Abstract
Achieving a good outcome for a person with Psoriatic Arthritis (PsA) is made difficult by late diagnosis, heterogenous clinical disease expression and in many cases, failure to adequately suppress inflammatory disease features. Single-centre studies have certainly contributed to our understanding of disease pathogenesis, but to adequately address the major areas of unmet need, multi-partner, collaborative research programmes are now required. HIPPOCRATES is a 5-year, Innovative Medicines Initiative (IMI) programme which includes 17 European academic centres experienced in PsA research, 5 pharmaceutical industry partners, 3 small-/medium-sized industry partners and 2 patient-representative organizations. In this review, the ambitious programme of work to be undertaken by HIPPOCRATES is outlined and common approaches and challenges are identified. It is expected that, when completed, the results will ultimately allow for changes in the approaches to diagnosing, managing and treating PsA allowing for better short-term and long-term outcomes.
Plain language summary
Psoriatic Arthritis (PsA) is a form of arthritis which is found in approximately 30% of people who have the skin condition, Psoriasis. Frequently debilitating and progressive, achieving a good outcome for a person with PsA is made difficult by late diagnosis, disease clinical features and in many cases, failure to adequately control features of inflammation. Research studies from individual centres have certainly contributed to our understanding of why people develop PsA but to adequately address the major areas of unmet need, multi-centre, collaborative research programmes are now required. HIPPOCRATES is a 5-year, Innovative Medicines Initiative (IMI) programme which includes 17 European academic centres experienced in PsA research, 5 pharmaceutical industry partners, 3 small-/medium-sized industry partners and 2 patient representative organisations (see appendix). In this review, the ambitious programme of work to be undertaken by HIPPOCRATES is outlined and common approaches and challenges are identified. The participation of patient research partners in all stages of the work of HIPPOCRATES is highlighted. It is expected that, when completed, the results will ultimately allow for changes in the approaches to diagnosing, managing and treating PsA allowing for improvements in short-term and long-term outcomes.
Introduction
Psoriatic arthritis (PsA) is a clinically heterogenous condition which complicates cutaneous psoriasis (PsO) in approximately 30% of cases and which is characterized by the variable expression of psoriatic skin and nail lesions, arthritis, enthesitis, dactylitis and/or axial disease. 1 Over time, and particularly if inflammation is not adequately suppressed, many patients will progress to articular, peri-articular and axial damage which is irreversible and which can lead to significant functional and psychosocial disability. These consequences of uncontrolled disease in turn have broader societal and economic impact. 2
There are several challenges or unmet needs in diagnosing and managing PsA, 3 some of which are addressed in this review. Unmet needs in PsA are the focus of the Innovative Medicines Initiative (IMI) HIPPOCRATES project (see: www.hippocrates-imi.eu, accessed on 20th September 2022). 4 The areas of unmet need which are addressed by the HIPPOCRATES project are 1 :the development of an algorithm for early diagnosis of PsA 2 ,the identification of clinical/molecular risk factors for PsA in people with cutaneous Pso 3 , the ability to predict which patients with PsA will progress to joint damage and 4 the development of a precision medicine approach to treating PsA based on an individual’s clinical/molecular profile. In this review, we describe the approach which is being taken by the HIPPOCRATES consortium to address these unmet needs and we outline what we hope will be the outcomes of our research efforts. We also anticipate the next steps which will be required to achieve better outcomes for patients with PsA.
Development of an algorithm for early diagnosis
While there are no diagnostic criteria or diagnostic tests for PsA, the ClASsification of Psoriatic ARthritis (CASPAR) criteria 5 are generally used for research purposes to classify PsA and to ensure some uniformity for enrolment in clinical trials or studies. The CASPAR criteria include clinical, laboratory and radiological features. However, a clinical diagnosis of PsA is made by rheumatologists based largely on clinical features. Differentiating PsA from other forms of inflammatory arthritis and PsO alone remains challenging.
It is assumed that PsO patients developing PsA may progress through different phases of ‘psoriatic disease’, starting with aberrant activation of the interleukin (IL) 17-IL-23 axis, followed by a ‘silent’ inflammatory phase, often visible by imaging such as ultrasound or magnetic resonance imaging (MRI), passing to a transition phase characterized by arthralgia and fatigue, ending with clinically evident PsA.4,6 It remains unclear if all patients developing PsA will go through these stages and which events may trigger the transition from one phase to another. Given that irreversible bone erosion and osteoproliferation can be detected early in the inflammatory process, an approach for early diagnosis of PsA that includes molecular biomarkers is of high importance.7,8
Different types of biomarkers have been assessed to see if they differentiate PsA from PsO. Several genetic variants that distinguish PsA from PsO have been identified, including specific HLA B alleles, an IL23R promotor and the 5q31 genomic region.9,10 Gene transcripts like serum micro-RNA (miRNA) may also aid diagnosis; in one study, miR-221-3p, miR-130a-3p, miR-146a-5p, miR-151-5p, miR-26a-5p and miR-21-5p were not only associated with a PsA diagnosis, but also with treatment response, independent of treatment type, making serum miRNA a potential diagnostic and prognostic marker in PsA. 11 In terms of proteins, serum concentrations of the chemokine C-X-C motif chemokine ligand 10 (CXCL10) were found to be significantly higher in PsO patients developing PsA compared to those who remain as PsO 12 and a combination of proteins, including Integrin, ß5 (ITGß5), Human Mac-2-binding Protein (M2BP) and C-Reactive Protein (CRP), has been reported to distinguish PsA patients among PsO and healthy controls. 13 One study of serum metabolites identified dysregulation of bile acids and inflammatory lipid mediators in patients with PsA. 14 In a systematic literature review, Mulder and colleagues reviewed the published studies reporting clinical, laboratory or genetic markers for the development or presence of PsA in patients with PsO. While these reports identified candidate biomarkers, the studies were largely performed in single centres, the cohorts were small and none of the candidate biomarkers were validated in large, independent datasets.
Immunophenotyping studies have revealed that circulating T-helper (Th)17 and Th22 cells are increased in both cutaneous PsO and early PsA compared to heathy controls, but the number of circulating memory Th cell differs. 15 Multiplex immunohistology, which allows visualization of an unlimited number of antibodies on the same tissue slice, allows single-cell phenotyping in biopsies of PsO and PsA patients. Detection and quantification of a wide-panel of cellular markers opens the possibility of characterizing PsO- and PsA-specific cellular networks comprising both immune and non-immune cells, including keratinocytes and blood vessels; indeed, dysregulated angiogenesis has been identified as a possible target for diagnostics in early stages of the disease. 16
Once symptoms of PsA have begun, it is also important to distinguish PsA from other forms of inflammatory arthritis. Differences in the genetic architecture of rheumatoid arthritis (RA), ankylosing spondylitis (AS) and PsA are known to exist, but further work is required to test whether this may aid diagnosis in early undifferentiated inflammatory arthritis. One study found that micro RNAs miR-146b-3p, miR-26a-2–3p, miR-485-3p and let7d-5p were useful to distinguish between RA, AS and PsA as the reason for inflammatory arthritis. 17 The serum lipidomic and metabolomic profiles show differences depending on the underlying inflammatory rheumatic disease making them potential biomarkers to discriminate between PsA and RA. 18 Furthermore, using proteomics and multivariate machine learning (ML), a panel of protein biomarkers was recently identified to separate early-onset PsA from RA. 19 Other accessible biosamples such as synovial fluid and urine may provide further information needed to improve the diagnosis of PsA. 20 Ultimately, a combination of molecular features from genomics, proteomics, metabolomics and lipidomics established and integrated with emerging artificial intelligence (AI) and ML approaches may be required to achieve a reliable diagnostic tool for early PsA. 3
Identifying those people with PsO at high risk of developing PsA and how we can develop strategies aimed at possible prevention?
Studies have shown that 15–30% of people with psoriasis develop PsA 21 but it is difficult to accurately predict in which person with psoriasis that PsA will emerge. As the majority (80–90%) of people present with psoriasis prior to showing features of musculoskeletal involvement, people with PsO represent a ‘population at risk’. Delaying treatment for PsA for as little as 6 months is known to contribute to worse radiographic outputs and functional disability. 8 Previous observational and small randomized controlled studies have raised the potential for preventing or delaying the onset of arthritis.22 –24 To consider such disease-prevention studies, the clinical and molecular factors which identify psoriasis patients at high risk of progressing to PsA need to be identified.
As a first step, we need to validate the existing clinical risk factors, such as nail dystrophic change or body mass index, 25 and investigate whether there might be other clinical predictive markers of PsA. With this in mind, we plan to examine large primary care datasets in the United Kingdom and Spain.26,27 In preparing for these studies, we have initially updated the 2021 systematic literature review looking for clinical, laboratory and genetic markers for the development of PsA in people with PsO. 28
To convince people with psoriasis that they are at risk of PsA and that they should consider early intervention therapies, there is a need to identify which people will progress with a high degree of specificity. The risk of PsA needs to be substantial and backed up by robust evidence, including from prospective studies. While it may yet be too early to conduct disease-prevention studies, plans to establish acceptability thresholds for interventional treatments from the patient perspective and to design optimal studies to evaluate potential prevention strategies are underway within HIPPOCRATES.
A key issue in designing future interventional studies is establishing when the intervention is acceptable to patients. This acceptability will be variable from person to person but is likely to depend on the quantified risk for the development of arthritis and any risks associated with the intervention, for example side effects or burden of treatment administration/adherence. To investigate this further, the HIPPOCRATES Prospective Observational Study (HPOS) investigators have partnered with members of the IMI-funded PREFER consortium who have expertise in patient choice experiments (see: www.imi-prefer.eu, accessed on 1st October 2016). In collaboration with PREFER and patient research partners (PRPs), we are planning a dynamic choice experiment to explore the balance of benefits and risks among a wide group of people with PsO.
While there are a small number of prospective studies of psoriasis patients that have followed/monitored subjects for the development of PsA, most of these studies are of small cohorts and insufficient to allow for robust verification and ultimately validation of findings. We are therefore proposing a new, highly innovative prospective study, HPOS. In HPOS, we propose to establish a European patient-driven prospective observational cohort of 25,000 adults who have PsO but not PsA. We anticipate that 2.7% per annum, 29 equivalent to 675 participants, will develop PsA. We plan to record demographic and clinical features and selectively collect biosamples from participants. At each follow-up timepoint, participants will complete a validated screening questionnaire for PsA and if they screen positive, they would be advised to seek medical review locally. In year 3 of the study, biosamples will be collected from study participants who have developed PsA and in a PsO subgroup with clinical risk factors for progression to PsA compared to those PsO patients with no such risk factors. Patient centric biosample collection will be achieved using self-administered blood sampling kits. This will potentially allow us to validate any previously identified candidate genetic and molecular markers of PsA
With access to the largest collection of liquid and tissue (synovium and skin) biopsy samples from deeply phenotyped PsO cohorts in Europe, HIPPOCRATES seeks to identify and evaluate genetic and molecular markers of PsA. Cutting-edge analytical technologies will be used to provide new molecular data from genomics, epigenomic chromosome conformation signatures (EpiSwitch 3D-genetics platform), mass spectrometry-based unbiased discovery and targeted proteomics, affinity-based proteomics metabolomics and lipidomics with tissues being used for single cell analysis, including Cytometry by time of flight/Epigenetic Time of Flight. 30 The data obtained here will be analysed alongside similar data obtained using samples from PsO patients with early musculoskeletal symptoms and/or with subclinical imaging changes. Each of the data acquisition studies are being undertaken with careful and rigorous attention the to study design, including statistical considerations to minimize or account for pre-analytical variables and analytical biases. Efforts will also be made to link data from genome-wide association studies to assess the contribution of genetics to the circulating systemic proteome, metabolome and lipidome. Such analyses will provide opportunities to gain insight into disease mechanisms and/or a more informed use of serum/plasma molecular data.
For integration in a central database, data will be collated and cleaned to ensure that all of the collected data can be made interoperable and analysed in a standardized and reproducible way: European Health Data and Evidence Network (EHDEN). 2022; Available from: https://doi.org/10.3030/806968). The clinical data will be transformed according to the OMOP CDM format: Observational Medical Outcomes Partnership Common Data Model, an open community data standard designed to standardize the structure and content of observational data. 31 The biological samples will be standardized according to the Minimum Information About BIobank data Sharing (MIABIS) standard. 32 The central database platform will provide a unique resource to permit interrogation of aligned polyomic datasets with advanced AI/informatics analyses. By combining clinical and molecular data using ML/AI, HIPPOCRATES will establish whether combinations (signatures) of biomarkers can provide new clinically useful and much needed tests. Access to large numbers of patient cohorts within HIPPOCRATES and at associated centres internationally will support independent evaluation and validation of biomarker signatures associated with risk for the development of PsA in people with PsO.
Predicting joint damage
Structural joint damage in PsA is linked to reduced quality of life, physical function and increased risk of mortality.33,34 Erosive and osteoproliferative joint changes can occur rapidly with worse radiographic outcomes and functional decline if patients are not treated promptly and appropriately. 8 However, predicting patients at risk of irreversible joint damage is difficult, since in some patients, damage occurs early and progresses rapidly, whereas in others no damage is observed even after prolonged disease. In addition, it remains a challenge to detect joint damage at a very early stage of PsA, as the majority of PsA patients experience a considerable diagnostic delay ranging from months to years. 35 Therefore, identifying patients at increased risk for developing PsA and establishing reliable predictors of progression in patients with established PsA could lead to a more targeted treatment with better outcomes. Predictors of damage progression defined to date include relatively non-specific factors such as the number of painful and swollen joints or the burden of systemic inflammation measured by soluble markers such as CRP. 36 To date, however, there are no more specific and validated biomarkers or clinical algorithms that predict which patients with PsA will develop bone or joint damage and differentiate slow radiographic progressors from fast progressors.
The damage progression work within the IMI project HIPPOCRATES therefore focuses on identifying and validating clinical, imaging and molecular biomarkers of early joint damage. The goal is to find reliable markers for progression of damage to allow an earlier treatment initiation in PsA patients at high risk for poor outcomes in a targeted manner, thus preventing the development or worsening of structural damage. To achieve this, the HIPPOCRATES partners are focussing their research efforts on combining innovative imaging techniques with state-of-the-art multi-omic approaches on tissue (synovial, entheseal) and liquid biopsies from PsA patients participating in multiple cohorts throughout Europe and worldwide. Considering the extreme variability of the phenotype and endotype of PsA, it is unlikely that a single specific and universally accessible biomarker can be identified. Therefore, the development of new reliable diagnostic algorithms is being driven by implementing AI-based technologies to integrate multidimensional clinical, imaging and molecular data generated by genomics, transcriptomics, proteomics and metabolomic approaches into a single diagnostic model.
Modern imaging techniques such as ultrasound, MRI, positron emission tomography (PET) – computed tomography (CT) and high-resolution peripheral quantitative CT (HR-pQCT) are diagnostic tools with high resolution and accuracy for the assessment of synovio-entheseal damage and are major assets for the current diagnostic approach to psoriatic disease.37 –41 The first results of HIPPOCRATES in this field were promising, showing that these imaging technologies can be further developed42 –44 to allow assessment of damage progression in PsA at a new level. The use of molecular approaches for damage characterization in PsA is a growing area of research and one of the main focuses of HIPPOCRATES. To date, techniques such as transcriptomics, proteomics and metabolomics have been effective in finding single soluble markers or signatures that can identify PsA in selected cohorts. 45 Examples include CXC-L10 and secreted phosphoprotein 1 (SPP-1).12,46 Furthermore, novel techniques such as lipidomics have revealed specific signatures in PsO and PsA.18,47 Within HIPPOCRATES, several standardization projects have been started 48 and are underway to apply multi-omic approaches not only to provide the means for an earlier PsA diagnosis, but also to predict damage progression in established disease.
Improving our knowledge of the clinical, imaging and molecular basis of different stages of damage progression will enable us to have a deeper understanding of optimized targeted treatment strategies. In turn, the development of stage-specific biomarker signatures will undoubtedly move the treatment paradigm from a focus on established and progressive disease to one focused on much earlier intervention and possibly damage prevention.
Development of a precision medicine approach to treating PsA
Over the past two decades, there has been a revolution in the understanding of the pathophysiology of psoriatic disease, leading to the successful development of targeted therapies. There are now a wide range of effective therapeutic options available in the clinic for the treatment of PsA, including conventional synthetic Disease-Modifying AntiRheumatic Drugs (DMARDs) (in this case methotrexate), biologic cytokine inhibitors targeting Tumour Necrosis Factor (TNF) and the IL-23/IL-17 pathway and targeted synthetic DMARDs (in the form of Janus Kinase (JAK) inhibitors).49,50 While welcome, this plethora of options presents a dilemma for clinicians when choosing the best therapy for an individual with PsA, as treatment responses are not uniform between patients or even between domains in the same individual. A significant proportion of patients with PsA do not respond to the administered therapeutic agent. 3 Moreover, those that do respond often only have a partial response or a response that is limited to certain disease domains (e.g. skin but not joints). 51 Head-to-head studies in PsO indicate a clear hierarchy of efficacy in the skin (with IL-17 and IL-23 p19 inhibitors more effective than TNF inhibitors), 52 however, this is not the case in the musculoskeletal component of PsA. This aspect of disease in PsA is more heterogenous and treatment does not generally result in the same level of response as those seen in the skin. While there are some extra-articular features (such as the severity of psoriasis or presence of inflammatory bowel disease) that can help guide therapeutic choice, for most patients this largely remains an empirical trial-and-error process, driven by population-level data and guidelines, as well as cost considerations. This is a highly unsatisfactory situation for both patients and their treating clinicians. Thus, there is an urgent need for biomarkers to help stratify patients and inform clinical treatment decisions to increase the likelihood of response to a given therapeutic. Unfortunately, despite decades of research, particularly in RA, there are still no validated and clinically useful theragnostic biomarkers for PsA beyond clinical assessment and gestalt. While the reasons for this are multiple, we believe that a focus on single technologies for biomarker discovery in small cohorts without validation in independent cohorts is a major contributory factor, while advances in synovial tissue biomarkers have facilitated delivery of large biopsy-based randomized controlled trials. 53 Our proposal, as part of the HIPPOCRATES IMI consortium, is to combine robust, validated signatures identified by several technologies, and linked to clinical data, to better reflect the heterogenous mechanisms that are likely to underpin treatment response in PsA. 54
Our proposed work will leverage data from existing cohorts across the consortium partners to identify potential theragnostic biomarkers using several technologies, including genomics, epigenomics, proteomics and metabolomics (see Figure 1). The initial discovery research will focus on commonly used therapies and the extremes (e.g., remission versus primary non-response) of response in the musculoskeletal domain in PsA. These findings will then be extended to evaluate a wider range of therapeutic responses and outcome measures. It is likely that due to tissue-specific mechanisms, 54 identified biomarkers will not reflect responses in all domains, and thus the primary focus will be the musculoskeletal domain (joints, enthesitis and dactylitis), as this encompasses the most unmet clinical need in PsA. Importantly, access to a range of cohorts, including phase III studies from industry partners and observational real-world cohorts from clinical academic partners, will enable the validation of any potential biomarker signatures in independent patient groups and the assessment of how they perform in these settings and their likely utility in the clinic. In addition to evaluating these signatures individually, advanced analytical algorithms will be applied to combine datasets to enable the evaluation of mixed polyomic and clinical signatures for the generation of optimized biomarker signatures that can predict likely response or even non-response to a specific class of treatment. Any signatures identified by these processes should also be prospectively tested in future studies, to confirm their clinical utility, cost effectiveness and impact on clinical outcomes.
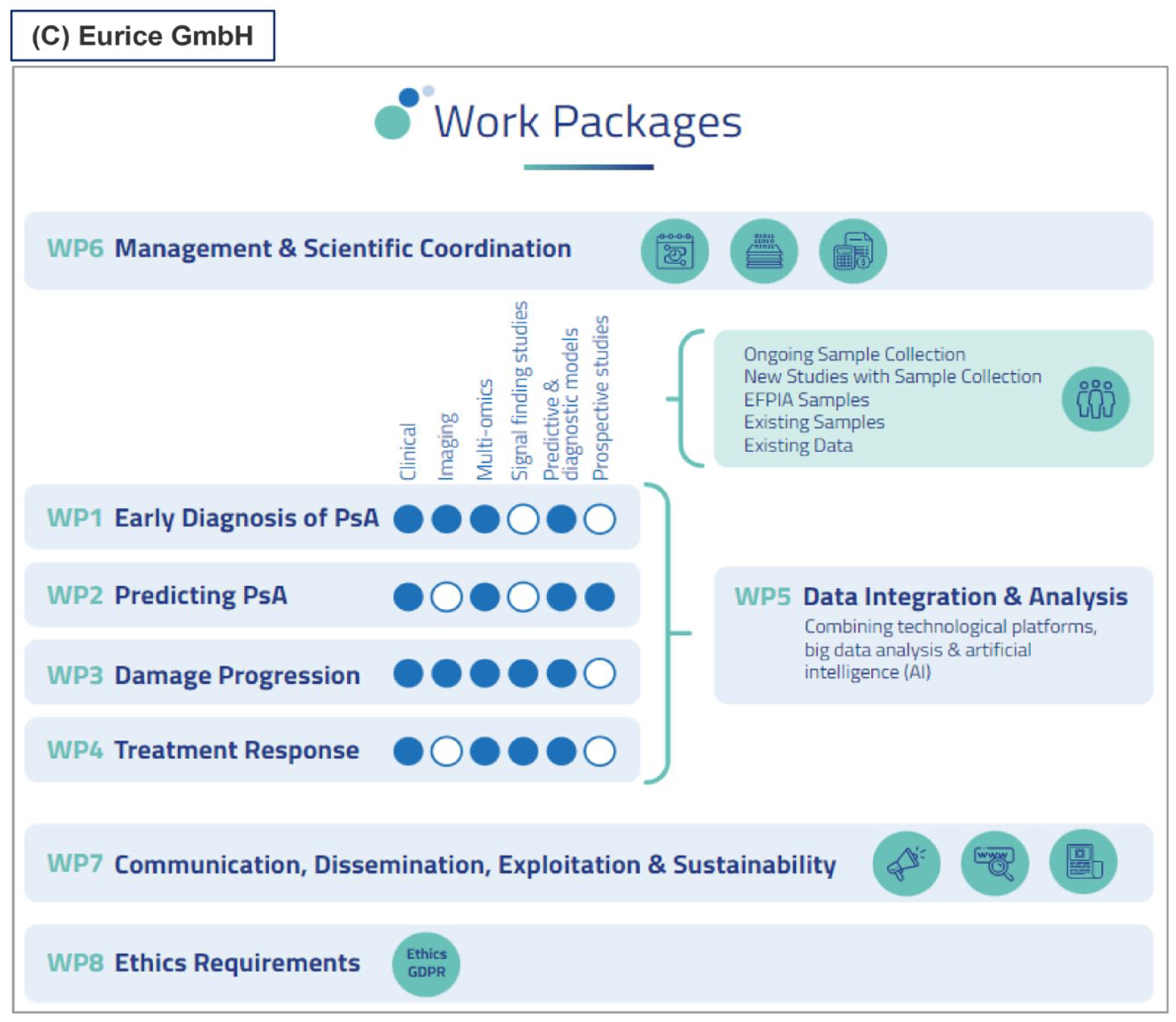
HIPPOCRATES project overview.
While individualized therapy for PsA remains unlikely in the foreseeable future, precision stratification and treatment of patients has the potential to deliver on the true promise of these and future therapies, to transform outcomes for people with PsA. Large collaborative consortia, bringing together key stakeholders, including patient partners, employing multi-omic approaches and sophisticated analytic algorithms in multiple clinically relevant patient cohorts offer the best opportunity to deliver this.
Collation of large datasets and potential for AI approaches
The full potential of AI can best be realized when we are able to train algorithms on large, structured databases. In the field of medical sciences, relying on smaller datasets from clinical studies can lead to suboptimal results. This issue can be addressed by introducing centralized medical databases. 55 This method offers the assessment of large sample sizes and many data points from different medical specialists. Furthermore, recording information digitally allows for a more standardized assessment than written medical reports. While in most countries, medical data cannot be collected centrally due to privacy laws, the HIPPOCRATES project is implementing an alternative solution: a comprehensive database including multiple international cohorts of patients with psoriasis and PsA who contributed data in the scope of clinical observational studies. The goal is to mimic the bandwidth of data that centralized medical databases offer. To achieve this goal, we are employing a standard data model, OMOP. 31 Recent trends are towards federated approaches to ML and AI to circumvent inherent issues preventing centralization, such as data ownership, privacy and security. Such approaches either use a central server for learning model parameters, or model parameters are learned through secure node to node communication as in Swarm learning. 56 Such approaches are extremely powerful for building AI models utilizing multiple data resources when ethical or legal considerations make it impossible for the data to be centralized. Indeed, for the future of clinical data research, especially for imaging data, 57 this approach holds much promise as it is scalable, limits data traffic and avoids having multiple copies of data in different locations. However, setting up such systems can be both technically challenging and costly.
For projects with multiple diverse datasets relevant to a specific disease area such as PsA, centralization has key advantages: First, data can be fully harmonized and quality controlled by specialists having access to the full dataset; second, AI models can be learned on different partitions of the full dataset without the complexity of learning across a network of smaller decentralized datasets; and third, since all data is on a single server, assessment of outliers and potential biases affecting the AI models can be easily addressed. A wealth of information including medical history, demographic data, omics and so on are assembled to create optimal preconditions for ML and AI applications. As pointed out, 43 a key to meaningful and generalizable results is to test results on subsets of the data that were excluded at the stage of training. Both subsets must be rich enough to represent the general population. The main goal is the development of innovative algorithms to predict disease progression and treatment response. Within these goals, different priorities can be set depending on the final application. One category of outcome are tools for supporting clinical decision-making as well as informing patients. Such tools should depend on easily obtainable data and be highly interpretable. By contrast, algorithms serving a more scientific investigation and deeper understanding of the disease itself can be more complex and should above all be highly accurate. The steps of building the dataset, developing intelligent methods and interpreting the results require close collaboration between interdisciplinary groups.
Discussion
In this review, we have focused on four areas of unmet need in PsA. These relate to identification of risk factors for PsA in people with PsO, the development of a diagnostic test or algorithm, the early identification of PsA patients likely to experience joint damage and the development of a precision medicine approach to treatment. All these areas are being addressed by the IMI-Industry funded HIPPOCRATES consortium, which comprises 27 European-based partners from academia, pharmaceutical industry, small-/medium-sized industries and patient-representative organizations. Other areas of unmet need which are not addressed in this review include 1 : agreement on which clinical outcome measures are best used in clinical trials and routine care settings to accurately reflect disease activity 2 ; the development of a molecular biomarker which reflects disease activity. This biomarker should have performance characteristics which are substantially superior to CRP. This is because in PsA, CRP is not elevated frequently in those who, on clinical grounds, appear to experience a considerable inflammatory burden3,58 with >50% of PsA patients having three or more features of the metabolic syndrome, 59 we need to better understand the two-way contribution of the cardio-metabolic dysfunction to PsO and PsA.
While each area of unmet need focus includes innovative approaches, there are several common threads throughout the work plan. Firstly, HIPPOCRATES has succeeded in involving key European investigators who have agreed to provide access to clinical and imaging data together with appropriate biosamples from some of the best-characterized and largest PsO and PsA cohorts in Europe. Data from these cohorts will be combined to provide sufficient patient numbers to allow us to address the key clinical needs at scale. Secondly, we primarily (but not exclusively) will work with liquid biopsy samples, serum, plasma and whole blood (DNA). While disease tissue samples may provide information on more disease-relevant markers and potentially identify new targets for therapy, 54 accessing such samples is invasive and the expertise required for obtaining such biopsies is not routinely available. Thus, to be applicable in routine care, the use of liquid biopsies is much more feasible. In addition, there are examples of successful multi-centre, biopsy-based clinical trials and consortia in RA53,60 and also a new US-based consortium, Accelerating Medicines Partnership® Autoimmune and Immune-Mediated Diseases (AMP® AIM); (The Foundation for the National Institutes of Health, North Bethesda, MD) which will focus on tissue-related studies in PsA, making the work of HIPPOCRATES nicely complementary. Thirdly, each of these liquid biopsy samples will be subjected to a range of omic platform analyses providing data which will be initially interrogated locally but will ultimately be combined and subjected to machine learning/AI approaches. 30 This will result in the identification of multi-omic biomarker panels or algorithms which can then be applied to other HIPPOCRATES cohorts for initial verification of findings. Ultimately, large-scale prospective validation studies will be required to introduce these panels into routine practice. Fourthly, and most importantly, there is embedded patient involvement in each of the areas of clinical need in HIPPOCRATES. Working together with other partners, the PRPs ensure that any proposals for change in clinical practice are likely to be both feasible and acceptable to patients. It will also be important to ensure that opinions from healthcare providers, both in primary and secondary care, are sought for and incorporated early in the process.
PRP involvement in HIPPOCRATES has been significant since the initial stages of the project, when the consortium was being set up and the research focus decided upon. PRPs are represented on the HIPPOCRATES executive team and on each of the work packages. In addition, work-package 7 which focuses on communication, dissemination, sustainability and exploitation is being co-led by a PRP. At a work-package level, PRP involvement has been particularly important in the design of the HPOS study where their input has ensured that the data collected are appropriate, non-repetitive and does not over-burden study participants.
The implementation of the ambition of HIPPOCRATES is likely to be met by several challenges. To share and combine the data, a robust data sharing agreement (DSA) needs to be developed and agreed among all the partners. This DSA needs to address European General Data Protection Regulation (GDPR) and any individual country GDPR variation. The DSA also needs to protect patient confidentiality (data protection) while at the same time allowing for patient-level data to be analysed and related to molecular data emerging from the detailed omic analysis of liquid biopsy samples (data utility). Once the DSA has been agreed, a glossary of terms used to describe the clinical features of PsA will have to be agreed to allow data to be transformed into a format which will permit combination with other similar datasets.
At first glance, exploring and combining data obtained from routinely collected biosamples does not seem overly problematic but what is critical is to ensure that any differences observed do not relate to differences in sample collection, storage, or processing. Each of these steps should be recorded in standard operating procedure documents and where there are differences, these then can be assessed to see if they have any impact on results obtained. To limit any variation which might occur because of differences in approaches to omic technologies, HIPPOCRATES took the decision to limit the technological evaluation to identified centres with a track record of expertise.
These are but some of the challenges which are being addressed by the HIPPOCRATES consortium, and more, no doubt, will emerge. As we move forward with the exciting prospect of engaging in collaborative research and work which could not be undertaken in single-centre studies, the focus needs to be set firmly on the goal of the research programme and the need to improve outcomes in PsA by identifying disease early and treating each patient based on their combined clinical and molecular profiles. For the first time, the HIPPOCRATES consortium presents an opportunity which really has the potential to change the management of PsA, to alleviate the disease burden and to reduce the wider societal and economic impact.
Footnotes
Appendix
List of HIPPOCRATES partners.

| No. | Shortened partner name for project | Partner |
|---|---|---|
| 1 | UCD | University College Dublin |
| 2 | UGLA | University of Glasgow |
| 3 | UERL | Universitätsklinikum Erlangen |
| 4 | VIB | VIB VZW |
| 5 | FhG-CIMD | Fraunhofer Gesellschaft Zur Foerderung Der Angewandten Forschung E.V. |
| 6 | KUL | Katholieke Universiteit Leuven |
| 7 | UMAN | The University of Manchester |
| 8 | UOXF | The Chancellor, Masters and Scholars of The University of Oxford |
| 9 | ATT | Atturos Limited |
| 10 | EURICE | EURICE European Research and Project Office GMBH |
| 11 | FCRB | Fundacio Clinic Per A La Recerca Biomedica |
| 12 | RH | Region Hovedstaden |
| 13 | UCSC | Universita Cattolica Del Sacro Cuore |
| 14 | GEU | Stichting Group For Research and Assessment of Psoriasis and Psoriatic Arthritis EU |
| 15 | EULAR | European League Against Rheumatism |
| 16 | SIB | SIB Institut Suisse De Bioinformatique |
| 17 | IHD | The European Institute for Innovation Through Health Data |
| 18 | AMC | Academisch Medisch Centrum Bij De Universiteit Van Amsterdam |
| 19 | OBIO | Oxford Biodynamics PLC |
| 20 | KTH | Kungliga Tekniska Hoegskolan |
| 21 | KCL | King’s College London |
| 22 |
|
|
| 23 | NVS | Novartis Pharma AG |
| 24 | UCB | UCB Biopharma |
| 25 | BMS | Bristol Myers Squibb Company Corp |
| 26 | PFIZER | Pfizer Limited |
| 27 | Trajan | Trajan Scientific and Medical |
| 28 | AbbVie | AbbVie Inc. |
Acknowledgements
None.
