Abstract
Introduction
Immunotherapy benefits gastrointestinal tumor patients. But traditional biomarkers like TMB and MSI can’t precisely identify beneficiaries. Phosphatidylinositol - 3,4,5 - triphosphate - dependent Rac exchange factor 2 (PREX2) plays a complex role in tumorigenesis.
Methods
In a retrospective study of 1764 patients (1385 colorectal, 379 gastric), NGS of 639 genes and PD - L1 staining were done.
Results
In colorectal cancer, PREX2 mutations were associated with increased TMB, MSI, TMB-H, and MSI-H. Mechanistically, this is related to an increased number of tumor pathway mutations, higher PD - L1 expression, increased immune infiltration, and immune - related pathway enrichment. Cetuximab and Bortezomib sensitivity was higher in PREX2 - mutated colorectal cancer. In gastric cancer, there are no established immune associations with PREX2 mutations.
Conclusion
PREX2 mutations may serve as a novel predictive biomarker for immunotherapy in CRC, potentially enhancing antitumor immunity via microenvironment modulation, but lack predictive value in GC. These findings highlight PREX2's role in refining patient stratification for immune checkpoint inhibitors.
Introduction
Immunotherapy has become increasingly prominent in the treatment of gastrointestinal tumors. However, traditional biomarkers such as Tumor Mutational Burden (TMB) and Microsatellite Instability (MSI) have proven insufficient for accurately identifying patients who may benefit from these therapies. 1 While TMB is associated with the efficacy of immune checkpoint inhibitors (ICIs) in certain cancer types, its predictive capability lacks universal applicability. For instance, combining TMB with PD-L1 expression has shown better predictive outcomes, but high TMB alone does not always correlate with immunotherapy benefits. 2 Furthermore, the lack of standardized determination methods and threshold criteria for TMB restricts its clinical application. 3 Similarly, MSI demonstrates predictive efficacy in some cancers, such as colorectal cancer, but its scope is limited. Tumors with high MSI often exhibit high TMB, which may explain their sensitivity to immunotherapy. However, MSI does not encompass all genotypes sensitive to immunotherapy, potentially excluding certain patients. 4 Therefore, the exploration of novel biomarkers is crucial to improve patient prognosis and optimize treatment strategies.
Phosphatidylinositol-3,4,5-triphosphate-dependent Rac exchange factor 2 (PREX2) has emerged as a protein closely linked to tumorigenesis and progression.5–7In hepatocellular carcinoma (HCC), the S1113R mutation in PREX2 promotes cell migration, activates the AKT pathway, and impairs the HectH9-mediated ubiquitination process. 5 Furthermore, this gain-of-function mutation enhances PREX2 protein stability, promotes cell proliferation, and correlates with increased invasiveness in HCC. In melanoma, PREX2 functions as an upstream activator of the RAC1-GTPase pathway. Although not essential for melanoma initiation and progression, loss of PREX2 renders tumors more sensitive to clinically relevant therapies targeting the MAPK pathway. Combined targeting of the MAPK pathway and the PREX2/RAC1/PI3Kβ axis has been shown to provide significant therapeutic benefits, exceeding those of MAPK inhibitor monotherapy in BRAF-mutant melanoma. 8 These diverse functions underscore PREX2's multifaceted impact on tumor biology and the regulation of the tumor microenvironment. However, the specific mechanisms of PREX2 mutations in gastrointestinal cancers are still unclear.
Recent research highlights the role of PREX2 in radiotherapy resistance in colorectal cancer (CRC). PREX2 inhibits the cGAS/STING/interferon (IFN) pathway, suggesting its potential as a biomarker and therapeutic target for radiation resistance. 6 Thus, PREX2 is important in regulating the tumor microenvironment and the response to radiotherapy. Understanding this mechanism helps identify patient populations that may benefit and also offers new insights for future clinical treatments. In addition, research shows that PREX2 mutations decrease the effectiveness of immunotherapy in gastric cancers (GCs). The median progression - free survival (PFS) for patients with wild - type PREX2 was 5.0 months (95% CI, 2.0–8.0 months), while for those with PREX2 mutations it was 2.4 months (95% CI, 0.5–4.3 months; p = 0.037). 9 However, no relevant studies on CRCs have been reported so far.
To investigate the role of PREX2 in gastrointestinal tumors, especially in immunotherapy, we retrospectively analyzed the genetic testing data of patients with gastrointestinal tumors. Then, we combined these data with mRNA data to analyze the molecular characteristics of patients with PREX2 mutations and explore potential molecular mechanisms. Our study will offer new insights into the role of PREX2 in the precision treatment of gastrointestinal tumors.
Methods
Patients and sample characteristics
This study, conducted at Shanghai Second Affiliated Hospital of Naval Medical University from April 2021 to April 2024, enrolled a total of 1385 patients with pathologically diagnosed colorectal cancer (CRC) and 379 patients with pathologically diagnosed gastric cancer (GC). Each participant had to undergo a pathological diagnosis and was required to provide both tumor tissue and paired blood samples. Cancer diagnoses were initially made based on clinical evaluations and X - ray findings and were later confirmed by histological analysis of tumor biopsies. The exclusion criteria for this study were as follows: cases where CRC or GC was not pathologically confirmed, and cases in which tissue or blood samples were not provided or the cell blocks contained less than 20% of tumor cells. Clinical data like age and gender were obtained from the medical records.
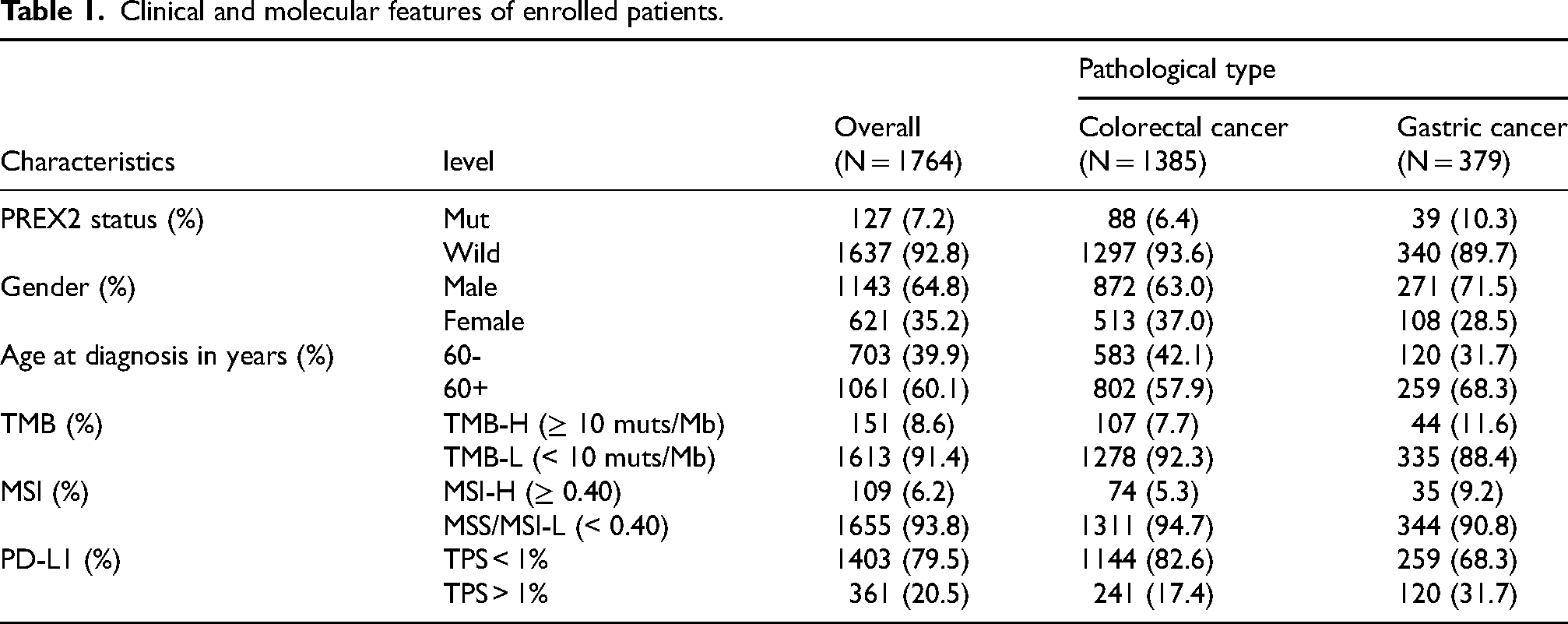
A summary of the enrolled patients is shown in Table 1, and detailed information for each patient can be found in Table S1. All participants provided written informed consent, and the study was approved by the institutional review board of our hospital (20211218).
Clinical and molecular features of enrolled patients.
DNA extraction and library construction
According to the manufacturer's protocols, the following procedures were carried out. First, tumor DNA and blood genomic DNA were extracted separately. The human tissue DNA extraction kit (Shanghai YunYing) was used for tumor DNA extraction, and the human blood genomic DNA extraction kit (Shanghai YunYing) was used for blood genomic DNA extraction. Then, the DNA was eluted in an elution buffer, and its concentration and purity were evaluated by a NanoDrop spectrophotometer. After that, the DNA was stored at −20 °C until use. Next, library preparation was performed with the VAHTS Universal DNA Library Prep Kit for Illumina. Subsequently, target enrichment was carried out using Shanghai YunYing's optimized probes, which target the exons and some introns of 639 cancer-related genes. Finally, sequencing was performed on an Illumina NextSeq500 platform following the manufacturer's protocols.
Next-generation sequencing (NGS)-based assay and bioinformatics analysis
FastQC software (version 0.11.2) and a customized Python script were utilized to screen FASTQ files. During this process, adaptor sequences and sequences with a quality (Q) value below 30 were removed. Subsequently, clean reads were mapped to the reference human genome GRCh37/hg19 by means of BWA (Burrows - Wheeler Aligner version 0.7.7). Next, BAM files were realigned and recalibrated using GATK3.5, which was also employed to detect mutations. To reduce potential polymerase chain reaction bias, Picard MarkDuplicates (version 1.35) was used to remove duplicate sequences. VarScan (version 2.3.2) was then used to select single nucleotide variations (SNVs) that met the following criteria: a depth of at least 100, a read count of at least 10, and an allele frequency of at least 5% (or at least 1% if in a hotspot). Finally, Pindel (version 0.2.5b8) was used for insertion or deletion (indel) detection with default parameters, requiring at least 5 unique reads.
Compared with matched normal samples, we named and functionally annotated somatic SNVs and InDels of tumors using MuTect v.1.1.4 and Varscan2 v.2.3.9 software. Mutations with a variant allele frequency of ≥ 5% were defined as high - confidence mutations (≥ 1% for hotspots). We calculated the tumor mutational burden (TMB) as follows: First, we used the number of all somatic, coding, base substitution, and indel mutations per megabase (including synonymous mutations). Then, we divided the total number of mutations counted by the size of the coding region of the targeted territory (1.36 Mb of the coding genome) to obtain the TMB per megabase. All samples’ microsatellite instability (MSI) scores were calculated using MSIsensor 10 with default parameters. MSIsensor is a software tool for quantifying MSI in genome sequencing data using tumor - only or paired tumor - normal samples. For the MSI detection of tumor - only patterns, we used 29 microsatellite sites as input files. The MSI score was defined as the percentage of unstable microsatellites among all microsatellites used. According to the characteristics of the MSIsensor model and the characteristics of YunYing's 639 - gene sequencing panel, samples with an MSI score greater than 0.40 were defined as MSI - H, and those less than 0.40 were defined as MSS/MSI - L. Each microsatellite site had at least 20 spanning reads and single - nucleotide mutations. Gene copy number gains were interpreted along with the amplicon sequencing data using the oncoCNV method as described in a previous study. 11
PD-L1 expression test
The PD - L1 expression level of each patient was determined through the Dako 22C3 pharmDx system (Agilent Technologies Inc., Santa Clara, CA, USA) assay, and the results were presented as a tumor proportion score (TPS) 12 and combined positive score (CPS). 13
Data collection and statistical analysis
All data of 625 CRCs (colorectal cancers) and 415 GCs (gastric cancers) with gene mutation and mRNA expression data used in this research were obtained from the public database, cbiportal (https://www.cbioportal.org/datasets). The R package “limma” was used for GSEA (gene set enrichment analysis). In GSEA, enrichment was performed using all differentially expressed genes, with a false discovery rate (FDR) < 0.05 (adjusted using the Benjamini-Hochberg method), and an absolute log2 fold-change (|log2FC|) > 0.” GO (Gene Ontology) and KEGG (Kyoto Encyclopedia of Genes and Genomes) enrichment analyses were conducted via the R package “clusterProfiler”. Pathways with adjusted p-values (Benjamini-Hochberg method) < 0.05 were considered significantly enriched. IOBR, 14 a computational tool for immune tumor biology research, was used. Based on the expression profile (bulk RNA-seq data), the ESTIMATE and MCPcounter methods were utilized to calculate the tumor immune infiltrating cell score.
Oncopredict (v1.2), an R package designed for drug response prediction and drug - gene association prediction based on data from the Sanger's Genomics of Drug Sensitivity in Cancer (GDSC v1) database, 15 was used to predict the drug sensitivity of samples to 367 drugs. 16 We utilized this package to predict the sensitivity of samples from CRCs and GCs to Cetuximab and Bortezomib (targeting plasma cells).
The R package “maftools” 17 was used to visualize the prevalence and distribution of genomic alterations. The boxplots and stacked bar charts were drawn using the R package “ggplot2”. Subsequently, the nonparametric Wilcox test was used to test for the significance of the difference in means between the two populations. For exploratory post hoc comparisons, multiple testing adjustments were performed using the Benjamini-Hochberg method to control the false discovery rate.
Results
Patient characteristics
The present study included a total of 1764 patients, with 1385 having colorectal cancer (CRC) and 379 having gastric cancer (GC). The mean age at diagnosis for the study participants was 61.71 years (ranging from 13–90 years, with a median of 63 years). Table 1 summarizes additional patient cohort characteristics, and more detailed information can be found in Table S1. All participants completed the targeted sequencing, which encompassed all exons and partial introns of the 639 genes listed in Table S2.
Higher TMB and MSI values occurred in PREX2 mutated CRCs
TMB and MSI are crucial biomarkers for gastrointestinal tumors. Our study aimed to explore the impact of PREX2 on TMB and MSI. To this end, we analyzed the differences in TMB and MSI between groups with and without PREX2 mutations.
As depicted in Figure 1, PREX2 has a notable effect on TMB and MSI in colorectal cancer (CRC). In CRC patients, those with PREX2 mutations had significantly higher TMB (Figure 1A) and MSI (Figure 1C) values compared to those without PREX2 mutations. Specifically, the mean TMB values were 28.71 muts/Mb and 5.33 muts/Mb respectively (p < 0.0001), and the mean MSI values were 0.17 and 0.07 respectively (p < 0.001). Moreover, within the distribution, patients with PREX2 mutations also had a higher proportion of high TMB (TMB - H) (Figure 1B) and high MSI (MSI - H) (Figure 1D), with the proportions being 40% vs. 6% for TMB - H (p < 0.0001) and 22% vs. 4% for MSI - H (p < 0.0001).
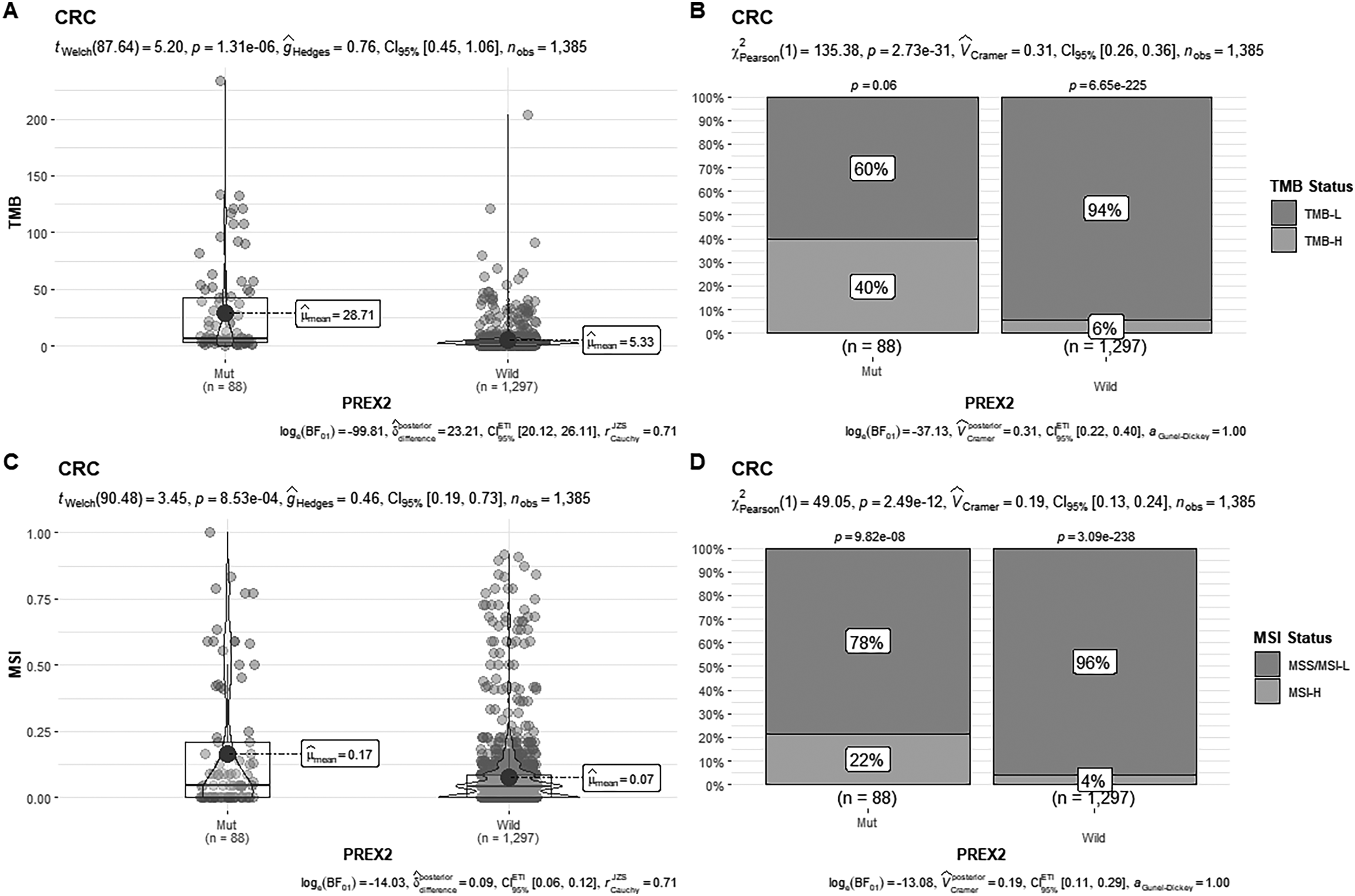
The effect of PREX2 mutation in TMB and MSI in CRC.
In contrast, for gastric cancer (GC) (Figure S1), although PREX2 showed a similar trend regarding TMB and MSI as observed in CRC, no significant differences were detected (Figure S1A and Figure S1C). However, in GC patients with PREX2 mutations, the proportion of TMB - H (Figure S1B) and MSI - H (Figure S1D) was also significantly higher, with the proportions being 40% vs. 6% for TMB - H (p = 0.02) and 22% vs. 4% for MSI - H (p = 0.16).
Consequently, we conclude that the effect of PREX2 mutations on TMB and MSI is significantly greater in CRC than in GC.
Different mutation profiles and higher frequency of tumor signaling pathway alterations in PREX2 mutated CRCs
To understand the mechanism underlying the differential impact of PREX2 mutations on tumor mutational burden (TMB) and microsatellite instability (MSI), we studied the differences in gene mutations between colorectal cancer (CRC) and gastric cancer (GC).
Firstly, we analyzed the effects of PREX2 mutations on other mutations in CRC and GC. The results are shown in Figures 2A and 2D. There were significant differences in the mutation profiles of PREX2 - mutant CRC and PREX2 - mutant GC. In the PREX2 - mutant GC group, the top 20 mutated genes were APC, TP53, KRAS, LRP1B, KMT2C, FAT1, PIK3CA, PRKDC, BRCA2, ATM, KMT2A, ROS1, BRAF, RANBP2, SPTA1, FBXW7, NF1, PTEN, ATR, and SETD2. In contrast, in the PREX2 - mutant CRC group, the top 20 mutated genes were TP53, LRP1B, ARID1A, ERBB2, KMT2C, PIK3CA, SPTA1, ATM, CDH1, PBRM1, ALK, ERBB3, FAT1, KMT2D, PHGR1, PTCH1, RNF43, SMAD4, TSC1, and EGFR. The detailed information of top 20 mutated genes of PREX2 mutation across CRC and GC as shown in Table S3.
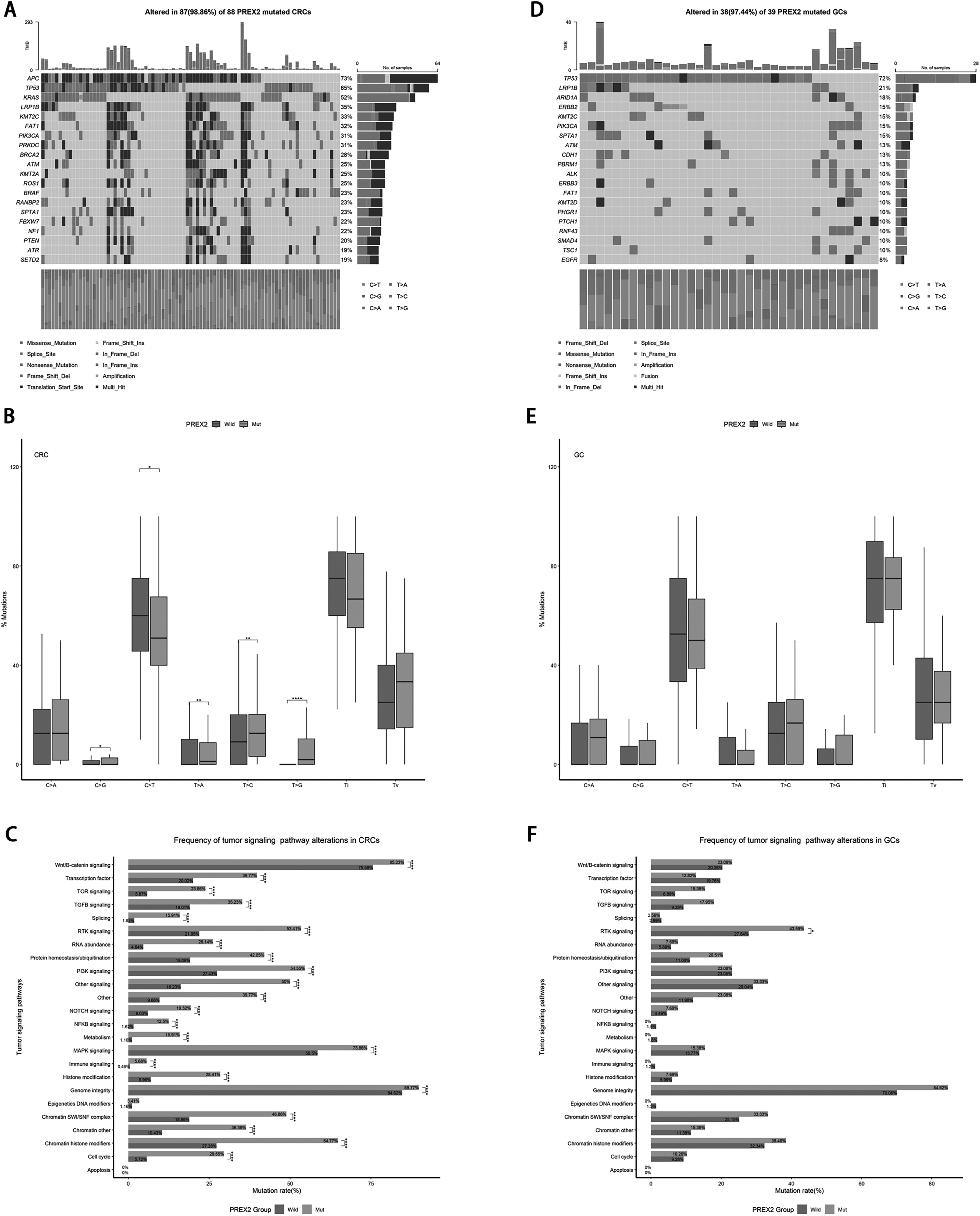

The effect of PREX2 mutation in gene mutation in CRC and GC.
To further understand the impact of PREX2 mutations on single nucleotide variants (SNPs), we used the titv function from the R package “maftools” to analyze the PREX2 groups in CRC and GC, followed by differential analysis. The results showed that in CRC, the proportion of SNPs C > G (p < 0.05), T > A (p < 0.01), T > C (p < 0.01), and T > G (p < 0.0001) was significantly higher in the PREX2 - mutant group than in the wild - type, while C > T was significantly lower in the PREX2 - mutant group (p < 0.05) (Figure 2B and Table S4). Although the overall trend was consistent in GC, no significant differences were observed (Figure 2E and Table S5).
In view of the differences in the impact of PREX2 mutations on gene mutations and SNPs in CRC and GC, we further investigated the influence of PREX2 mutations on tumor signaling pathways. In CRC, PREX2 mutations, apart from having no significant effect on Apoptosis and Epigenetics DNA modifiers, significantly increased the mutation rates of 22 signaling pathways (Figure 2C and Table S4), including the immune signaling pathway (5.68% vs. 0.48%, p < 0.0001) and other tumor signaling pathways such as Wnt/B - catenin signaling (85.23% vs. 75.58%, p < 0.0001), RTK signaling (53.41% vs. 21.95%, p < 0.0001), and PI3K signaling (54.55% vs. 27.43%, p < 0.0001). In contrast, in GC, PREX2 mutations only significantly elevated the mutation rate of the RTK signaling pathway (43.59% vs. 27.84%, p < 0.05), while other pathways showed similar trends but did not reach significance (Figure 2F and Table S5). The detailed information of frequency of tumor signaling pathway alterations of PREX2 mutation across CRC and GC as shown in Table S3.
Immune-related pathways enriched in PREX2 mutated CRCs based on differential expressive gene analysis
Based on the above results, we further conducted differential gene expression analysis to understand the impact of PREX2 mutations on tumor - related pathways. As depicted in Figure 3A, in comparison to CRC patients without PREX2 mutations, those with PREX2 mutations had 2157 upregulated genes and 5309 downregulated genes, and the corresponding gene expression profile is illustrated in Figure 3B.
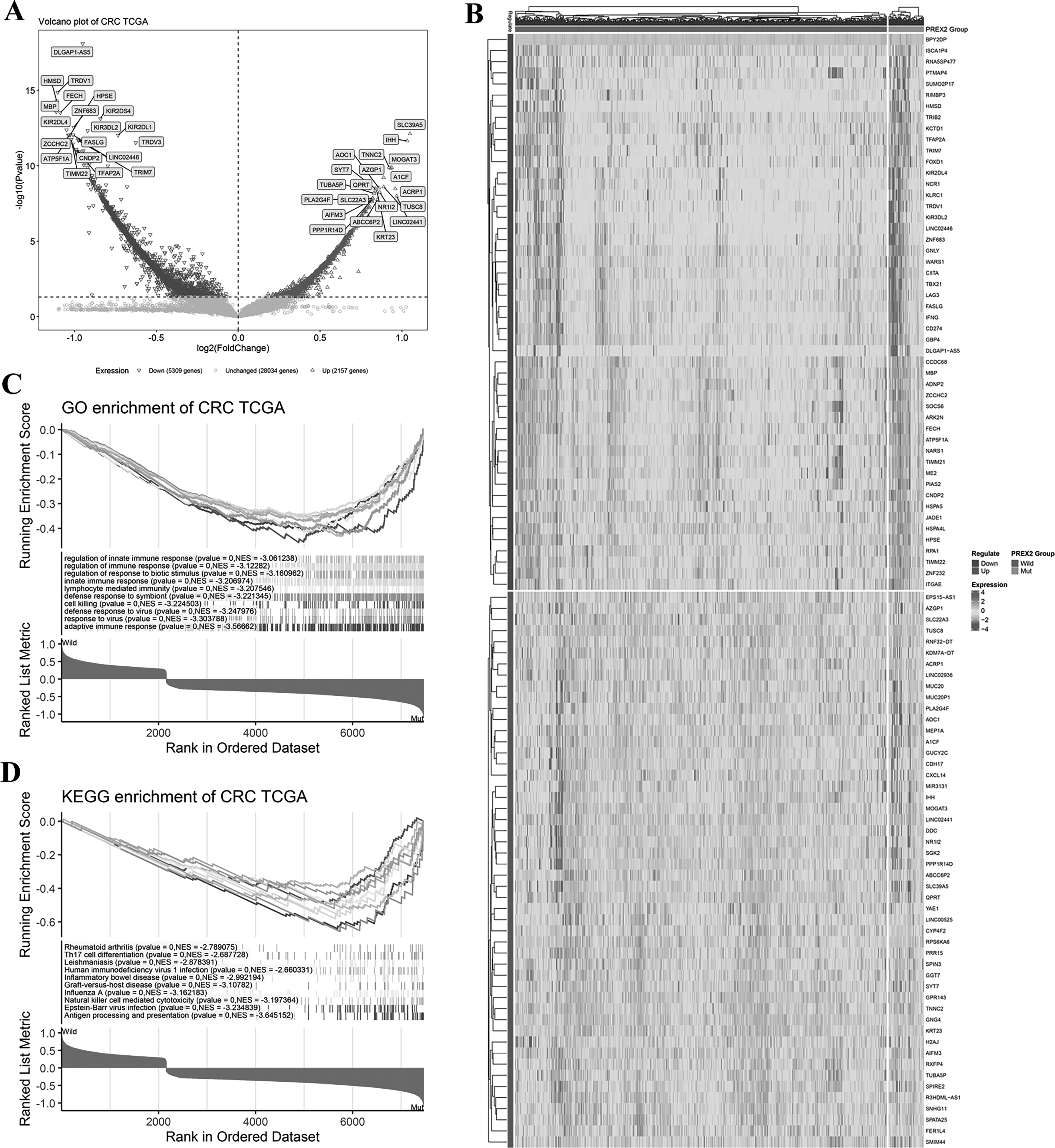
Differential expressive gene analysis based on PREX2 status in CRC.
To understand the pathways influenced by these differential genes, we then utilized the gene expression data to carry out GO and KEGG enrichment analysis (as presented in Figures 3C and 3D). In the GO enrichment analysis, among the top 10 enriched pathways, several immune - related pathways exhibited significant positive correlations with PREX2 mutations. These pathways included adaptive immune response (NES = −3.56662), lymphocyte - mediated immunity (NES = −3.207546), innate immune response (NES = −3.206974), regulation of immune response (NES = −3.12282), and regulation of innate immune response (NES = −3.061238). In the KEGG enrichment analysis, within the top 10 enriched pathways, pathways such as Antigen processing and presentation (NES = −3.645152), Natural killer cell - mediated cytotoxicity (NES = −3.197364), and Th17 cell differentiation (NES = −2.687728) also had significant correlations.
However, in GC, differential gene expression analysis was carried out based on PREX2 status. The results showed that, compared with GC patients without PREX2 mutations, those with PREX2 mutations had 3263 upregulated genes and 1332 downregulated genes (Figure S2A), and the corresponding gene expression profile is presented in Figure S2B. GO and KEGG enrichment analysis (Figure S2C and Figure S2D) did not show any immune - related pathways in the top 10 enriched pathways.
Higher PD-L1 expression and immune infiltration level in PREX2 mutated CRCs
Based on the GESA analysis, we explored the impact of PREX2 mutations on PD - L1 expression. Figures 4A and S3A present the PD - L1 staining results of four representative CRC patients and GC, showing both high and low expression levels. As shown in Figure 4B, in CRC, patients harboring PREX2 mutations exhibited a significantly higher proportion of high PD-L1 expression compared to wild-type counterparts under various thresholds: TPS ≥ 1% (26% vs. 17%, p = 0.03), TPS ≥5% (18% vs. 8%, p = 0.000794), and TPS ≥ 10% (11% vs. 5%, p = 0.00945) (Table S6; Figures S4A and S4B). In contrast, in the gastric cancer (GC) cohort, no significant difference was observed under the TPS ≥ 1% threshold (Figure S3B). However, when the PD-L1 threshold was set to CPS ≥ 5, the PREX2 mutation group showed a significantly lower proportion of high PD-L1 expression compared to the wild-type group (10% vs. 27%, p = 0.02) (Table S7; Figure S4C).
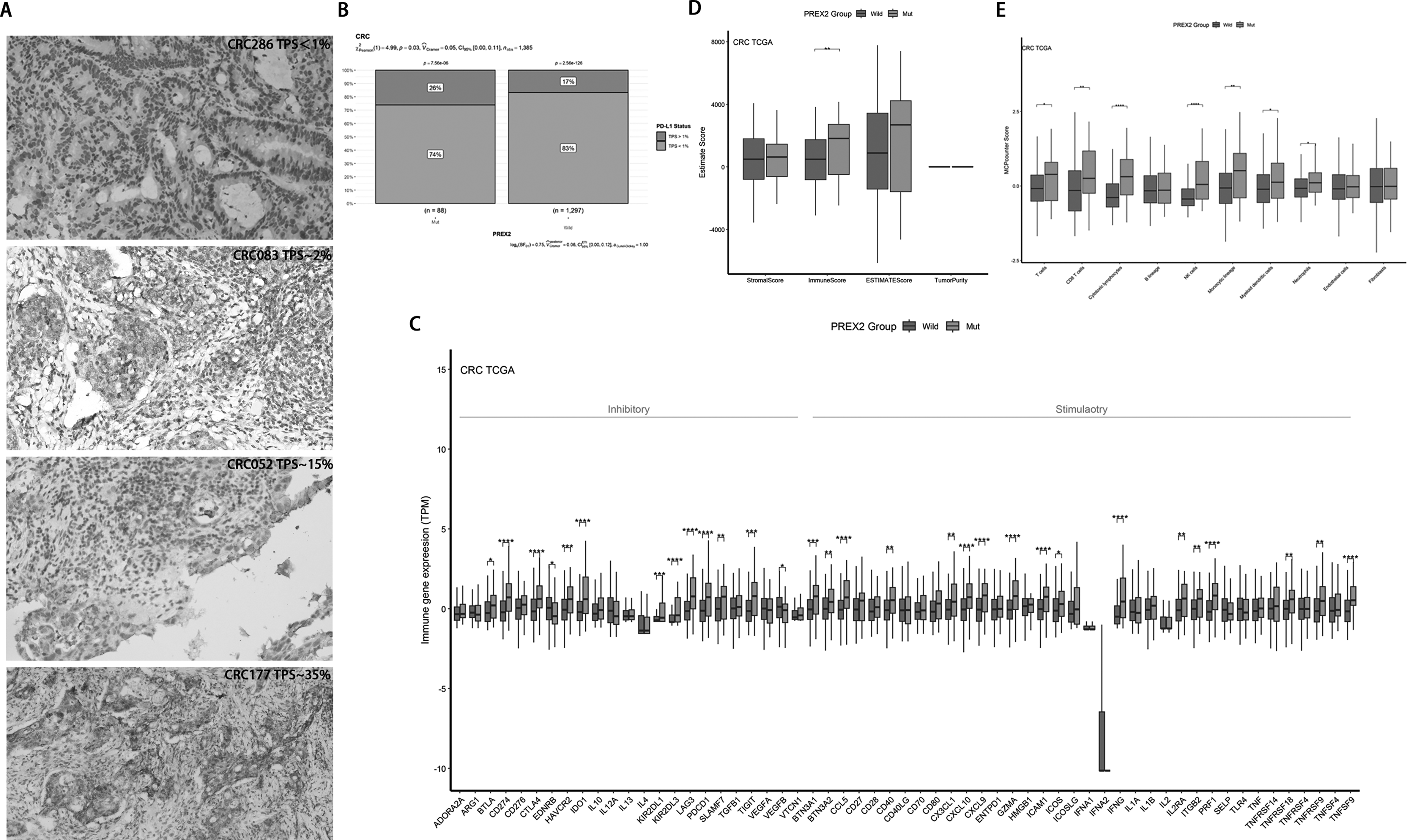
The effect of PREX2 mutation in PD-L1 expression and immune infiltration in CRC.
We identified 59 immune checkpoints according to a previous study 18 and calculated the relative percentages of immune infiltrating cell infiltration and checkpoints. In CRC (Figure 4C and Table S8), the analysis of immune checkpoint expression differences revealed that compared to non - mutant patients, PREX2 mutant patients had significantly higher expression levels of immune inhibitory genes, except for VEGFB (p < 0.05), EDNRB (p < 0.05), BTLA (p < 0.05), CD274 (p < 0.0001), CTLA4 (p < 0.0001), HAVCR2 (p < 0.001), IDO1 (p < 0.0001), KIR2DL1 (p < 0.001), KIR2DL3 (p < 0.0001), LAG3 (p < 0.0001), PDCD1 (p < 0.0001), SLAMF7 (p < 0.01), and TIGIT (p < 0.001). Additionally, the expression levels of immune stimulatory genes, including BTN3A1 (p < 0.001), BTN3A2 (p < 0.01), CCL5 (p < 0.0001), CD40 (p < 0.01), CX3CL1 (p < 0.01), CXCL10 (p < 0.0001), GZMA (p < 0.0001), ICAM1 (p < 0.0001), ICOS (p < 0.05), IFNG (p < 0.0001), IL2RA (p < 0.01), ITGB2 (p < 0.01), PRF1 (p < 0.0001), TNFRSF18 (p < 0.01), TNFRSF9 (p < 0.01), and TNFSF9 (p < 0.0001), were also significantly higher in PREX2 mutant patients. In GC (Figure S3C and Table S9), compared to non - mutant patients, PREX2 mutant patients had significantly lower expression levels of immune inhibitory genes ADORA2A (p < 0.05), EDNRB (p < 0.001), and VTCN1 (p < 0.05), while only ENTPD1 (p < 0.01), and immune stimulatory genes CD70 (p < 0.05), CXCL10 (p < 0.05), GZMA (p < 0.05), HMGB1 (p < 0.05), and IFNG (p < 0.05) had significantly higher expression levels.
The immune infiltration analysis indicated that in CRC, the immune score of PREX2 mutant patients was significantly higher than that of unmutated patients (Figure 4D, p < 0.01), but no significant results were observed in GC (Figure S3D). The immune infiltrating cell analysis showed that in CRC, compared to unmutated patients, PREX2 mutant patients had significantly higher infiltration levels of T cells (p < 0.05), CD8T cells (p < 0.01), cytotoxic lymphocytes (p < 0.0001), NK cells (p < 0.0001), monocytic lineage cells (p < 0.01), myeloid dendritic cells (p < 0.05), and neutrophils (p < 0.05) (Figure 4E). In contrast, in GC (Figure S3E), only the infiltration levels of endothelial cells (p < 0.01) showed significant differences between the groups, with PREX2 mutant patients having significantly lower infiltration levels than unmutated patients. The detailed information is provided in Table S8 and Table S9.
Higher predicted sensitivity of immunotherapy agents in PREX2 mutated CRCs
We further analyzed how PREX2 mutations impact the sensitivity to immunotherapy agents. Given that the GDSC (v1) database only contains Cetuximab and Bortezomib as tumor - immune drugs, we specifically carried out a drug sensitivity prediction analysis for these two drugs. As reported in Figure 3A of the previous study, 16 a higher predicted drug sensitivity score means greater resistance to the drug. The results, presented in Figure 5A and Figure 5B, show that PREX2 mutations significantly enhance the sensitivity of CRC patients to Cetuximab (Sensitivity Score, 6.03 vs. 6.11, p < 0.01) and Bortezomib (Sensitivity Score, - 5.98 vs. - 5.86, p = 0.03). However, in GC patients, no significant results were observed for Cetuximab (Sensitivity Score, 6.09 vs. 6.10, p = 0.66) (Figure 5C) or Bortezomib (Sensitivity Score, - 5.87 vs. - 5.88, p = 0.93) (Figure 5D).
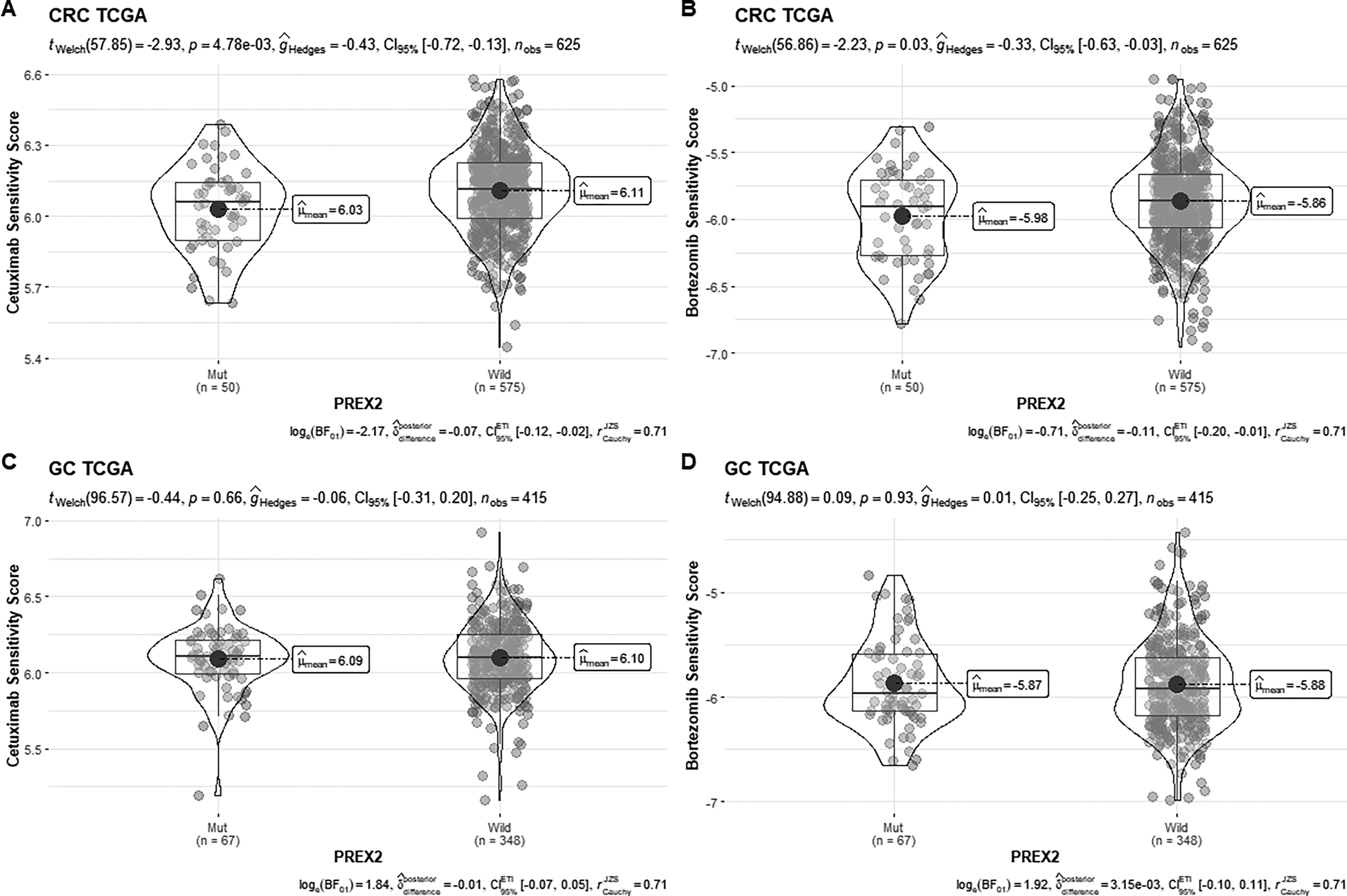
The effect of PREX2 mutation on sensitivity to Cetuximab and Bortezomib in CRC and GC.
Discussion
Many clinical studies have highlighted the significance of TMB and MSI as biomarkers for immunotherapy in gastrointestinal tumors. For instance, a phase 2 basket trial (NCT02091141) demonstrated a 60% objective response rate (ORR) in TMB-high MSS metastatic colorectal cancer (mCRC) patients treated with atezolizumab, compared to 0–2% in the general MSS mCRC population. Similarly, a phase 1b/2 trial (NCT02915432) found that high TMB significantly predicted better outcomes for advanced gastric cancer (GC) patients treated with toripalimab, with an ORR of 33.3% versus 7.1% for low-TMB patients (P = 0.017). 19 Additionally, MSI-H/dMMR has been established as a pan-cancer indicator for pembrolizumab in advanced solid tumors (involving advanced gastrointestinal cancers) refractory to standard therapies, based on results from five KEYNOTE trials (NCT01848834, NCT02054806, NCT01876511, NCT02628067, and NCT02460198). In this study, we first found that PREX2 mutations are associated with TMB and MSI in CRC, but not in GC. We also found that PREX2 mutations correlate with PD - L1 expression levels in CRC from our cohort, while no such correlations were observed in GC. Moreover, immune checkpoint analysis shows that CD274 (encoding PD - L1), PDCD1 (encoding PD - 1), and other immune checkpoints are highly expressed in PREX2 mutant CRC rather than GC. Cancer immunotherapy, which is largely based on programmed cell death - 1 (PD - 1)/PD - ligand 1 (PD - L1) immune checkpoint blockade, has made remarkable progress in treating human cancers over the past decade,20,21 including some gastrointestinal malignancies. 22 This suggests that, compared to GC, patients with PREX2 mutations in CRC may benefit more from immunotherapy. The result of the predicted drug sensitivity analysis also confirms this. This difference may be related to the differences in the tumor microenvironment between colorectal cancer and gastric cancer.
The biological role of PREX2 varies considerably across different tumor types. Although PREX2 functions as an upstream activator of the RAC1-GTPase pathway, its impact on melanoma appears to be limited. However, loss of PREX2 has been shown to sensitize melanoma tumors to MAPK pathway-targeted therapies, suggesting a potential tumor-suppressive role in this specific context. 8 In hepatocellular carcinoma, the S1113R mutation of PREX2 not only activates the AKT signaling pathway but also suppresses HectH9-mediated ubiquitination, thereby enhancing the proliferative and migratory capacities of tumor cells. 5 Current research indicates that PREX2 might inhibit the radiotherapy - induced immunogenicity of colorectal cancer via the cGAS/sting/IFNs pathway. However, the impact of PREX2 mutations on the tumor immune microenvironment has not been thoroughly studied. Studies have shown that in colorectal cancer treated with immunotherapeutic agents such as bevacizumab, tumor - infiltrating CD8 + cytotoxic T cells are associated with better prognoses. 23 There are other examples that emphasize the antitumor immune response, mainly the density of CD8 + cytotoxic tumor - infiltrating lymphocytes, which is correlated with improved outcomes in colorectal cancer.24–26 Our research shows that PREX2 mutations significantly increase the infiltration levels of CD8+ T cells and other immune cells in CRC. This provides a mechanistic basis for the potential benefits of immunotherapy in CRC rather than in GC patients with PREX2 mutations. It also theoretically explains the fact that GC patients with PREX2 mutations have poorer immunotherapy efficacy compared to those without PREX2 mutations. 9
Despite the promising insights, our study has several limitations. First, the single-center design and reliance on public databases introduce potential sample selection bias, limiting the generalizability of our findings to broader populations with inherent heterogeneity. Moreover, the absence of external validation through multi-center cohorts and prospective studies weakens the robustness and clinical applicability of our results. Additionally, the lack of longitudinal clinical outcome data restricts our ability to assess the predictive and prognostic value of PREX2 mutations over time. The findings, derived from a cohort without pathological staging data, may not be applicable to patients with specific disease stages. Furthermore, the small sample size and the high molecular heterogeneity of gastric cancer further limit our understanding of the role of PREX2 mutations in this disease. Future studies should address these issues by expanding the sample size, incorporating multi-center cohorts, and performing stratified analyses based on molecular subtypes. Such approaches would provide a deeper understanding of the biological significance and therapeutic potential of PREX2 mutations. Additionally, the lack of gene expression data for our samples, along with the use of publicly available data for immune infiltration analysis and computational drug response predictions, may introduce biases. Lastly, while our findings suggest that PREX2 mutations could serve as novel targets for immunotherapy in colorectal cancer, direct experimental validation and a systematic comparison with established biomarkers, such as TMB and MSI, are still needed. Future research should aim to address these gaps and further investigate the impact of PREX2 mutations on immunotherapy.
Conclusion
In conclusion, our findings reveal that PREX2 mutations in CRC are closely linked to tumor TMB, MSI status, and increased PD-L1 expression, suggesting their role in shaping the tumor immune microenvironment. Mechanistically, PREX2-mutant CRCs show enhanced immune-related pathway activation and distinct immune cell infiltration patterns, highlighting a functional immunogenic phenotype. While PREX2 mutations also occur in GC, their immune-related effects differ between the two cancers, emphasizing the context-dependent nature of this mutation. These results position PREX2 mutations as promising biomarkers for immunotherapy in CRC, enabling better patient stratification for immune checkpoint inhibitors and other immunotherapeutic approaches. To validate these findings and refine precision oncology, further multi-center clinical trials are needed to confirm the predictive value of PREX2 mutations and explore their role in therapeutic response.
Supplemental Material
sj-doc-1-cbm-10.1177_18758592251380454 - Supplemental material for Unraveling the role of PREX2 mutations as a biomarker for immunotherapy response in colorectal cancer
Supplemental material, sj-doc-1-cbm-10.1177_18758592251380454 for Unraveling the role of PREX2 mutations as a biomarker for immunotherapy response in colorectal cancer by Huan Peng, Pengmin Yang, Xintao Wang, Xiaokai Zhao, Jieyi Li, Ziying Gong, Daoyun Zhang and Zhiguo Wang in Cancer Biomarkers
Supplemental Material
sj-xlsx-2-cbm-10.1177_18758592251380454 - Supplemental material for Unraveling the role of PREX2 mutations as a biomarker for immunotherapy response in colorectal cancer
Supplemental material, sj-xlsx-2-cbm-10.1177_18758592251380454 for Unraveling the role of PREX2 mutations as a biomarker for immunotherapy response in colorectal cancer by Huan Peng, Pengmin Yang, Xintao Wang, Xiaokai Zhao, Jieyi Li, Ziying Gong, Daoyun Zhang and Zhiguo Wang in Cancer Biomarkers
Footnotes
Acknowledgements
We would like to express our gratitude to Huaqing Liu from the Molecular Experiment Department and Qing Su from the Pathology Experiment Department of Jiaxing Yunying Medical Inspection Co., Ltd for their assistance in sample handling and experiments. We also acknowledge Xiaozhi Wang and Jianle Wang from the Bioinformatics Analysis Department for their contributions to the processing and analysis of the experimental data.
Ethical considerations
Ethical approval to report this study was obtained from the institutional review board of Shanghai Second Affiliated Hospital of Naval Medical University (20211218).
Written informed consent was obtained from the patients for their anonymized information to be published in this article.
Author contributions
All authors contributed to the manuscript revision, and read, and approved the submitted version.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (grant number. 81000845); the Key Research and Development Program of Zhejiang province [grant number 2023C03057]; the Jiaxing Science and Technology Program-Social Development Special Project [grant number 2025BS009]; and the Key Technology Innovation Projects of Jiaxing [grant number 2024BZ20002].
Declaration of conflicting interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Pengmin Yang, Xintao Wang, Jieyi Li, Xiaokai Zhao, Daoyun Zhang, and Ziying Gong were employed by the Jiaxing Yunying Medical Inspection Co., Ltd, and the Shanghai Yunying Biopharmaceutical Technology Co., Ltd The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Data availability statement
The data supporting this study's findings are available on request from the corresponding author. The data are not publicly available due to privacy or ethical restrictions.
Supplemental material
Supplemental material for this article is available online.
