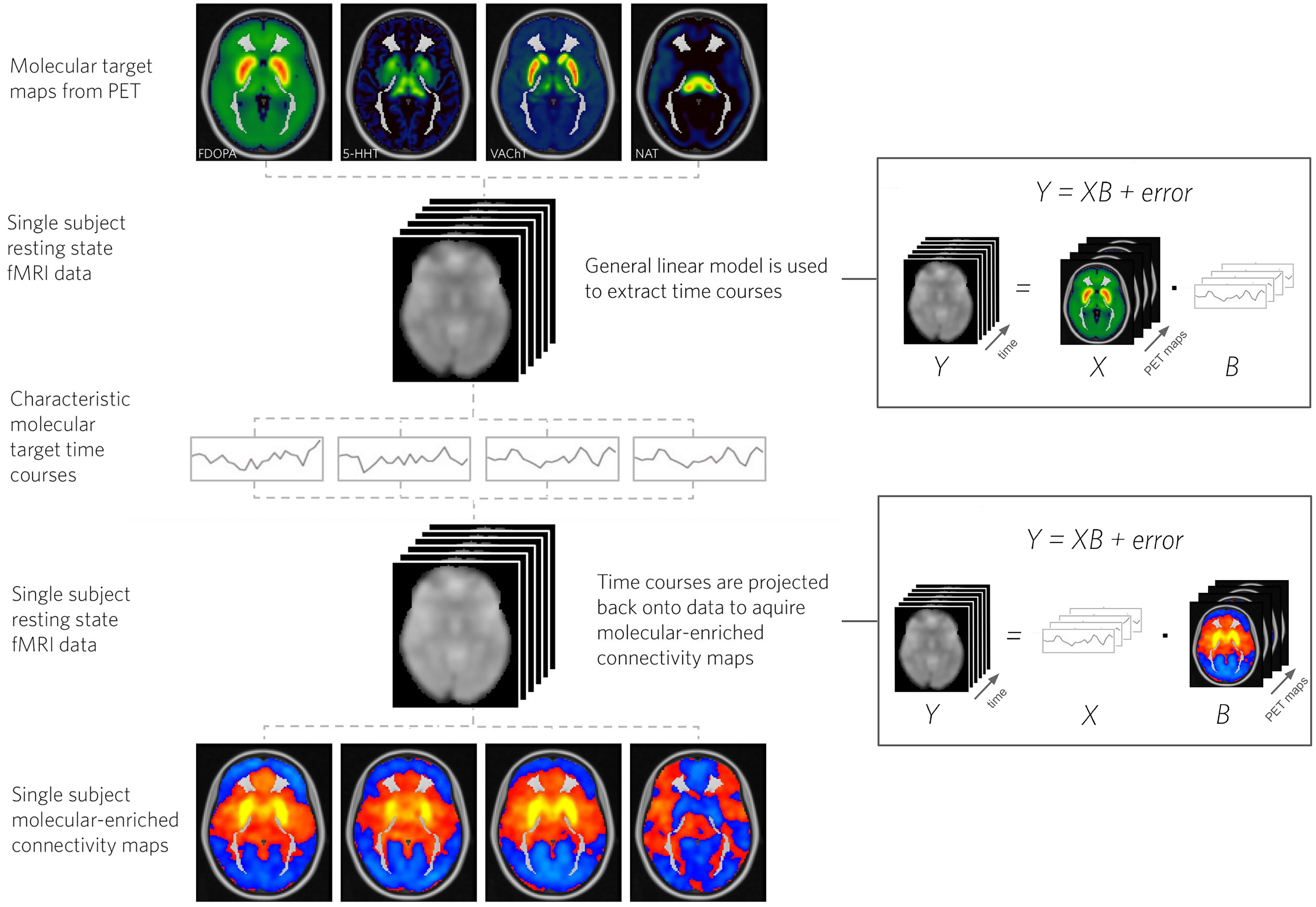
Multimodal PET-fMRI REACT pipeline. Molecular target maps are used to extract characteristic molecular target time courses from the rs-fMRI data. These time courses are then used to acquire molecular-enriched connectivity maps.
Group-level contrasts mirrored the seed-to-voxel analyses and were tested using FSL's randomise with 5000 permutations and TFCE, with Bonferroni correction applied across the molecular maps. The same regression-based voxelwise pattern analysis framework was then applied to each map to assess whether exercise-related changes tracked baseline PD–HC differences, with specificity confirmed via permutation testing and Bonferroni correction used for multiple comparisons.
Results
Seed-based voxelwise connectivity analysis
Conventional voxelwise contrasts did not yield clusters surviving TFCE correction; however, uncorrected maps (Figure 2) suggested altered striato-motor connectivity patterns in PD at baseline, with broadly opposite changes following exercise. To capture distributed effects that voxelwise analyses may miss, regression-based pattern analysis was applied, confirming a significant relationship across voxels: exercise-related changes were negatively correlated with baseline PD–HC differences (r = –0.246, p = 0.041; Figure 3), indicating that exercise shifted connectivity in a direction opposite to disease-related alterations.
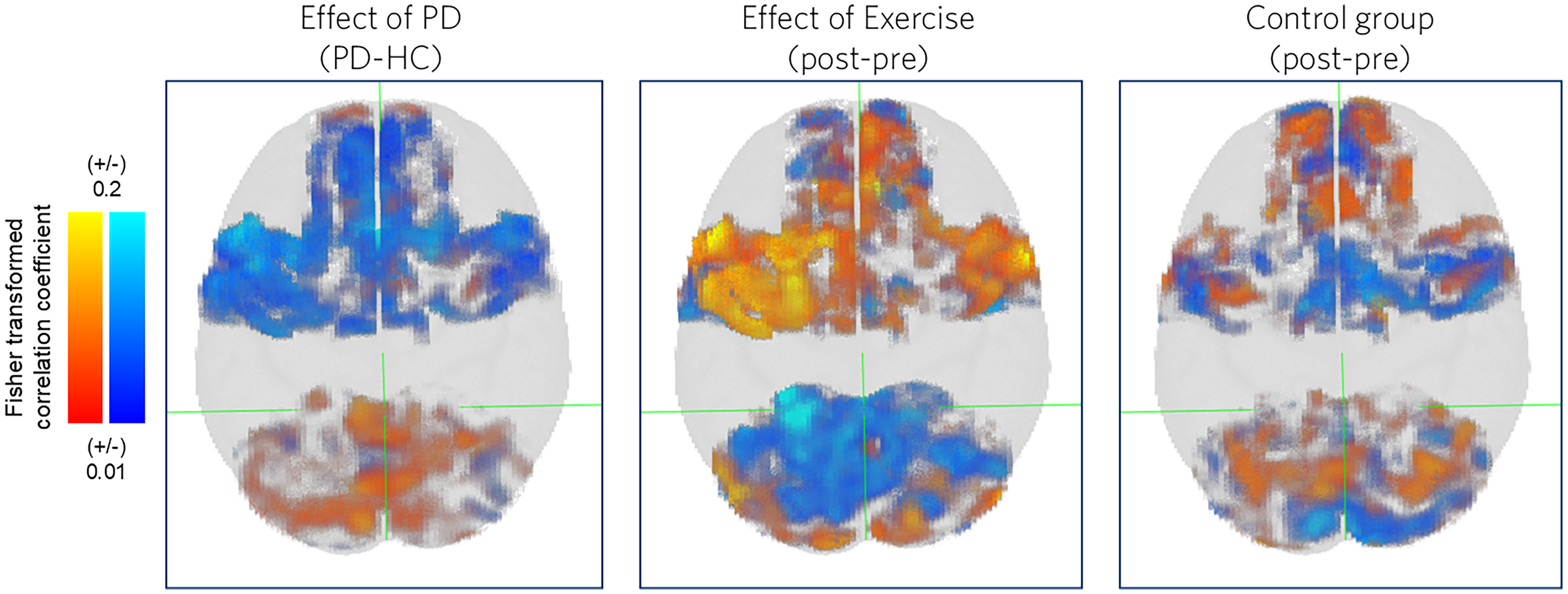
Voxelwise functional connectivity showing Fisher transformed correlation coefficients. Functional connectivity was assessed using the left (more affected) posterior putamen as the seed, with cortical and cerebellar motor regions as voxelwise targets. Group contrasts are shown for: (left) baseline PD vs. HC, (middle) longitudinal change in the PD exercise group, and (right) longitudinal change in the PD control group. Superior view is shown. No clusters survived TFCE correction; uncorrected maps are displayed to illustrate the spatial distribution pattern of altered striato-motor connectivity.
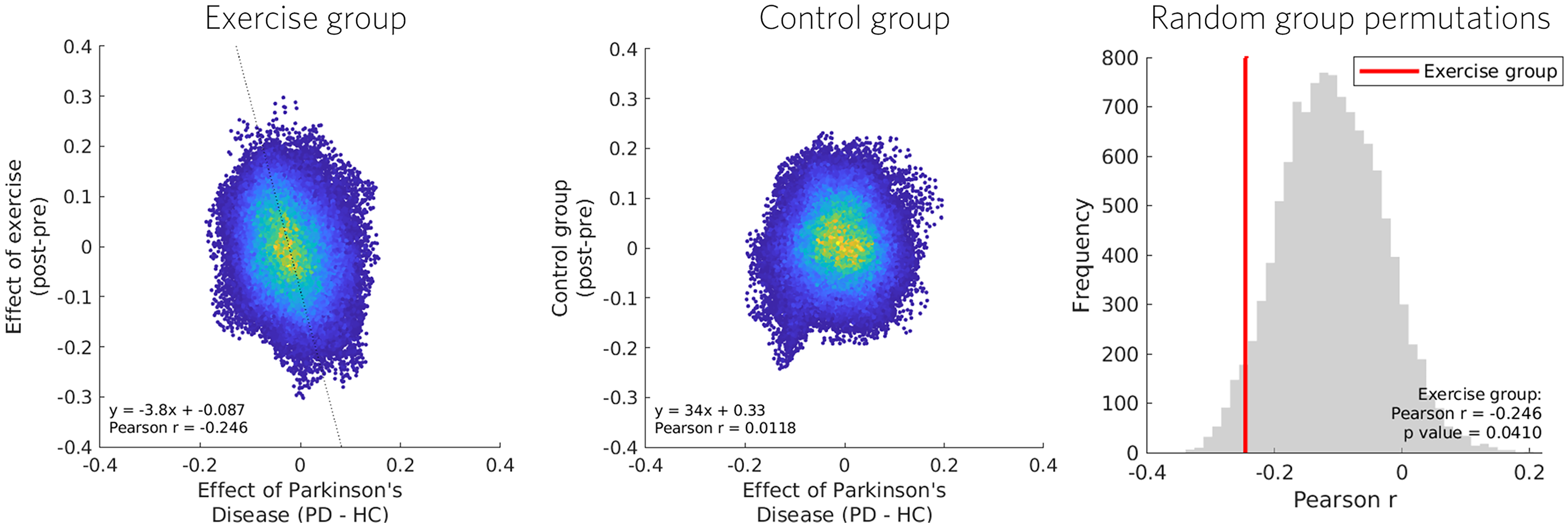
Voxelwise regression-based pattern analysis of exercise effects. Longitudinal connectivity changes in the PD exercise and control groups were regressed against baseline PD–HC differences to test whether exercise-related modulation tracked disease-related alterations. To avoid regression-to-the-mean bias, PD data contributing to each post–pre comparison were excluded from the corresponding baseline map. The observed negative correlation indicates that exercise shifted connectivity opposite to baseline disease differences. Significance was assessed using exhaustive permutation testing of group labels.
ROI-to-ROI-based connectivity analysis
Similar to the voxelwise analysis, conventional ROI-to-ROI contrasts did not yield individual connections surviving strict multiple-comparison correction. However, network-level changes (Figure 4) indicated altered striatal–motor connectivity in PD at baseline, with broadly opposite changes following exercise. Regression-based pattern analysis again revealed a robust relationship between exercise-related connectivity changes and baseline PD–HC differences (r = –0.594, p = 0.0105; Figure 5).
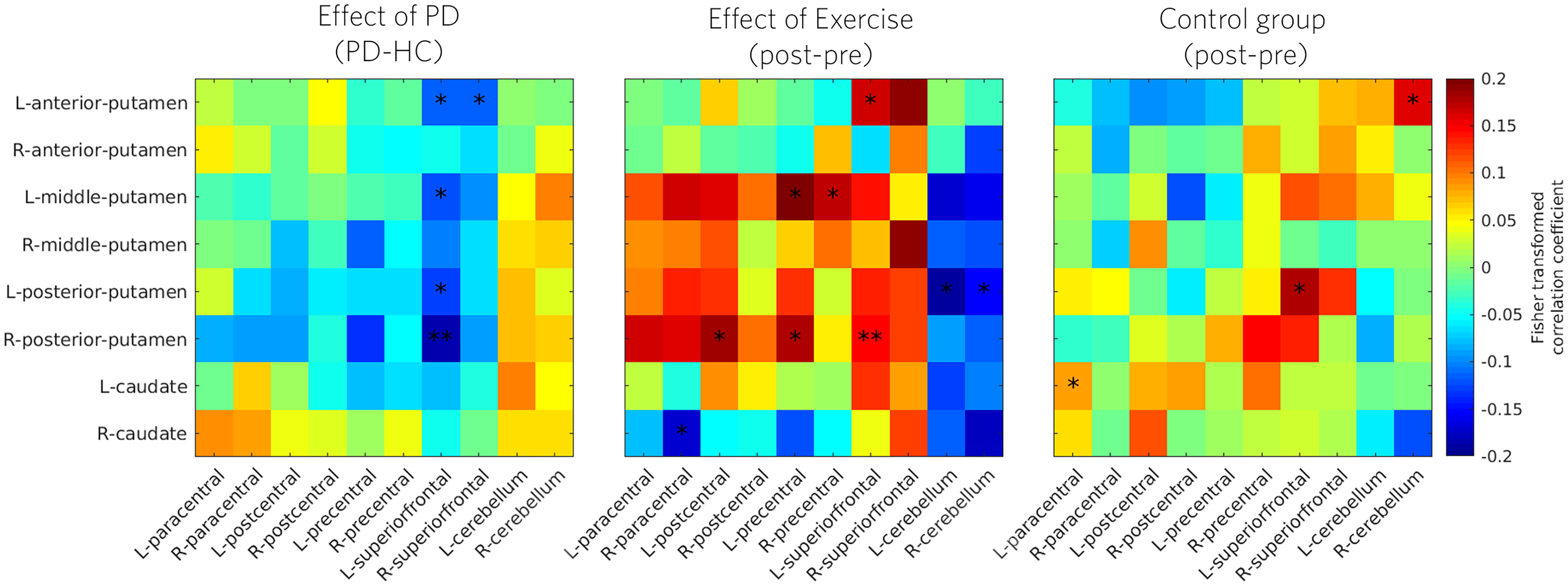
ROI-to-ROI connectivity matrices showing Fisher transformed correlation coefficients between striatal seeds (rows) and cortical/cerebellar motor targets (columns). Group contrasts are shown for: (left) baseline PD vs. HC, (middle) longitudinal change in the PD exercise group, and (right) longitudinal change in the PD control group. Asterisks indicate group-level differences (p < 0.05 and p < 0.01, uncorrected for multiple comparisons).
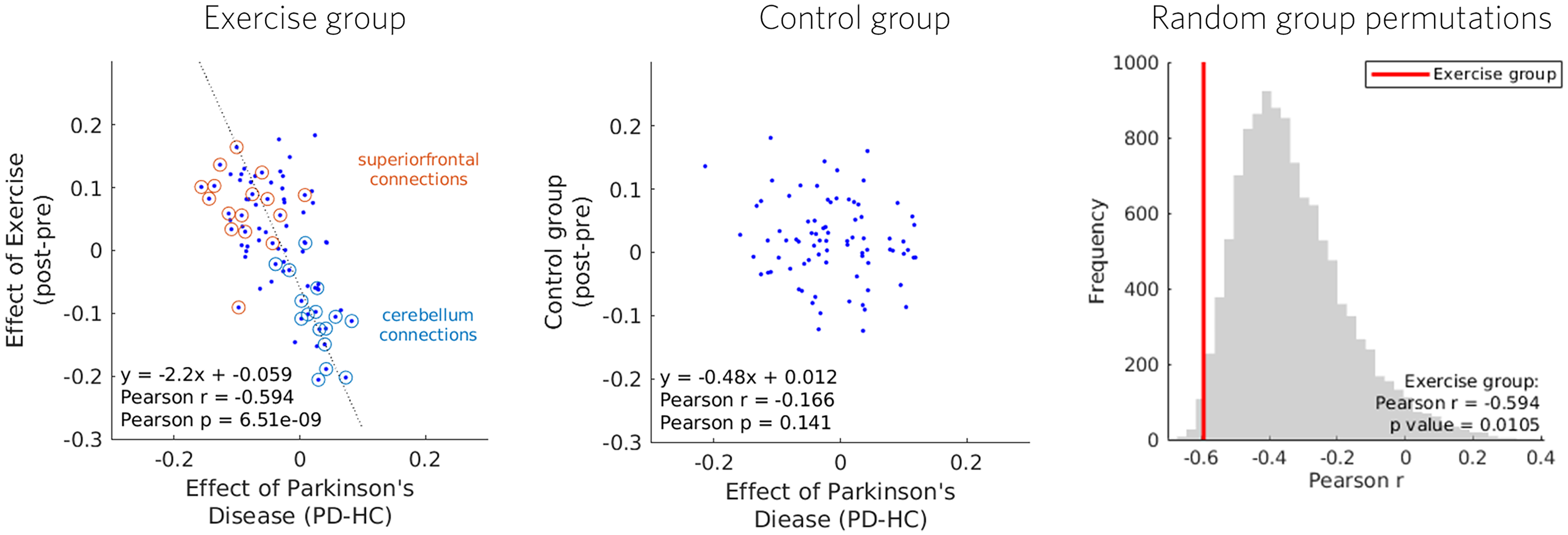
ROI-to-ROI regression-based pattern analysis of exercise effects. Group-level longitudinal connectivity changes (exercise PD group and control PD group) were regressed against baseline PD–HC differences to test whether exercise-related modulation tracked disease-related alterations. To avoid regression-to-the-mean bias, PD data contributing to each post–pre comparison were excluded from the corresponding PD–HC map. The observed negative correlation indicates that exercise shifted network connectivity opposite to baseline disease-related differences. The superior frontal cortex and cerebellum are noted as potential key hubs driving this relationship. Significance of the correlation coefficient was assessed using exhaustive permutation testing of group labels.
REACT analysis
Of the six PET-derived molecular maps initially considered, two maps exhibiting the highest multicollinearity were iteratively removed until all remaining maps had variance inflation factors (VIFs) below 5. The four retained maps (FDOPA, 5-HTT, VAChT, and NAT) represented each neurotransmitter system of interest and were used for subsequent molecular-enriched analyses, ensuring stable and interpretable estimation within the GLM (Table 2).
Variance inflation factors for PET-derived molecular maps retained for REACT analysis.
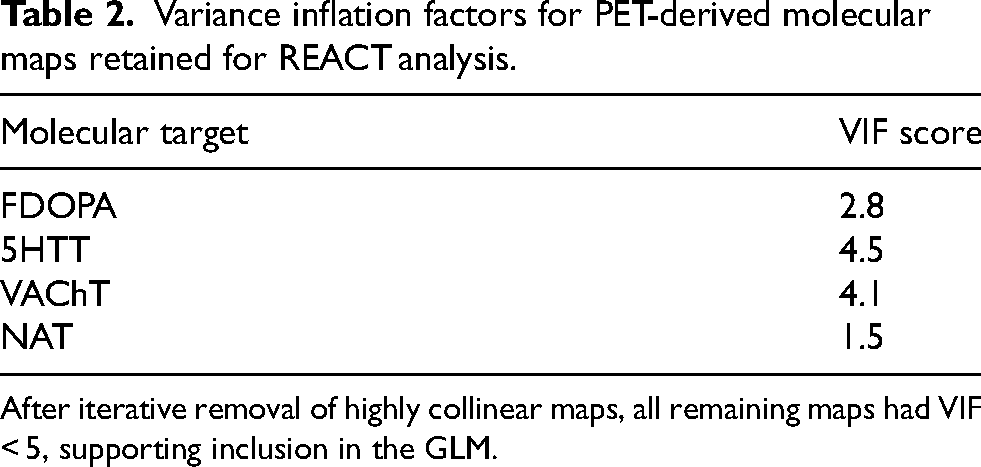

After iterative removal of highly collinear maps, all remaining maps had VIF < 5, supporting inclusion in the GLM.
The selected PET-derived molecular maps and their corresponding molecular-enriched functional connectivity maps, averaged across healthy controls, are shown in Figure 6. Across these maps, conventional voxelwise contrasts did not yield clusters surviving TFCE and Bonferroni correction. However, regression-based pattern analysis revealed that exercise-related changes were significantly aligned with baseline PD–HC differences for the dopaminergic- and cholinergic-enriched networks (FDOPA: r = −0.289, p corrected = 0.0058; VAChT: r = −0.332, p corrected = 0.026; Figure 7). No significant relationships were observed for the serotonergic (5-HTT) or noradrenergic (NAT) related networks.
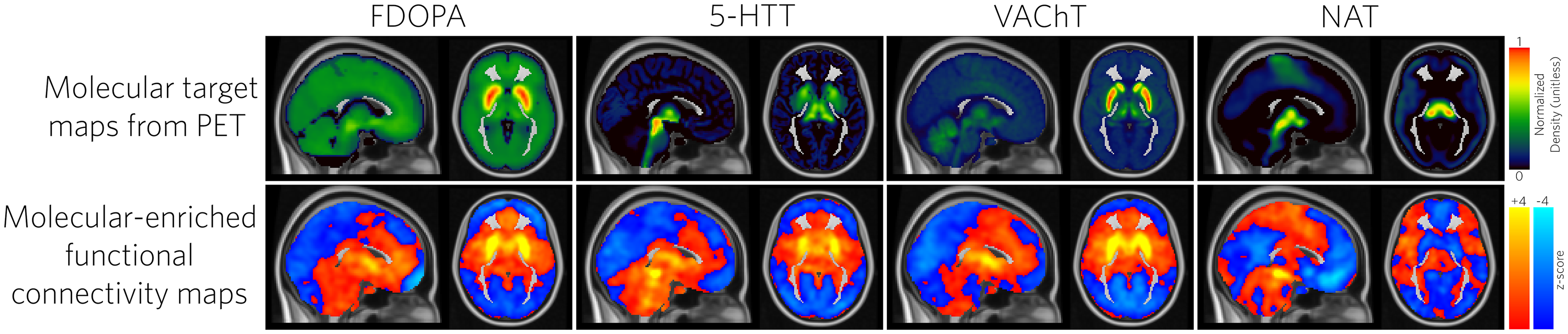
Top: PET-derived molecular target maps included in the REACT analysis (FDOPA, 5-HTT, VAChT, and NAT). Bottom: Corresponding molecular-enriched functional connectivity maps, averaged across healthy controls (HCs), showing the characteristic large-scale network profiles of each neurotransmitter system. These maps serve as reference spatial patterns for the subsequent group-level contrasts and regression-based pattern analyses.
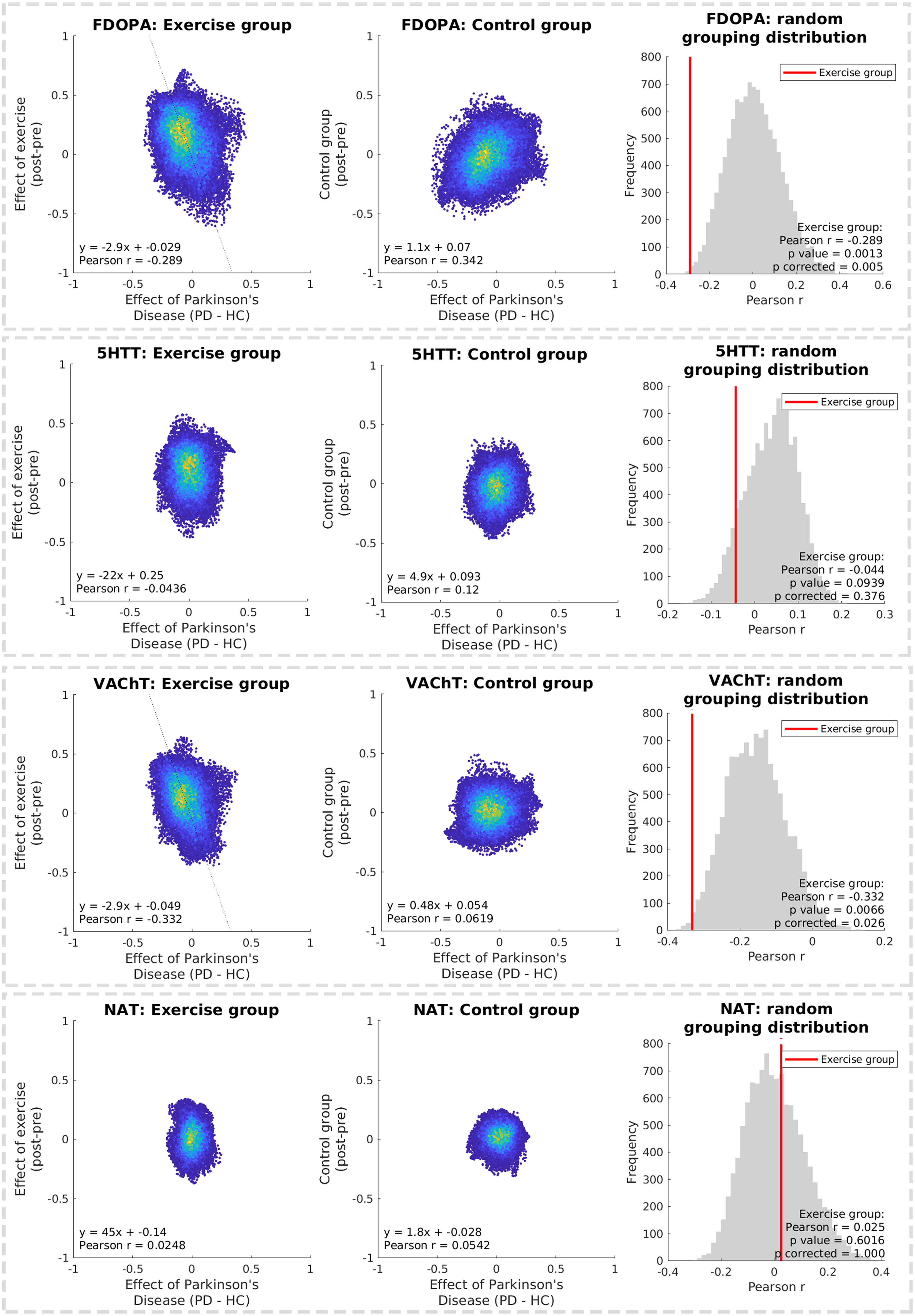
Voxelwise regression-based pattern analysis of molecular-enriched connectivity. Longitudinal connectivity changes (exercise PD group and control PD group) were regressed against baseline PD–HC differences separately for each molecular-enriched functional connectivity map (FDOPA, 5-HTT, VAChT, NAT). To avoid regression-to-the-mean bias, PD data contributing to each post–pre comparison were excluded from the corresponding PD–HC map. Significant negative correlations in the FDOPA- and VAChT-enriched networks indicate that exercise shifted connectivity in the opposite direction to baseline disease-related differences. Statistical significance was assessed using exhaustive permutation testing with Bonferroni correction across molecular maps.
Discussion
This study investigated the neurobiological mechanisms by which aerobic exercise influences brain function in PD, using a combination of conventional resting-state fMRI analyses and the molecular-enriched REACT framework. Our main findings are threefold. (1) While conventional seed-based and ROI-to-ROI analyses did not reveal clusters or connections surviving stringent multiple-comparisons correction, regression-based pattern analyses demonstrated that exercise-related changes systematically tracked baseline group differences between PD patients and HCs. Importantly, this regression framework emphasizes the global spatial relationship rather than isolated effects, yet specific regions such as the superior frontal cortex and cerebellum were noted as consistent hubs, contributing strongly to the observed disease-normalizing pattern. (2) This effect was selective to dopaminergic- and cholinergic-enriched networks, with no significant modulation observed in the serotonergic or noradrenergic related networks. (3) Exercise-induced changes were inversely correlated with PD–HC baseline differences, suggesting that aerobic training reorganizes connectivity in a compensatory manner, working towards a normalization of PD-altered network profiles. Together, these findings provide new insight into the systems-level substrates of exercise in PD and highlight the utility of regression-based pattern analyses for detecting subtle intervention effects that may be overlooked by conventional approaches.
Dopaminergic and cholinergic contributions
Our results suggest that the effects of exercise in PD may be linked to dopaminergic (FDOPA-enriched) and cholinergic (VAChT-enriched) systems, whereas no linkage to the serotonergic and noradrenergic systems was found.
The dopaminergic findings align with the well-established view that PD pathology is driven by degeneration of nigrostriatal dopamine neurons, leading to abnormal corticostriatal connectivity. Preclinical studies have demonstrated that exercise enhances striatal dopamine release and modulates dopamine receptor expression in striatal and prefrontal circuits, with parallel observations in human studies.44–47 Our findings contribute supporting evidence that aerobic training promotes dopaminergic network reorganization at the systems level.
The cholinergic findings are particularly noteworthy, given that cholinergic dysfunction in PD contributes not only to motor impairments but also to gait, balance, and cognitive symptoms.48,49 In PD animal models, repeated exercise has been shown to help preserve signaling that supports motor and cognitive function. 50 Our results therefore provide supporting evidence for potential mechanisms by which exercise may influence broader neurotransmitter systems involved in motor control and cognition, warranting future studies that link these network changes to behavioral outcomes, and whether they correspond to subject-specific changes in cholinergic signaling, for example via VAChT or AChE PET imaging.
Normalization of pathological connectivity patterns
An important observation is that exercise-induced changes were inversely correlated with baseline PD–HC differences. Across voxelwise, ROI-to-ROI, and molecular-enriched analyses, connectivity alterations in PD patients in the exercise group tended to shift toward the healthy control pattern after intervention. Despite one exercise participant appearing as an outlier, the exhaustive permutation testing used in our pattern analyses accounts for inter-subject variability, including such outliers, by generating a null distribution across all possible label assignments. Notably, removing this participant strengthened the observed effects, but the direction and statistical significance of the results remained robust in the full dataset, indicating that the reported trends reliably capture exercise-induced network reorganization.
These findings suggest that aerobic exercise may not simply increase or decrease connectivity, but rather reorganize networks in a compensatory direction, working toward normalization of disrupted communication within motor circuits. Such “restorative” effects align with theoretical models of neuroplasticity in PD, in which interventions enhance compensatory recruitment of intact networks or restore balance across neurotransmitter systems. 51 Johansson et al. reported a compensatory “shift” in striato-cortico motor connectivity following exercise, in which connectivity increases in the anterior putamen appeared to offset lost connectivity in the posterior putamen. 7 In our ROI-to-ROI analysis, we observed a qualitatively similar pattern, with the left (more affected side) middle putamen showing greater connectivity increases than the left posterior putamen, although we did not quantify this effect. While these observations must be interpreted very cautiously, they may reflect a comparable compensatory mechanism, in which the more affected posterior putamen—the region most impacted by PD—may be less able to fully recover, leading to greater reliance on more anterior regions.
Methodological contributions
This study also advances methodological practice in molecular-informed fMRI analysis by leveraging ROI-based REACT. In PD, many regions of interest are small subcortical areas, which may be further reduced due to disease-related atrophy. We found that extracting receptor-enriched time courses using subject-specific ROI parcellations was crucial for capturing meaningful signal changes, highlighting a practical approach to improve sensitivity when targeting these regions. While REACT has been increasingly applied, prior studies have not emphasized ROI-based extraction for small, disease-relevant areas. By integrating ROI-to-voxel and REACT approaches, this work demonstrates a scalable framework for investigating neuromodulatory network dynamics, particularly in challenging subcortical regions.
Limitations
Several limitations should be acknowledged. (1) A very important limitation is the modest sample size for the exercise subgroup, limiting power to detect subtle effects. No formal a priori power analysis was performed due to the pilot nature of the study. This may explain why conventional voxelwise contrasts did not yield significant clusters, whereas a pattern-based approach was able to capture more broad systematic, network-level changes. Due to the modest sample size, our analyses mainly focused on motor circuits, which limited the ability to detect potential exercise-related changes in extra-motor networks such as prefrontal, parietal, or associative regions. (2) PET-derived molecular target maps were generated from healthy populations rather than patient-specific receptor maps. However, while absolute receptor density alterations in PD are not directly captured, REACT still weights regions typically dense for each neurotransmitter, allowing subject-specific functional connectivity disruptions in these key areas to be reflected in the molecular-enriched maps. (3) The single follow-up prevented assessment of dose–response effects or long-term stability. (4) The exercise intervention was open-label, and the control group did not participate in a matched social or group-based activity, so some effects may partly reflect psychosocial or motivational influences associated with exercise participation (placebo and/or Hawthorne effects). Future studies with larger cohorts, patient-specific PET, and active control conditions (e.g., stretching or social engagement) would help to validate and extend these findings.
Conclusions
In summary, our findings suggest that aerobic exercise may reorganize dopaminergic- and cholinergic-enriched networks in a compensatory direction, working toward counteracting PD-related connectivity alterations. This provides supporting evidence for mechanistic links between exercise and symptomatic improvement, highlights its multisystem effects, and reinforces the value of exercise as a promising intervention for PD, while demonstrating the utility of molecular-enriched fMRI for uncovering neurotransmitter-specific mechanisms.
Supplemental Material
sj-docx-1-pkn-10.1177_1877718X261420080 - Supplemental material for Exercise-Induced modulation of molecular-enriched functional connectivity in Parkinson's disease
Supplemental material, sj-docx-1-pkn-10.1177_1877718X261420080 for Exercise-Induced modulation of molecular-enriched functional connectivity in Parkinson's disease by Erik Reimers, Connor Bevington, Jordan U Hanania, Sahib Dhaliwal, Jess McKenzie, Ryan Stein, Teresa Liu-Ambrose, A Jon Stoessl and Vesna Sossi in Journal of Parkinson's Disease
Footnotes
Acknowledgements
The authors would like to thank Brittney Brescian for leading the aerobic exercise sessions in the challenging context of the COVID-19 pandemic. The authors also acknowledge contributing work by additional members of the UBC PET/MRI team — Ju-Chieh (Kevin) Cheng, Carolyn English, Ellen Chen, Jaemin Chung, Elham Shahinfard, Nasim Vafai, and Meng-Chen (Rebecca) Wu — for their assistance with data acquisition and technical expertise. All authors declare that they have no known conflicts of interest. This work was supported by Pacific Parkinson's Research Institute [grant number GR014738, 2018] and the Natural Science and Engineering Research Council [grant number GR010121, 2017].
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Pacific Parkinson's Research Institute [grant number GR014738, 2018]; and the Natural Science and Engineering Research Council [grant number GR010121, 2017].
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
The datasets generated during this study are not publicly available. Custom software used for data analysis is available from the corresponding author upon reasonable request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
