Abstract
Background
Reactive astrocytes are one of the pathological features of Parkinson's disease (PD) and are associated with neuroinflammation and neuronal damage.
Objective
To explore the causal relationship between reactive astrocyte-related genes and PD through the summary data-based Mendelian randomization (SMR).
Methods
We combined these reactive astrocyte-related Quantitative Trait Loci (QTLs) data with PD genome-wide association study (GWAS) statistics. Using SMR, we explored causal links between gene expression, methylation and protein levels (pQTL) with PD, which were validated through colocalization analysis, replication cohorts and substantia nigra tissue data. The study also explored the causal relationship between DNA methylation and gene expression.
Results
SMR analysis identified 95 mQTLs (corresponding to 44 genes), 9 eQTLs, and 7 pQTLs nominally associated with PD (P-SMR_multi < 0.05 & P-SMR < 0.05, and P-HEIDI > 0.01). There was still a significant causal association between MAPK1 expression and Parkinson's disease risk after FDR correction (OR = 2.085, 95%CI = 1.463 to 2.972, P-HEIDI = 0.225, P-SMRFDR = 0.013), supported by strong colocalization (PPH4 = 0.987). Similarly, there was a significant association between corrected CTSB protein levels and Parkinson's disease risk (OR = 0.855, 95%CI = 0.791 to 0.925, P-HEIDI = 0.076, P-SMRFDR = 0.028). The methylation and expression of CLEC3B and PLAU were both nominally associated with the risk of PD. Further analysis revealed that there was also a causal relationship between their methylation and expression.
Conclusions
We identified the MAPK1 gene as a potential causative gene for PD. Its high expression was robustly causally associated with an increased risk of PD and was supported by strong colocalization evidence.
Plain Language Summary
Parkinson's disease (PD) is a serious condition that affects movement and is caused by the loss of certain brain cells. In this study, we looked at specific genes linked to reactive astrocytes to see how they might contribute to the development of PD. We used various approaches to analyze genetic data from large studies to find connections between these genes and Parkinson's disease.
Our research identified genetic signals related to potential changes in gene activity and protein levels that might be involved in the disease. Out of these, we found a strong link between MAPK1 and the risk of PD. This suggests that MAPK1 may play an important role in the PD development.
These findings point to the idea that genes involved in brain inflammation could be significant in understanding PD. They open up possibilities for further research to explore how targeting these genes could lead to new treatments. By learning how these genes function, we could improve the management or even prevention of PD in the future.
Introduction
As a debilitating neurodegenerative disorder, Parkinson's disease (PD) is currently treated with medications like levodopa, which only alleviate symptoms without addressing the underlying pathology.1,2 Emerging evidence suggests that astrocytes, which are activated in response to central nervous system (CNS) injury, play a crucial role in PD progression. 3 Understanding the mechanisms by which astrocytes contribute to PD could reveal novel therapeutic targets.
Reactive astrocytes represent an activated state of astrocytes playing a multifaceted role in inflammation, as well as potential neuroprotective or neurotoxic effects. 4 They release pro-inflammatory cytokines (such as IL-1β, TNF-α) under pathological conditions, initiating and perpetuating neuroinflammation, which may aggravate the damage to dopaminergic neurons. 5 Additionally, the reactive oxygen species (ROS) produced by astrocytes can induce oxidative stress, thereby damaging neurons. 4 The regulation of glutamate uptake and release by reactive astrocytes may lead to glutamate accumulation, causing excitotoxicity that triggers neuronal necrosis and apoptosis. 6 At the same time, astrocytes may change the integrity of the blood-brain barrier, increasing its permeability by affecting the expression of tight junction proteins, 7 potentially promoting the entry of harmful substances into the CNS. Recent studies have also highlighted that astrocyte senescence, mediated by pathways such as cGAS-STING, contributes to PD progression in mouse model. 8 Despite these findings, the specific genes regulating astrocyte function in PD remain unclear, limiting our understanding of PD pathogenesis and the identification of new therapeutic targets.
Mendelian randomization (MR) is a statistical method that employs genetic variants as instrumental variables to evaluate causal associations, serving as a powerful tool for exploring the etiology of complex diseases. 9 In recent years, the MR approach has been widely applied to investigate the causal associations between complex diseases and genes. The summary data-based MR (SMR) method integrates genome-wide association studies (GWAS) data with blood gene expression quantitative trait locus (eQTL), methylation quantitative trait locus (mQTL) and protein quantitative trait locus (pQTL) data, exploring the causal associations between genes and diseases across different levels. 10 This method can solve the limitations of existing research methods, such as avoiding the influence of confounding factors and reverse causality, as well as identifying the regulatory mechanisms of gene expression.
This study aims to explore the causal associations between genes related to reactive astrocytes and PD using an SMR approach. Colocalization analysis and the integration of multi-omics data will further validate the potential causal roles of these genes. The objective is to reveal key molecular events in the pathogenesis of PD and to provide new targets for future therapeutic strategies.
Methods
Study design
This study followed the STROBE-MR reporting guidelines 11 and used a SMR approach to investigate potential causal associations between reactive astrocyte-related genes and PD. The approach included the examination of genetic data to assess the association between these genes and PD using SMR. In addition, we performed colocalization analyses to identify shared genetic variants. The study design and analysis strategy are shown in Figure 1. Ethical approval was not required as this study was based on publicly available GWAS data.
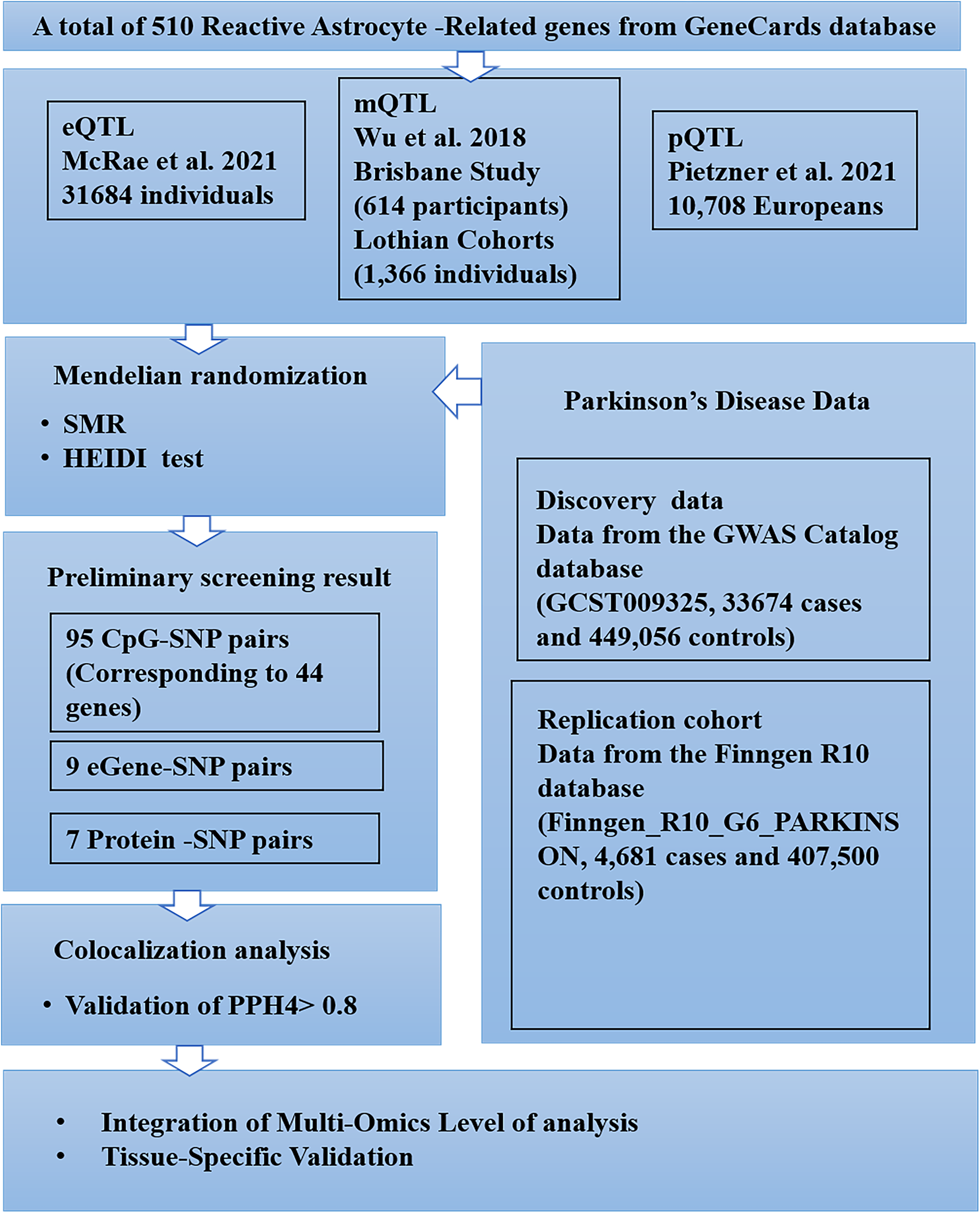

Study design flow chart.
Data sources
Using the search term of “reactive astrocytes”, we retrieved genes related from the GeneCards database (Version 5.20: 10 April 2024). By applying filters to select genes categorized as “protein coding”, we identified a total of 510 reactive astrocyte-related genes (Supplemental Table 1). The primary discovery dataset for PD was from the GWAS Catalog dataset (GCST009325, including 33674 cases and 449056 controls). 12 We also performed validation using Finngen R10 datasets, with sample sizes of 4681 cases and 407,500 controls.
Blood eQTL summary data for reactive astrocyte-associated genes were also obtained from eQTLGen, which included data from 31,684 individuals (https://molgenis26.gcc.rug.nl/downloads/eqtlgen/cis-eqtl/2019-12-11-cis-eQTLsFDR-ProbeLevel-CohortInfoRemoved-BonferroniAdded.txt.gz). 13 Blood mQTL summary data were obtained from meta-analyses of two European cohorts: the Brisbane Systems Genetics Study with 614 participants and the Lothian Birth Cohorts with 1366 individuals (https://yanglab.westlake.edu.cn/data/SMR/LBC_BSGS_meta.tar.gz). 14 Blood pQTL summary data were from the Pietzner et al., including 10,708 Europeans (https://www.synapse.org/Synapse:syn51761394/files/). 15
To assess tissue-specific gene expression and explore its potential causal impact on PD, we used tissue-specific expression eQTL data from the GTEx database. In the context of PD, we used eQTL data from GTEx_Brain_Substantia_nigra.
SMR analysis
This study used version 1.3.1 of the SMR software tool to conduct SMR and HEIDI tests. The objective was to investigate the association between the gene methylation, expression, protein abundance of reactive astrocytes, and PD. 10 In our analysis, we focused specifically on cis-QTL loci that showed significant associations in two large independent cohorts. Compared to traditional MR analysis, the SMR analysis based on these top-related cis-QTL loci exhibited higher statistical power. In selecting the top-related cis-QTL loci, we used a region centered on the corresponding gene with a 1000 kb window on each side and set a P-value threshold at 5.0 × 10−8. 16 In addition, we excluded SNPs with allele frequency differences exceeding 0.2 between different datasets, including LD reference samples, QTL summary data, and outcome summary data. We allowed a maximum proportion of SNPs with allele frequency differences of 0.05, which was achieved using the parameter –diff-freq-prop 0.05.
Building on the SMR analysis, we further employed a multi-SNP-based SMR analysis method (–SMR-multi), which considers multiple SNPs at QTL site. We focused on the results obtained from this method and used the heterogeneity in dependent instruments (HEIDI) test to screen for results without pleiotropy, with a P-HEIDI value greater than 0.01. We considered results with p_SMR < 0.05, p_SMR_multi < 0.05, and P-HEIDI > 0.01 as nominally significant associations. To further control the False Discovery Rate (FDR) due to multiple testing, we applied the Benjamini-Hochberg method to adjust the P-values.
The SMR method was not only used to explore the causal associations between QTLs (including mQTL, eQTL, and pQTL) and PD, but also to further investigate the cascading effects between these QTLs. Specifically, we examined the causal relationship between mQTL and eQTL by treating mQTL as the exposure and eQTL as the outcome, and similarly, the causal link between eQTL (as the exposure) and pQTL (as the outcome). 17
Colocalization analysis
To identify common causal variants between reactive astrocyte-related cis-QTLs (including eQTLs, mQTLs, pQTLs) and PD, we used the R package coloc for colocalization analysis. 18 In the colocalization analysis, we considered five different posterior probabilities corresponding to five exclusive hypotheses 19 : H0 (no genetic association between the trait and SNP), H1 (only trait 1 is genetically associated with the SNP), H2 (only trait 2 is genetically associated with the SNP), H3 (both traits are associated with the SNP but through different causal variables), and H4 (both traits are associated with the SNP and share a common causal variable). Finally, we conducted colocalization analysis on SNPs within a 1000 kb window around the top cis-QTLs colocalization region, using the coloc parameter P12 = 5 × 10− 5 . 20 For the posterior probability of colocalization, PPH4 values exceeding 0.8 were deemed to provide strong evidence of colocalization.20,21
All statistical analyses were performed using R (v4.3.0). The R package “forestplot” was used for forest plot drawing. The plotting codes for SMR Locus Plot and SMR Effect Plot were obtained from Zhu et al. 22
Enrichment analysis of astrocyte-related genes
Gene Ontology (GO) enrichment analysis was conducted to explore the biological processes (BP), molecular functions (MF), and cellular components (CC) of the astrocyte-related genes. In addition, Kyoto Encyclopedia of Genes and Genomes (KEGG) was utilized to investigate the pathway in which these genes were involved. The package “Clusterprofiler” was used for GO and KEGG analysis and visualization.
Result
Integration of pd GWAS and blood reactive astrocytes-related mQTL data
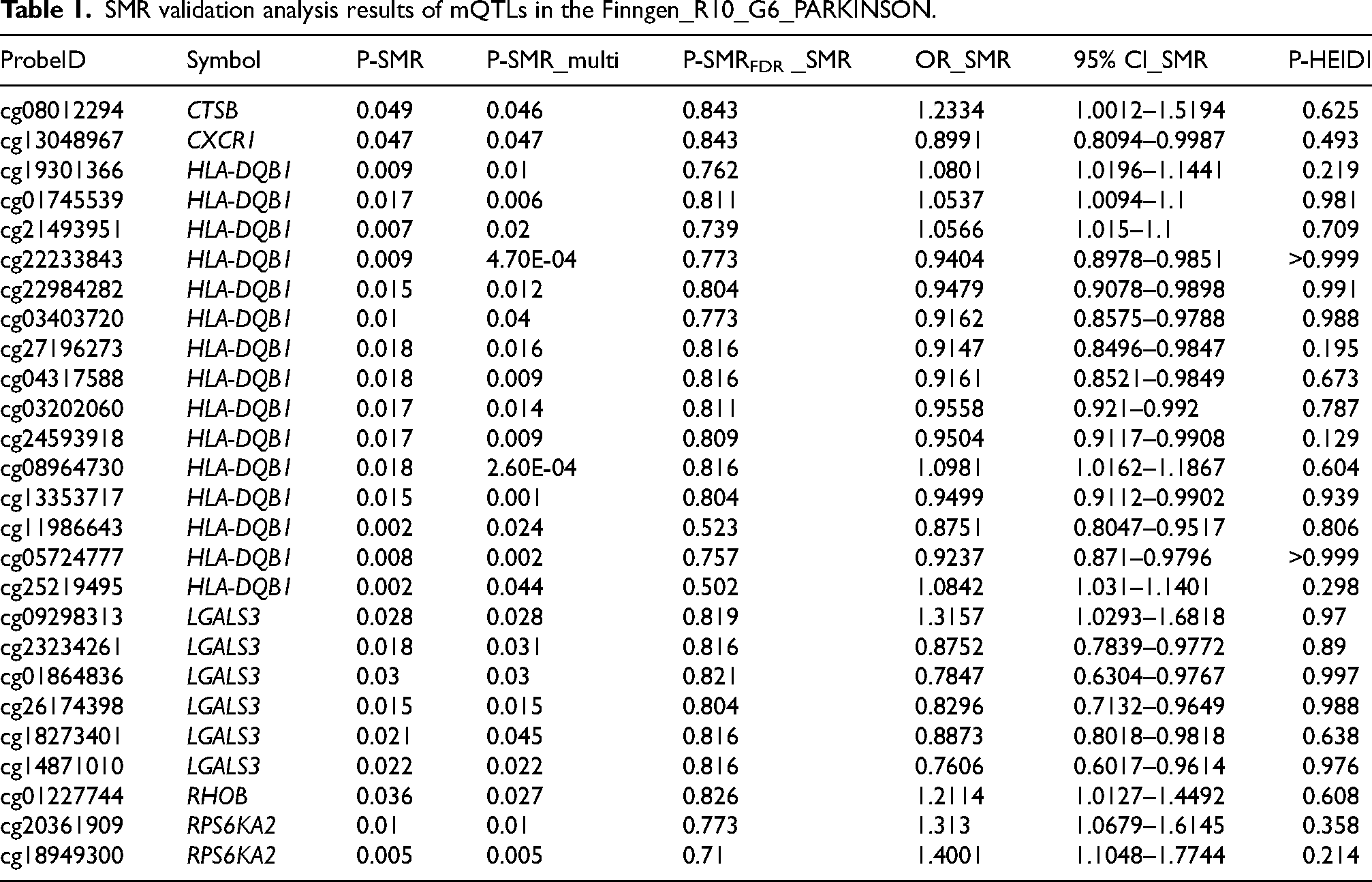
The SMR analysis identified 95 methylation sites (corresponding to 44 genes) with a nominally causal association with PD (Supplemental Table 2, P-SMR_multi < 0.05 & P-SMR < 0.05, and P-HEIDI > 0.01). Among these, 14 sites (corresponding to 5 genes) had colocalization evidence (Figure 2(A), PPH4 > 0.8) and 3 sites (corresponding to 3 genes) passed the FDR correction (P-SMRFDR < 0.05). Among 95 methylation sites, 26 methylation sites corresponding to 6 genes were validated in the Finngen_R10_G6_PARKINSON cohort (Table 1, Supplemental Table 3, P-SMR_multi < 0.05 & P-SMR < 0.05, and P-HEIDI > 0.01).
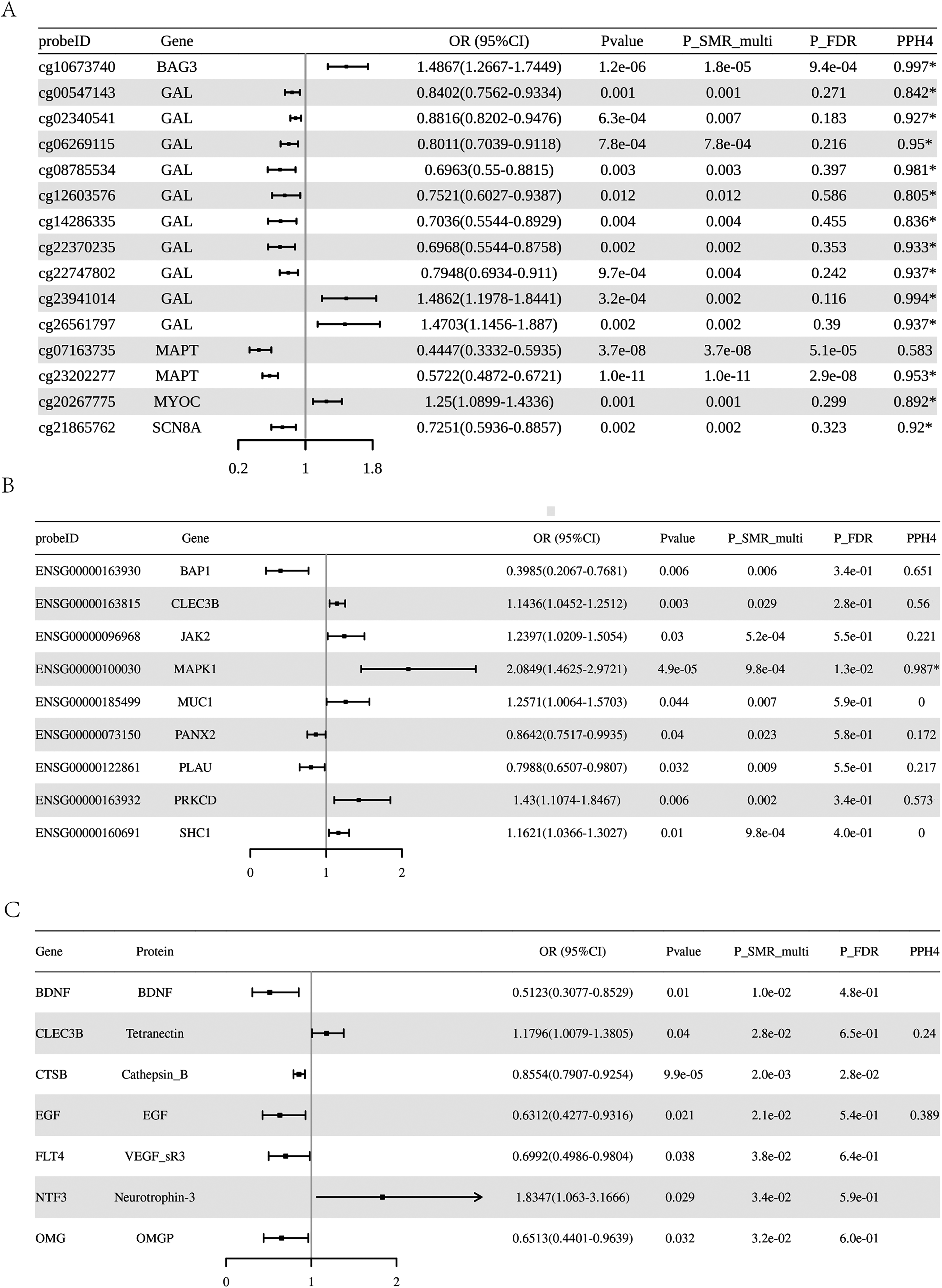
Forest plot. Forest plot depicting the association between (A) Those methylation sites that have a nominal association with Parkinson's disease and have been co-localized or passed the FDR correction (B) gene expressions that have a nominal association with PD, (C) protein abundance and PD risk. * indicated causal associations supported by colocalization evidence.
SMR validation analysis results of mQTLs in the Finngen_R10_G6_PARKINSON.
Integration of PD GWAS and blood reactive astrocytes-related eQTL data
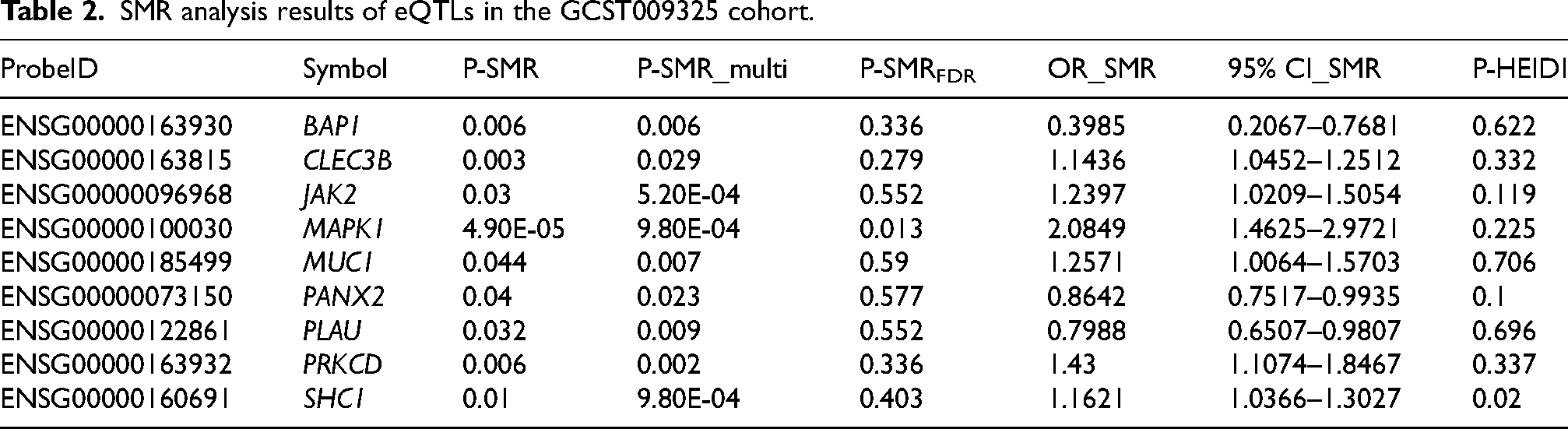
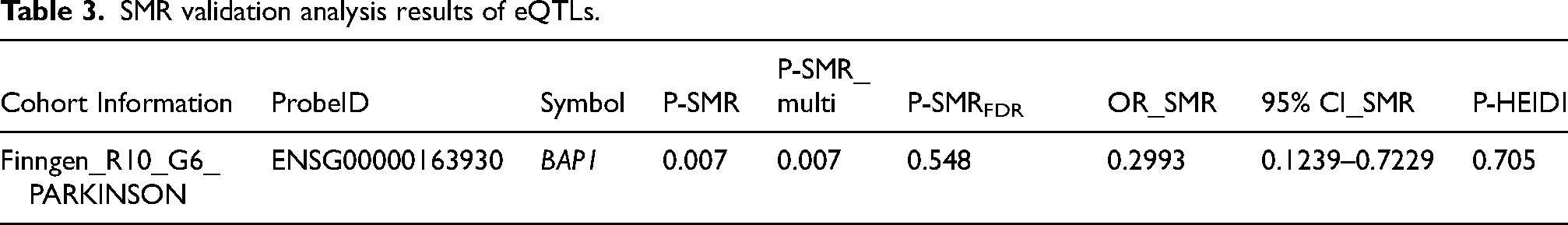
The SMR analysis identified 9 genes (BAP1, CLEC3B, JAK2, MAPK1, MUC1, PANX2, PLAU, PRKCD, SHC1) nominally associated with PD (Table 2, Supplemental Table 4, P-SMR_multi < 0.05 & P-SMR < 0.05, and P-HEIDI > 0.01). High expression levels of 6 genes (CLEC3B, JAK2, MAPK1, MUC1, PRKCD, SHC1) were positively associated with PD risk, while high expression levels of the remaining genes were negatively associated. Notably, the SMR analysis identified MAPK1 still as significant associated with PD after FDR correction (OR = 2.085, 95%CI = 1.463–2.972, P-SMR = 4.90 × 10−5, P-SMR_multi = 9.80 × 10−4 & P-SMRFDR =0.013, and P-HEIDI = 0.225) (Figure 2(B), Table 2, Supplemental Table 4). Moreover, colocalization analysis revealed that its expression was colocalized with PD risk (PPH4 = 0.987) (Supplemental Figure 1A). Figure 3(A) displays the SMR effect plot for the causal association between MAPK1 expression (eQTL) and Parkinson's disease (PD GWAS). Figure 3(B) provides a locus plot illustrating the colocalization of genetic signals for both traits within a 500 kb genomic region, supporting a shared causal variant. Among 9 genes, BAP1 was validated in the Finngen_R10_G6_PARKINSON cohort (Table 3, Supplemental Table 5, P-SMR_multi < 0.05 & P-SMR < 0.05, and P-HEIDI > 0.01). However, the evidence of co-localization of BAP1 expression with Parkinson's disease is relatively low (Supplemental Figure 1B, PPH4 = 0.651).
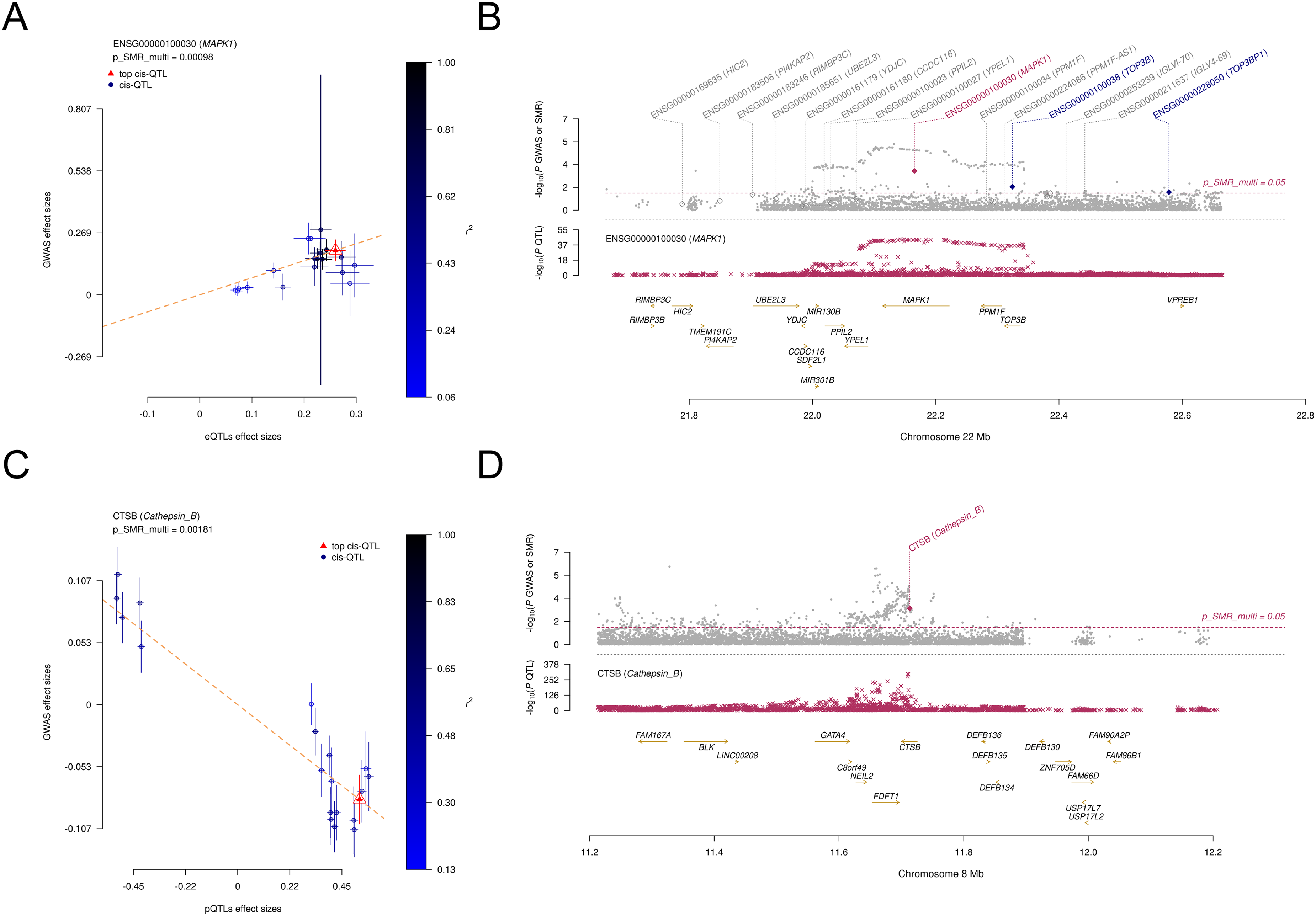
SMRLocusPlot and SMREffectPlot. (A) SMR Effect Plot of MAPK1 in eQTL. (B) The SMR locus plot for MAPK1 in eQTL. (C) SMR Effect Plot for CTSB in pQTL. (D) The SMR locus plot for CTSB in pQTL.
SMR analysis results of eQTLs in the GCST009325 cohort.
SMR validation analysis results of eQTLs.

Integration of PD GWAS and blood reactive astrocytes-related pQTL data
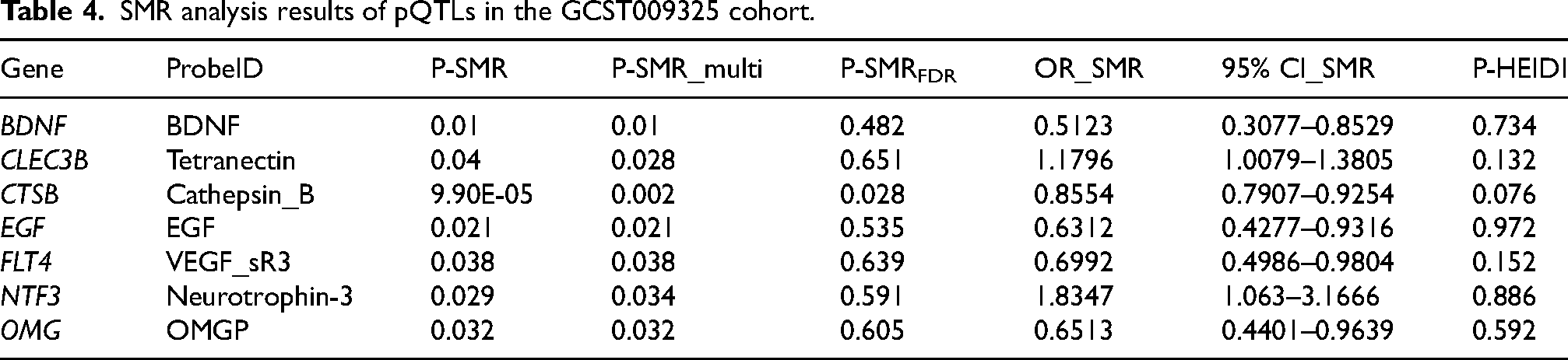
The SMR analysis identified 7 proteins (BDNF, CLEC3B, CTSB, EGF, FLT4, NTF3, OMG) nominally associated with PD (Table 4, Supplemental Table 6, P-SMR_multi < 0.05 & P-SMR < 0.05, and P-HEIDI > 0.01). Protein abundance of CLEC3B and NTF3 was positively associated with PD risk, while the abundance of proteins encoded by the remaining genes were negatively associated. The SMR analysis identified CTSB still as significantly associated with PD risk after FDR correction (OR = 0.855, 95%CI = 0.791–0.925, P-SMRFDR =0.028). Figure 3(C) presents the SMR effect plot for CTSB expression and PD risk. The regional correlation diagram in Figure 3(D) shows that their genetic signals have a high degree of overlap within a 500 kb window. No colocalization evidence was found (PPH4 > 0.8). These results were not validated in the Finngen_R10_G6_PARKINSON cohort (Supplemental Table 7).
SMR analysis results of pQTLs in the GCST009325 cohort.
Integration of multi-omics level evidence
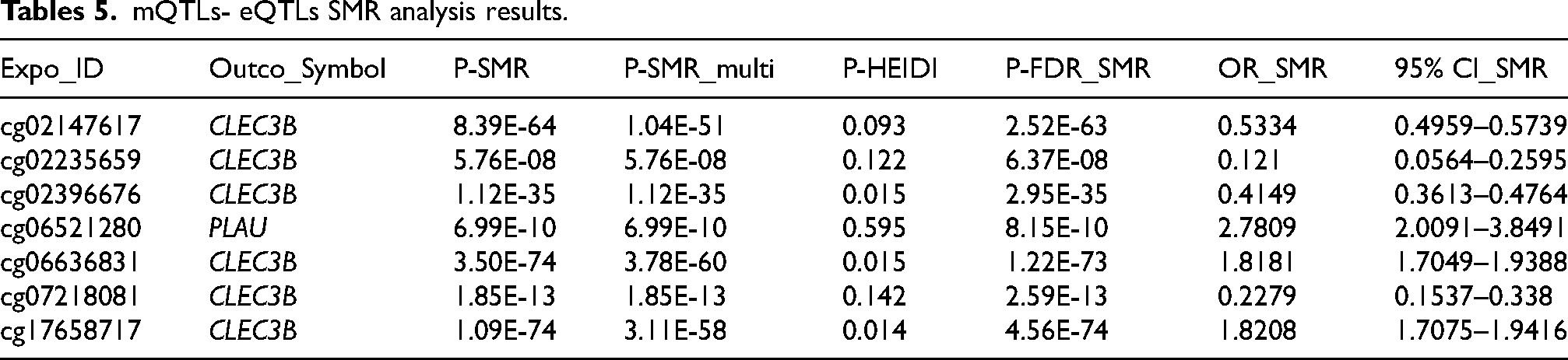
Integrating the results of SMR analysis based on mQTLs and eQTLs, there were 4 common genes (CLEC3B, PANX2, PLAU, PRKCD) in the mQTL and eQTL GWAS results. By integrating blood mQTL and eQTL data, we performed SMR with the methylation loci of these genes as the exposure and their expressions as the outcome. At a stringent criterion (P-SMR_multi < 0.05, P-SMR < 0.05, P-SMRFDR < 0.05 and P-HEIDI > 0.01), 7 methylation loci and 2 genes (CLEC3B and PLAU) were causally associated (Table 5, Supplemental Table 8). Notably, we identified that methylation loci near CLECL3B showed diverse causal associations with its expression, with cg02147617 (OR = 0.5334, 95% CI: 0.4959–0.5739), cg02235659 (OR = 0.121, 95% CI: 0.0564–0.2595), cg02396676 (OR = 0.4149, 95% CI: 0.3613–0.4764) and cg07218081 (OR = 0.2279, 95% CI: 0.1537–0.338) positively associated with its expression, and cg06636831 (OR = 1.8181, 95% CI: 1.7049–1.9388) and cg17658717 (OR = 1.8208, 95% CI: 1.7075–1.9416) exhibiting the opposite (Table 5).
mQTLs- eQTLs SMR analysis results.
Further exploration was conducted into the regulation of blood gene expression on the expression of key reactive astrocyte proteins in PD. However, since the gene's effect on protein expression did not pass the HEIDI test, no positive results were observed (Supplemental Table 9).
Tissue-specific validation
To further investigate the causal association between the expression of identified genes in tissue and PD, SMR analysis was conducted using the GTEx_Brain_Substantia_nigra dataset. The results obtained from the integrated analysis were not validated in the SMR analysis of tissue eQTLs (Supplemental Table 10, P-SMR_multi < 0.05 & P-SMR < 0.05, and P-HEIDI > 0.01).
GO and KEGG enrichment analysis
Based on the findings from mQTL eQTL and pQTL data, a total of 15 signals (BAP1, CLEC3B, JAK2, MAPK1, MUC1, PANX2, PLAU, PRKCD, SHC1, BDNF, CTSB, EGF, FLT4, NTF3 and OMG) were found to be causally associated with PD risk. Functional enrichment analysis was performed on the identified gene set to explore their potential roles in the context of reactive astrogliosis and PD. The analysis revealed significant enrichment of Gene Ontology terms and KEGG pathways strongly relevant to the characteristics of reactive astrocytes (Supplemental Figure 2). Enriched Biological Process terms included regulation of protein kinase and transferase activities, protein phosphorylation, and cellular responses to stimuli, processes vital for astrocytic activation and adaptation. Cellular Component enrichment highlighted vesicle-related structures (e.g., secretory, cytoplasmic, endosome lumen), suggesting altered vesicular transport and secretion. Enriched Molecular Functions involved receptor binding (e.g., cytokines, growth factors, neurotrophins) and kinase/phosphatase activities, consistent with prominent receptor-mediated signaling in reactive astrocytes. Furthermore, KEGG pathway analysis identified significant enrichment in key signaling cascades implicated in reactive astrogliosis, including the PI3K-Akt, MAPK, Ras, Neurotrophin, and Chemokine signaling pathways.
Discussion
The causal association between PD and reactive astrocytes has been explored through SMR analysis, which identified MAPK1 and CTSB as causal signals associated with PD risk even after multiple testing correction. This indicated that these two signals may be potentially implicated in PD pathogenesis.
MAPK1 (Mitogen-Activated Protein Kinase 1), a central molecule in the MAPK/ERK pathway, plays a significant role in cellular stress responses. In PD, abnormal activation or inhibition of the MAPK1/ERK pathway can affect mitochondrial morphology and function, leading to dysregulation of the autophagic process, which aggravates cellular damage and neuronal death. 23 Additionally, the MAPK1/ERK pathway is involved in regulating neuroinflammatory responses, and its abnormal activation can lead to increased expression of inflammatory factors, further aggravating neuronal injury. 24 Studies have found that certain neuroprotective factors can exert neuroprotective effects by activating the MAPK1/ERK signaling pathway, 25 suggesting that this pathway may be a potential therapeutic target for PD. In this study, the expression of MAPK1 was found to be causally associated with PD risk even after multiple testing correction. This suggested the potential involvement of MAPK1 in PD pathogenesis. Further investigation into the specific molecular pathways through which MAPK1 exerts its effects may provide valuable insights into the underlying mechanisms of PD and identify potential therapeutic targets for intervention.
The CTSB gene encodes Cathepsin B, a lysosomal cysteine protease involved in intracellular protein degradation and turnover. 26 In the context of PD, CTSB has been identified as a significant genetic risk factor. For example, variants in the CTSB gene are associated with an increased risk of developing PD. 27 However, Cathepsin B can cleave alpha-synuclein, and this cleavage may generate truncated forms of the protein that have a higher propensity to aggregate, suggesting a complex role in either promoting clearance or potentially enhancing pathology. 27 Studies have demonstrated that increasing Cathepsin B function promotes the clearance of pathogenic alpha-synuclein aggregates, reducing their toxic accumulation.27,28 Thus, higher CTSB levels have been reported to correlate with decreased PD risk. 28 These insights suggest that Cathepsin B and its modulation of lysosomal function and alpha-synuclein turnover are promising targets for potential therapeutic strategies in PD. Consistently, our findings revealed that CTSB protein levels in the blood were negatively associated with PD risk, even though no colocalization evidence was found. This suggested that Cathepsin B and its modulation of lysosomal function and alpha-synuclein turnover are promising targets for potential therapeutic strategies in PD.
The SMR approach used in this study allows for the integration of data from different biological levels, providing a powerful tool for exploring genetic causal associations in complex diseases. By integrating GWAS data with blood mQTL, eQTL, and pQTL data, we were able to avoid the influence of confounding factors and reverse causality, while also identifying regulatory mechanisms of gene expression. 10 Although MR methods have advantages, they also have some limitations. First, MR analysis relies on the validity of genetic instrumental variables; if the selected SNPs do not accurately represent the causal variants, it may lead to incorrect causal inferences. Second, this study primarily analyzed blood samples, which may not fully reflect the regulatory situation of genes in brain tissue. Furthermore, although we have validated our findings in multiple independent cohorts, more research is needed to replicate and validate our discoveries. Fourth, other brain regions, such as the basal ganglia, are also implicated in PD, but due to limitations in data availability, we selected only the substantia nigra eQTL data for tissue-level validation.
In this study, we revealed the potential causal association between reactive astrocyte-associated genes and PD by a multi-omics Mendelian randomization approach. In particular, MAPK1 and CTSB were significantly associated with PD, providing a new perspective for understanding the molecular mechanisms of PD and laying the foundation for future research and development of therapeutic strategies.
Supplemental Material
sj-docx-1-pkn-10.1177_1877718X251395514 - Supplemental material for Reactive astrocyte-related pathogenic genes in Parkinson's disease: A multi-omics Mendelian randomization study
Supplemental material, sj-docx-1-pkn-10.1177_1877718X251395514 for Reactive astrocyte-related pathogenic genes in Parkinson's disease: A multi-omics Mendelian randomization study by Huixi Wang, Jiahao Hu, Bin Mo, Junju Li, Haixin Cai and Qingzhi Li in Journal of Parkinson's Disease
Supplemental Material
sj-xlsx-2-pkn-10.1177_1877718X251395514 - Supplemental material for Reactive astrocyte-related pathogenic genes in Parkinson's disease: A multi-omics Mendelian randomization study
Supplemental material, sj-xlsx-2-pkn-10.1177_1877718X251395514 for Reactive astrocyte-related pathogenic genes in Parkinson's disease: A multi-omics Mendelian randomization study by Huixi Wang, Jiahao Hu, Bin Mo, Junju Li, Haixin Cai and Qingzhi Li in Journal of Parkinson's Disease
Footnotes
Acknowledgements
The authors have no acknowledgments to report.
Ethical considerations
Not applicable
Consent to participate
Not applicable
Consent for publication
Not applicable
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
The data supporting the findings of this study are available within the article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
