Abstract
Pyroptosis is a programmed cell death, which garners increasing attention by relating to immune and therapy response. However, the role of pyroptosis in colorectal cancer (CRC) remains unclear. Our study mainly to explore the role of pyroptosis in CRC. The mRNA expression data and corresponding clinical information of CRC patients were achieved from The Cancer Genome Atlas (TCGA). Pyroptosis-related genes (PRGs) were identified using DESeq2 R package and biological function was analyzed using cluster Profiler R package. A PRGs-based prognosis model was constructed by a univariate Cox and LASSO regression analyses. Then, the affecting of risk signature to clinicopathological characteristics, immune status and infiltrated immune cells, immune checkpoint and chemotherapy sensitivity was analyzed. qRT-PCR and IHC were performed for the expression level of PRGs. Moreover, a nomogram predict model was constructed. Total 57 PRGs were identified between 500 CRC samples and 44 normal samples. Those PRGs mainly enriched in immune-related and pyroptosis-related pathways. GABRD, NADK, TMEM240, RER1, AGRN, UBE2J2, CALML6, PLCH2, TMEM88B have been identified as gene signature and a prognostic model was constructed and validated. CRC patients with high-risk score showed poor survival, high TMB score, high proportion of CD4 + memory T cells, common lymphoid progenitors, cancer associated fibroblasts, mast cells, and neutrophils. The immune checkpoint related genes, CD160, CD200R1, CD244, CD28, CD40LG, CD44, CD48, CD80, CD86, HHLA2, ICOS, IDO1, TIGIT, TNFRSF25, TNFRSF4, TNFRSF9, TNFSF15, TNFSF18 also increased in high-risk score group. CRC patients with high-risk score more sensitive to docetaxel and rapamycin but resistance to gemcitabine and mitomycin. Moreover, a predictive nomogram for 1-, 3-, 5-year for CRC patients was established and validated. In the study, a PRGs-based prognostic model and a predictive model were constructed. These models are effective and robust in prediction the 1-, 3-, and 5-year survival of CRC patients.
Introduction
Colorectal cancer (CRC) ranks third in terms of incidence, and is the second leading cause of cancer related deaths in the World. The latest study found the incidence of CRC in humans under the age of 50 was increasing. 1 In the US, an estimated 149 500 new cases of CRC occurred and 52 980 deaths in 2021. 2 Therefore, it remains a major health burden and brings huge economic burden to the society. Although colonoscopy has provided a better screening method for the diagnosis of early colorectal cancer, many people can't accept this kind of examination for its price, people's psychological pressure, related risks and so on. 3 The result is 80% of the patients are found in the middle and late stage, and nearly half of the patients have a survival time of no more than 5 years. At present, CRC can be treated by surgery, chemotherapy, radiotherapy and biotherapy regimens, et al, however, the resistance to chemotherapeutic drugs and postoperative recurrence make it more difficult to treat CRC. 4 Thus, it is urgent to explore new treatments improve the clinical prognosis of patients with CRC, which make the establishment of a novel prognostic model become particularly important.
Pyroptosis, which is also called inflammatory necrosis of cells, is a newly discovered mode of programmed cell death (PCD). It is characterized by dependence on inflammatory caspases (mainly caspase-1, 4, 5, 11) and accompanied by the release of a large number of proinflammatory factors, such as interleukin (IL) −18 and IL-1β. 5 It was found that microbial infections can initiate pyroptosis via binding to pattern recognition receptors (PRRs). 6 The complex of ligand and receptor activates the body's immune system and inflammasomes, so as to release a large number of inflammatory factors and eliminate these pathogens. Some studies also found that mucosa-associated microbes can drive different kinds of immune cells, such as IL-17 and IL-22-producing cells, T helper 17 (Th17) cells, dendritic cells(DCs), mast cells and so on to strengthen gut barrier function.7,8 Our previous studies have found that microorganisms and their products play a key role in immune and inflammatory responses, 9 and these responses also play a crucial role in the occurrence and development of tumors. 10 Since Pyroptosis is related to the immune response of microorganisms and the host, the specific mechanism in immunity and tumors needs to require further elucidation.
Some important genes have been proved to be closely related to pyroptosis. Researchers found that the gasdermin family which includes ; gasdermin -A (GSDMA), GSDMB, GSDMC, GSDMD, GSDME (also known as DFNA5) and PJVK (also known as DFNB59) plays a vital role in the occurrence of pyroptosis. 11 It was found that GSDMD is a common substrate protein of all inflammatory caspases (including caspase1/4/5/11). 12 A recent study found Caspase 3 specifically cleaves activated GSDME, which can punch holes in the membrane and trigger pyroptosis. CRC has a high recurrence and progression rate due to its biological invasion and resistance to radiotherapy and chemotherapy. Many studies have revealed that pyroptosis plays a crucial role in the pathogenesis and progression of CRC. Ma et alfound that GSDMD were revealed to be associated with lymph node and distant metastases in patients with CRC, and may prove valuable in the future diagnosis and prognosis of CRC. 13 Secoisolariciresinol diglucoside induces pyroptosis by activating caspase-1 to cleave GSDMD in colorectal cancer cells. 14 And recent study has found that membranous GSDMD expression is closely related to immune response, and that cytoplasmic GSDMD correlates with tumor immune microenvironment and improves patient prognosis; but that expression of GSDMD in the cancer cell nucleus promotes tumor invasion and metastasis. 15 Researchers have found that GSDME mediates lobaplatin-induced pyroptosis downstream of the ROS/JNK/Bax-mitochondrial apoptotic pathway and caspase-3/-9 activation in CRC cells, which indicated that GSDME-dependent pyroptosis is an unrecognized mechanism by which lobaplatin eradicates neoplastic cells. 16 Moreover, it has also shown that pyroptosis patterns play a vital role in immunoregulation and the formation of tumor microenvironment (TME) formation in CRC. 17 Whether there are other pyroptosis related genes that affect the occurrence and progression of colorectal cancer remains unclear, and whether these genes will affect the immune microenvironment and chemotherapy resistance of colorectal cancer still needs further research.
Thus, in order to further explore the mechanism and impact of pyroptosis in the occurrence and prognosis of CRC, we used bioinformatics to find prognostic markers based on the expression profile of pyroptosis related genes in TCGA-CRC cohort and to construct a pyroptosis related prognostic model for CRC patients. Our findings indicate the potential connection between pyroptosis, prognosis, the immune microenvironment, and the response to chemotherapy in CRC patients.
Material and Methods
Data Sources and Processing
The mRNA expression data and corresponding clinical data (Table S1) were downloaded from The Cancer Genome Atlas (TCGA) database (https://portal.gdc.cancer.gov/), which including 500 CRC tumor samples and 44 normal samples. All CRC tumor samples were randomly divided into a training set (70%, n = 352) and a test set (30%, n = 148), and the training cohort was used to construct a prognostic risk mode, and test cohort was used to validate the reliability and accuracy of the model. Further, a total 57 pyroptosis-related genes (PRGs) were obtained from previous articles, 18 and shown in Table S2.
Screening Differentially Expressed Genes
The PRGs in CRC were obtained by intersecting 57 PRGs and expressed genes in all samples (559 tumor samples and 44 normal samples). The differentially expressed PRGs in CRC were screened using DESeq2 by setting criteria with | log2 (fold change, FC) |> 1 and P < .05 (Table S3).
GO and KEGG Pathway Enrichment Analysis
Cluster filer R package was used to perform the Gene Ontology (GO), including biological process (BP), cellular component (CC), and molecular function (MF), and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment (Table S4-5).
Construction and Validation of A Pyroptosis-Based Gene Signature
Univariate Cox regression analysis was performed using coxph function of survival R package and used to identify the prognosis-related PRGs for patients with CRC in training cohort. A threshold P < .05 was set to identify the prognostic variables. A least absolute shrinkage and selection operator (LASSO) penalized Cox proportional hazards regression further to determine the PRG-based prognosis gene signature. The optimal values of the penalty parameter (λ) were determined by 10-fold cross-validation based on minimum criteria. Subsequently, the optimal genes were screed to construct the risk model, which was established by multivariate Cox regression analysis. The individual risk score of each sample was calculated and all samples were distribution into high-risk and low-risk score groups according to the median of risk score. subsequently, Kaplan-Meier curves and receiver operating characteristic (ROC) score were calculated to evaluate the prognostic value of risk model both in training cohort and test cohort.
Tissue Samples
A retrospective study was performed on 14 patients with colorectal cancer. CRC and nearby non-tumor tissues were collected from CRC patients and confirmed via pathological examination. The samples were embedded into paraffin blocks or preserved at −80°C till use. Written informed consents were offered by all individuals included in this study. The study was permitted by the Ethics Committee. Related tumor tissue information was shown in Table S6.
RNA Isolation and Quantitative Real-Time Polymerase Chain Reaction PCR (RT-qPCR)
Total RNA was extracted from CRC patient tissues using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). Complementary DNA (cDNA) was synthesized using the total RNA and a PrimeScript RT reagent kit (Takara). SYBR-Green assays (Takara) were used to perform the RT-qPCR on a CFX-96 instrument (Bio-Rad Laboratories, Inc., USA). The data were copulated through the 2−ΔΔCt strategy, normalizing with ACTB. The primer sequences used for qRT-PCR in this study are listed in Table S7.
Immunohistochemistry (IHC)
Immunohistochemistry for 6 PRGs was performed as follows: slides were deparaffinized and rehydrated in graded xylenes and alcohols. After blocked with skim milk, GABRD (cat.no.47823-1;1:100 dilution; Signalway antibody), NADK (cat.no. 15548-1-AP; 1:100 dilution; Proteintech), AGRN (cat.no. SAB4301032; 1:100 dilution; SIGMA), UBE2J2(cat.no.433663-1;1:100 dilution; Signalway antibody), CALML6 (cat.no.22274-1-AP; 1:100 dilution; Proteintech), PLCH2(cat.no.DF9032; 1:100 dilution; Affinity biosciences), primary antibodies were incubated with slides for overnight at 4 °C. After that the application of secondary antibody was for slides at room temperature for 30 min. Slides were then developed using horseradish peroxidase DAB Quanto kit (Thermal Scientific). Photomicrographs were taken with a Leica microscope (Leica Microsystems). The pathologic diagnoses of these cases were microscopically confirmed by two researchers. We assigned two scores as described before 19 : a 5-tier scale for proportion of cells with positive staining as 0, 1%–25%, 26%–50%, 51%–75%, and >75% (graded 0-4) and a 4-tier scale for intensity of staining (graded 0-3). The scores were multiplied providing a range of 0-12 which was subsequently grouped into negative (score = 1-3), moderate (score = 4-6), and strong (score = 8-12; scores 5, 7, 10, and 11 do not exist in this scheme).
Construction of a Nomogram Predicated Model
A predicated nomogram was developed by including clinical characteristics (age, gender, risk score, tumor stage, and T/N/M stages), which was used to predict the 1-, 3-, 5-year overall survival (OS).
Analysis of the Tumor Immune Status and Infiltrating Immune Cells
The immune score of each sample was calculated by ESTIMATE algorithm using estimate package (Table S8). And the fraction of immune cells between high-risk and low-risk score groups was assed by xCell algorithm using immunedeconv package. The difference between high-risk and low-risk score groups was detected using a Wilcoxon rank-sum test. The expression of immune checkpoint related genes also detected using student's t-test.
Exploration of Chemotherapy Sensitive
We explored the chemotherapy sensitive based on the gene expression levels using pRRophetic package. 20 The chemotherapeutic response was determined by the half maximal inhibitory concentration (IC50) of each sample based on the Genomics of Drug Sensitivity in Cancer (GDSC) database (Table S9).
Statistical Analysis
Statistical Analysis Mann–Whitney U Test or Student's t-test was used to compare continuous variables in clinical features, the immune cell infiltration and immune pathway activation between the two groups when appropriate. The ANOVA method was applied to compare the continuous variables among the three groups. The receiver operating characteristic (ROC) curve was used to assess the specificity and sensitivity of the risk score and to determine the area under the curve (AUC). Kaplan–Meier survival analysis was performed using the log-rank test. All analyses in this study were performed with R software version 3.5.2. And the P value < .05 was considered as the significance if without specifically mentioned the cut-off criterion.
Results
Identification of the Differentially Expressed Pyroptosis-Related Genes (PRGs) in CRC
The process and workflow of this study were shown in Figure 1. Firstly, we obtained the mRNA expression data of 500 CRC tumor samples and 44 normal samples from TCGA database, and 57 PRGs from previous articles. After intersecting the expressed genes from TCGA and the 57 PRGs, then 44 PRGs were identified in CRC with criteria of P value < .05, including 23 upregulated and 21 downregulated PRGs (Figure 2, Table S4).
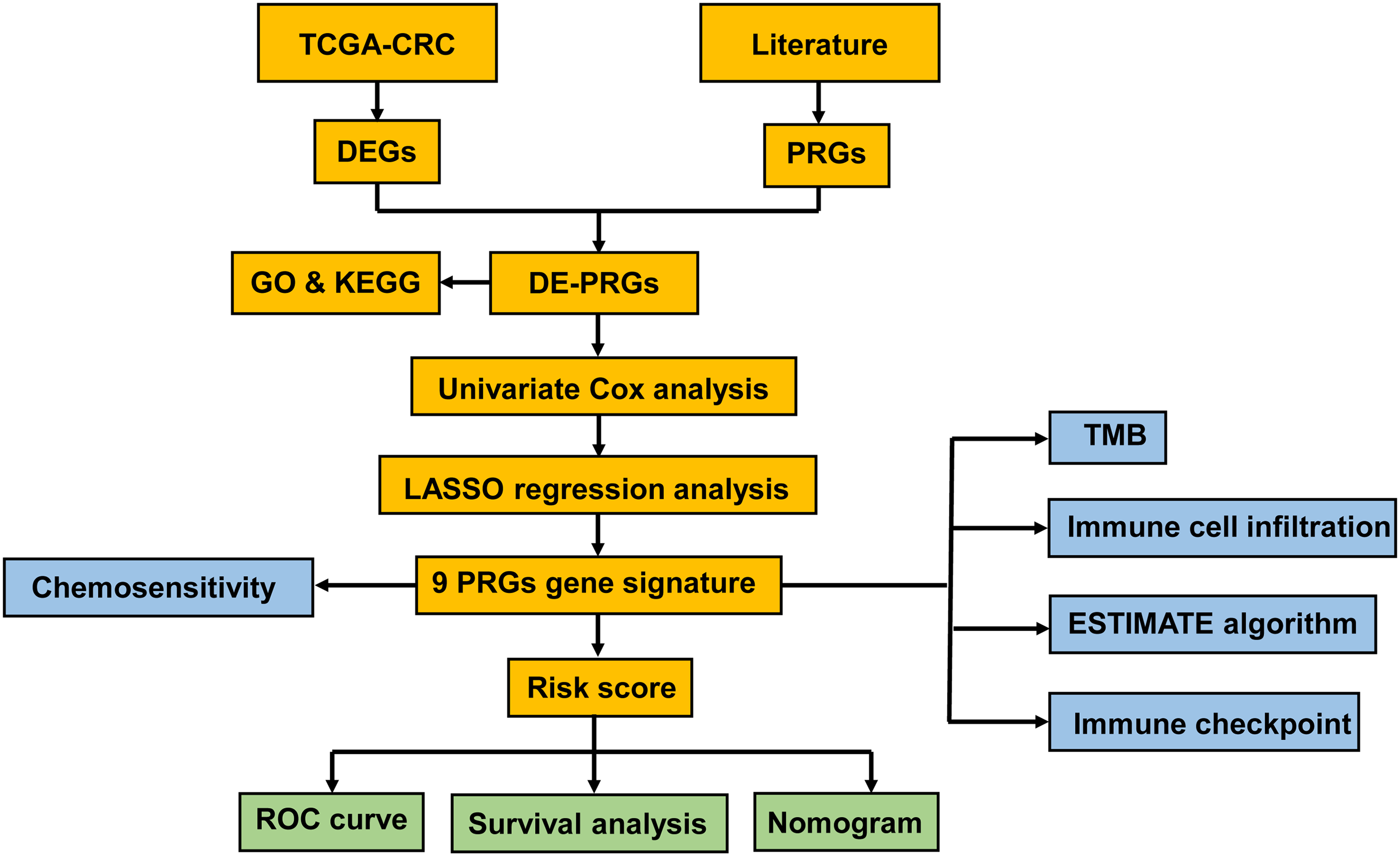
Flowchart of the analytic processes of this study.

Heatmap of the differentially expressed pyroptosis-related genes (PRGs) in CRC.
Functional Enrichment Analysis of PRGs in CRC
To clarify the biofunction of PRGs in CRC, GO and KEGG pathway enrichment were performed. And we found that most of PRGs involved in positive regulation of cytokine production, regulation of cysteine-type endopeptidase activity involved in apoptotic process, pyroptosis, positive regulation of interleukin-1 beta (IL-1β) production, positive regulation of IL-1 production, inflammasome complex, ESCRT III complex, ESCRT complex, multivesicular body, cysteine-type endopeptidase activity involved in apoptotic process and pathways, and cysteine-type endopeptidase regulator activity involved in apoptotic process, peptidase activator activity, CARD domain binding in GO analysis (Figure 3A, Table S5). And the KEGG enrichment indicated that most of PRGs involved in the salmonella infection, necroptosis, pertussis, NOD-like receptor signaling pathway, pathogenic Escherichia coli infection, shigellosis (Figure 3B, Table S6). It can be seen that the PRGs are mainly enriched into the biological functions related to pyroptosis and the related pathways of pyroptosis.

Functional enrichment analysis of PRGs in CRC. (A) The bubble plot of the Gene Ontology (GO) enrichment, including biological process (BP), cellular component (CC), molecular function (MF). The size of circle indicated the number of genes enriched. (B) The bar plot of the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment.
Construction of a PRGs-Based Prognostic Model for CRC Patients
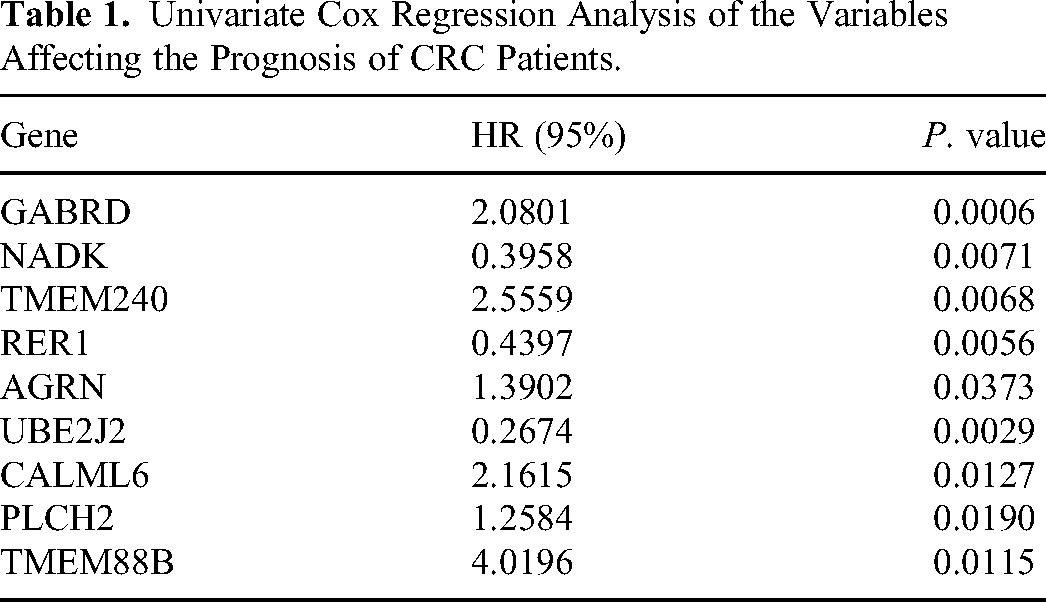
To construct a PRG-based prognosis model, Univariate Cox analysis firstly performed to identify the prognosis-related PRGs, as shown in Table 1, 9 PRGs, including GABRD, NADK, TMEM240, RER1, AGRN, UBE2J2, CALML6, PLCH2, TMEM88B, were screened. Except of NADK, RER1, UBE2J2, other PRGs with HRs > 1. Then, we conducted LASSO analysis based the expression of 9 prognostic PRGs (Figure 4A-B). Subsequently, these PRGs, GABRD, NADK, TMEM240, RER1, AGRN, UBE2J2, CALML6, PLCH2, TMEM88B, were identified as the prognostic gene signature and used to construct the risk model. The collinearity test and univariate Cox analysis indicated that CALML6 and TMEM88B significantly affects the prognosis of CRC patients (Figure 4C-D). The Kaplan-Meier curve indicated that CRC patients with high-risk score showed a worse survival time than the patients with low-risk score both in training set and test set ((Figure 4E-F). ROC curves also revealed that the AUC value of 1-, 3-, 5-years in training set were 0.75, 0.69, and 0.75, and the AUC value of 1-, 3-, 5-years in test set were 0.72, 0.7, and 0.54 (Figure 4G-H). These results indicated that the risk model had a good prognostic effect in CRC.

Construction of a PRGs-based prognostic model for CRC patients. (A) LASSO coefficient of the nine PRGs. (B) Plots of the ten-fold cross-validation error rates. (C) The diagram indicated the collinearity analysis of the nine PRGs. (D) Forest plot indicated the results of the multivariate Cox analysis of the nine PRGs. (E)-(F) Kaplan-Meier curves showed OS of CRC patients in high-risk and low-risk groups both in training and test sets. (G)-(H) Time-dependent ROC curves indicated the 1-, 3-, 5-years OS of CRC patients in high-risk and low-risk groups both in training and test sets.
Univariate Cox Regression Analysis of the Variables Affecting the Prognosis of CRC Patients.
The PRGs’ mRNA Expression in Human Colorectal Tissues
We compared the mRNA expression levels between CRC and adjacent normal tissues and found that the expression levels of most PRGs were consistent with TCGA. TMEM240(P = .0045), TMEM88B(P = .0010), NADK(P = .0058) and GABRD (P = .0246) were expressed at lower levels in CRC samples compared to those in adjacent normal colorectal samples (Figure 5A-B, E and H), while CALML6 and RER1were significantly elevated in CRC samples (Figure 5D and F), suggesting that these PRGs might play an important role in the oncogenesis of CRC. However, some PRGs, such as UBE2J2, PLCH2, AGRN showed no difference between adjacent normal and CRC samples and RER1 showed upregulated mRNA expression in tumor samples, which is inconsistent with the TCGA database. It is possible that the sample size is not large enough to be significant or skewed in some way. The results of our analysis showed a significant difference in expression levels of PRGs between CRC and control samples, indicating the latent function of PRGs in CRC oncogenesis.

The 9 PRGs’ expression in human colorectal tissues. (A-I) The expressions of 9 PRGs between Pare-Carcinoma and Carcinoma. *P < .05,**P < .01.
The PRGs’ Protein Expression in Human Colorectal Tissues
The 6 PRGs’ protein expression was analyzed in 14 pairs of colorectal cancer and their corresponding adjacent normal tissues. The UBE2J2 staining was scored as 8 (H-score groups, strong), NADK staining was scored as 6 (H-score groups, moderate) in adjacent normal tissues and the GABRD staining was scored as 6 (H-score groups, moderate) in adjacent normal tissues which were statistically highly different in CRC tissues. The protein expression of CALML6 (scored as 6, moderate) in CRC tissues were significantly increased compared with adjacent normal tissues. However, the protein expression of PLCH2 and AGRN showed no significance between CRC and adjacent normal tissues (Figure 6). It is possible that the protein is expressed too low or the antibody is not sensitive. Due to the fact that the TMEM240, TMEM88B and RER1 primary antibodies are not in stock the purchase cycle is too long, the other three genes have not been tested by IHC. But we found these three gene expression images from the Human Protein Atlas (HPA) database and put them in the Supplemental Figure 1. The protein expression of RER1 in adjacent normal tissues were significantly increased compared with CRC tissues, which is consistent with the TCGA database. However, TMEM240 and TMEM88B showed no difference between adjacent normal and CRC tissues.

The PRGs protein expression as analyzed by IHC.
Establishment of a Nomogram Predicated Model for CRC Patients
Considering the clinical characteristics, multivariate Cox analysis was performed to determine age, gender, risk score, tumor stage, T/N/M stages were independent variables affecting the prognosis of CRC patients (Figure 7A). Moreover, tumor stage and risk scores were significantly correlated with prognosis (P < .001). Subsequently, a nomogram model was constructed and revealed that the 1-, 3-, 5-years overall survival rates was high prognostic accuracy (Figure 7B).

Establishment of a nomogram predicated model for CRC patients. (A) Forest plot revealed the results of the results of multivariate Cox analysis of the clinical characteristics (age, gender, and tumor stage) and risk score. (B) A nomogram to predict the 1-, 3-, 5-year OS rate of CRC patients.
High Risk Score Associated to Different Immune Status, Immune Cell Infiltration and Immune Checkpoint-Related Genes
To investigate the role of PRGs in the Tumor microenvironment(TME) of CRC, We also evaluated the TME score by using the ESTIMATE package. Then we assessed the correlations between the PRGs-based risk score groups and human immune cell subsets. And we found high-risk score patients revealed high immune score than the low-risk score patients (Figure 8A, Table S8). And the fraction of CD4 + memory T cells, common lymphoid progenitors, cancer associated fibroblasts, mast cells, and neutrophils were obvious differences between high-risk and low-risk score groups (Figure 8B). Then, we also detected the expression of immune checkpoint related genes, and we found that CD160, CD200R1, CD244, CD28, CD40LG, CD44, CD48, CD80, CD86, HHLA2, ICOS, IDO1, TIGIT, TNFRSF25, TNFRSF4, TNFRSF9, TNFSF15, TNFSF18 increased in high-risk score group than low-risk score group (Figure 7C-D). These results indicated that significant correlation between PRGs and tumor immune infiltrating.

High risk score associated to different immune status, immune cell infiltration and immune checkpoint-related genes. (A) The violin plot of the immune score between high-risk and low-risk groups. (B) The violin plot of the different immune cells between high-risk and low-risk groups. (C)-(D) The violin plot of the expression of the immune checkpoint related genes.
High Risk Score Associated the Chemosensitivity
We next investigated the correlation between risk score and drugs sensitivity using pRRophetic R package based on the Genomic of Drugs Sensitivity in Cancer (GDSC) database. Interestingly, we found that the patients in the high risk score group had lower IC50 value for docetaxel and rapamycin (9A, 9D, Table S9), while the patients in the low risk score group had lower IC50 value for gemcitabine and mitomycin (Figure 9B-C, Table S9). Together, these results showed that PRGs were related to drug sensitivity. Interestingly, these data suggested that the CRC patients with high-risk score more sensitive to docetaxel and rapamycin than patients with low-risk score. On the contrary, the CRC patients with high-risk score more resistant to gemcitabine and mitomycin. Together, these results showed that PRGs were related to drug sensitivity.

High risk score associated the chemotherapy sensitivity. (A)-(D) Boxplots of the IC50 values of the docetaxel, gemcitabine, mitomycin, and rapamycin between the high-risk and low-risk score groups.
Discussion
Emerging evidence of pyroptosis is related to the occurrence, development and immune microenvironment of tumors.21,22 However, its role in CRC is less studied, especially with the specific pyroptosis-related genes (PRGs) in CRC and their impact on the prognosis, immune pathways, immune checkpoint, and chemoresistance of CRC. Therefore, construction of a PRGs-based prognostic model and a predictive model can not only predict the survival rate of patients, but also provide deeper insights into PRGs-mediated immune response and chemoresistance, which will provide novel ideas and methods for pathogenesis and treatment of CRC.
Pyroptosis is a new type of programmed cell death, which plays a dual role in the occurrence and treatment of tumors. Firstly, normal cells can transformed into tumor cells by releasing a large amount of inflammatory mediators of pyroptosis; secondly, tumor cells can be inhibited by pyroptosis, so as to achieve the purpose of treatment. 23 Moreover, PRGs play a major role in this biofunction.24,25 Since PRGs play an important role in the occurrence and treatment of tumors, the effect of the prognosis and biofunction of CRC is still unclear. In the current study, 44 PRGs were identified significantly in CRC, including 23 upregulated and 21 downregulated PRGs. And we found that these PRGs involved in positive regulation of cytokine production, regulation of cysteine-type endopeptidase activity involved in apoptotic process, pyroptosis, positive regulation of interleukin-1 beta (IL-1β) production, positive regulation of IL-1 production, and so on. And they also involved in the salmonella infection, necroptosis, pertussis, NOD-like receptor signaling pathway, pathogenic Escherichia coli infection, shigellosis. We then screened 9PRGs (GABRD, NADK, TMEM240, RER1, AGRN, UBE2J2, CALML6, PLCH2, TMEM88B) and found that it expressed differently in CRC and normal tissue in TCGA. At the same time, we compared the mRNA expression levels between CRC and normal tissues and found that the expression levels of most PRGs were consistent with TCGA. TMEM240, TMEM88B and NADK were expressed at lower levels in CRC samples compared to those in normal colorectal samples, while CALML6 and RER1 were significantly elevated in CRC samples, suggesting that these PRGs might play an important role in the oncogenesis of CRC. However, some PRGs, such as UBE2J2, PLCH2, AGRN showed no difference between normal and CRC samples and RER1 showed upregulated mRNA expression in tumor samples, which is inconsistent with the TCGA database.
It is possible that the sample size is not large enough to be significant or skewed in some way. In addition, we found that the protein expression levels of the PRGs by immunohistochemistry were consistent with the database. The results of our analysis showed a significant difference in expression levels of PRGs between CRC and control samples, indicating the latent function of PRGs in CRC oncogenesis. Moreover, we also found that theses PRGs could predict overall survival (OS) in CRC patients and it closely relates tumor immune infiltrating and chemotherapy sensitivity.
In the present study, Gamma-aminobutyric acid type A receptor subunit delta (GABRD) was one of the PRGs identified. Our results are consist with the results of Yan et al, 26 who demonstrated the higher the expression of GABRD, the worse the prognosis of CRC patients, and it could be used as a biomarker for the prognosis of patients with CRC. NADK plays a key role in maintaining the stability of intracellular environment, which is related to cell metabolism, redox state, oncogenic KRAS and a phosphatidylinositol 3-kinase (PI3 K)/Akt mediated signal pathway. Inhibition of NADK may refer to KRAS-driven colon cancer and PI3K-driven breast cancers. 27 Our work found NADK is related to Pyroptosis, which may provide a novel biomarker for the further study of the pathogenesis and treatment of CRC. It was found that hypermethylation of TMEM240 was associated with the prognosis of CRC, and circulating free methylated TMEM240 could be used as a biomarker for the prediction of early colorectal cancer. 28 We found that TMEM240 affects the prognosis of CRC through pyroptosis, which has become another meaningful point to study the relationship between TMEM240 and CRC. Retention in endoplasmic reticulum 1 (RER1) was first identified in yeast and plays a key role in regulating Notch signaling pathway. And it can be regulated by hypoxia-inducible factor (HIF)-1α to inhibit tumor cell's proliferation and diminished migratory and invasive abilities under hypoxic condition. 29 Our former research has proved that HIF-1α plays an essential role in the pathogenesis of CRC and referred to the chemoresistance, 30 which indirectly indicate the role of RER1 in CRC. Heparan sulfate proteoglycan AGRN is a basement membrane component, which mainly organizes postsynaptic differentiation at the neuromuscular junction. Recently, it has been reported that AGRN is up-regulated in hepatocellular carcinoma (HCC) and other types of cancer. AGRN is secreted from human hepatic stellate cells after being activated by platelet-derived growth factor (PDGF), and transmits mechanically sensitive signals to cells to regulate focal adhesion kinase and promote epithelial–mesenchymal transition (EMT), proliferation, migration and invasion. Moreover, AGRN can also promote the expression of ZEB-1 in pancreatic ductal adenocarcinoma (PDAC). 31 And our results first showed the relationship between AGRN and CRC by induce pyroptosis. Researchers showed that the E2 binding enzyme UBE2J2 can promote the polyubiquitination and degradation of evi. UBE2J2 decreased evi ubiquitination and increased evi homeostasis. Since evi protein levels are often maladjusted in cancer, targeting the regulatory UBE2J2 component may be a useful method of therapeutic intervention.32,33 Some researchers reported that EF hand protein calmodulin 6 (CALML6) directly binds to the serine rich phosphorylation (SR) region of IRF3 and impairs its dimerization and nuclear translocation. Overexpression of CALML6 inhibits IFN-β induced by virus infection while knocking down CALML6 has the opposite effect. Wang et al 34 found that CALML6 is a negative regulator of the NF-kB signaling pathway, which is important for maintaining the balance of the innate immune response. The occurrence and development of tumors are related to innate immunity. These studies have timely its possible role in tumor. Our current research found that CALML6 affects the occurrence of CRC cells by inducing pyroptosis, which provides a novel direction for the study of tumor immune microenvironment. It was found that one of the phospholipase C superfamily called PLCH2, was involved in the cleavage of membrane phospholipids, thereby resulting in the production of second messengers inositol 14,5-trisphosphate (PtdIns(4,5)P2) and 1,2-diacylglycerol (DAG). The messengers are important for G protein coupled receptor activation. 35 It may provide essential information for tumorigenesis. There are few studies about TMEM88B and tumors. We first proposed the role of TMEM88B in immune cell infiltration and chemosensitivity of colorectal cancer, which provides a new idea for further study of its mechanism in the future.
Infiltrating immune cells are the main cells in tumor tissues and play an important role in tumor biology. 36 In this study, we investigated the immune status in PRGs-based risk score groups. And we found high-risk score patients revealed high immune score than the low-risk score patients. And the fraction of CD4 + memory T cells, common lymphoid progenitors, mast cells, and neutrophils were obviously decreased in high-risk score group, while cancer associated fibroblasts were significantly increased in high-risk score group. These results were consisted with the former study. 37 The low level of key anti-tumor infiltrating immune cells indicates that the immune function of the high-risk group in the TCGA cohort is generally impaired. Then, we also detected the expression of immune checkpoint related genes, and we found that CD160, CD200R1, CD244, CD28, CD40LG, CD44, CD48, CD80, CD86, HHLA2, ICOS, IDO1, TIGIT, TNFRSF25, TNFRSF4, TNFRSF9, TNFSF15, TNFSF18 increased in high-risk score group than low-risk score group. Based on these findings, the poor survival outcome of high-risk CRCs may be caused by decreased levels of antitumor immunity. According to our results of GO and KEGG analyses, it is reasonable to speculate that pyroptosis can regulate the composition of the tumor immune microenvironment.
The research on the sensitivity of chemotherapeutic drugs has always been a hot spot for scientists. We have also previously studied the relationship between some signaling pathways and chemoresistance in colon cancer. 29 Wu et al found that LPS enhances the chemosensitivity of oxaliplatin in CRC via GSDMD-mediated pyroptosis. 38 And our current research found that the CRC patients with high-risk score more sensitive to docetaxel and rapamycin than patients with low-risk score. On the contrary, the CRC patients with high-risk score more resistant to gemcitabine and mitomycin. These studies provide new ideas and biomarkers for selection of reasonable chemotherapeutic drugs and the further study of chemotherapeutic drug resistance.
So far, pyroptosis has not been fully clarified, especially the specific mechanism in CRC. Our present research identified 9 genes that have the ability to regulate pyroptosis. However, However, our validation efforts are mainly based on data from public databases and some clinical trials, the research on the deeper mechanisms is very limited. How they play a further role by regulating the pyroptosis pathway in CRC deserves further in-depth studies.
Conclusion
In conclusion, our study shows that pyroptosis is closely related to CRC, the score generated based on the risk characteristics of 9 PRGs was an independent risk factor for predicting OS in TCGA. The DEGs between the low and high-risk groups were associated with tumor immunity and chemotherapeutic drugs sensitivity. Our study provides a novel gene signature for predicting the prognosis of CRC patients and offers a significant basis for future studies of the relationships between pyroptosis-related genes, immunity and chemoresistance in CRC.
Supplemental Material
sj-tif-1-tct-10.1177_15330338241277584 - Supplemental material for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer
Supplemental material, sj-tif-1-tct-10.1177_15330338241277584 for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer by Linjing Li, Yuyi Li, Junyi Lin and Wenjing Pang in Technology in Cancer Research & Treatment
Supplemental Material
sj-xlsx-2-tct-10.1177_15330338241277584 - Supplemental material for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer
Supplemental material, sj-xlsx-2-tct-10.1177_15330338241277584 for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer by Linjing Li, Yuyi Li, Junyi Lin and Wenjing Pang in Technology in Cancer Research & Treatment
Supplemental Material
sj-xlsx-3-tct-10.1177_15330338241277584 - Supplemental material for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer
Supplemental material, sj-xlsx-3-tct-10.1177_15330338241277584 for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer by Linjing Li, Yuyi Li, Junyi Lin and Wenjing Pang in Technology in Cancer Research & Treatment
Supplemental Material
sj-xlsx-4-tct-10.1177_15330338241277584 - Supplemental material for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer
Supplemental material, sj-xlsx-4-tct-10.1177_15330338241277584 for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer by Linjing Li, Yuyi Li, Junyi Lin and Wenjing Pang in Technology in Cancer Research & Treatment
Supplemental Material
sj-xlsx-5-tct-10.1177_15330338241277584 - Supplemental material for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer
Supplemental material, sj-xlsx-5-tct-10.1177_15330338241277584 for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer by Linjing Li, Yuyi Li, Junyi Lin and Wenjing Pang in Technology in Cancer Research & Treatment
Supplemental Material
sj-xlsx-6-tct-10.1177_15330338241277584 - Supplemental material for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer
Supplemental material, sj-xlsx-6-tct-10.1177_15330338241277584 for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer by Linjing Li, Yuyi Li, Junyi Lin and Wenjing Pang in Technology in Cancer Research & Treatment
Supplemental Material
sj-xlsx-7-tct-10.1177_15330338241277584 - Supplemental material for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer
Supplemental material, sj-xlsx-7-tct-10.1177_15330338241277584 for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer by Linjing Li, Yuyi Li, Junyi Lin and Wenjing Pang in Technology in Cancer Research & Treatment
Supplemental Material
sj-xlsx-8-tct-10.1177_15330338241277584 - Supplemental material for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer
Supplemental material, sj-xlsx-8-tct-10.1177_15330338241277584 for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer by Linjing Li, Yuyi Li, Junyi Lin and Wenjing Pang in Technology in Cancer Research & Treatment
Supplemental Material
sj-xlsx-9-tct-10.1177_15330338241277584 - Supplemental material for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer
Supplemental material, sj-xlsx-9-tct-10.1177_15330338241277584 for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer by Linjing Li, Yuyi Li, Junyi Lin and Wenjing Pang in Technology in Cancer Research & Treatment
Supplemental Material
sj-xlsx-10-tct-10.1177_15330338241277584 - Supplemental material for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer
Supplemental material, sj-xlsx-10-tct-10.1177_15330338241277584 for A Pyroptosis-Related Gene Signature Predicts Prognosis and Tumor Immune Microenvironment in Colorectal Cancer by Linjing Li, Yuyi Li, Junyi Lin and Wenjing Pang in Technology in Cancer Research & Treatment
Footnotes
Abbreviations
Acknowledgements
We would like to acknowledge the TCGA, GTEx and the GEO (GSE140082) network for providing data.
Author Contributions
Li L, Li Y, and Lin J performed experiments, analyzed data. Li L reviewed the manuscript. Pang W supervised experiments and prepared the manuscript.
Data Availability Statement
The original contributions presented in the study are included in the article/Supplemental Material, further inquiries can be directed to the corresponding author.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
This study obtained approval from the independent Ethics Committee of Shanghai Ninth People's Hospital, Shanghai Jiao Tong University School of Medicine on July 6th, 2022. The ethics number is SH9H-2021-T284-1 and all patients had signed informed consents.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by research grants of Cross disciplinary Research Fund of Shanghai Ninth People's Hospital, Shanghai JiaoTong University School of Medicine(YG2022QN046). Cross disciplinary Research Fund of Shanghai JiaoTong University(YG2022QN046). And Fundamental research program funding of Ninth People's Hospital affiliated to Shanghai JiaoTong University School of Medicine (JYZZ148).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
