Abstract
Background
Hepatocellular carcinoma (HCC) is a major global health burden, with limited tools for early diagnosis and prognosis. This study explores serum-derived exosomal miR-1275 as a potential non-invasive biomarker for HCC.
Methods
Exosomes were isolated from serum samples of 50 HCC patients and 50 matched healthy controls. miR-1275 expression was quantified by qRT-PCR and compared with traditional biomarkers (AFP, CEA, CA199, DCP, AFP-L3%). Diagnostic performance was evaluated using ROC curves. Bioinformatic analyses, including TCGA pan-cancer data, target gene prediction, and pathway enrichment, were performed to explore regulatory mechanisms.
Results
Exosomal miR-1275 levels were significantly reduced in HCC patients and correlated with advanced clinical stage, tumor burden, and metastasis. miR-1275 showed strong diagnostic value (AUC = 0.869), outperforming CEA and CA199, and improved significantly when combined with other biomarkers (AUC = 0.982). UBE2V1 was identified as a key miR-1275 target involved in cancer-related pathways, including ubiquitination and mTOR signaling.
Conclusion
Serum exosomal miR-1275 is a promising biomarker for early diagnosis and prognosis of HCC. Its integration into multimarker panels could enhance clinical decision-making and patient management.
Introduction
Hepatocellular carcinoma (HCC) is one of the most common cancers worldwide. According to 2022 global cancer statistics, 1 HCC ranks sixth in incidence globally and is the third leading cause of cancer-related deaths, particularly in regions with high rates of chronic hepatitis infection and cirrhosis. Early detection is crucial for improving survival rates, and the five-year survival rate for patients with early-stage HCC exceeds 70%. However, for patients with advanced HCC, this rate decreases to less than 12.5%. 2 Despite advancements in treatment, including immunotherapy and targeted therapies, the prognosis for HCC patients remains poor, primarily due to late-stage diagnosis and the limited effectiveness of these current treatment options. 3 Therefore, developing more effective screening methods that allow earlier detection of HCC represents an important research focus.
However, existing early diagnostic tools3,4—including imaging and traditional serum biomarkers such as alpha-fetoprotein (AFP), carcinoembryonic antigen (CEA), carbohydrate antigen 19-9 (CA19-9), alpha-fetoprotein-L3% (AFPL3%), and des-γ-carboxy-prothrombin (DCP)—often lack sufficient sensitivity and specificity, especially in early-stage disease. Due to the high rate of missed diagnoses, there is an urgent clinical need for novel, non-invasive biomarkers that can facilitate early diagnosis and provide valuable prognostic information.5–7
Recent studies have highlighted exosomes as promising tools for non-invasive cancer diagnosis.8,9 Exosomes are small extracellular vesicles, approximately 30–150 nm in diameter, that facilitate intercellular communication by transporting biomolecules such as proteins, lipids, and nucleic acids, including microRNAs (miRNAs).10,11 These vesicles are released by various cell types, including cancer cells, and are readily detectable in bodily fluids, making them promising candidates for the development of non-invasive biomarkers. 12
Among the various miRNAs, miR-1275 has gained attention for its involvement in multiple oncogenic processes across different cancer types. The gene encoding miR-1275 is located on the short arm of chromosome 6 and contains only one exon; the mature miR-1275 sequence is 17 nucleotides in length. 13 It was found that miR-1275 plays a vital role in regulating various cellular signaling pathways like the PI3K/AKT, ERK/JNK, MAPK, and Wnt signaling pathways, and the dysregulation of this miRNA has been shown to contribute to the development of multiple cancer types such as cancers of the liver, breast, lung, gastrointestinal tract and genitourinary tract.14–16 In our previous study, 16 we reported that downregulated levels of miR-1275 may be associated with a higher risk of HCC, suggesting that it could be used as a biomarker for personalized diagnosis and treatment. Our preliminary studies involving 6 HCC patient samples confirmed that exosome-derived miR-1275 expression is significantly downregulated in HCC through high-throughput sequencing. These results suggest that miR-1275 may play a crucial role in tumor progression and could serve as a potential biomarker for the diagnosis and prognosis of HCC. Detecting serum exosomal miR-1275 might offer a more accurate and noninvasive method for diagnosing HCC and assessing patient prognosis. However, to validate exosomal miR-1275 as a potential biomarker, more detailed clinical evidence is needed to establish the relationship between exosomal miR-1275 levels and the clinical characteristics of HCC.
This study aims to evaluate serum-derived exosomal miR-1275 levels in HCC patients, assess its diagnostic efficacy compared to traditional biomarkers, and explore its prognostic value through integration with clinical features and bioinformatics analysis, including pan-cancer comparisons and enrichment analyses of target pathways. Ultimately, establishing exosomal miR-1275 as a clinical biomarker could lead to improved early detection, enhanced prognostic stratification, and personalized treatment approaches in HCC management.
Materials and methods
Patient and clinical samples
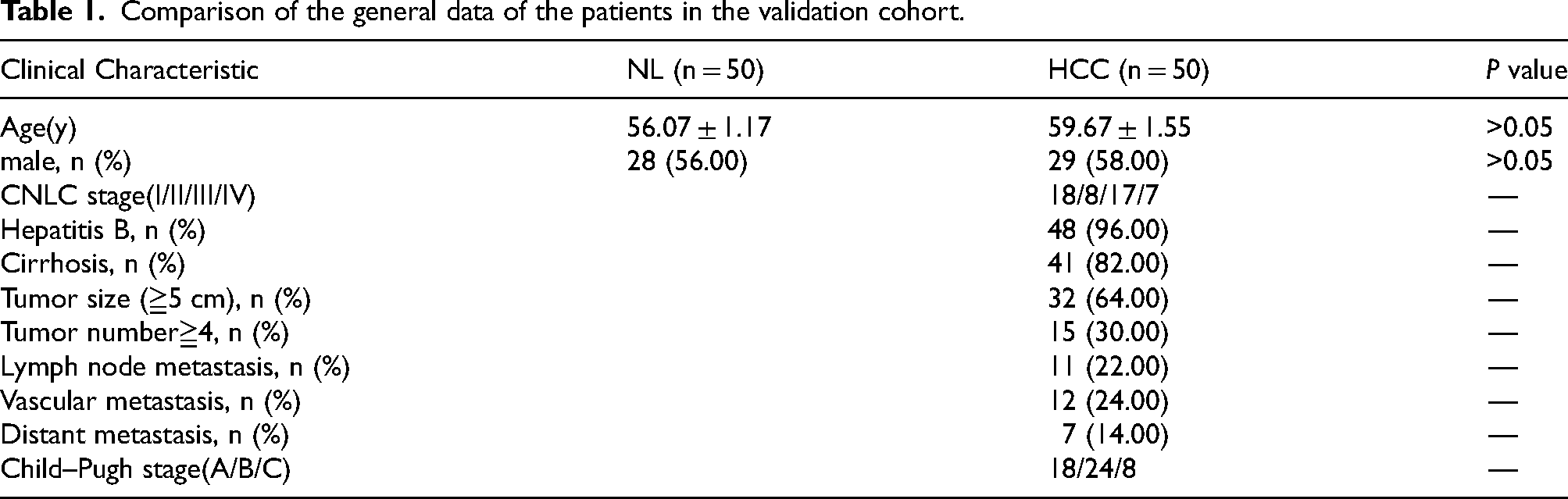
This study was conducted at the Affiliated Hospital of Guilin Medical University. Serum samples were collected from 50 patients with histopathologically confirmed HCC and 50 age- and sex-matched healthy individuals between February 2022 and February 2024. Eligible patients were adults (≥18 years) with a confirmed diagnosis of HCC who had not received prior treatment, and who had complete clinical and pathological data, including tumor characteristics, CNLC stage, liver function, and HBV infection status. Patients with other primary malignancies, severe systemic comorbidities, concurrent autoimmune or infectious diseases, or poor-quality serum/RNA samples were excluded. Healthy controls were recruited based on normal liver function, absence of liver disease, viral hepatitis, malignancy, or major chronic illness. The clinical stage of HCC was determined according to the China Liver Cancer (CNLC) staging system, which classifies tumors into four stages (I–IV, with substages a/b) based on tumor size, lesion number, vascular invasion, extrahepatic spread, and liver function. The clinical characteristics of the subjects are listed in Table 1.
Comparison of the general data of the patients in the validation cohort.
TCGA pan-cancer data acquisition and analysis of miR-1275 expression
The expression profile of miR-1275 across various human cancers was retrieved from the TCGA Pan-Cancer (PanCanAtlas) dataset using the UCSC Xena Browser (https://xenabrowser.net/). Log2-normalized RNA-seq data (log2[RPM + 1]) were used to assess the differential expression of miR-1275 between tumor and normal tissues. For hepatocellular carcinoma (LIHC), miR-1275 expression levels were compared between tumor and adjacent normal liver tissues, and statistical significance was assessed using the Wilcoxon rank-sum test. Additionally, the expression of miR-1275 across HCC clinical stages (I–IV) was analyzed using the Kruskal–Wallis and pairwise Wilcoxon tests. Copy number alteration (CNA) data for miR-1275 were also analyzed from the TCGA PanCanAtlas database to investigate genomic alterations across cancer types.
Prediction of miR-1275 target genes and enrichment analysis
The target genes of miR-1275 were predicted using three publicly available databases for miRNA-target interactions: TargetScan, miRDB, and miRTarBase. Genes that were consistently predicted by both miRDB and miRTarBase were considered for further analysis. A total of 28 overlapping genes were selected for downstream investigation. Protein–protein interaction (PPI) networks were constructed using the STRING database (https://string-db.org/), and the resulting network was visualized using Cytoscape software (v3.9.1). Functional enrichment analyses of Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways were performed using the clusterProfiler R package. Enriched terms with adjusted P values < 0.05 were considered statistically significant.
Isolation of exosomes from serum samples
Peripheral blood was freshly collected and allowed to stand at room temperature for 30 min. The samples were then centrifuged at 3000 rpm for 10 min at 4°C to obtain 1.5 mL of serum, which was transferred to an EP tube. Serum exosome purification and RNA extraction from 1.5 mL of serum were performed via a MinuteTM High-Efficiency Exosome Precipitation Reagent Kit (Invent Biotechnologies, MN, USA). After 750 µL of Exosome Precipitation Reagent was added to each serum tube, the samples were incubated for 60 min at 4°C. The mixture was centrifuged at 10,000 × g for 15 min at 4°C, and the supernatant was discarded. The pellet was washed with PBS and centrifuged at 10,000 × g for 1 min at 4°C. The final pellet was resuspended in 100 µL of PBS to obtain serum exosomes.
Identification of exosomes from serum samples
Exosome identification was performed using transmission electron microscopy (TEM) and nanoparticle tracking analysis (NTA). The purified exosomes from the serum of HCC patients and nontumor donors were negatively stained with 2% phosphotungstic acid on a copper grid for 2 min and then air-dried for 10 min. The exosomes were examined and imaged using transmission electron microscopy (TEM) (Tecnai G2 Spirit FEI, Thermo Fisher Scientific, Waltham, MA, USA). The core facility platform of Zhejiang University operated the TEM. NTA was performed using a NanoSight NS300 instrument (Malvern, UK) according to the manufacturer's instructions to measure the hydrodynamic diameter distribution of the exosomes.
The surface markers of the exosomes (CD9, CD63, and CD81) were detected via flow cytometry. Immunofluorescence staining was performed via the following FITC-labeled antibodies purchased from BD Pharmingen™: mouse anti-human CD9 (Clone M-L13), CD63 (Clone H5C6), and CD81 (Clone JS-81). The isolated exosome samples (30 µL) were resuspended in 60 µL of PBS, and 20 µL of each FITC-labeled antibody was added to reach an approximate particle concentration of 1 × 1010 particles/mL. These mixtures were incubated at 37°C for 30 min. The samples were washed twice with PBS at 10,000 × g for 30 min at 4°C. The pellet was resuspended in 100 μL of PBS for analysis using NanoFCM (N30E, NanoFCM Inc., China).
Serum miRNA quantification/miR-1275 quantification
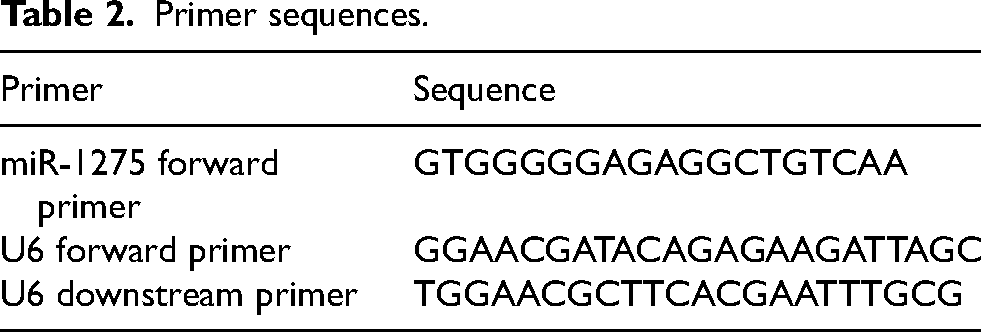
Total RNA was extracted from serum exosome samples using the EZB-exo-RN1 Exosome RNA Purification Kit, and RNA concentration and purity were measured with a NanoDrop spectrophotometer. Sequence information for miR-1275 was obtained from miRBase. U6 snRNA was selected as the internal reference for normalization, as it is widely used in exosomal miRNA studies and showed stable expression across all samples in this study. The forward primers for miR-1275 and U6 were synthesized by Shanghai Shenggong Bioengineering Co., Ltd., with the universal downstream primer for miR-1275 provided by Takara Bio. Primer sequences are presented in Table 2.
Primer sequences.
Real-time quantitative fluorescent RT–PCR (qRT-PCR) was performed via the TB Green® Premix Ex Taq™ II (Tli RNaseH Plus) Kit on an ABI 7500 instrument (Thermo Fisher Scientific, Waltham, MA, USA). qRT-PCR was performed using the following cycling conditions: an initial denaturation step at 95°C for 30 s, followed by 40 cycles of 95°C for 5 s and 60°C for 34 s. The qRT-PCR data were obtained from three replicates.
Tumor biomarker detection
AFP, CEA, and CA199 were detected with a Cobas Elecsys 801 analyzer (Roche, Germany). AFP-L3% and DCP were detected via a C2000 chemiluminescent analyzer (Hotgen Biotech, China). All original matching reagents and quality control materials were used.
Statistical analysis
Statistical analyses and graphical visualizations were performed using IBM SPSS Statistics (version 26.0), GraphPad Prism (version 10.0), and R software (version 4.4.3).
Continuous variables were expressed as the means ± standard errors (SE) if normally distributed, and medians with interquartile ranges (IQR) if non-normally distributed. Differences between the two groups were analyzed using the Mann–Whitney U test, and multiple-group comparisons were conducted using the Kruskal–Wallis H test, followed by pairwise tests as necessary. Diagnostic performance was evaluated using receiver operating characteristic (ROC) curves, with the area under the curve (AUC) calculated. Binary logistic regression analysis was used to assess the combined diagnostic efficiency of miR-1275 and traditional biomarkers. Statistical significance was considered at a two-tailed P < 0.05.
miR-1275 and UBE2V1 expression data were obtained from the TCGA PanCanAtlas dataset via UCSC Xena. Expression differences between tumor and normal tissues were analyzed using the Wilcoxon rank-sum test, and the Kruskal-Wallis test examined stage-specific differences. The Wallis test and pairwise Wilcoxon rank-sum tests were also used. Survival analysis was conducted using Kaplan–Meier curves with statistical significance determined by the log-rank test. Functional enrichment analyses (GO and KEGG pathways) were performed using the clusterProfiler R package, and significantly enriched terms were identified based on adjusted P values < 0.05.
Results
Differential expression of miR-1275 in TCGA pan-cancer database
To investigate the distribution of miR-1275 in pan-cancer, we analyzed data from the TCGA RNA-seq database. We conducted a more in-depth analysis of miR-1275 expression in various types of cancer by examining the TCGA PanCanAtlas data (Figure 1(a)). In the TCGA PanCanAtlas dataset, copy number alterations (CNA) of miR-1275 were analyzed across various tumor types. In Liver Hepatocellular Carcinoma (LIHC), most samples showed shallow deletions or diploid status, while amplifications were rare. miR-1275 is significantly dysregulated in several other cancer types in addition to HCC(eg. Lung adenocarcinoma(LUAD), Stomach adenocarcinoma(STAD), Pancreatic adenocarcinoma(PAAD), Colorectal adenocarcinoma(COAD),Glioblastoma multiforme(GBM), Kidney renal clear cell carcinoma(KIRC)).The distribution of CNA log2 values in LIHC centered around zero or negative values, indicating a tendency toward neutral or reduced MIR-1275 dosage. These findings support a possible tumor-suppressive role of miR-1275 in HCC.
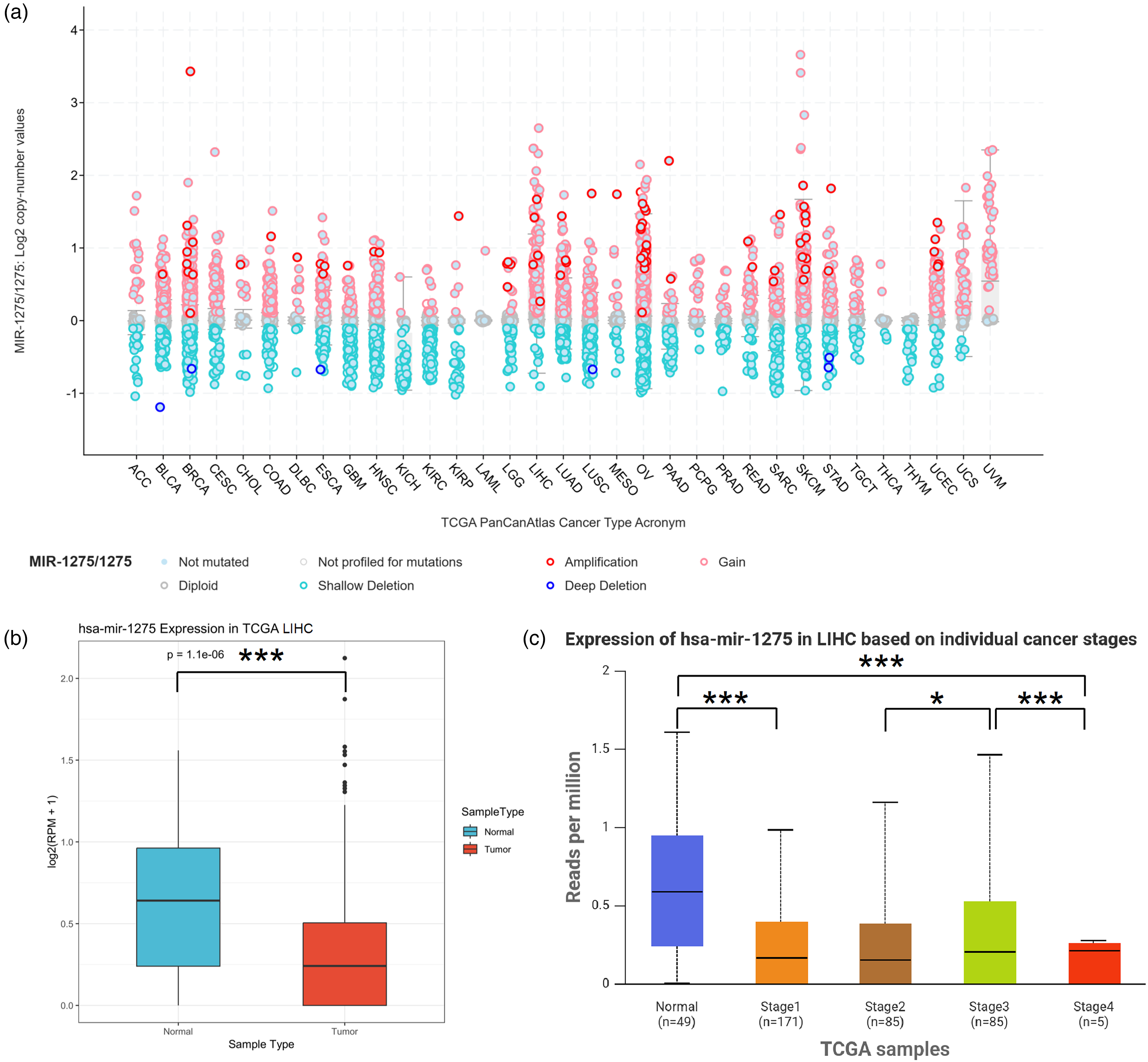
Genomic alteration and expression analysis of miR-1275 across TCGA cancer types and specifically in hepatocellular carcinoma (LIHC). (a) Pan-cancer analysis illustrating copy number variations (CNVs) of miR-1275 across various cancer types, including amplification, gain, shallow deletion, deep deletion, and diploid states. (b) Differential expression of miR-1275 between LIHC tumor and normal tissues in the TCGA dataset, showing significantly decreased expression in tumor tissues (***P < 0.001). (c) Expression levels of miR-1275 in LIHC according to individual cancer stages, demonstrating significantly lower expression in cancer tissues at all stages compared to normal tissues (*P < 0.05, ***P < 0.001).
After comparing the expression of miR-1275 in TCGA LIHC (Liver hepatocellular carcinoma) database, the box plots showed a significant downregulation of miR-1275 in in liver tumor tissues compared to liver normal tissues (Figure 1(b)). This difference is statistically significant (P = 1.1e-06), indicating that hsa-miR-1275 may play a role in liver tumorigenesis and could be considered as a potential diagnostic biomarker.
Furthermore, we analyzed the prognostic value of miR-1275 in different stages of HCC. The expression level of hsa-miR-1275 gradually decreases from Stage I to Stage IV. This pattern may indicate that hsa-miR-1275 expression inversely correlates with tumor progression. There is no significant difference between stage I and II (P = 0.475), but there is a substantial difference between stage II and III (P = 0.0433). Stage IV data show significant variability compared to Stage III (P = 5.28e-03), but are limited due to a small sample size (n = 5), which may affect the statistical reliability of this group.
Enrichment analysis of miR-1275 in HCC
It was predicted that miR-1275 is involved in genetic and pathway regulation, providing a direction for exploring the pathogenesis of HCC through bioinformatics. To investigate the target genes of miR-1275, we predicted its target genes using three online databases: TargetScan, miRDB, and miRTarBase. The prediction results are shown in Figure 2(a). Notably, the predicted targets from TargetScan partially overlapped with those of miR-4665-5p, resulting in a considerable discrepancy compared to the results from miRDB and miRTarBase. To enhance prediction confidence, we selected 28 target genes that both miRDB and miRTarBase consistently predicted, and performed protein–protein interaction (PPI) analysis using the STRING database and Cytoscape software. The results are presented in Figure 2(b).
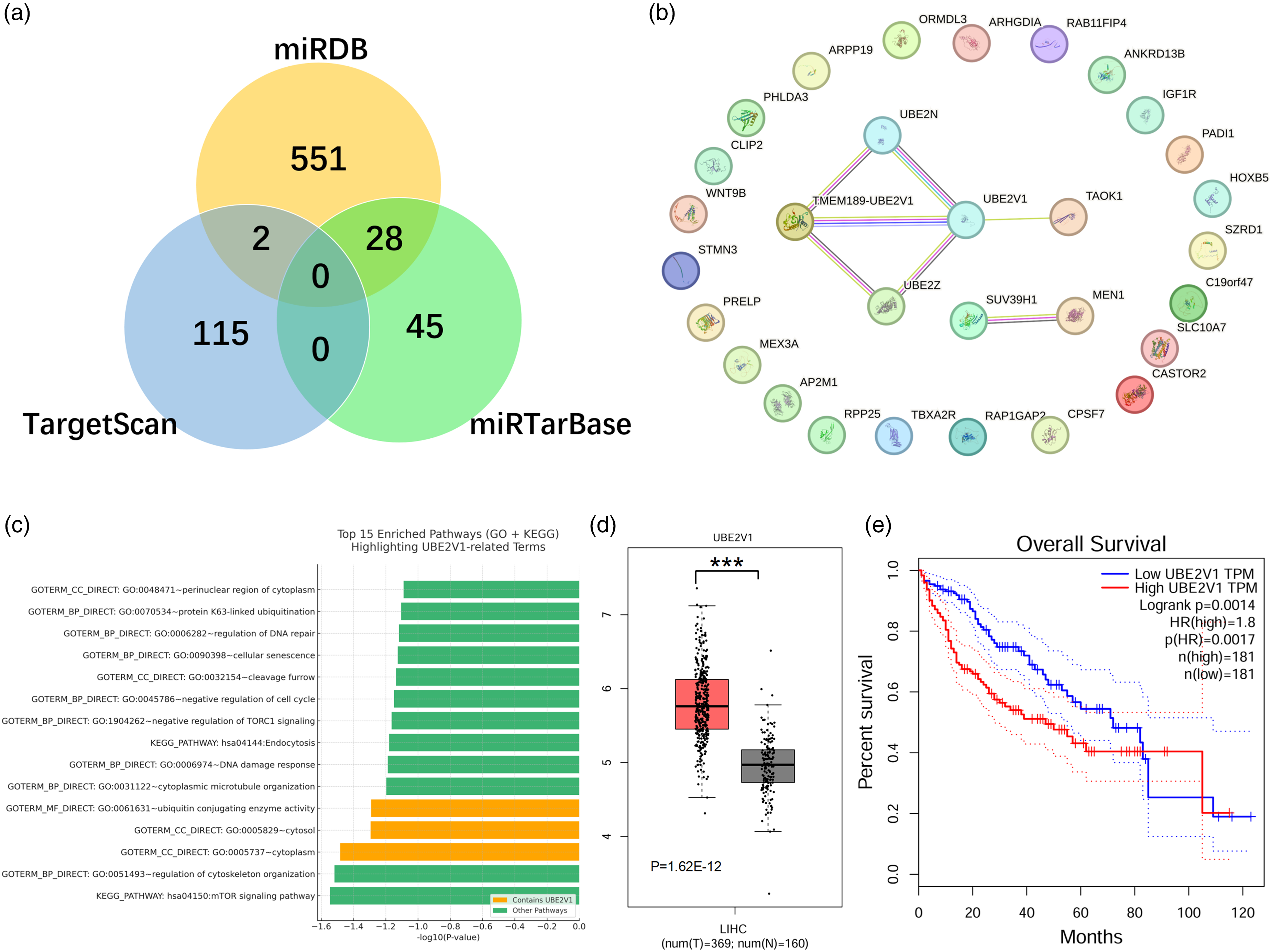
Top 15 enriched pathways identified from DAVID functional annotation analysis, including gene ontology (GO) and KEGG pathway terms. The x-axis represents statistical significance as -log10(P-value). Pathways containing the gene UBE2V1 are highlighted in orange, while other pathways are shown in green. The analysis was based on differentially expressed or miRNA-targeted genes associated with miR-1275. This visualization highlights the involvement of UBE2V1 in key biological processes and signaling pathways, such as ubiquitin-conjugating enzyme activity and cytosolic localization.
The PPI network analysis revealed that UBE2V1, TMEM189-UBE2V1, and UBE2N are key genes potentially regulated by miR-1275. Further GO and KEGG pathway enrichment analyses (Figure 2(c)) demonstrated that the differentially expressed genes are mainly involved in DNA repair, ubiquitination regulation, negative regulation of the cell cycle, and cytoskeleton organization. Among them, UBE2V1 was enriched in several critical functional categories, including ubiquitin conjugating enzyme activity, the cytosol, and regulation of cytoskeleton organization, suggesting that UBE2V1 may play a crucial role in maintaining cellular homeostasis, protein degradation, and signal transduction. Additionally, the significant enrichment of mTOR signaling and endocytosis pathways in the KEGG analysis further supports the hypothesis that UBE2V1 may participate in tumorigenesis by modulating these key signaling cascades. The significantly elevated expression of UBE2V1 in HCC, along with its association with poor overall survival, suggests that UBE2V1 may act as an oncogenic driver and serve as a potential prognostic biomarker and therapeutic target in hepatocellular carcinoma.
Characterization of isolated serum exosomes
Transmission electron microscopy revealed vesicular structures with a saucer- or cup-like shape, consistent with the morphology of exosomes. On average, the particle diameter was 118.1 ± 2.8 nm, and 1-2 such structures of varying sizes were visible per field of view (scale bar: 100 nm) (Figure 3(a)). After a 500-fold dilution of the exosome sample, the results of the particle tracking analysis revealed that the concentration and size range of the detected particles matched those of the exosomes (Figure 3(b)). Flow cytometry analysis showed that the HCC and NL groups expressed the exosomal surface marker proteins CD9, CD63, and CD81 (Figure 1(c)). These results confirmed that the extracted samples were exosomes.
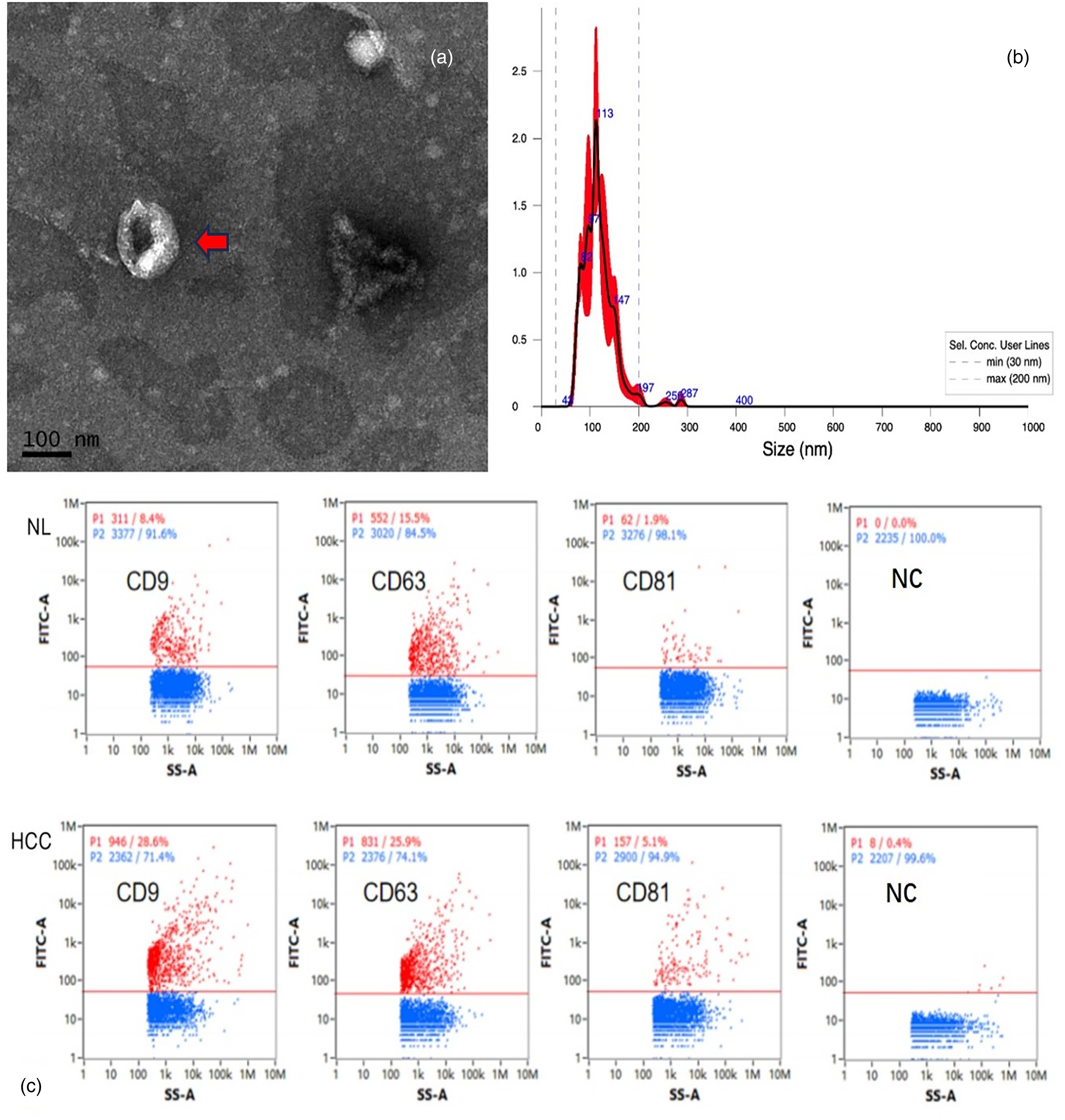
Images of isolated serum exosome identification: (a, c) Transmission electron microscopy of exosomes. (b) Particle size analysis of exosomes. Schematic diagram of the fluorescence detection of exosome marker proteins.
Serum exosomal miR-1275 expression in patients with HCC is significantly decreased
PCR was used to verify the differential expression of miR-1275 between patients with liver cancer and healthy individuals. The results revealed that the relative expression level of miR-1275 in serum exosome samples from 50 HCC patients was significantly lower than that in 50 healthy individuals (P < 0.0001) (Figure 4).
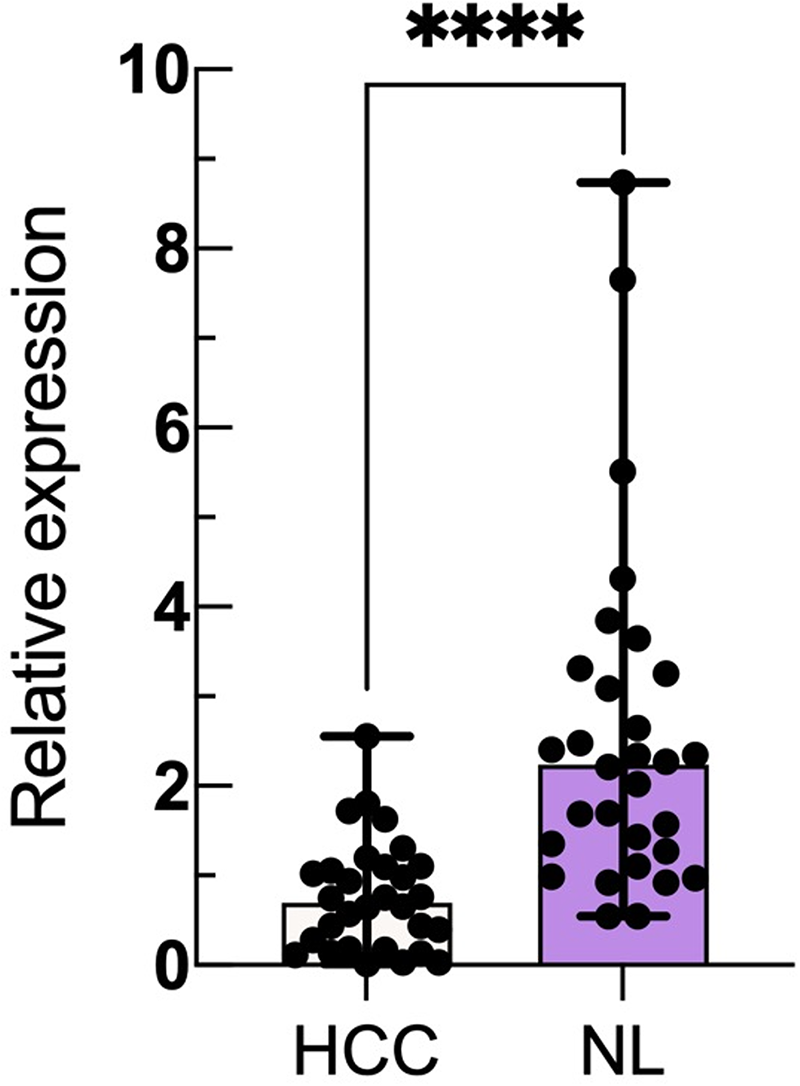
Comparison of exosomal miR-1275 expression levels between the HCC and NL groups; 2−△△Ct represents the relative expression level of miR-1275 in HCC patients and the NL. The expression of miR-1275 in each exosome sample is displayed in scatter plots and histograms, and the median and interquartile range are indicated. (**** P < 0.0001).
Relationships between exosomal miR-1275 expression levels and the clinical characteristics of HCC patients
In the serum samples of 50 HCC patients, we compared exosome-derived miR-1275 levels with the tumor CNLC stage, the presence of liver cirrhosis, tumor size, the number of tumors, lymph node metastasis, vascular invasion, distant metastasis, and the Child-Pugh classification. The results revealed that lower levels of exosomal miR-1275 are associated with more advanced stages of HCC and a poorer prognosis. (Figure 5):
In terms of clinical stage, the expression level of miR-1275 in stage I CNLC patients was significantly higher than in stage II-IV patients (P < 0.05). The expression levels in stages II and III were more significant than in stage IV, but the differences were not statistically significant (P > 0.1). The level of miR-1275 in patients with cirrhosis was lower than that in patients with HCC without cirrhosis, but the difference was not statistically significant. In terms of tumor diameter, patients with an MRI-indicated tumor size of less than 5 cm had greater serum exosome-derived miR-1275 levels than patients with a tumor size of 5 cm or greater, but the difference was not statistically significant. According to the 2022 “Guidelines for the Diagnosis and Treatment of Primary Liver Cancer” from the CNLC staging criteria, patients with ≥4 tumors are no longer considered candidates for curative surgery. A comparison of miR-1275 levels between patients with fewer than 4 and 4 or more tumors revealed statistically significant differences (P < 0.05), indicating that patients with 4 or more lesions had significantly lower miR-1275 expression than those with fewer than 4 lesions. In the presence or absence of lymph node metastasis, miR-1275 levels were significantly lower in patients with lymph node metastasis compared to those without lymph node metastasis (P < 0.01). The results indicated that HCC patients with vascular metastasis had lower miR-1275 expression than did those without vascular metastasis (P < 0.05). A comparison of patients with and without distant metastasis revealed that patients with distant metastasis had significantly lower miR-1275 expression (P < 0.01). Patients were divided into three groups according to the Child–Pugh classification. A comparison of miR-1275 expression between each pair of groups revealed a decreasing trend from Child–Pugh A to C, but none of the differences were statistically significant.
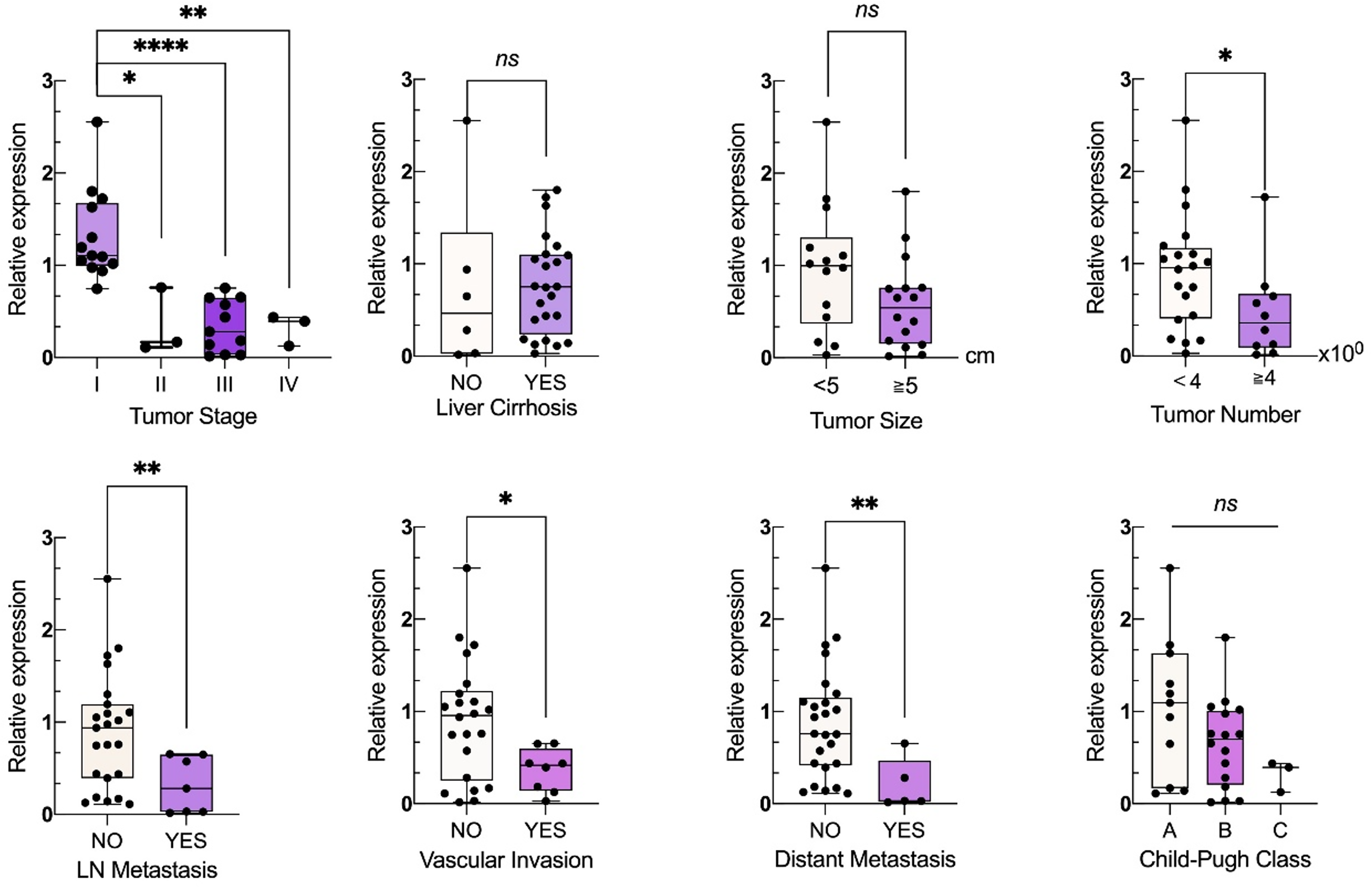
Relationships between exosomal miR-1275 levels and the clinical characteristics of HCC patients. 2−△△Ct represents the relative expression level of miR-1275 in HCC patients, and the expression levels of exosomal miR-1275 in different groups are shown in boxplots and scatter plots. (* : P < 0.05, **: P < 0.01, ***: P < 0.001, ns:no statistical significance).
Value of miR-1275 as a diagnostic biomarker for HCC
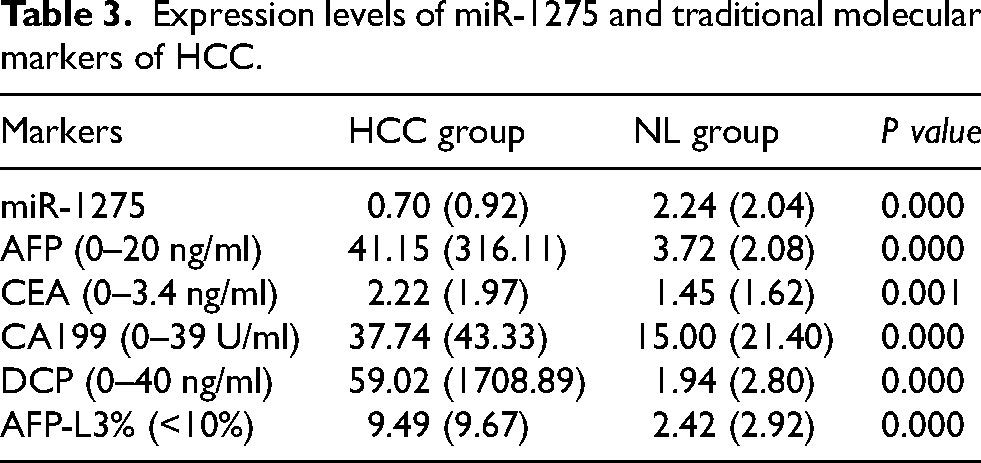
We analyzed the expression levels of commonly used liver cancer biomarkers, including AFP, CEA, CA199, DCP, and AFP-L3%, in both the HCC and NL groups, along with the relative expression of miR-1275 measured by qPCR (Table 3). Significant differences in the expression levels of these biomarkers were observed between the HCC and NL groups (P < 0.05). We then performed a receiver operating characteristic (ROC) analysis to evaluate the diagnostic efficiency of each biomarker for HCC, calculating the area under the curve (AUC). The results indicated that each of the biomarkers had good diagnostic value for liver cancer (P < 0.001), with the AUC values being as follows: miR-1275 (AUC = 0.869), AFP (AUC = 0.902), CEA (AUC = 0.745), CA199 (AUC = 0.808), DCP (AUC = 0.925), and AFPL3% (AUC = 0.898). The diagnostic performance of miR-1275 alone was superior to that of CEA and CA-199, but still inferior to that of AFP, DCP, and AFP-L3% %.
Expression levels of miR-1275 and traditional molecular markers of HCC.
When the optimal cutoff value (0.755) was used, the sensitivity of miR-1275 was 86.7%, and the specificity was 73.3%. Among all the biomarkers tested, the sensitivity of miR-1275 was the second highest for AFP, indicating a high ability to diagnose liver cancer patients accurately. Its specificity ranked fifth, indicating a slightly higher false-positive rate (Table 4, Figure 6(a)).
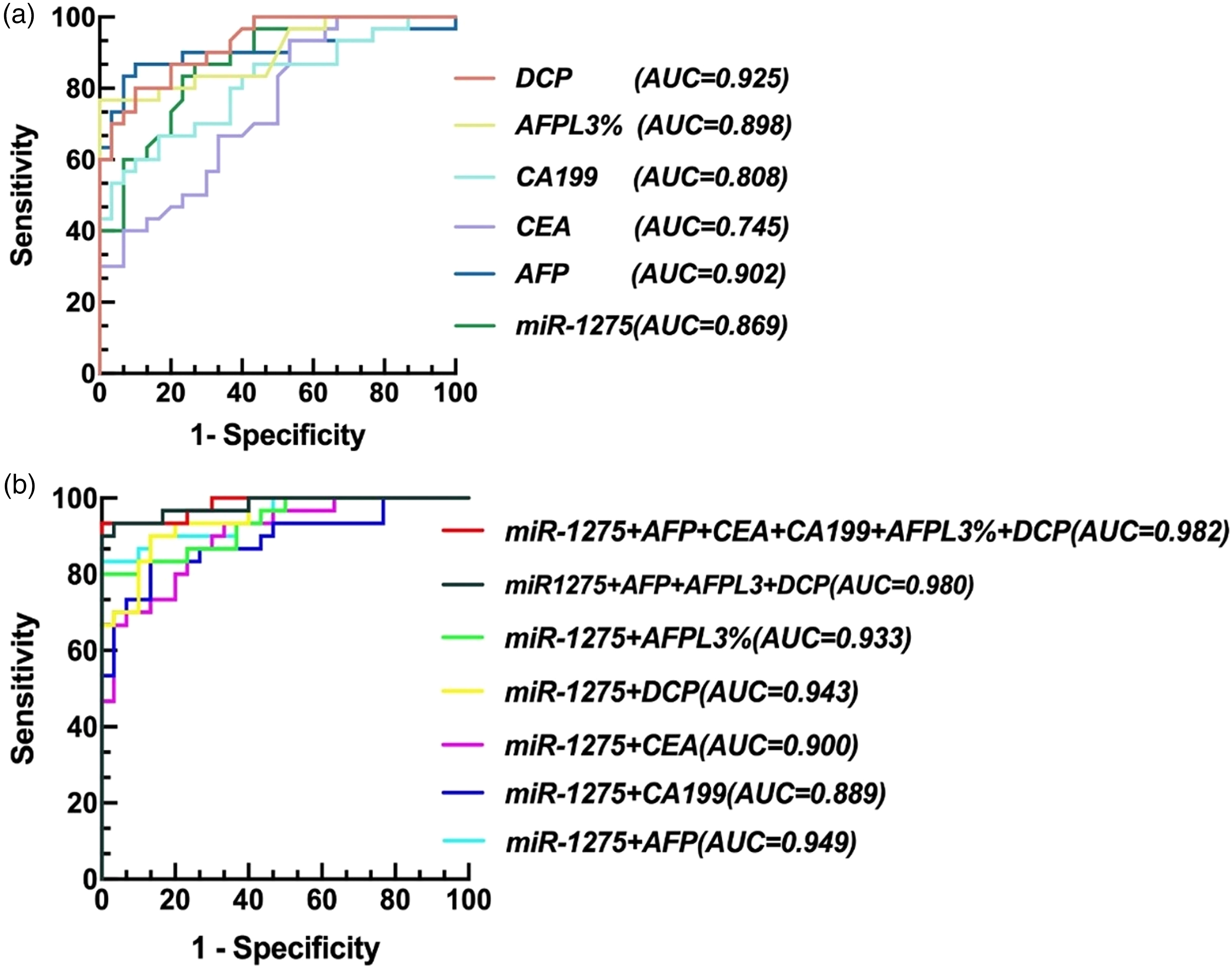
ROC analysis of miR-1275 and traditional molecular markers of HCC: A. ROC analysis of single detection of miR-1275 and traditional molecular markers of HCC. B. ROC analysis of the combined detection of miR-1275 and traditional molecular markers of HCC.
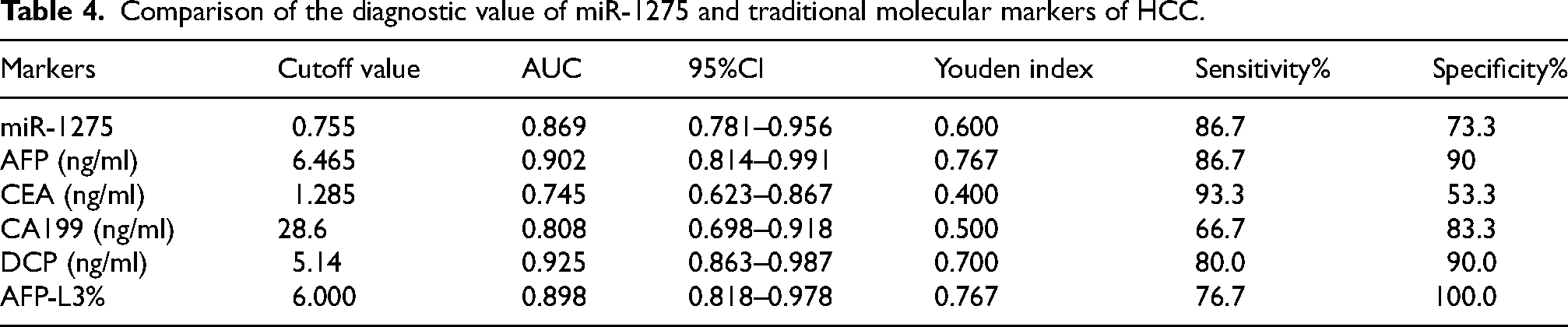
Comparison of the diagnostic value of miR-1275 and traditional molecular markers of HCC.
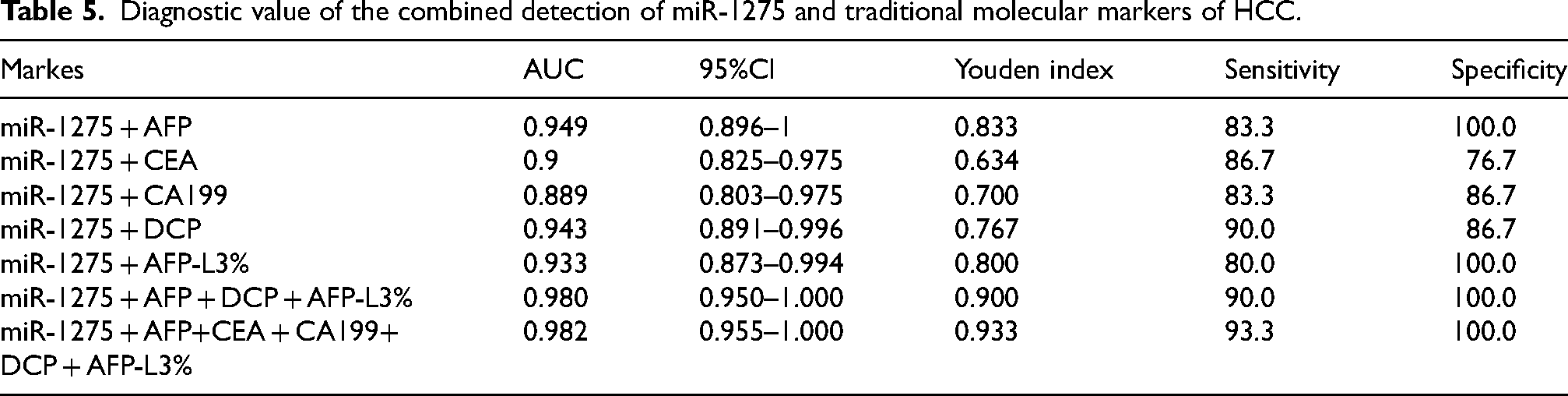
We also evaluated the diagnostic efficiency of miR-1275 in combination with traditional HCC biomarkers. The results indicated that combining miR-1275 with AFP, CEA, CA199, DCP, or AFPL3 improved diagnostic performance compared with the use of these markers individually. When all six markers were tested together, the diagnostic efficiency was the highest (AUC = 0.982), with a sensitivity of 93.3% and a specificity of 100.0%. Notably, the diagnostic efficiency of the combined detection of miR-1275, AFP, DCP, and AFPL3% was almost equivalent to that of the combined detection of all six markers (Table 5, Figure 6(b)).
Diagnostic value of the combined detection of miR-1275 and traditional molecular markers of HCC.
According to the CNLC staging criteria, the 50 HCC patients were divided into early-stage (stages Ia–IIa, n = 24) and intermediate-to-late-stage (stages IIb–IV, n = 26) groups for comparative analysis. In early-stage HCC, exosomal miR-1275 demonstrated moderate diagnostic performance (AUC = 0.784), which was higher than that of CA199 (AUC = 0.754) and CEA (AUC = 0.674), but lower than DCP (AUC = 0.939), AFP-L3% (AUC = 0.939), and AFP (AUC = 0.909) (Figure 7(a)). Interestingly, in intermediate-to-late-stage HCC, exosomal miR-1275 achieved excellent diagnostic value (AUC = 0.946), outperforming conventional biomarkers, among which CEA (AUC = 0.563) and DCP (AUC = 0.661) showed only moderate predictive capacity for metastasis (Figure 7(b)). The combined detection of miR-1275 with AFP, DCP, AFP-L3%, CEA, and CA199 yielded the highest diagnostic accuracy (AUC = 1.000), achieving 100% sensitivity and specificity (Figure 7(c)). These findings support the potential clinical utility of exosomal miR-1275 as a complementary biomarker for early detection and as a robust indicator of disease progression in advanced HCC.
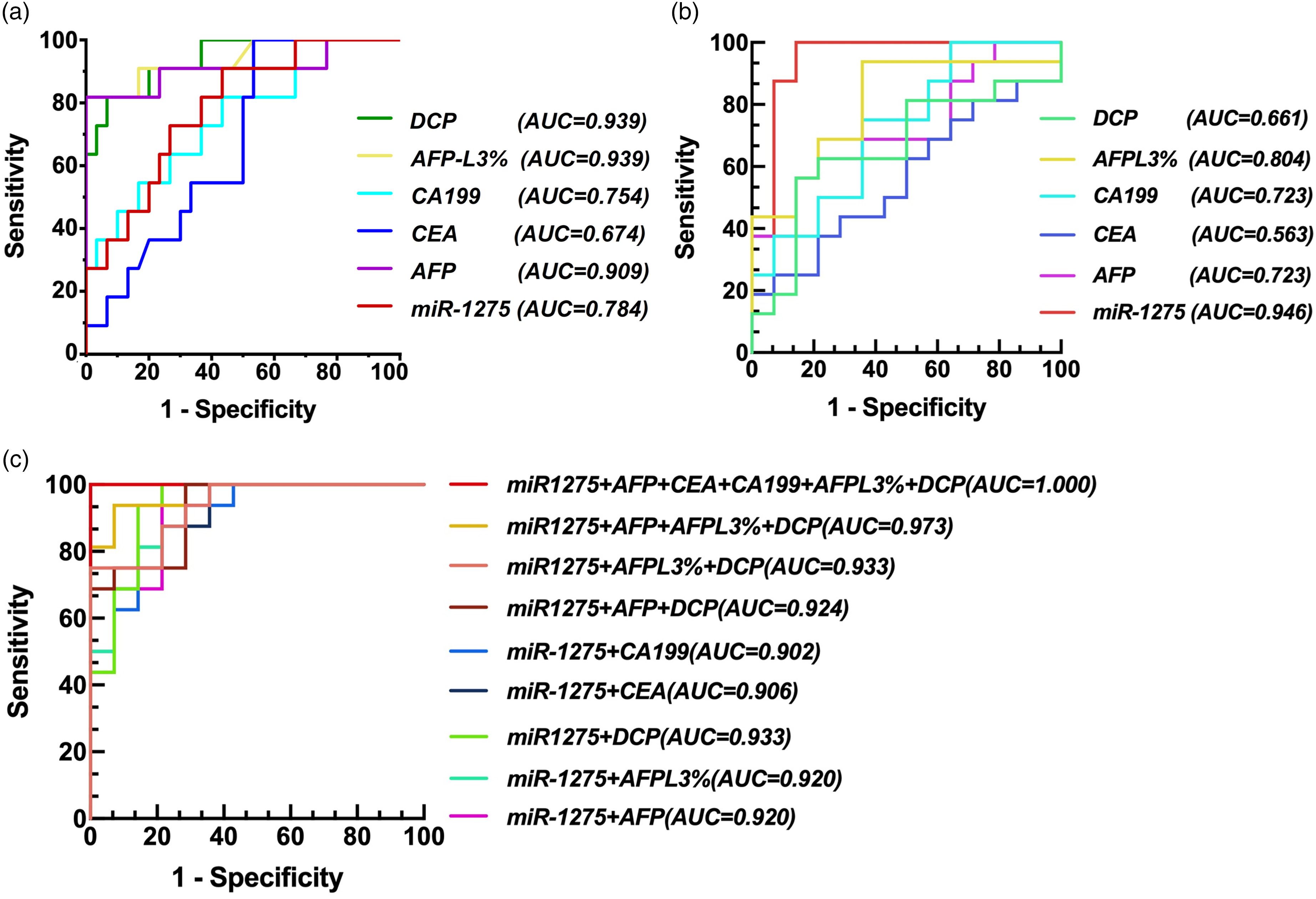
ROC analysis of miR-1275 and traditional molecular markers of HCC. (a) Diagnostic value of miR-1275 and traditional molecular markers of HCC in HCC early diagnosis. (b) Diagnostic value of miR-1275 and traditional molecular markers of HCC in predicting HCC metastasis. (c) Diagnostic value of the combined detection of miR-1275 and traditional molecular markers of HCC in predicting HCC metastasis.
Discussion
HCC remains a global health challenge with limited early diagnostic options and poor prognosis for advanced-stage patients. 17 Early diagnosis and treatment can significantly improve HCC patient survival rates. 18 In this study, we identified serum-derived exosomal miR-1275 as a promising diagnostic and prognostic biomarker. miR-1275 was significantly downregulated in HCC patients compared with healthy controls, consistent with its putative tumor-suppressive role. Analysis of the TCGA PanCanAtlas dataset confirmed reduced expression in HCC tissues, with progressive declines across clinical stages, reinforcing its association with disease progression.
The role of miR-1275 in cancer remains controversial, with reports describing it as both tumor-promoting and tumor-suppressive across different cancer types.14,15,19,20 Previous studies have noted increased miR-1275 levels in serum from liver cancer patients, 20 while others have documented its tumor-suppressive function.21,22 Our findings of reduced exosomal miR-1275 contrast with the report by Pascut et al., who observed elevated serum miR-1275 in DAA-treated HCV patients that later developed HCC in their discovery cohort. This discrepancy may reflect differences in etiology and treatment context (HBV-related, treatment-naïve patients vs. HCV after DAA therapy), analyte compartment (exosomal vs. total serum miRNAs), and study design (discovery microarray vs. validated qRT-PCR). Notably, miR-1275 did not remain significant in their validation phase, suggesting a context-limited effect. These differences highlight the importance of standardized workflows and multicenter validation to reconcile divergent results.
Bioinformatics analysis revealed frequent shallow deletions and neutral copy number status for miR-1275 in HCC, suggesting genomic mechanisms for its reduced expression. Predicted target genes were enriched in pathways related to DNA repair, ubiquitination, and cell cycle regulation. Protein–protein interaction (PPI) analysis further highlighted UBE2V1, TMEM189-UBE2V1, and UBE2N as key targets of miR-1275, implicating them as critical mediators in miR-1275-regulated pathways. Notably, UBE2V1 exhibited significantly elevated expression in HCC tissues within the TCGA dataset. It was strongly associated with poorer patient survival outcomes, suggesting its potential role as an oncogenic driver and prognostic marker.
UBE2V1 (Ubiquitin-Conjugating Enzyme E2 V1), also known as UBE2V or CROC-1, is a ubiquitin-conjugating enzyme variant that lacks intrinsic catalytic activity but functions as a critical regulatory component in the ubiquitination cascade. 23 UBE2V1 primarily functions by forming complexes with UBE2N, facilitating the synthesis of lysine 63-linked polyubiquitin chains that regulate key signaling pathways, including NF-κB, DNA damage repair, and inflammation.24,25 Recent evidence suggests that UBE2V1 contributes to tumorigenesis by modulating PI3K/AKT and MAPK/ERK signaling, further supporting its oncogenic role.26,27 Notably, UBE2V1 overexpression has been associated with a poor prognosis in multiple cancers, suggesting its potential as an oncogenic driver.28,29 However, its upstream miRNAs and context-dependent mechanisms remain incompletely characterized, warranting further investigation to elucidate its therapeutic relevance in disease pathophysiology. Our analysis suggests that downregulation of miR-1275 in HCC may lead to derepression of UBE2V1, a regulator of ubiquitination and NF-κB/PI3K-AKT/mTOR signaling. These pathways are central to tumor proliferation, survival, and immune escape. Thus, loss of miR-1275 could promote HCC progression through dysregulated ubiquitination cascades, highlighting a potential mechanistic link that warrants further validation.Nonetheless, further experimental studies are necessary to elucidate these regulatory mechanisms.
Since miR-1275 is highly susceptible to degradation in peripheral blood, whereas exosome-encapsulated miR-1275 remains stable and participates in intercellular communication, the level of exosomal miR-1275 secreted by hepatocellular carcinoma (HCC) cells holds promise as a potential biomarker. 30 In our study, we successfully isolated and identified exosomes from the serum of HCC patients. The RT–PCR results revealed that serum-derived exosomal miR-1275 was significantly downregulated in HCC patients compared with healthy individuals.
The diagnostic performance of exosomal miR-1275 (AUC = 0.869) surpasses that of traditional markers like CEA and CA-199, although it trails AFP and DCP. However, its integration into multimarker panels (e.g., miR-1275 + AFP + DCP + AFP-L3%) achieves near-perfect diagnostic accuracy (AUC = 0.982, sensitivity = 93.3%, specificity = 100%). This synergy highlights the importance of combining exosomal miRNAs with protein-based biomarkers to improve early detection, particularly for small or radiologically occult tumors.
We further analyzed exosome-derived miR-1275 levels according to HCC stage and found that its diagnostic and prognostic utility varied across disease progression. In early-stage patients, miR-1275 demonstrated moderate diagnostic performance, outperforming CEA and CA199 but remaining inferior to AFP, DCP, and AFP-L3%. In contrast, in intermediate-to-late-stage disease, miR-1275 exhibited excellent diagnostic accuracy (AUC = 0.946), clearly surpassing conventional biomarkers such as CEA and DCP. These findings suggest that exosomal miR-1275 may serve as a valuable complementary biomarker for early detection and as a robust indicator of advanced disease and metastatic potential. Clinically, serum exosomal miR-1275 could be integrated into liquid biopsy protocols to aid in patient stratification, surveillance, and therapeutic decision-making. Moreover, its combination with established protein biomarkers achieved near-perfect diagnostic accuracy, underscoring the added value of a multimarker approach that leverages the complementary strengths of both nucleic acid– and protein-based indicators.
Despite promising findings, the present study has several limitations, including a modest sample size, which may restrict the broader applicability of these results. Validation in more extensive, multicenter cohorts is essential to confirm the clinical utility of miR-1275. Future studies should address these limitations by validating findings in larger, multicenter cohorts with HBV and cirrhosis controls, performing mechanistic experiments, and conducting longitudinal monitoring to evaluate the predictive value of exosomal miR-1275 for recurrence and treatment response.
In summary, this study demonstrates that serum exosomal miR-1275 is a valuable biomarker for HCC diagnosis and prognosis. Its integration into multimarker panels markedly enhances diagnostic accuracy, and its potential regulation of UBE2V1 provides mechanistic insights into tumor progression. Validation in diverse populations and functional studies are warranted to confirm its clinical and biological relevance, with the ultimate goal of incorporating exosomal miR-1275 into precision medicine strategies for HCC management.
Conclusion
In conclusion, serum-derived exosomal miR-1275 represents a highly promising biomarker for hepatocellular carcinoma (HCC), demonstrating significant potential for noninvasive early detection and prognosis assessment. The observed decrease in exosomal miR-1275 levels correlates with advanced disease stages and metastatic progression, highlighting its role as a reliable indicator of disease severity. Furthermore, the integration of exosomal miR-1275 into established biomarker panels significantly enhances diagnostic accuracy, underscoring its clinical utility in improving patient stratification and therapeutic decision-making. Future studies with larger cohorts and detailed mechanistic validations are warranted to confirm these findings and fully establish miR-1275 as a standardized biomarker in clinical practice.
Limitations
This study has several limitations that should be acknowledged. First, the sample size (50 HCC patients and 50 healthy controls) was relatively modest, which may reduce the statistical power and generalizability of the findings. Second, all controls were healthy individuals without liver disease, whereas most HCC patients had underlying HBV infection and cirrhosis. The absence of disease controls, such as patients with chronic HBV or cirrhosis without HCC, may have led to an overestimation of diagnostic specificity. Third, the identification of UBE2V1 as a key miR-1275 target was based on bioinformatic predictions without experimental validation, and functional assays such as luciferase reporter analysis or knockdown/overexpression studies are required to confirm this regulatory relationship. Fourth, the very high diagnostic performance (AUC up to 1.0 for metastasis prediction) may indicate potential overfitting, particularly in a relatively small cohort. Although we applied internal validation methods, additional external and multicenter validation is warranted. Finally, this study was cross-sectional in design; longitudinal studies are needed to determine the dynamic changes of exosomal miR-1275 during disease progression and treatment response.
Footnotes
Acknowledgment
We appreciate the support of many people. We thank Dr. Chen Jianming (Guilin Medical University) for providing guidance in exosome extraction, the Department of Laboratory Medicine, Affiliated Hospital of Guilin Medical University, and the sample donors.
Ethical approval and consent to participate
All the patients who participated in this study provided informed consent. The study was carried out according to the principles of the Declaration of Helsinki and was approved by the Ethics Committee of the Affiliated Hospital of Guilin Medical University (Approval No. 2022YJSLL-25).
Consent for publication
All the authors read and approved the final manuscript.
Authors’ contributions
All the authors contributed to the study's conception and design. YJ provided funding. XJ and KBAK designed the experiments. XJ, HY, and ZQ performed the experiments. HY, QT, ZQ, and SS analyzed the data. YJ, CY, KBAK, and MBM wrote the manuscript. All authors reviewed the manuscript. Yang Jun and Chen Yeng contributed equally to this work and should be considered co–first authors. Maslinda Binti Musa and Khalilah Binti Abdul Khalil were co-corresponding authors.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the Guangxi Natural Science Foundation of China under Grant No.2024JJH140492, the Guangxi Medical and Health Appropriate Technology Development and Promotion Application Project under Grant No.S2018018.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Proprietary interest statement
The authors declare no proprietary or commercial interest in any product, service, or company mentioned in this article.
Availability of data and materials
The datasets analyzed in this study include both original experimental data and publicly available datasets. The serum exosomal RT-qPCR data supporting the findings of this study are available from the corresponding author upon reasonable request.
The public miRNA expression data used for validation were obtained from The Cancer Genome Atlas (TCGA) database via the UCSC Xena platform (![]() ). Specifically, compressed files (TCGA.LIHC.sampleMap_miRNA_HiSeq_gene.gz, TCGA.PANCAN.sampleMap_miRNA_HiSeq_gene.gz, and TCGA.PANCAN.sampleMap_miRNA_GA_gene.gz) were used for bioinformatic analysis.
). Specifically, compressed files (TCGA.LIHC.sampleMap_miRNA_HiSeq_gene.gz, TCGA.PANCAN.sampleMap_miRNA_HiSeq_gene.gz, and TCGA.PANCAN.sampleMap_miRNA_GA_gene.gz) were used for bioinformatic analysis.
