Abstract
Gut microbiome alterations are increasingly linked to Parkinson's disease (PD), yet regional signatures remain underexplored. We performed shotgun metagenomic sequencing of stool samples from Egyptian PD patients and healthy controls. PD patients exhibited depletion of short-chain fatty acid-producing taxa, and enrichment of pathobionts. Our findings suggested a pro-inflammatory gut shift in PD and emphasized the need for geographically diverse microbiome studies. While limited in sample size (n = 7 PD patients and n = 6 controls), this pilot addressed a critical gap in African PD microbiome research.
Plain language summary
Gut bacteria may differ in Parkinson's disease: New insights from an Egyptian pilot study using stool DNA
Parkinson's disease (PD) is a condition that affects movement and the nervous system, but recent research has also pointed to a role for gut bacteria. These bacteria, known as the gut microbiome, can affect brain health through chemicals/proteins they produce and their influence on the immune system. However, most microbiome studies in PD have been done in Europe, Asia, or North America. Very little is known about the gut microbiome in PD patients from Africa. In this study, we examined stool samples from seven Egyptian PD patients and six healthy household controls. Using a method called shotgun metagenomic sequencing, which looks at all DNA in the stool, we identified and compared the gut bacteria present in each group. Interestingly, we observed clear differences in which bacterial species were more or less abundant. Specifically, beneficial bacteria that produce anti-inflammatory compounds called short-chain fatty acids (SCFAs) were reduced in PD patients. We also found lower levels of bacteria involved in bile acid metabolism, which may influence inflammation. On the other hand, PD patients had higher levels of certain bacteria like Bacteroides intestinalis, which may damage the gut lining and promote inflammation. These patterns suggested a shift toward a more pro-inflammatory gut environment in PD. While this study involved a small number of participants, it is one of the first microbiome studies of PD in the Middle East and North Africa, up to our current knowledge. These findings highlighted the importance of including diverse populations in PD research and may help guide future efforts to develop microbiome-based therapies or diagnostics.
The gut microbiome has emerged as a key player in Parkinson's disease (PD) pathogenesis through its role in neuroinflammation, intestinal permeability, and altered metabolism of short-chain fatty acids (SCFAs) via the microbiota-gut-brain axis. 1 While large-scale metagenomic studies have revealed microbial disruptions in PD cohorts from Europe, Asia, and North America,1–3 geographic heterogeneity and the underrepresentation of African populations remain major limitations in identifying universal or population-specific microbial signatures.
To address this gap, we conducted a pilot shotgun metagenomic study of stool samples from seven Egyptian PD patients and six matched household controls. Controls matched the patients age inclusion criteria as well, however, they were not strictly matched by sex. One control sample was excluded due to insufficient DNA concentration for sequencing. Diagnosis was confirmed using the UK Parkinson's Disease Society Brain Bank criteria, 4 and exclusion criteria included antibiotic or probiotic use, chronic illness, or neurological symptoms in controls. Informed consents were obtained upon study approval by the Institutional Review Board of the American University in Cairo. DNA was extracted using DNA was extracted using QIAamp Fast DNA Stool Mini Kit (Qiagen, Germany), following the manufacturer's instructions, and sequencing was conducted via the Illumina NovaSeq platform (150 bp PE reads). Taxonomic profiling used Kraken2, 5 and differential abundance was evaluated using DESeq2 6 (significance was considered at a cutoff of FDR adjusted p-value of < 0.05) and LEfSe 7 (taxa with p-value < 0.05 and |LDA score| > 2 were considered to show statistically significant enrichment).
Surprisingly, we observed non-significantly higher microbial alpha diversity, Shannon index (Mann-Whitney U test), in PD patients at both the genus and species levels, as presented in Figure 1A, consistent with recent studies (with statistical significant alpha diversity).1,3,8 This finding contrasted with the general expectation of decreased microbial diversity in disorders, suggesting compensatory expansion of rare taxa or ecological instability. Beta diversity (Bray-Curtis dissimilarity, PERMANOVA test) analysis did not reveal statistically significant separation between PD and control groups, indicating overlapping core microbiota. Nonetheless, specific compositional shifts were observed, as indicated in Figure 1B, supporting the hypothesis that PD may selectively target specific microbial groups rather than completely overhauling the gut ecosystem.
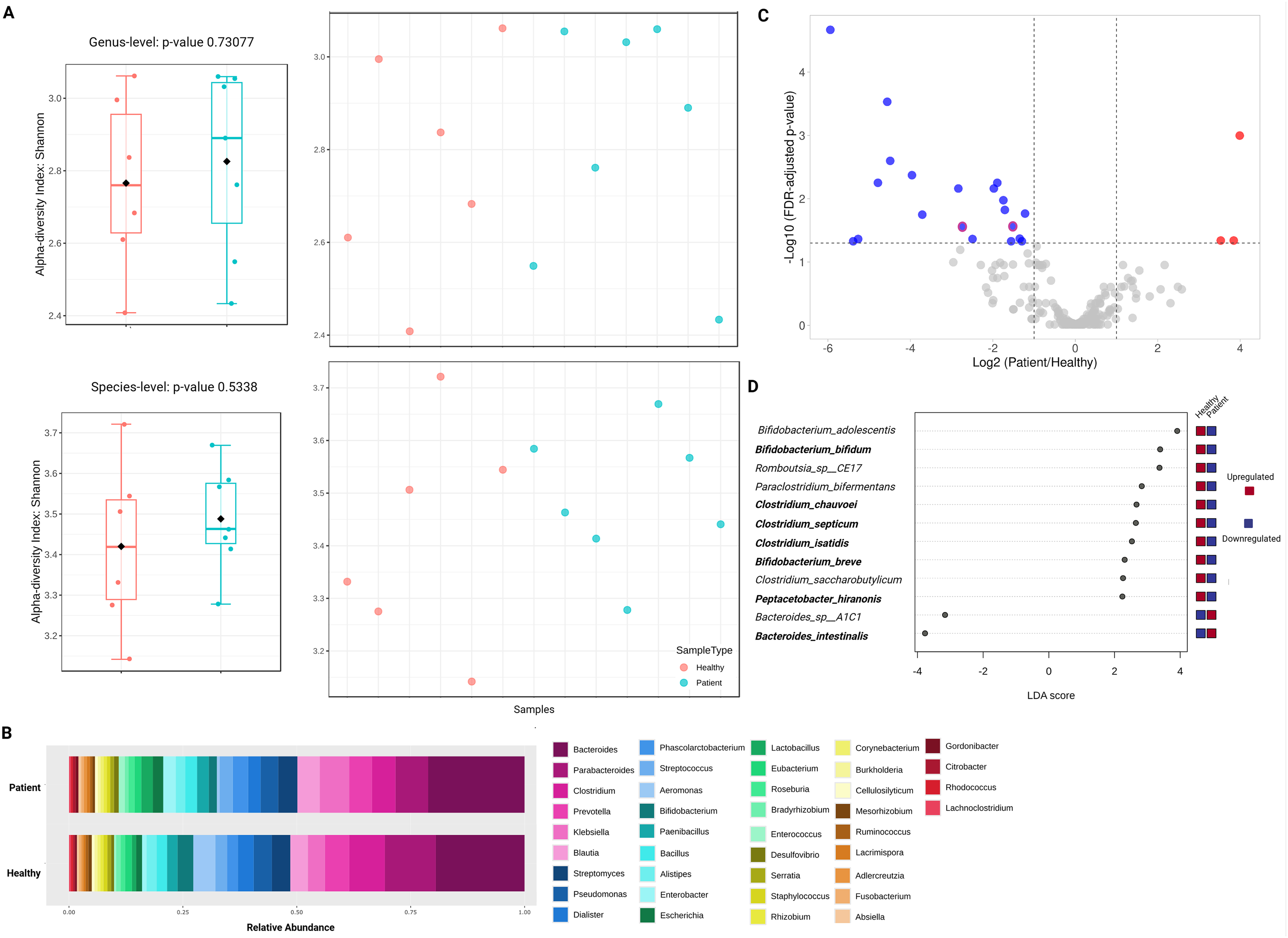
Microbial diversity and taxonomic shifts in Egyptian Parkinson's disease patients. (A) Alpha diversity measured by Shannon index at the genus and species levels showing slightly non-significantly higher diversity in PD patients compared to controls. The individual sample analysis as well as the overall diversity profile as boxplot were shown. (B) Relative abundance profiles at the genus level revealing dominant taxa shared across groups. (C) Volcano plot showing results from DESeq2-based differential abundance analysis between PD patients and controls at both levels, genus and species. Each point represents a bacterial taxon. The x-axis indicates the log2 fold change (PD vs. Control), while the y-axis shows the –log10 of the FDR-adjusted p-value, reflecting statistical significance. Outlined dots indicate taxa at the genus level. D) Species-level Linear Discriminant Analysis (LDA) Effect Size (LEfSe) identifying differentially significant (p-value < 0.05 and |LDA score| > 2) enriched species between PD patients and controls. Bold taxa were also identified as significantly different in DESeq2 analysis, indicating consistent differential abundance across both methods.
PD patients exhibited significant depletion of several beneficial taxa, including Bifidobacterium bifidum, B. breve, B. pseudocatenulatum, Clostridium chauvoei, C. septicum, Peptacetobacter hiranonis, Dialister massiliensis, and D. hominis (Figure 1C). Remarkably, reduction of Bifidobacterium contrasted with previous reports showing increased Bifidobacteriaceae abundance in PD cohorts from different geographical regions,3,9,10 highlighting potential regional, dietary, or clinical differences of this Middle Eastern cohort. From a mechanistic standpoint, the observed loss of SCFA-producing bacteria in PD supported a growing body of evidence implicating microbial-derived metabolites in neurodegeneration. Butyrate and propionate, in particular, have been shown to exert anti-inflammatory effects, promote regulatory T cells, and modulate microglial activity.1,11 Their depletion could potentiate neuroinflammatory cascades and compromise dopaminergic neuron survival. While some studies paradoxically reported that SCFA supplementation may worsen PD phenotypes, 12 the broader consensus supported a protective role for microbial SCFAs in maintaining gut-brain homeostasis.13,14
We also observed reduced abundance of Peptacetobacter hiranonis, a bile acid-transforming bacterium involved in the conversion of primary to secondary bile acids. Dysregulation of bile acid metabolism may lead to accumulation of pro-inflammatory intermediates such as deoxycholic and lithocholic acid, previously linked to PD pathology. 15 Thus, bile acid alterations may be a complementary axis of dysbiosis in PD. An unexpected finding was the depletion of Aeromonas in PD patients, a facultative environmental genus rarely reported in PD microbiome literature. While the clinical significance remained unclear, this may reflect environmental or dietary differences within this Egyptian cohort, yet further research is indispensable.
In contrast, pathobionts such as Bacteroides intestinalis was significantly enriched in PD patients. Members of this genus were capable of mucin degradation and lipopolysaccharides release, potentially disrupting epithelial integrity and contributing to endotoxemia, a proposed trigger for α-synuclein aggregation in the enteric nervous system. 16 Yet, Bacteroides species are also known as SCFA producers, and hence, whether they postulate a dysbiosis hallmarks should be further confirmed.
Importantly, differential abundance results were consistent across both DESeq2 and LEfSe analyses, reinforcing the robustness in the observed taxa shifts. LEfSe confirmed key discriminatory species included B. bifidum, B. breve, C. chauvoei, C. septicum, C. isatidis, P. hiranonis, and Bacteroides intestinalis as shown in Figure 1D.
In summary, our findings highlighted gut microbial signatures in Egyptian PD patients that parallel, yet diverge in key aspects, from those reported in other global populations. The loss of beneficial SCFA- and bile acid-associated taxa, along with the enrichment of inflammatory bacteria, supported the hypothesis that PD is associated with ecological shifts favoring pro-inflammatory and potentially neurotoxic states. While limited by small sample size, our study underscored the feasibility and value of performing shotgun metagenomics in underrepresented regions. Inclusion of diverse populations is crucial to refining microbiome-based biomarkers and unraveling environmental or dietary factors that shape disease risk and progression. Further multi-omic studies across longitudinal cohorts in Africa and the Middle East are warranted to validate these preliminary findings and clarify their relevance for therapeutic development.
Footnotes
Author contributions
All authors have read and approved the final manuscript version.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: M.S. received funding from the Bartlett Fund for Critical Challenges-2021 (Agreement Number: 2—Cycle 3) and the American University in Cairo Faculty Support Grant-2021 awarded to. And Academy of Scientific Research and Technology ASRT, JESOR program, ID: 3086.
The funders were not involved in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
Anonymized genotyping and clinical data can be provided to bone fide researchers upon request from the corresponding author.
