Abstract
Background:
Accurate documentation of tumours presents significant opportunities for advancing cancer research and improving patient care, yet it also poses challenges for healthcare management.
Objective:
This study aimed to assess the effectiveness and resource implications of source data verification (SDV) in enhancing the quality of tumour documentation data, focusing on accuracy, completeness and correctness.
Method:
Using tumour documentation data from a large German University Hospital, an SDV was conducted by an external audit group (group RE), comparing the data initially documented by the centre’s tumour documentalists (group TD) to available source documents for the years 2016–2020. The analysis set comprised 240 cases, with exemplary data fields strategically selected across various organ entities and other tumour features. Identified errors were cross-validated by a third group (group CO).
Results:
Visualisations depicted error frequencies by diagnosis year and organ entity. Potential errors were identified, providing feedback to the tumour documentation unit. However, uncertainties in error identification raised questions about the efficacy of SDV.
Conclusion:
While effective in identifying errors, SDV faced challenges due to ambiguous source data and potential bias from external auditors, as well as being deemed uneconomical. The study suggests SDVs suitability for small sample validation but questions its scalability for large datasets.
Implications for health information management:
Alternative methods, such as data exchange interfaces to subsystems or plausibility checks, are recommended for enhancing data quality. This study emphasises the need to explore alternatives for improving data quality in tumour documentation.
Keywords
Introduction
In medical research and clinical practice, accurate and reliable documentation of tumour-related data can serve as a prerequisite for a better understanding of cancer and improved patient care (Kourou et al., 2021). As medical data are increasingly converted to digital formats, data quality challenges become more relevant (Raghupathi and Raghupathi, 2014). Ensuring the integrity, completeness and accuracy of tumour documentation data is not only essential for research and clinical decision-making but also has the potential to significantly impact patient outcomes and treatments (Zarour et al., 2021). In this context, source data verification (SDV) has emerged as a potential strategy to improve data quality by validating different data quality indicators of source data against their documented representation (Nonnemacher and Stausberg, 2006).
In many countries, such as the United States (based on the Surveillance, Epidemiology, and End Results programme (Cronin et al., 2014), the United Kingdom (based on the Nation Cancer Registration and Analysis Service) (Henson et al., 2020) or Germany (based on the Oncological Base Dataset) (Basisdatensatz, 2021), standardised tumour documentation has been established, driven by certification requirements or legal mandates.
Tumour documentation data are derived from routinely collected information. The utilisation of routinely collected data carries a risk highlighted by the “garbage-in, garbage-out” phenomenon, which posits that the quality of output is contingent upon the quality of input within any given system (Kilkenny and Robinson, 2018). Additionally, transcription errors represent another potential source of inaccuracies in data entries, manifesting as errors that occur during the manual transcribing of information from unstructured sources into structured tumour documentation systems (Feng et al., 2020). Theoretically, the obligation to document should ensure the absence of data gaps; however practically, regulations in terms of data quality are often missing. While there is an awareness of the importance of high-quality data, the establishment of a structured and sustainable data quality measurement and enhancement process to ensure reliable data for health research, audits, certifications and broader quality assurance efforts is notably lacking (Jacke et al., 2012).
Borner et al. (2022) addressed this problem by conducting a study about a data quality analysis with a SDV for 2014 and 2015 at the Ludwig Maximilian University (LMU) Hospital, based on Comprehensive Cancer Centre (CCC) (Comprehensive Cancer Centre Munich, 2023) tumour documentation data. The term “SDV” entails the examination of collected data against original documents, such as laboratory reports, to ensure accuracy and reliability (Andersen et al., 2015).
The results in Borner et al.’s (2022) experiment were largely positive and indicated that SDV might serve as an effective tool for detecting errors in tumour documentation. However, retrospectively, a potential methodological problem was identified in which the local tumour documentalists themselves carried out the SDV within their own context. Consequently, it is possible that systematic errors that were considered correct within the local team of tumour documentalists might have been assessed as incorrect by an external auditor.
Motivated by this, the current analysis team embarked on a reassessment of SDV for the years 2016–2020, on the LMU-Hospital’s tumour documentation data focusing on selected data fields and utilising an external audit group not being part of the team of tumour documentalists, for the verification process. The aim was to re-evaluate the effectiveness and resource implications of SDV as a means of enhancing data quality in tumour documentation. The key areas of concentration included the data quality indicators accuracy, completeness and correctness, as well as the cost-effectiveness of the method.
Method
Research design
The methodological framework employed in this study encompasses a SDV as its primary approach and quantifying the data quality indicators of completeness, correctness and accuracy within a subset of data extracted on 9 February 2021 from the LMU Hospital’s CREDOS system (Cancer Retrieval Evaluation and Documentation System), a local clinical cancer registry that is not publicly accessible (Nasseh et al., 2020; Voigt et al., 2010).
While the local tumour documentation system was relatively extensive, encompassing over 50,000 cases in the year 2022, and comprising more than 100 tables and 2000 data fields, a deliberate reduction in data scope to a sample (hereinafter the ‘analysis set’) was undertaken for the purposes of evaluation (1 February 2021). This reduction led to a final dataset of 240 cases. It is essential to underscore that the cases exclusively pertained to primary cases, as defined by Onkozert (2021), a certification-based designation best described as cases that received their primary treatment at one of the local cancer centres. In our context, these centres were affiliated with the CCC Munich. The selection of cases for inclusion in this analysis set was conducted with an emphasis on an equitable distribution across the years of diagnosis 2016 to 2020.
Additionally, the dataset was stratified based on the following organ entities, with equal representation for each: breast, colon, gynaecological, head and neck, lung, neurological, pancreas and prostate tumours. Furthermore, an additional selection criterion was applied, leading to the inclusion of a total of six cases per year and organ entity based on their initial therapy, where two cases received either surgery, radiation therapy or systemic therapy. This selection criterion was adopted to ensure a focus on representative data fields in the domains of diagnosis, therapy and progression, as shown in Box 1. All cases with the given data contents were exported into a study database.
Description of selected fields. Each field belongs to one of the three categories: first assessment; therapy; progression.
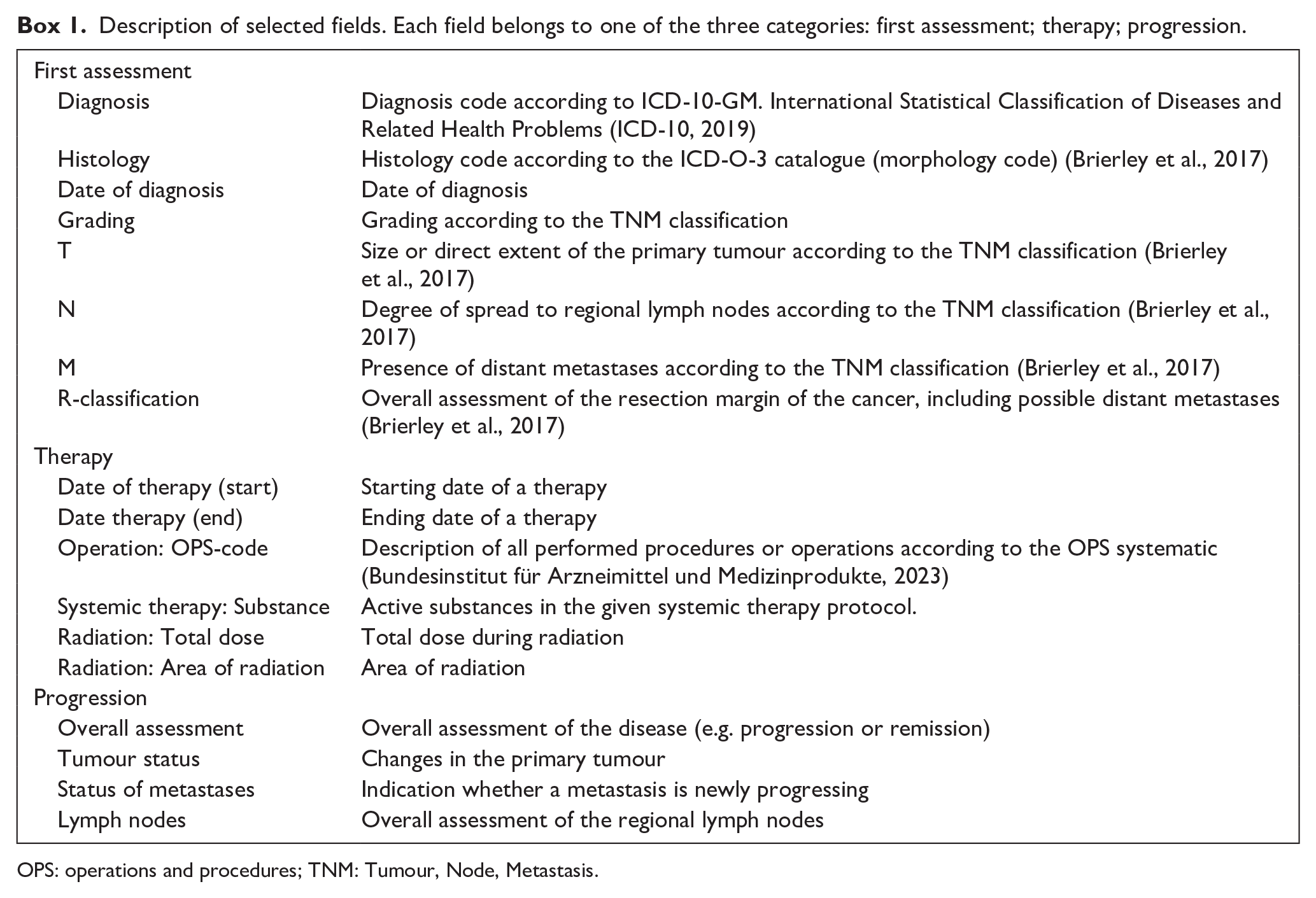
OPS: operations and procedures; TNM: Tumour, Node, Metastasis.
The execution of this study was entrusted to two groups, group RE (re-evaluation) and group CO (control), both freshly affiliated with the CCC in terms of the study, albeit external to the tumour documentation team, but experienced in the field of medicine. This strategic choice was deliberate, serving the purpose of ensuring an absence of bias associated with team internal regulations governing tumour documentation.
Group RE was assigned the responsibility of re-documenting the provided sample in the absence of access to the preceding tumour documentation. Group RE was given a blank version of the research database, only containing patient identification numbers and Supplemental Material about the year of diagnosis, entity and the required documentation in terms of therapeutic interventions, encompassing surgical procedures, systemic therapy or radiation treatments. Like the tumour documentalists on the internal team, group RE then got access to all source data, such as doctors’ notes and radiation therapy reports. This allowed group RE to fill in the missing parts of the study database. The initial data collection process was straightforward, focusing only on the fields selected for the study. For therapy-related fields, only those associated with the first therapy were documented. In this process, it was revealed that the operations and procedures (OPS) code consistently exhibited full correctness, as the information was semi-automatically imported from a SAP i.s.h.med-based surgery subsystem (Hoeksma, 2009). Consequently, contrary to the initial plan, this data field was excluded from the analysis.
Regarding the patient’s progress, it was initially decided to document only the fields of the most recent progress in relation to the export date of the sample. However, this approach was abandoned during the data collection phase (a decision we discuss in the subsequent discussion section of this article).
Prior to data collection, a copy of the database containing exports from the tumour documentation system was created and isolated from group RE. Later, this enabled a computational comparison between the original data collection from the tumour documentation conducted by group TD (tumour documentalists) and the independent data collection by group RE. During data reconciliation, three types of errors were distinguished:
Completeness errors: Documentation for a data field was present for either group RE or group TD but not both.
Correctness errors: Documentation was present in both datasets but exhibited significant discrepancies (e.g. ICD-10 diagnosis codes C34 and C55).
Accuracy errors: Documentation was available in both datasets, but the discrepancy was considered an inaccuracy based on analysis specific rules (e.g. ICD-10 diagnosis C34.9 instead of C34.3). A listing of our accuracy rules can be found in Box 2.
Description of accuracy errors and how it was applied.
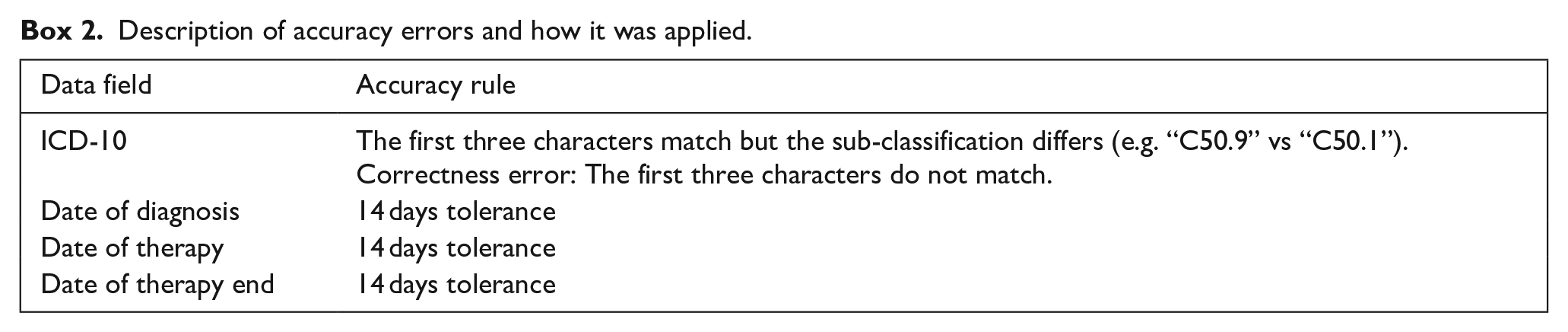
A pitfall observed in Borner et al.’s (2022) work was the attribution of errors solely to the detriment of the tumour documentarians (group TD). In reality, it is plausible that errors could have been made by group RE as well. While our study regarded concurrence in errors as genuine agreements, all cases of disagreement were re-evaluated by a third individual (group CO), who also had access to source data and previous documentation but without the knowledge of its origin (group RE or group TD). Group CO thus adjudicated responsibility for errors. There were scenarios where group CO did not concur with the findings of either group RE or group TD. In these instances, the results from group CO had to be neglected due to ambiguity. The complete workflow is illustrated in Figure 1.

SDV process conducted by group RE and group CO.
The following error classes were defined for the final evaluation (see Box 3). Using these error classes, the analysis results were visualised using Python v. 3.8.8 (Python, 2021). If group RE committed an error, it was annotated as ACC_RE, COR_RE, and COM_RE. For group TD as ACC_TD, COR_TD, COM_TD. In the event that group CO determined the documentations of both groups to be incorrect, the labels ACC_CO, COR_CO, and COM_CO were applied.
Error categorisation for accuracy, correctness and completeness in documentation for groups RE, TD and CO, including uncertainty (U) for group CO.
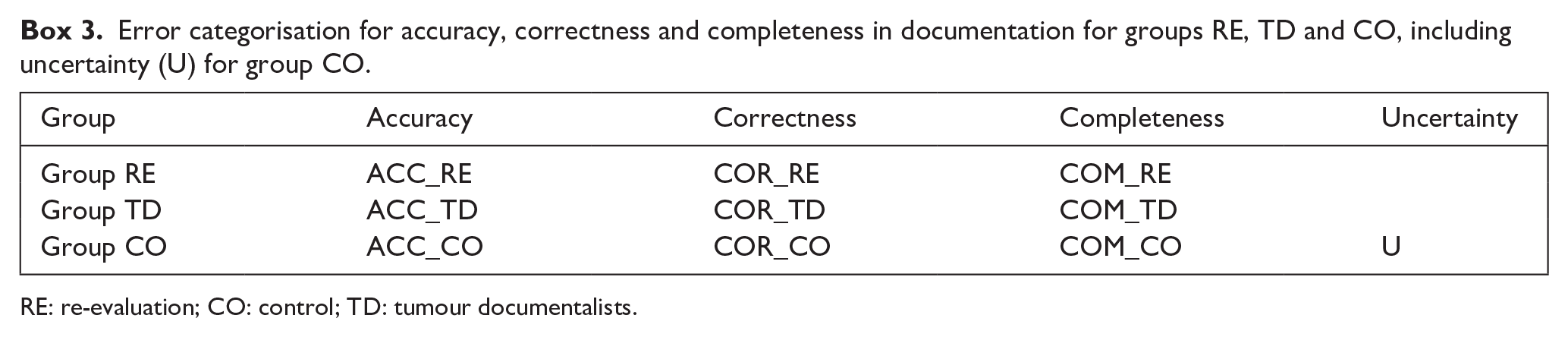
RE: re-evaluation; CO: control; TD: tumour documentalists.
Ethics approval
Formal ethics approval from a Human Research Ethics Committee (HREC) was considered unnecessary as the data were retrospective, de-identified and aggregated for publication.
Results
Figure 2 portrays the relative frequency of errors uniquely made in the original tumour documentation, focusing on the distribution of errors related to correctness, completeness and accuracy within the individual audit fields. This chart essentially represents the result of the SDV process, evaluated by group RE and validated by group CO. Errors or uncertainties of both groups TD and RE according to group CO, due to ambiguity, were excluded from this graph. It should be noted again that the audit fields in the domain of progressions were not used due to methodological difficulties (see ‘Discussion’ section).
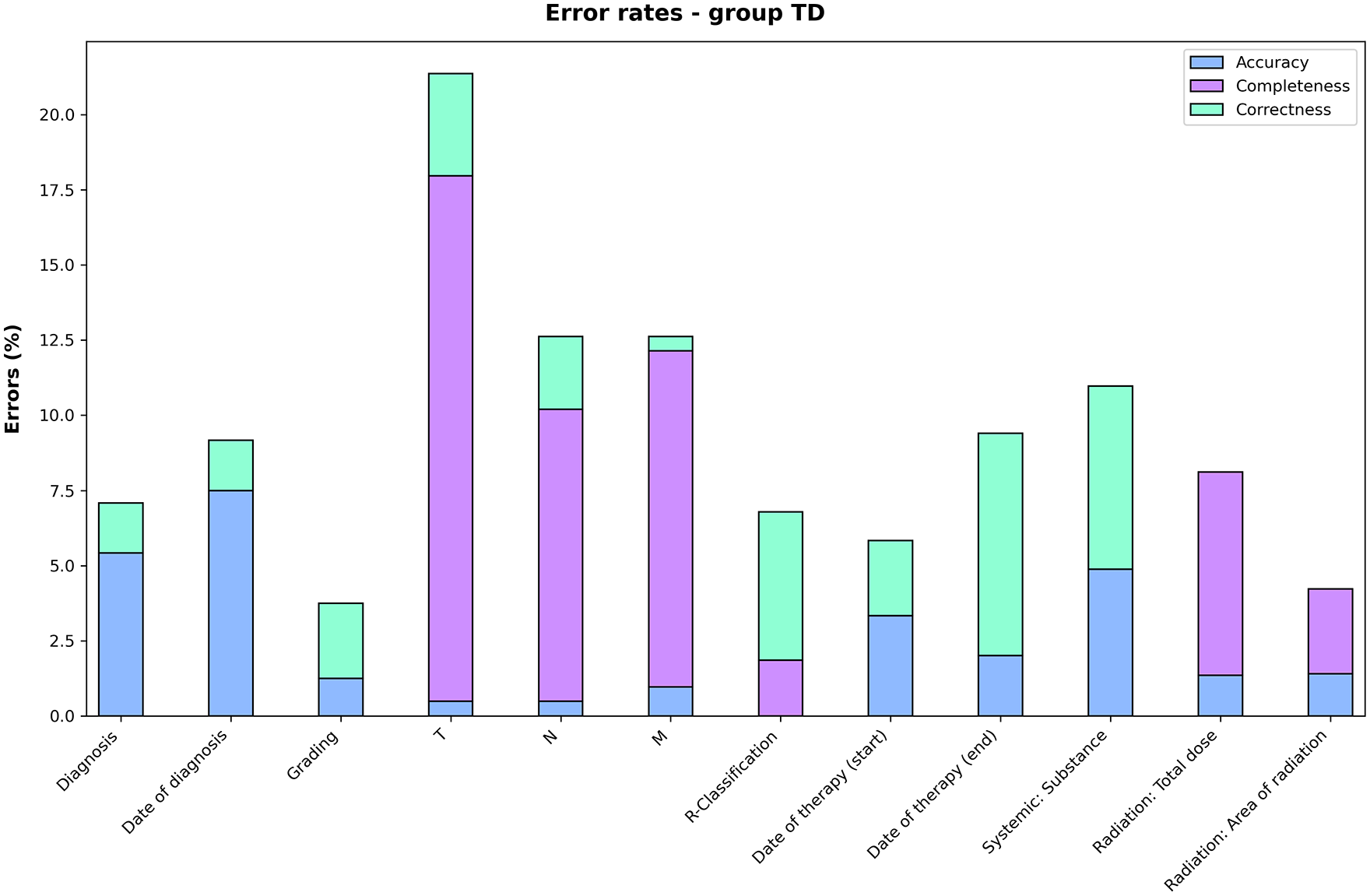
Individual error rates of group TD.
Figure 3(a and b) (first assessment) and Figure 4(a and b) (therapies), on the other hand, compares the distribution of the earlier introduced error classes also concerning uncertainties or errors made by both group TD and RE (according to group CO). In these graphs column TD again displays the errors attributed to group TD, while column RE shows the errors attributed to group RE by group CO. The last column represents the fields where neither group RE nor group TD, according to group CO, were deemed correct, or cases in which group CO was uncertain (degree of ambiguity). ‘
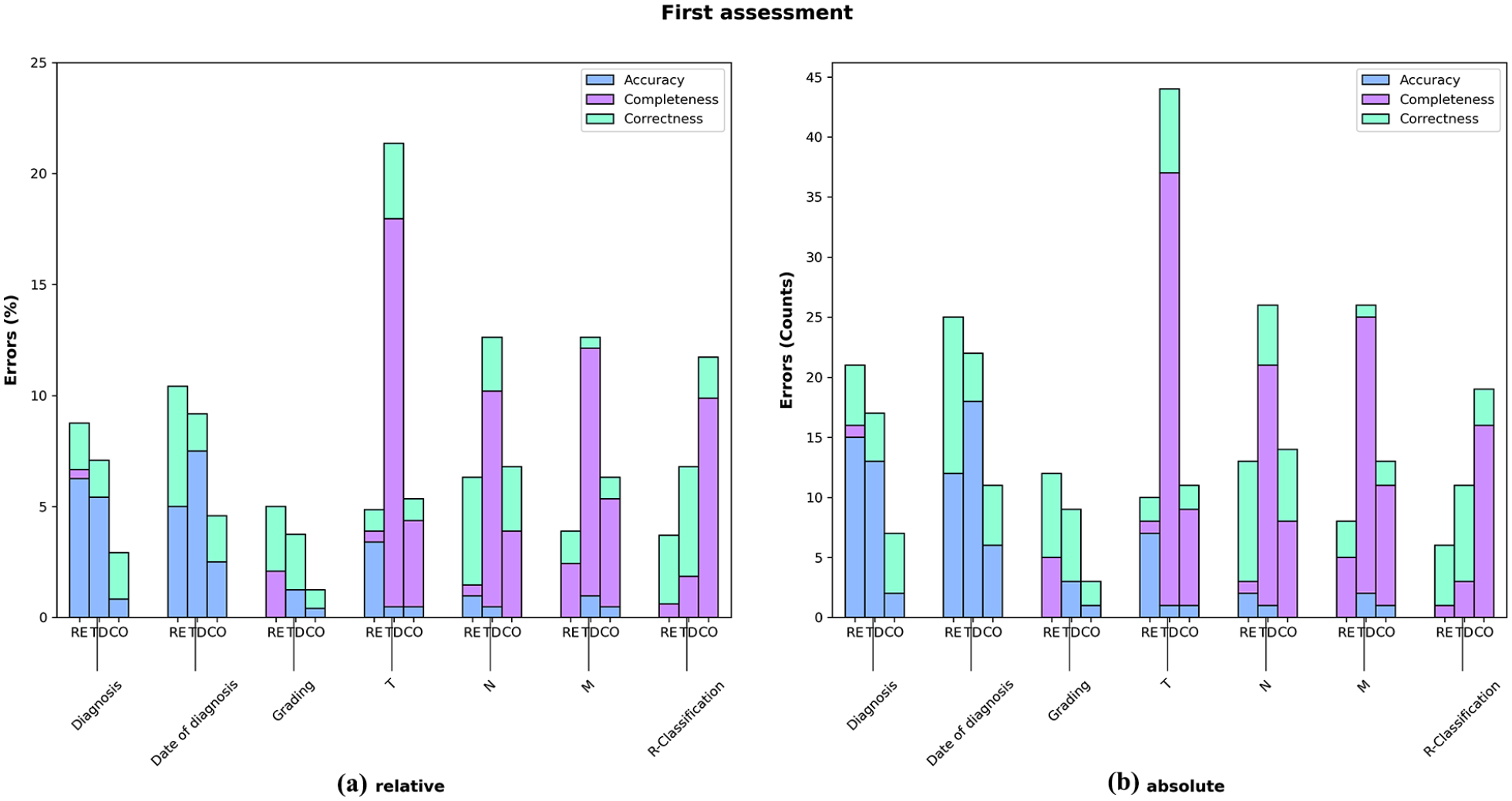
(a and b) Potential error classes of fields related to first assessment grouped by group RE, group TD or ambiguity (CO) with graph A showing relative error rates and graph B showing absolute numbers.
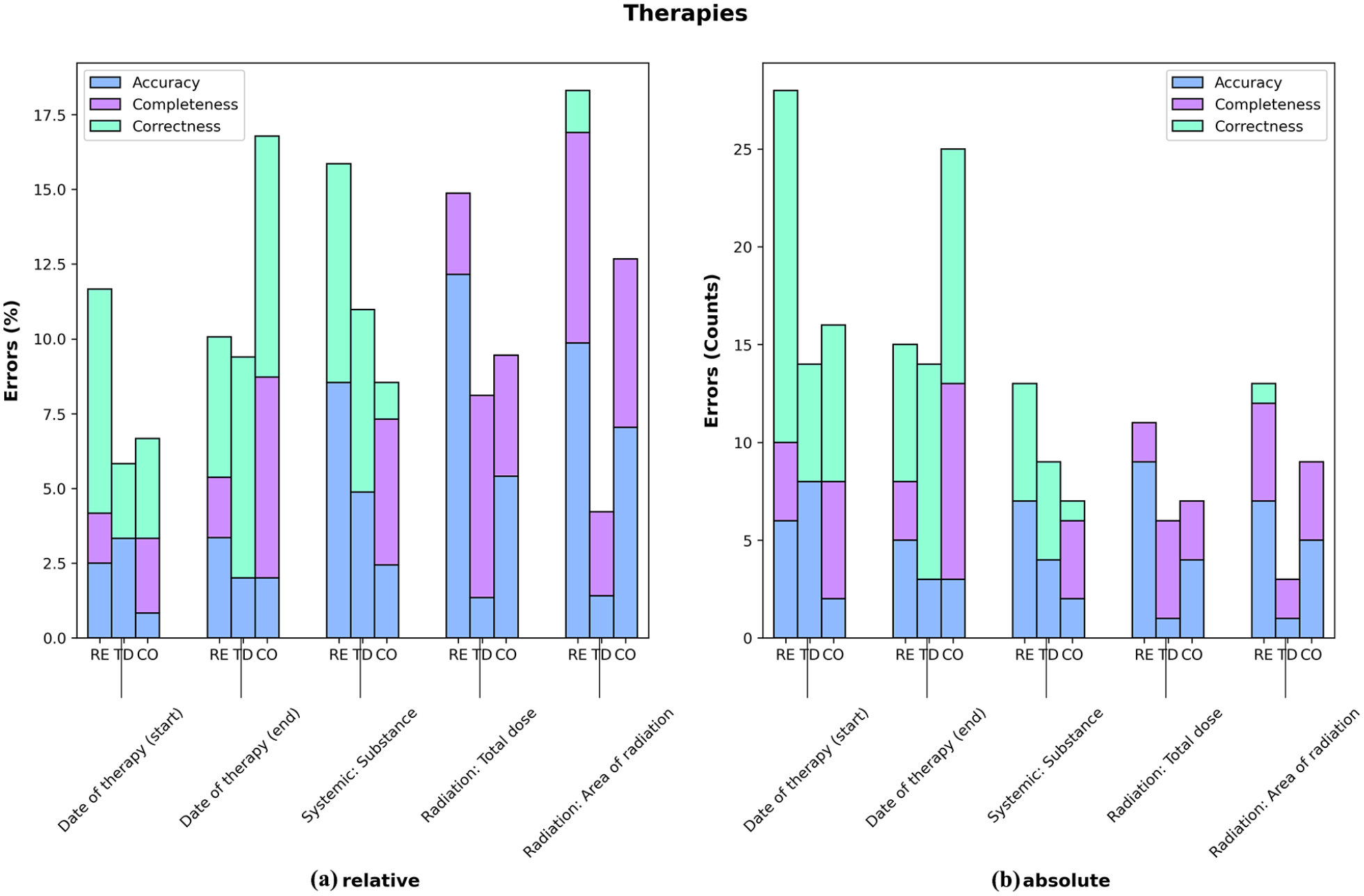
(a and b) Potential error classes of fields related to therapies grouped by group RE, group TD or ambiguity (CO) with graph A showing relative error rates and graph B showing absolute numbers.
The errors, categorised by type (correctness, completeness and accuracy), uniquely made by Group TD were also analysed over the years, providing insights into potential trends in data quality (see Figure 5). Figure 6 provides insights into the level of accuracy in date fields, as these are often suspected to exhibit accuracy errors, rather than errors related to correctness or completeness. A substantial proportion of discrepancies remain within a narrow margin from the original date (14 days), a range that is unlikely to have a significant impact on research outcomes and is therefore classified as an accuracy error. Most deviations beyond this threshold are typically within approximately 50 days. However, a small number of extreme outliers are evident, representing clear and significant errors that may warrant closer examination.
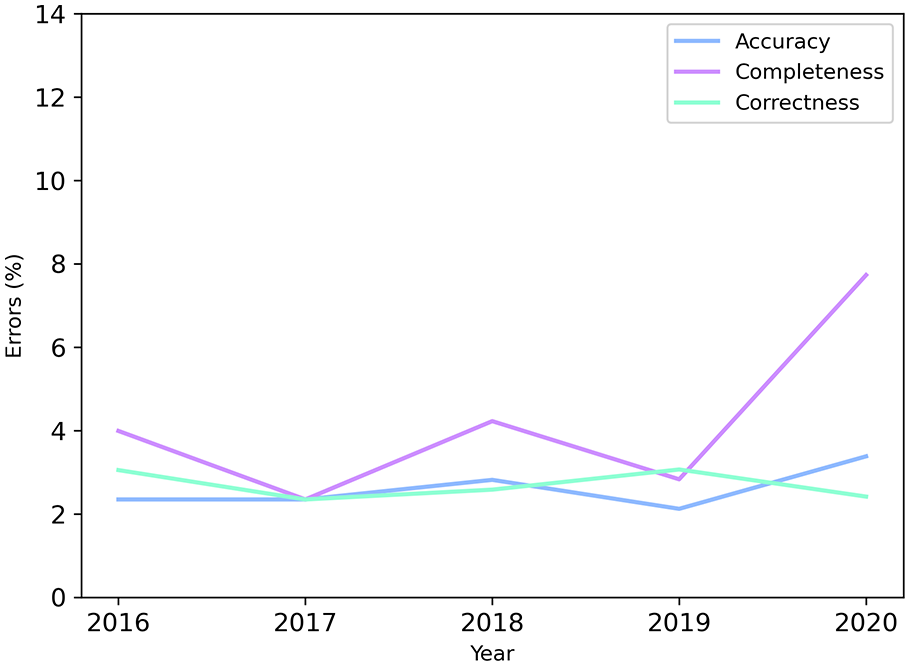
Potential error rates over the years of group TD.
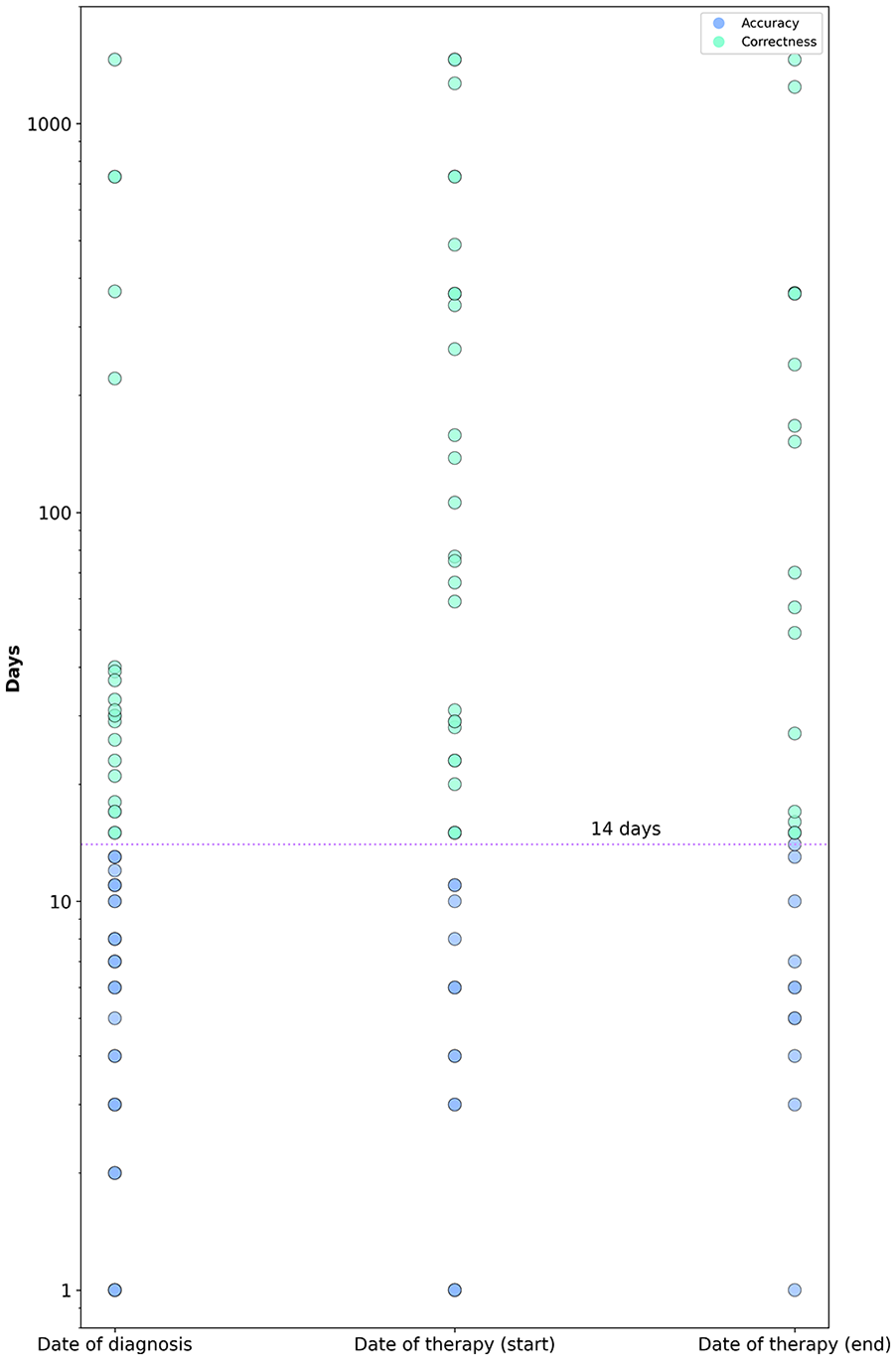
Accuracy error assessment (date fields): The blue region (14 days) delineated signifies the interval where date errors are exclusively evaluated as accuracy errors and do not carry the classification of correctness errors.
Next, the performances of group RE and group TD were re-evaluated (excluding those errors made by both according to group CO). Heat maps were generated to represent error frequencies, stratified by entity (Figure 7) and by diagnosis year (Figure 8). The entities/organs were anonymised in the statistics to prevent inferences about individual documentalists. While the majority of combinations of entity and data field (Figure 7), as well as year of diagnosis and data field (Figure 8), are entirely error-free, certain areas emerge as potential sources of concern, with darker colours indicating more critical issues. Moreover, it becomes apparent that group RE (positioned at the top of each figure) and group TD (positioned at the bottom of each figure) exhibit issues in distinct and different areas.
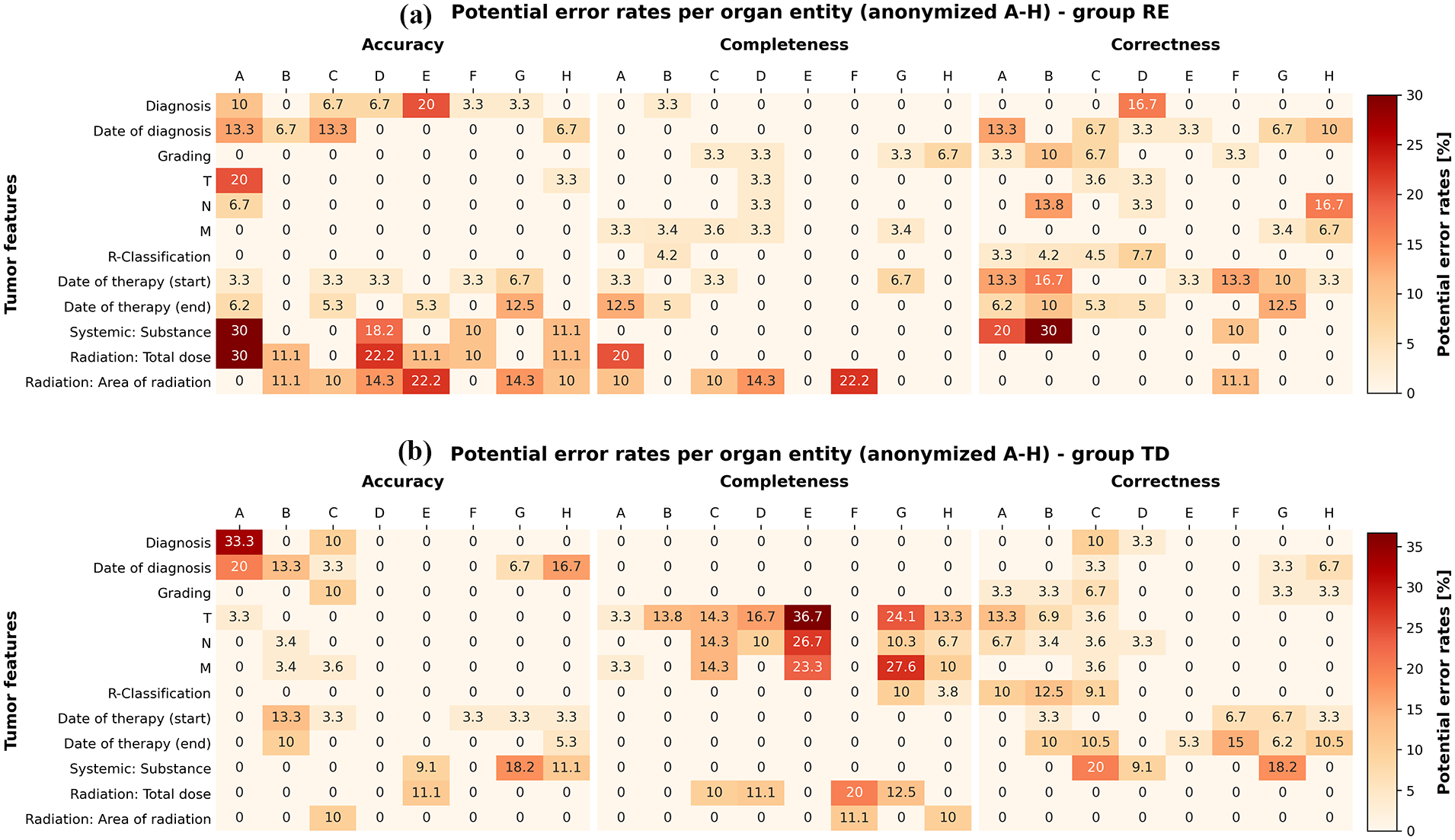
Potential errors by group RE (a) and group TD (b) per anonymised organ entity (A–H) versus individual data fields.
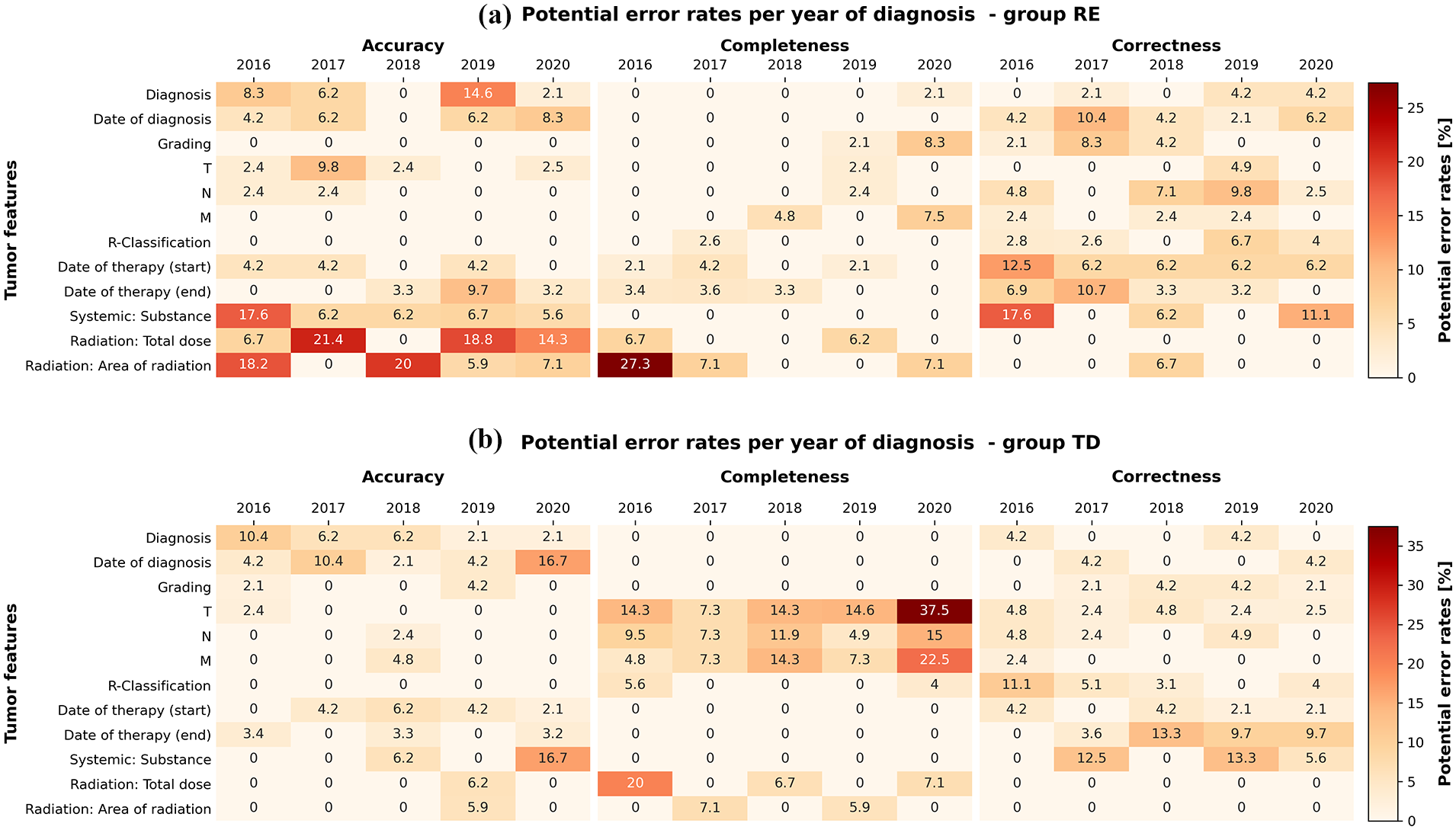
Potential errors by group RE (a) and group TD (b) per diagnosis year versus individual data fields.
Regarding the whole set of analyses fields (according to group CO), group RE achieved a total (total amount of errors/total amount of fields) correctness rate of 96.4%, while group TD achieved 97.4%. Completeness rates were 98.7% for group RE and 95.8% for group TD, respectively, with accuracy rates of 96.7% for group RE and 97.4% for group TD.
In the overall process, group RE invested 300 hours examining the source data and re-documenting the relevant data items to set up the analysis database. The examination of our complete set of fields per case was projected to demand around 60 minutes for both group RE and CO. The TD group required approximately 30 minutes for rectifying individual errors (feedback).
Discussion
High-quality health data, such as tumour documentation, play a crucial role in tasks like clinical decision-making, research advancements or external certifications. However, uncertainties, inconsistencies or errors can undermine trust in its reliability, underscoring the importance of robust mechanisms to ensure transparency and data quality (Lighterness et al., 2024; Zarour et al., 2021). This study sought to address these concerns by evaluating the practical utility of SDV within the context of tumour documentation. By comparing the performance of internal tumour documentalists with that of an external audit group, we aimed to not only assess the reliability of tumour documentation but also to explore the potential biases introduced by different methodological setups. With this new approach, we believe our findings provide a more realistic perspective on the strengths and limitations of SDV in this specific setting. The results revealed that the quantification of correctness and completeness (group TD) appears to be lower compared to Borner et al.’s (2022) analysis conducted a few years ago (correctness 97.3% vs 99.2%, completeness 95.8% vs 99.0%) even though, a portion of correctness errors in the current study has been classified as accuracy errors (accuracy: 97.4%).
However, we posit that the observed discrepancy may not necessarily indicate a deterioration in local data quality. Rather, it might suggest that the decrease of data quality has been affected by the new experimental setup. An aspect is that the evaluation team in the previous study (Borner et al., 2022) were integral members of the internal team of tumour documentalists. As such, they controlled their own data through SDV, potentially leading to an underestimation of errors simultaneous with an overestimation of data quality due to internal rules and regulations. Furthermore, it is essential to consider that the analysed data fields between the two studies varied. In our current study, certain fields, such as the OPS code, were excluded from the analysis due to its inherent nature of being semi-automatically integrated, leading to full correctness and completeness in regard to transcription errors.
While bias introduced through internal verification has been successfully eliminated in the current setup, a novel concern has emerged compared to the experiment conducted by Borner et al. (2022). The team of documentalists possesses a higher level of training within their respective field compared to the group RE. Group RE included a dentist with a completed degree, a family history of cancer and relevant medical experience, which fostered a strong interest in oncology. While possessing general knowledge in the medical domain, the participant lacked formal training in tumour documentation. To address this, introductory training related to the fields under investigation was provided, and additional support was offered by the CCC staff, including experienced documentalists, when challenges in documentation arose. While external auditors help mitigate bias, it would have been advantageous if the auditors themselves were also external tumour documentalists. Ideally, the experimental design would have facilitated an exchange of tumour documentalists between multiple university hospitals or cancer registries, ensuring that all auditors possessed comparable levels of training and expertise in tumour documentation. However, this optimal setup was not feasible due to funding constraints. It is important to note, though, that even among regular tumour documentalists, there is typically no standardised national training programme, and professionals in this field often come from diverse educational and professional backgrounds.
Nonetheless, the educational background of the auditors could be perceived as the main limitation of this study. However, it is crucial to note that the findings of group RE underwent validation by a separate group, group CO, which also possesses medical expertise (Master’s students specialising in Medical Information Management and Nutrition) and received similar support from the CCC staff when needed. What emerged from this arrangement was that group CO, in fact, identified a comparable number of errors in the disagreements of TD and RE. Additionally, it should be noted that group CO also, at times, encountered difficulties in discerning who was at fault. Due to this ambiguity, when measuring the error rates of group TD, we mainly focused on errors where group RE and group CO agreed. Still, these problems display that even in those cases, a certain degree of ambiguity remains and it is crucial to emphasise that the errors identified may not be definitive errors; instead, they may serve as indications or markers of potential errors. This outcome raises questions regarding the efficacy of SDV as a standard tool for improving data quality for tumour documentation.
SDV proves efficient when information is strictly confined to specific source documents ensuring the absence of ambiguity. However, tumour documentation faces a difficulty: the source of documented data lacks traceability (as the source itself is usually not documented), and varying sources for the same data field are present at different time points. This issue is being addressed in many studies where source documentation is often available, detailing which information originates from which source (Bargaje, 2011). However, this is usually not the case in tumour documentation. For example, information about the diagnosis code may originate from both medical reports or pathology findings. It might even be possible that, at the time of documentation, certain information may not have been available. Logically, the individuals documenting later in time (in this case, group RE documented after group TD) possess more information/source documents since the tumour documentation process aims to record information as promptly as possible, usually within 3 months. As described in Giusti et al. (2023) there is usually a trade-off between timeliness and both completeness and validity in data quality.
In our study, the issue of ambivalent source data has been particularly pronounced in the monitoring of disease progression and the identification of the original source document reached a level where the evaluation became devoid of meaningful interpretation. Ultimately, this led us to exclude the examination of progress fields from the methodology.
If we focus solely on potential errors within the context of group TD, we see a decline in completeness in the year 2020 (refer to Figure 5). Upon examining the heatmaps, it becomes evident that especially the fields related to “Tumour, Node, Metastasis” (TNM) are the cause, as illustrated in Figure 8. While subject to interpretation, we posit that this phenomenon may be attributed to site specific internal changes in the documentation of TNM. Based on our own experiences, TNM appears to be one of the most inconsistently documented fields in Germany, lacking semantic clarity on how TNM should be documented. Therefore, the error may also stem from the fact that group RE and group CO, despite operating independently, interpreted the documentation of TNM differently than group TD, suggesting that these are potential rather than definitive errors.
Furthermore, it is noteworthy that certain issues manifest only in specific organ entities, indicating a non-uniform occurrence of the phenomenon. For instance, in Entity A (refer to Figure 7(b)), we observe a 33% error rate in accuracy, implying that group RE was able to provide a more accurate diagnosis than group TD in these cases. This discrepancy could be attributed to the timeliness of the source documents, indicating that group RE and CO had access to more recent and precise data, or it may signify an individual problem specific to a particular tumour documentalist. Another example is the recurring issue of poor completeness in TNM, particularly notable in organ E.
A distinctive aspect of our study was the nuanced categorisation of deviations not solely as correctness errors but also differentiated as accuracy errors based on our own defined criteria (see Box 2). This methodology carries medical significance, particularly in instances where subtle variations in factors like diagnosis dates have negligible effects on calculations such as survival curves. Considering the roughly equal prevalence of correctness and accuracy errors in our results, we advocate for the adoption of this refined distinction in other potential SDV projects.
Despite the challenges encountered, as common after an SDV, we can report potential errors as feedback, or at least as error indicators to the tumour documentation unit. Feedback mechanisms play a crucial role in identifying and correcting errors, ensuring continuous improvement in both planning and execution phases (Oyebode, 2013). The feedback would allow the TD team to rectify potential errors. However, since these are only potential errors, the tumour documentation unit would essentially also need to undertake unnecessary work, including re-examining valid cases.
Ultimately, the data quality is certain to benefit from such a measure; however, there is a need to scrutinise the time investment and economic viability associated with it. In our assessment, the benefits of regular SDV interventions in the realm of tumour documentation, aimed at rectifying errors, do not justify the expended effort. In our study, the time investment required for the SDV process was systematically recorded without any predefined assumptions regarding its expected duration. Although we only verified selected fields rather than entire cases, we found that this still required nearly full review of the respective cases, as the necessary information was often distributed across multiple sources. Even for this targeted sample-based approach, the effort was substantial. Identifying errors in specific fields thus often resembled the workload of complete re-documentation. This level of effort makes widespread implementation of SDV economically and practically unfeasible.
However, it appears to be suitable for uncovering individual systematic problems, such as (in our study) incompleteness in TNM staging by specific staff members, which could be re-trained accordingly after receiving feedback. For more sensitive and wide spread error detection and quality improvement, we suggest exploring alternative approaches. An alternative method for enhancing data quality appears to be the direct integration of subsystems (instead of using source documents), as exemplified in the current scenario with the surgery system. Pathology and pharmacy systems, in particular, offer potential for this approach. However, it is technically demanding, albeit effective in preventing transcription errors entirely. Other options could involve implementing plausibility checks, standardisation of tumour documentation education, or refining the semantics of data standards.
Finally, it is important to emphasise that this work is not intended to cast a negative light on tumour documentation. Rather, the errors observed in group RE highlight the difficulty in achieving completely error-free documentation. Additionally, due to the case of unclear source documentation, it may not always be precisely clear what constitutes an error. Therefore, the profession should be valued accordingly.
Conclusion
In conclusion, our findings suggest that, based on the employed methodology, SDV is most suitable for validating small samples and identifying systemic errors in tumour documentation data. The time investment required for a thorough examination of the entire dataset is not economically justifiable and does not yield a favourable cost–benefit outcome. The obstacle of data quality, especially in tumour documentation, poses a challenge to trust. Hence, it is advisable to also explore alternative methods, including automated data transfer from subsystems, conducting plausibility checks, and making semantic enhancements to existing standards, to effectively enhance data quality in this particular context.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship and/or publication of this article.
