Abstract
Background
15-Hydroxyprostaglandin dehydrogenase (15-PGDH, gene symbol HPGD) is considered a tumor suppressor, and its expression is often proportional to the anticancer response. However, the clinical significance of HPGD/15-PGDH in predicting immune response and its diagnosis and prognosis value in cervical cancer remains unclear.
Objective
This study aims to explore the clinical significance of HPGD/15-PGDH in predicting carcinogenesis, prognosis, and sensitivity to immuno- and chemotherapy in cervical cancer.
Methods
A comprehensive evaluation of the diagnostic, treatment-sensitive, and prognostic value of HPGD/15-PGDH in cervical cancer was conducted by bioinformatics analysis of public databases and validation of real cohort data.
Results
Bioinformatics analysis showed that HPGD expression was decreased in cervical cancer and did not independently predict patient prognosis. Low HPGD expression was linked to resistance to certain chemotherapies, potentially due to immunosuppression triggered by low HPGD levels. Validation in clinical samples from the local hospital confirmed the decreased 15-PGDH expression and increased COX-2 expression in HPV16-positive cervical cancer patients and increased immune suppression during cancer progression.
Conclusions
HPGD/15-PGDH is a potential biomarker for predicting the progression, immune response, and chemotherapy sensitivity of cervical cancer, with implications that it is of great value for the diagnosis and individual-based treatment of cervical cancer.
Introduction
Cervical cancer is the second most common malignancy in women worldwide. According to statistics of the Global Cancer Observatory (GLOBOCAN) 2020, there were an estimated 604 127 cervical cancer cases, accounting for about 6.5% of all new cancer cases in women, and approximately 84% occur in low- and middle-income countries.1,2 At present, the treatment of cervical cancer is mainly based on surgical resection, radiotherapy, and chemotherapy, while the efficacy is limited and easy to relapse. Novel immunotherapies and adoptive cell therapy (ACT) have been used in clinical practice, but still, the efficacy is unsatisfactory.3,4 Listed cervical cancer preventive vaccines cannot be treated for patients with cervical cancer and HPV infection.5,6 Therefore, it is urgent and necessary to explore new therapeutic targets and methods for cervical cancer.
High-risk human papillomavirus (HPV) infection is the main cause of cervical cancer.7,8 The HPV genome integrated into the host cell chromosome induces the destruction of the transcriptional regulation function of E2 and the persistent expression of HPVE6 and E7 oncoproteins, which interfere with the cell cycle and apoptosis regulation, leading to chromosome instability of host cells and immortalization and carcinogenesis. 9 Expression of HPVE6 and E7 directly alters host gene expression and affects host tolerance to infection and immune function. The main function of HPVE6 / E7 is to maintain the cell cycle of cutin cells and prevent the host anti-tumor and immune regulation of interferon-mediated effect.10,11 Thus, the immune status of the host is closely related to the occurrence and development of cervical cancer.12–14 Therefore, altering the tumor immune microenvironment is key to the success of antitumor immune therapy.
15-Hydroxyprostaglandin dehydrogenase (15-PGDH, encoded by HPGD) is a catabolic enzyme for the degradation of prostaglandins, including prostaglandin E2 (PGE2), PGD2, and PGF2α. As a tumor suppressor, 15-PGDH has been reported to inhibit proliferation and induce the differentiation of cancer cells, including colorectal, lung, and breast cancers. 15 Recent research shows that the depletion of 15-PGDH can effectively restore the tumor microenvironment. Down-regulation of HPGD promotes cell proliferation and migration in cervical cancer, 16 and inhibition of 15-PGDH causes tumor expansion in the pancreas. 17 Tumor microenvironmental 15-PGDH depletion promotes tumor formation and angiogenesis in pancreatic cancer. 18 While, overexpression of 15-PGDH inhibits cell growth and epithelial-mesenchymal transition in lung cancer, 19 and 15-PGDH expression may increase the anti-tumor immune response in gastric cancer. 20 However, the underlying mechanism of interconversion of suppressive or stimulating immune environments regulated by 15-PGDH remains unclear, and the predictive value of 15-PGDH in diagnosis, prognosis, and therapeutic intervention needs to be further elucidated. In the current study, we explore the clinical significance of 15-PGDH in the prediction of carcinogenesis, prognosis, and immuno- and chemotherapy sensitivity, as well as the underlying signal pathways associated with 15-PGDH involved in cervical squamous cell carcinoma and endocervical adenocarcinoma (CESC) with transcriptome profile, which further validated with our samples and cohorts.
Methods
TCGA data collection
TCGA database (https://portal.gdc.cancer.gov/) can provide gene expression data, miRNA expression data and copy number variation (CNV), DNA methylation, single nucleotide polymorphisms (SNPs), and other data. We downloaded raw and processed mRNA expression data and clinical information of the TCGA-CESC cohort. A total of 309 specimens were collected, including the normal group (n = 3) and the tumor group (n = 306), for the differential expression analysis of HPGD. Sample information is detailed in Supplemental Table 1.
Analysis of immune cells infiltration
We used the CIBERSORT algorithm (https://cibersort.stanford.edu/) to analyze the RNA-seq data of cervical cancer patients to infer the relative proportion of 22 immune infiltrating cells, and further analyze the correlation between gene expression and immune cell content. 21 We used the CIBERSORT function to calculate the immune cell content of the samples, with the perm (number of permutations) set to 100 and QN (quantile normalization) set to TRUE. A p-value of less than 0.05 was considered statistically significant.
GSVA analysis
Gene set variation analysis (GSVA) is a non-parametric and unsupervised method to assess the enrichment of transcriptome gene sets. 22 GSVA converts changes at the gene level into changes at the pathway level by comprehensively scoring the gene set of interest, to judge the biological function of the sample. According to the expression of core genes, patients were divided into high- and low- expression groups and the differences in signaling pathways between the two groups were further analyzed by GSVA. The gene sets were downloaded from version 7.0 of the Molecular Signatures Database (MsigDB: https://www.gsea-msigdb.org/gsea/msigdb/index.jsp), and each gene set was comprehensively scored using the GSVA algorithm to evaluate potential biological functional changes in different samples. 22
Analysis of HPGD gene co-expression network
The co-expression of the HPGD gene in cervical cancer data was analyzed, the filter condition of the correlation coefficient was set to 0.3 and the P-value was 0.05. After screening the genes with the most significant expression of HPGD, the “corrplot” and “circlize” of R packages were used to draw the circle map and heat map of HPGD correlation analysis.
Predictive analysis of response to immunotherapy based on TMB, MSI, and neo-antigen data
Tumor mutational burden (TMB) is defined as the total number of somatic gene coding errors, base substitutions, insertions, or deletions detected per million bases. In the current study, TMB was defined by calculating variant frequency and variant number/exon length for each tumor sample by dividing non-synonymous mutation sites by the total length of the protein-coding region. Microsatellite instability (MSI) refers to the phenomenon of changes in the length of microsatellite sequences caused by insertion or deletion mutations during DNA replication. The value of MSI for each TCGA patient was derived from a previously published study. 23 Neo-antigens were assessed in each patient using NetMHCpan v3.0. 24
Chemotherapy sensitivity analysis
Based on the largest cancer drug sensitivity genomics database, Genomics of Drug Sensitivity in Cancer (GDSC: https://www.cancerrxgene.org/), we use the R software package “pRRophetic” to predict the chemotherapy sensitivity of each tumor sample. 25 The estimated IC50 values of each specific chemotherapy drug were obtained by regression method, and the regression and prediction accuracy was verified by 10 cross-validations with the GDSC training set. Default values were selected for all parameters, including “combat” to remove batch effects and take an average of repeated gene expression.
Nomogram model construction
Nomogram is based on regression analysis, according to the expression of HPGD and clinical factors, using a scaled line segment, according to a certain proportion, drawing on the same plane, to express the relationship between each variable in the prediction model. 26 By constructing a multivariate regression model, according to the contribution of each influencing factor in the model to the outcome variable (the size of the regression coefficient), assign scores to each value level of each influencing factor, and then add up the scores to get the total score, to calculate the predicted value. 26
WGCNA Analysis
By constructing a weighted gene co-expression network, we search for the gene modules that are co-expressed and explore the association between the gene network and HPGD, as well as the key genes in the network. The WGCNA-R package was used to construct the co-expression network of all genes in the cervical cancer dataset, and the genes with the first 5000 variances were screened by the algorithm for further analysis, with the soft threshold set to 3. 27 The weighted adjacency matrix was transformed into a topological overlap matrix (TOM) to estimate the network connectivity, and the hierarchical clustering method was used to construct the clustering tree structure of the TOM matrix. Different branches of the clustering tree represent different gene modules, and different colors represent different modules. Based on the weighted correlation coefficient of genes, genes were classified according to the expression pattern, and genes with similar patterns were classified into a module. All genes were divided into multiple modules according to the gene expression pattern.
Functional enrichment analysis of gene modules
To obtain the biological functions and signaling pathways involved in WGCNA module genes, the Metascape database (www.metascape.org) was used for annotation and visualization, and module genes were analyzed by Gene Ontology (GO) analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis.
Patients
Twelve formalin-fixed paraffin-embedded patient tissue samples were obtained with ethics committee approval from the Shenzhen Guangming District People's Hospital. The tissues were used for immunohistochemistry (IHC) or multiple immunofluorescence (IF) labeling. Patient consent was approved by the institutional review board (IRB) of the Shenzhen Guangming District People's Hospital.
Immunohistochemistry
Both HPV16- normal and HPV16 + patients in the cervical cancer stage were recruited for IHC. The tissues were fixed in 4% formalin, then embedded in paraffin and serially sectioned into 4 μm. The tissue sections were deparaffinized and hydrated before antigen repair was performed by hyperbaric treatment with citrate buffer (pH 6.0, 10 Mm, Dako, Denmark) for 1–2 min. The incubation time of the primary antibody refers to the instruction manual of the antibody, which is adjusted according to different incubation temperatures (HPV16E7 antibody, 1:1000, Novus Biologicals; COX-2 antibody, 1:150, Abcam; 15-PGDH antibody, 1:500, Novus Biologicals, USA). The HRP-conjugated secondary antibody (Cell Signaling Technology, USA) was visualized using a 3,30-diaminobenzidine substrate (Vector Laboratories Inc., USA).
Multiplex immunofluorescence labeling
Our clinical cohort was graded from the cervical intraepithelial neoplasia (CIN) I-III to cervical cancer (CC). Multiplex fluorescence staining was performed with the Opal staining system and images were acquired using the Mantra platform (Perkin Elmer).28,29 Quantification of positively stained cells was performed using ImagePro Software and verified with manual counting from 6 to 10 random fields at 100×magnification. The proportion of the number of positive cells was calculated as the mean of all fields for each sample. The primary antibodies for CD4, CD8, FOXP3, CD68, and CD11b were purchased from Novus Biologicals (UK), and the working dilutions are 1:100 for all antibodies.
Statistical Analysis
All statistical analyses were conducted using R language (version 4.0). All statistical tests were bilateral, and p < 0.05 was statistically significant. Survival analysis was performed by the Kaplan-Meier method, and the high- and low-expression groups were divided according to the median value of gene expression.
Results
HPGD is a susceptible predictor but not a prognostic gene for cervical cancer
To verify whether HPGD expression is a prognostic factor for cervical cancer, we first downloaded and integrated the original mRNA expression data of CESC from the TCGA database, and compared the expression level of HPGD between normal and cancer tissue. There was a significant difference in the expression level of HPGD between normal and cancer tissues (Figure 1(A)). Then, we performed the survival analysis based on the grouping of high- and low-expression levels of the HPGD gene. The results showed no difference in overall survival between the two groups with the median expression value as the dividing point (Figure 1(B) and (C)).

HPGD is susceptible but not an independent prognostic risk factor for CESC.
Furthermore, we performed univariate and multivariate Cox regression analyses to investigate whether HPGD expression was a clinically independent prognostic factor for CESC patients. The risk scores of HPGD and clinicopathological characteristics, including age, histological grade, pathological T stage (referring to tumor size), and pathological N stage (referring to lymph node metastasis), were used as co-variates. The results revealed that none of these factors could be used as independent prognostic factors to predict the survival rate in CESC patients (Figure 2(D) and (E)). We further determine whether HPGD expression was correlated with each of the above clinicopathological characteristics. All patients were divided into the indicated subgroup. Then, stratification analysis was performed according to age, histological subtype, pathological T stage, and pathological N stage. The HPGD expression was not statistically significant with these clinicopathological characteristics, despite a slightly significant difference between the T3 and T2 subgroups (Figure 1(F) to (I)). These results suggest that HPGD may serve as a susceptible but not an independent prognostic factor in CESC.

The immune cell populations, GSVA, and co-expression network analysis of HPGD. A. The correlation between immune cell infiltration and HPGD. B. The GSVA analysis with high- and low-expression of HPGD in CESC patients. C. The heat map presents the top 10 co-expression genes of HPGD with a positive/negative correlation coefficient. D. The Circos plots show the correlation of the HPGD co-expression network.
The cellular and molecular mechanisms of HPGD underlying the progression of cervical cancer
The tumor microenvironment significantly affects the tumorigenesis, disease outcome, and clinical treatment sensitivity of various types of cancer. A recent study has shown that tumor-associated macrophages inhibit the expression of 15-PGDH in pancreatic cancer cells by secreting IL-1β, leading to a poor prognosis of pancreatic cancer. 17 To analyze the relationship between HPGD and tumor immune infiltration, we further explored the potential cellular and molecular mechanism of HPGD in the progression of cervical cancer. We first analyzed the TCGA dataset using the CIBERSORT algorithm to assess tumor-infiltrating immune cells correlated to HPGD expression. The results showed that HPGD expression was significantly positively correlated with “Mast and dendritic cells resting”, “B cells naïve”, and “T cell regulatory”, etc., and significantly negatively correlated with “B cells memory”, “Macrophages M0”, and “Mast cells activated”, etc. (Figure 2(A)). We further investigated the specific signaling pathways and molecular mechanisms associated with HPGD that underlie tumor progression of cervical cancer by GSVA analysis. The results showed that patients with high expression of HPGD were enriched in the pathway of KRAS signaling regulation, bile acid metabolism, estrogen response, apoptosis, etc. (Figure 2(B)). Patients with low expression of HPGD were enriched in MYC targets, DNA repair, and unfolded protein response (UPR) pathways. These results suggest that the expression of HPGD is associated with immune activation and regulation in cervical cancer.
In addition, we explored the co-expression network of HPGD according to the expression profile of the CESC cohort in the TCGA database, with a correlation coefficient filtering of 0.3 and a P-value less than 0.05. A total of 67 genes significantly related to HPGD expression (Table 1) were screened. The top 10 genes with positive/negative correlation coefficients are shown in the heat map (Figure 2(C), upper panel) and the co-expression correlation circle map (Figure 2(C), lower panel).
Co-expression gene network of HPGD in CESC.

The HPGD gene is an effective marker for the sensitivity of chemotherapy
To evaluate the role of the HPGD gene as an indicator of chemotherapy and immunotherapy sensitivity. We downloaded the processed SNP data of cervical cancer from the TCGA database and selected the top 30 genes with high mutation frequency as displayed. We compared the differences of mutated genes between high and low expression groups of HPGD with R package ComplexHeatmap (Figure 3(A)), which showed that the mutation in gene TTN, PIK3CA, and MUC16 was significantly increased in patients with high expression of HPGD. We further explored the relationship between the HPGD gene and common immunotherapy-related tumor markers. The results show that there is no difference in microsatellite instability (MSI), neoantigen, and tumor mutation burden (TMB) between the high- and low-expression groups of the HPGD in CESC patients (Figure 3(B) to (D)).
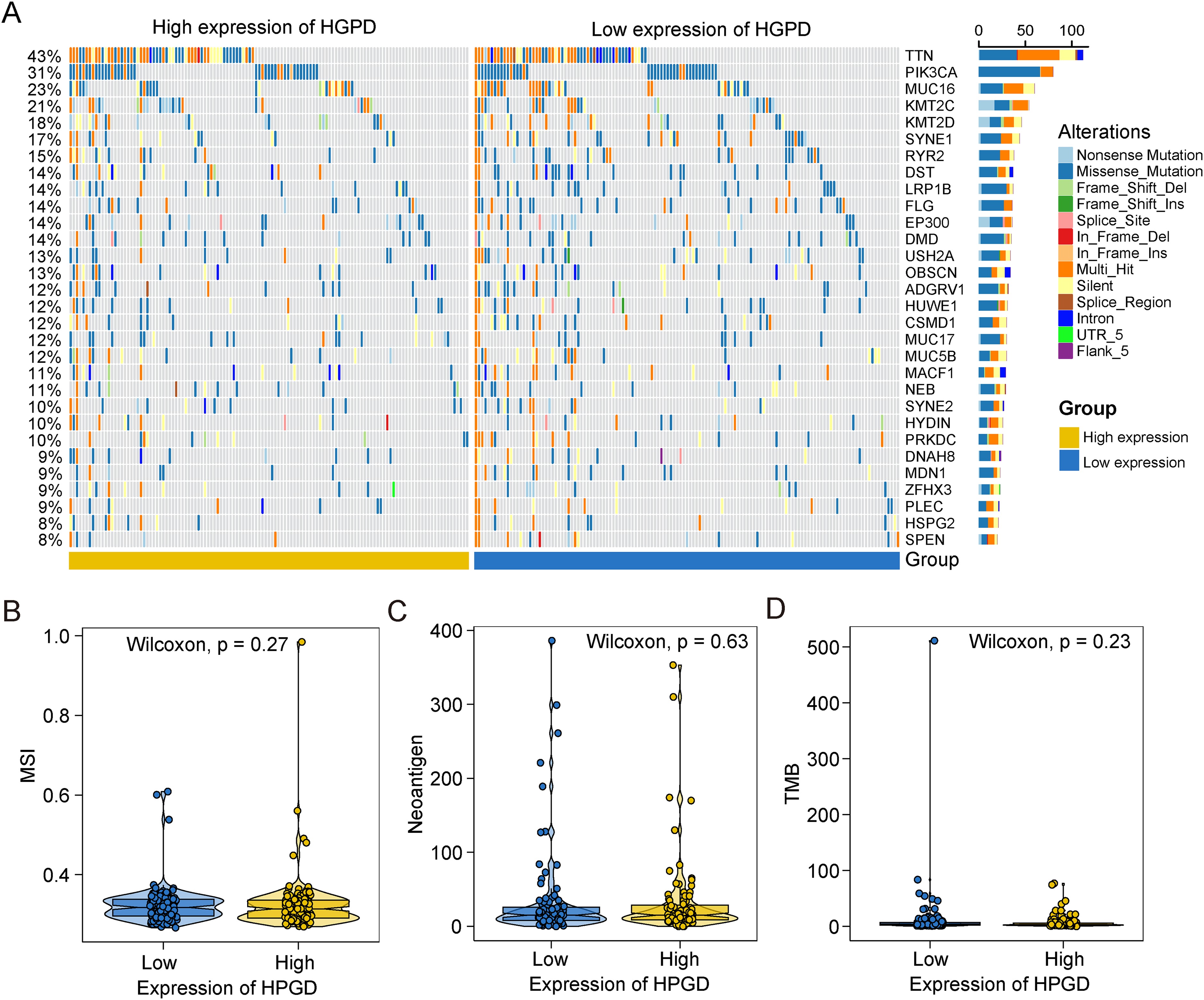
The value of HPGD in predicting sensitivity to immunotherapy treatment. A. The waterfall diagram of the top 20 driver genes between HPGD high- and low-expression subgroups of CESC patients. B–D. The violin diagram shows the correlations between HPGD expression and MSI (B), Neoantigen (C), and TMB (D) in CESC patients.
Early-stage cervical cancer is effective in the treatment of surgery combined with chemotherapy. 30 To further explore the sensitivity of HPGD and common chemotherapy drugs, we predicted the chemotherapy sensitivity of each tumor sample by the R package “pRRophetic” based on the drug sensitivity data of the GDSC database. Our results show that the IC50 scores of AZD6482, BIBW2992, and AKT.inhibitor.VIII, JNK.9L, and Lapatinib were significantly higher in the HPGD low-expression group, except for Axitinib. These results suggested that patients with low expression of HPGD may be resistant to AZD6482, BIBW2992, and AKT.inhibitor.VIII, JNK.9L, and Lapatinib treatment (Figure 4). These findings suggest that HPGD expression can be a predictor of the sensitivity of chemotherapy. The tolerance to chemotherapy in these patients may be influenced by immunosuppression triggered by low expression of HPGD.
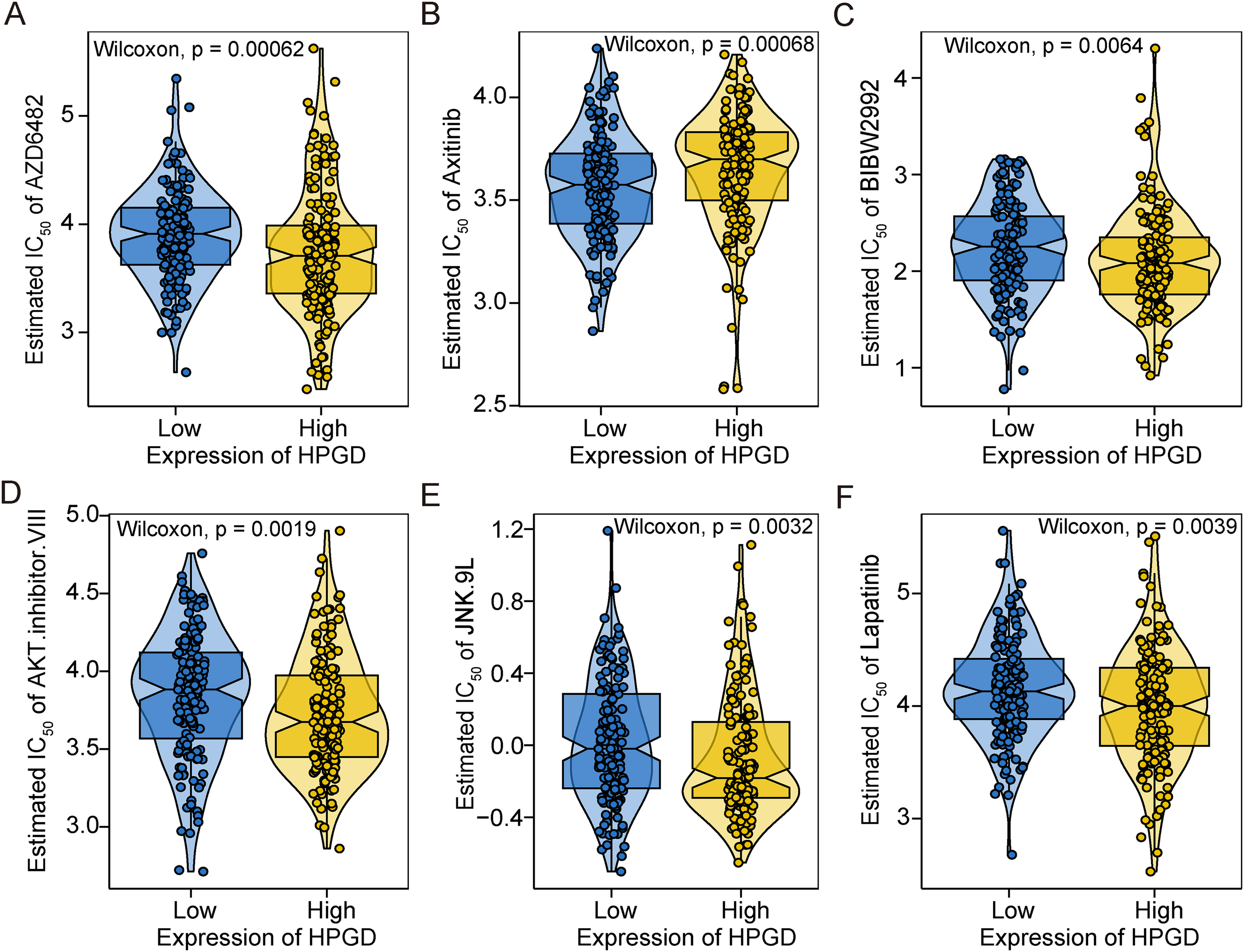
The value of HPGD in predicting sensitivity to chemotherapy treatment. A-F. The violin diagram shows the predicted value of half the maximal inhibitory concentration of common chemotherapy drugs in patients with different expression levels of HPGD. The scores were compared using the Wilcoxon test.
Establishment of Nomogram model and WGCNA regulatory network of HPGD
The Nomogram model allows the contribution of each factor to be taken together from multiple factors and converted into a score for each factor. We re-evaluate the impact of HPGD and other clinical characteristics on the patient's probability of survival by the Nomogram model. The results of regression analysis by HPGD expression quantity were presented in the form of a line chart, which showed that in all our samples, the values of different clinical indicators of cervical cancer and the distribution of HPGD expression had different degrees of contribution in the whole scoring process (Figure 5(A)). We further conducted prediction analysis on the OS of one year and three years (Figure 5(B)), and the results showed that the predicted OS was consistent with the observed OS, and the Nomogram model had good prediction efficiency.
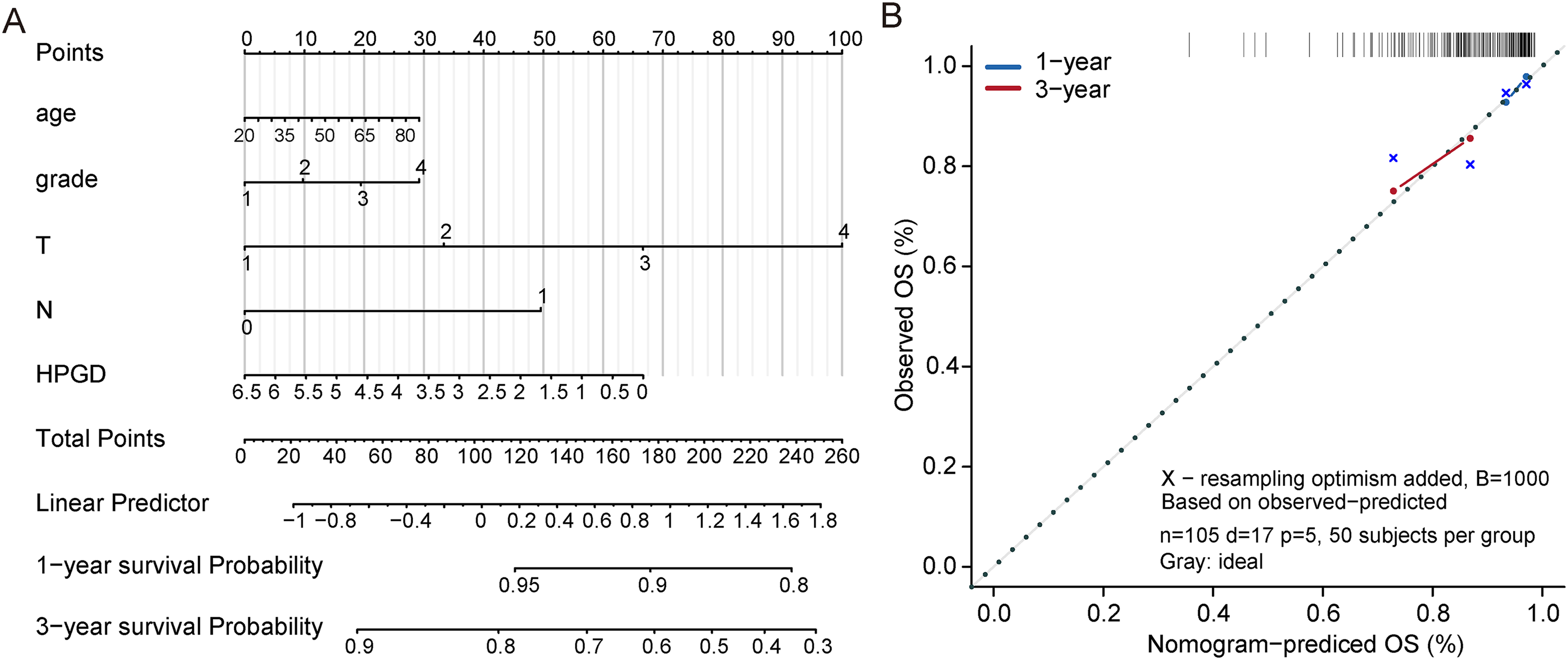
Nomogram to predict 1 and 3-year overall survival in the TCGA-CESC training set. A. Nomogram for predicting the 1- and 3-year overall survival time in patients. B. Calibration curve for the prediction of 1- and 3-year overall survival.
Furthermore, we constructed the WGCNA network to explore the regulatory network related to HPGD in CESC. The soft threshold β was set to 3 (Figure 6(A)), and the gene modules were detected based on the TOM matrix. A total of 12 gene modules were detected in cervical cancer (Figure 6(B)), including brown (423), blue (2468), cyan (72), green (298), salmon (76), tan (87), black (644), magenta (142), pink (238), midnight blue (68), green-yellow (116) and yellow (368). Through the analysis between modules and traits, it was found that the brown module had the highest correlation with HPGD (Figure 6(C)). Then the brown module gene was selected to conduct the subsequent pathway analysis, and the results showed that the brown module gene was mainly enriched in the pathways of the cornified envelope, cell-cell junction, desmosome, and intermediate filament cytoskeleton organization, etc. (Figure 6(D)). These results suggest that HPGD is likely to have tumor suppressor functions through modulation of the cornified envelope and intercellular communication to keep the epidermal barrier.

WGCNA revealed gene co-expression networks of HPGD expression in CESC patients. A. Analysis of the scale-free fit index and the mean connectivity for various soft-thresholding powers. B. Clustering dendrogram of differentially expressed genes related to HPGD expression in CESC. Dendrogram of all HPGD correlated genes clustered based on a dissimilarity measure. Dynamic tree cutting was applied to identify modules by dividing the dendrogram at significant branch points. C. Analysis of correlations between modules and HPGD expression. The brown and blue modules were significantly correlated with HPGD expression. D. Functional enrichment analysis of genes from brown modules. Bar plot of genes functional enrichment terms (upper panel). Network relationship plots among all terms (lower panel).
Validation of HPGD expression and immune cell infiltration in samples collected from the local hospital
PGE2 is an immunosuppressive factor of the tumor microenvironment, and its level is closely related to tumor progression, which is regulated by both COX-2 and 15-PGDH. We thus examined the protein expression of COX-2 and 15-PGDH in CESC samples with HPV16-positive staining. The immunohistochemistry results showed that COX-2 expression was significantly increased while 15-PGDH expression was significantly decreased in HPV16-positive tissues compared to HPV16-negative tissues (Figure 7(A)). We further confirmed the immune cell infiltration by multiplex immunofluorescence labeling in different subgroups based on tumor grade. Our results show that CD4-, CD8- and Treg-positive staining was significantly increased in CIN-II grade and decreased along with the cancer progress (Figure 7(B) and (C)). CD68 and CD11b showed consistent expression levels from CIN-I grade to cancer grade. These results are consistent with the bioinformatics analysis that 15-PGDH expression was decreased in CESC and suggests that a transformation of cervical cancer from the immune hot tumor into a cold tumor may occur between CIN-II to CIN-III grade transitions during the cancer progression, which may be associated with 15-PGDH expression.

Experimental validation from local cohort samples. A. The representative IHC images and expression levels of COX-2 and 15-PGDH in HPV16- normal and HPV16 + cancer tissues. n = 3, Student's t-test, *P < 0.05, ***P < 0.001 vs. normal group. B. Data statistics of multiplex immunofluorescence labeling from CESC patients graded from CIN I-III to CC. n = 3. One-way ANOVA, *P < 0.05, **P < 0.01 vs. CIN-II group. C. The representative images of immune profile at different grades of CESC.
Discussion
15-PGDH has been reported as the tumor suppressor and modulator of tumor microenvironment functioned in various cancers, while the role of 15-PGDH in regulating immune environments and its clinical significance in cervical cancer remains unclear. The current study investigated the landscape and predictive value of 15-PGDH in carcinogenesis, prognosis, and therapeutic sensitivity, as well as the immune infiltration and the underlying signal pathways in CESC patients. The public databases and cancer samples collected from local hospitals were used for bioinformatics analysis and experimental verification, respectively. Our study reveals that HPGD can serve as a valuable marker for predicting carcinogenesis, alteration of immune microenvironment, and response to chemotherapy in CESC patients. The major findings are three-fold: (1) HPGD/15-PGDH is a potential susceptible predictor of HPV16-associated cervical carcinogenesis. (2) The depletion of 15-PGDH in TAM portends the alteration of the immune microenvironment with a tendency toward a cold tumor environment. (3) The HPGD gene can be used as a sensitivity indicator for chemotherapy.
15-PGDH as a key enzyme of the PGE2 metabolism pathway, has been identified as a susceptibility biomarker for gastric and colorectal cancers.31,32 In the current study, we found the mRNA and protein expression levels of HPGD were decreased in CESC patients by analyzing data from the public database and our cohort, respectively (Figure 1(A) and Figure 7(A)). We also observed an expression pattern with high COX-2 and low 15-PGDH in the HPV16 positive sample which may favor high levels of tumor-promoting PGE2 in the tumor microenvironment (Figure 7(A)). The multiplex fluorescence results showed a transient immune activation in the CIN-II grade, which is consistent with the trend of 15-PGDH expression. These data support the idea that 15-PGDH may act as a tumor suppressor, and negative regulation of 15-PGDH may serve as a susceptibility biomarker for the diagnosis of cervical cancer. Given these findings, we further investigate whether HPGD was a prognostic gene for CESC patients. We found that HPGD was not a prognostic gene of CESC patients, despite a slightly significant difference between the T3 and T2 subgroups (Figure 1(B) to (I)). Our results showed that the Nomogram model, which consists of scores for clinical characteristics and 15-PGDH, predicted OS of one year and three years (Figure 5) was consistent with the observed, suggesting the Nomogram model had good prediction efficiency and can be further conducted for prediction analysis on the OS in CESC patients. Recent research reported that downregulation of 15-PGDH is associated with poor prognosis in ERα-positive breast and endometrial cancer,33,34 and HPGD is also shown as a significant risk factor in colorectal cancer. 35 Thus, the prognostic value of HPGD in cervical cancer needs to be validated in a larger cohort of samples to ascertain their possible clinical use.
It has been reported that oncolytic HSV-1 virus expressing 15-PGDH could change the immune tolerance status of TME, reverse tumor tolerance to oncolytic virus therapy, and enable tumors sensitive to other immunotherapies. 36 Therefore, we further evaluate the predictive value of the HPGD gene as a sensitivity indicator for immunotherapy and chemotherapy. We found neither MSI, neoantigen, nor TMB is associated with HPGD (Figure 3(B) to (D)). By analyzing the proportion of tumor-infiltrating immune cells, we found that HPGD expression was significantly positively correlated with “T cell regulatory” and negatively correlated with “Macrophages M0” and “Mast cells activated” (Figure 2(A)). These results are consistent with our multiplex immunofluorescence results which show the increases of CD68 and CD11b cells from CIN-I to cancer grade in cervical cancer (Figure 7(B)), and suggesting that the low expression of HPGD imply a positive mast cell activation and viral infections in CESC patients. It is worth noting that low expression of HPGD is resistant to AZD6482, BIBW2992, AKT.inhibitor.VIII, JNK.9L, and Lapatinib treatment by analyzing the GDSC database (Figure 4). Taken together, our results indicated the role of HPGD as a predictor for immune activation in tumor microenvironment, as well as a sensitivity indicator for chemotherapy in cervical cancer.
Although the current study has initially addressed the value of HPGD in the diagnosis, prognosis, and assessment of immuno- and chemotherapy which are promising for clinical application, there are still limitations. First, the prognostic value and immune regulator efficacy of HPGD need to be further validated in a broader clinical cohort. Secondly, the IHC data collected without distinguishing the different grades of cervical cancer, and the more in-depth mechanism and clinical transformation studies with in vivo and in vitro experiments need to be systematically elucidated. In summary, we conducted a comprehensive assessment of HPGD in the clinical significance and revealed its diagnostic, prognostic, and therapeutic-predictive value for CESC patients. In future studies, further investigation of the role and mechanism of HPGD in carcinogenesis and immune environment regulation will pave the way for the translation to clinical application.
Conclusion
This study identified HPGD/15-PGDH as a potential biomarker for predicting the progression, immune response, and chemotherapy sensitivity of cervical cancer, which provide critical implications for diagnosis and treatment strategies. These findings pave the way for identifying patients who are most likely to benefit from specific therapies and tailoring treatment approaches to individual patients. Further research that conducts validation studies in larger and more diverse clinical cohorts, distinguishing between different grades of cervical cancer to validate the findings is crucial for advancing the understanding and potential clinical application of HPGD/15-PGDH in cervical cancer.
Supplemental Material
sj-xlsx-1-cbm-10.1177_18758592241296277 - Supplemental material for Predictive role of HPGD gene in carcinogenesis and immune environment monitoring in human cervical cancer
Supplemental material, sj-xlsx-1-cbm-10.1177_18758592241296277 for Predictive role of HPGD gene in carcinogenesis and immune environment monitoring in human cervical cancer by Mingzhu Zhai, Weihua Yang, Chen Zou, Shan Du, Benqing Wu, Changshan Wang, Yuanzhi Lu and Yi Zheng in Cancer Biomarkers
Footnotes
Acknowledgments
This work was supported by the Science, Technology & Innovation Commission of Shenzhen Municipality (JCYJ20220530165407018, JCYJ20230807141405012, JCYJ20220531094815034) and the Open Research Fund Program of the State Key Laboratory of Virology of China (2022KF004).
Author contributions
Conception: M.Z., Y.Z., and Y.L.
Interpretation or analysis of data: M.Z., W.Y., C.Z., S.D., and B.W.
Preparation of the manuscript: M.Z., Y.Z., and Y.L.
Revision for important intellectual content: C.Z., S.D., B.W., and C.W.
Supervision: Y.L. and Y.Z.
Institutional review board statement
The study was conducted according to the guidelines of the Declaration of Helsinki, and approved by the Ethics Committee of the Shenzhen Guangming District People's Hospital (protocol code: LL-KT-2020014, date of approval: Apr. 9th, 2020).
Informed consent statement
Informed consent was obtained from all subjects involved in the study. Written informed consent has been obtained from the patients to publish this paper.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
Data will be made available upon request.
Supplemental Material
Supplemental material for this article is available online.
