Abstract
Enteric glia regulate gut functions in health and disease through diverse interactions with neurons and immune cells. Intracellular localization of traditional markers of enteric glia such as GFAP, s100b, and Sox10 makes them incompatible for studies that require antigen localization at the cell surface. Thus, new tools are needed for probing the heterogeneous roles of enteric glia at the protein, cell, and functional levels. Here we selected several cell surface antigens including Astrocyte Cell Surface Marker 2 (ACSA2), Cluster of differentiation 9 (CD9), lysophosphatidic acid receptor 1 (LPAR1), and Proteolipid protein 1 (PLP1) as potential markers of enteric glia. We tested their specificity for enteric glia using published single-cell/-nuclei and glia-specific translating mRNA enriched transcriptome datasets, immunolabeling, and flow cytometry. The data show that ACSA2 is a specific marker of mucosal and myenteric glia while other markers are suitable for identifying all subpopulations of enteric glia (LPAR1), glia and immune cells (CD9), or are not suitable for cell-surface labeling (PLP1). These new tools will be useful for future work focused on understanding specific glial functions in health and disease.
This study identifies astrocyte cell surface antigen 2 as a novel marker of myenteric glia in the intestine. This, in combination with other markers identified in this study, could be used for selective targeting of enteric glia.
Introduction
Enteric glia are non-myelinating peripheral glia intrinsic to the gastrointestinal tract that function to regulate gut homeostasis in health and disease (Grubisic & Gulbransen, 2017a; Seguella & Gulbransen, 2021). Enteric glia regulate essential gut functions such as gut motility and intestinal epithelial secretions through communication with the enteric neurons (Grubisic & Gulbransen, 2017b; McClain et al., 2014, 2015), and modulate the progress and outcomes of intestinal inflammation through interactions with enteric neurons (Brown et al., 2016) and immune cells (Chow et al., 2021; Grubisic et al., 2020). Enteric glia also contribute to the pathophysiology of common human conditions such as irritable bowel syndrome (IBS), inflammatory bowel disease (IBD), necrotizing enterocolitis (NEC), and chronic intestinal pseudo-obstruction (Delvalle et al., 2018; Grubisic et al., 2020; Kovler et al., 2021; Ahmadzai et al., 2022). Understanding the specific functions of enteric glia in health and disease is, therefore, important to identify new therapies for gastrointestinal disorders.
Despite some shared morphological and transcriptional characteristics with the glial cells in the central nervous system (CNS), enteric glia are unique in terms of their developmental origin, local environment, gene expression, and function (Grubisic & Gulbransen, 2017a). However, the characteristics of enteric glia in health and disease are only beginning to be understood. Current single-cell/-nuclei sequencing data suggest extensive complexity in glial functions and glial heterogeneity (Drokhlyansky et al., 2020; Progatzky et al., 2021; Zeisel et al., 2018). The ability to conduct glial sorting for subsequent cellular profiling currently relies on expressing fluorescent reporters in glia, which complicates sorting glia in other transgenic lines and more complex disease models. Traditional markers of enteric glia including GFAP, Sox10, and s100b are intracellular proteins that are not suitable for studies where cell surface markers are required such as flow cytometry and fluorescence-activated cell sorting. In the CNS, this issue has been addressed by identifying Astrocyte Cell Surface Marker 2 (ACSA2) as a promising cell surface marker that specifically recognizes the plasma membrane marker ATPase Na+/K+ Transporting Subunit Beta 2 (ATP1B2) expressed by astrocytes (Batiuk et al., 2020). However, whether ACSA2 is expressed by enteric glia or if other cell surface markers are more suitable is unknown.
Here we characterize several potential surface markers of enteric glia using RNA expression datasets, immunohistochemistry, and flow cytometry. Based on these studies, ACSA2 and lysophosphatidic acid receptor 1 (LPAR1) emerged as good candidates for cell surface markers of enteric glia. LPAR1 in the intestines is expressed by all subpopulations of enteric glia, but its transcription is reduced in acute intestinal inflammation. ACSA2 has stable transcription in health and inflammation and is more selective as a marker of mucosal and myenteric glial cells.
Materials and Methods
Animals
All experimental protocols were approved by the Institutional Animal Care and Use Committees (IACUC) at Michigan State University (MSU), USA. Mice were maintained in a temperature-controlled environment (Innovive, San Diego, CA; Innocage system with ALPHA-dri® bedding) on a 12-h light: dark cycle with access to acidified water and food ad libitum. Transgenic mice were generated on the C57Bl/6 genetic background. Sox10CreERT2 transgenic mice were a gift from Dr. Vassilis Pachnis (The Francis Crick Institute, London, England) (Laranjeira et al., 2011) and have been validated as a driver line for enteric glia in prior work (Delvalle et al., 2018; Grubisic & Gulbransen, 2017b; McClain & Gulbransen, 2017). Transgenic mice expressing tdTomato in enteric glia (hereafter referred to as Sox10CreERT2; tdTomoto) were generated in-house by crossing Sox10CreERT2+/− mice with B6.Cg-Gt(ROSA)26Sortm14(CAG−tdTomato)Hze/J (Jackson Laboratory stock# 007914; RRID: IMSR_JAX:007914) or B6;129S6-Polr2aTn(pb−CAG−GCaMP5g,−tdTomato)Tvrd/J (Jackson Laboratory stock# 024477; RRID: IMSR_JAX:024477). Cre recombinase activity was induced by feeding animals tamoxifen citrate in the chow (400 mg/kg) for 2 weeks before experiments. All mice (including controls) were fed with a tamoxifen diet for the same duration to control for the potential effects of tamoxifen. Genotyping was performed by Transnetyx, Inc (Cordova, TN).
Whole-Mount Immunohistochemistry (IHC)
Whole-mount preparations of mucosal and myenteric plexus-longitudinal muscle (LMMP) were prepared by microdissection from mouse distal colon fixed overnight in Zamboni's fixative or 4% paraformaldehyde at 4°C. Immunolabeling was conducted as previously described (Grubisic et al., 2020) with antibodies summarized in Table 1. Whole mounts were rinsed three times (10 min each) in phosphate-buffered saline (PBS) or PBS containing 0.1% Triton X-100 (PBST) followed by a 30 min incubation in blocking solution [containing 4% normal goat serum (or 5% normal donkey serum), 0.1 or 0.4% Triton X-100 and 1% bovine serum albumin]. Primary antibodies were applied overnight at room temperature (RT) before being rinsed three times with PBS. Secondary antibodies were applied for 2 h at RT. Whole mounts were rinsed with 0.1 M phosphate buffer and mounted on slides with bicarbonate-buffered glycerol (consisting of a 1:3 mixture of 142.8 mM sodium bicarbonate and 56.6 mM carbonate to glycerol).
Details of Primary and Secondary Antibodies Used in This Study.
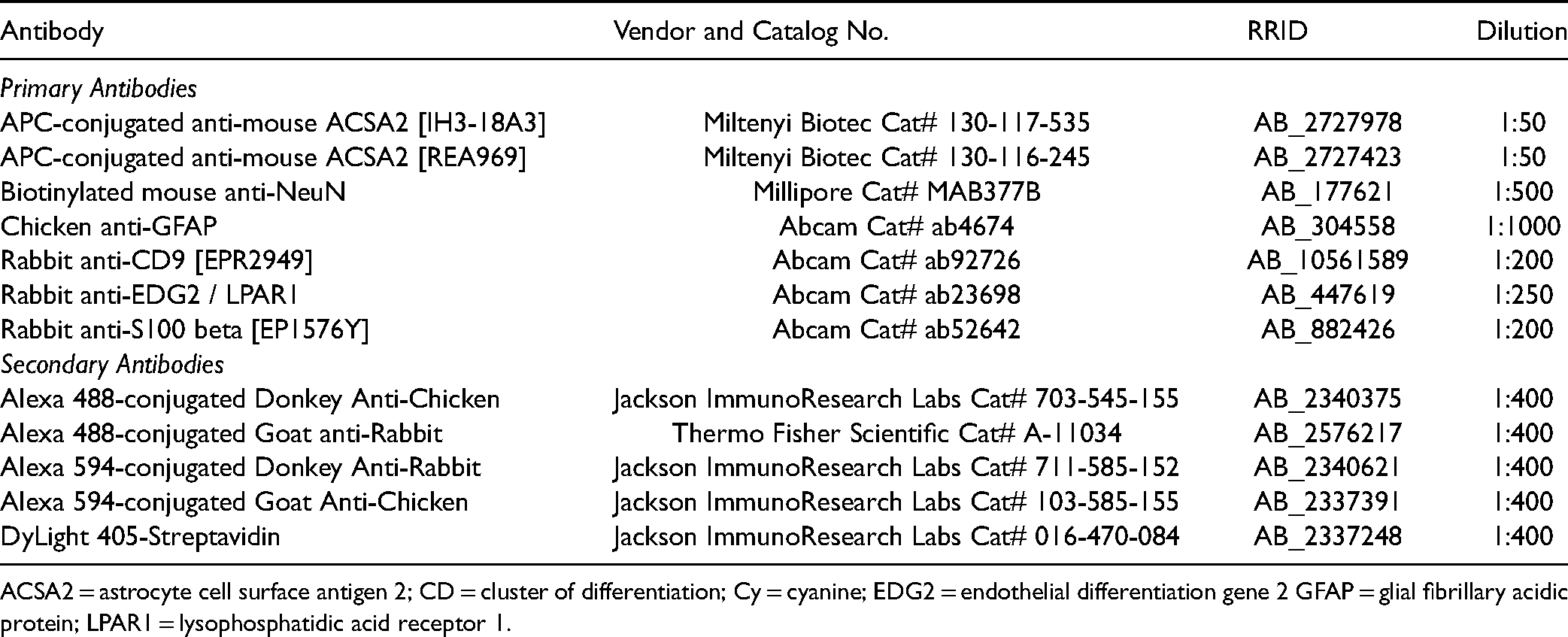
ACSA2 = astrocyte cell surface antigen 2; CD = cluster of differentiation; Cy = cyanine; EDG2 = endothelial differentiation gene 2 GFAP = glial fibrillary acidic protein; LPAR1 = lysophosphatidic acid receptor 1.
Free-Floating Brain Slices IHC
Mouse brains were fixed by immersion in 4% paraformaldehyde with continuous shaking for 3 days at room temperature. The brains were then rinsed 3 times in PBS and immersed in 30% sucrose (dissolved in water) for several days at 4°C (until sunken to the bottom of the 15 ml tubes). At this juncture, 35 µm brain slices were prepared and stored in PBS at 4°C. All IHC steps were performed at room temperature. Following three 15-min washes in PBS containing 0.1% Triton X-100 (PBST) and an hour incubation in blocking solution (4% normal donkey serum, 0.1% Triton X-100, and 1% bovine serum albumin), the slices were incubated with primary antibodies overnight. Then, the slices were rinsed 3 times with PBST for 15 min and immersed with secondary antibodies for 2 h. Lastly, the slices were washed twice in PBST and twice in PBS, mounted on slides with Fluoromount-G® (SouthernBiotech, 0100-01), and imaged.
Image Acquisition and Processing
Epifluorescence images were acquired through the 10x [Plan, 0.25 numerical aperture (n.a.)], 20x (PlanApo, 0.75 n.a.), and 40x (PlanFluor, 0.75 n.a.) objectives of an upright epifluorescence microscope (Nikon Eclipse Ni, Melville, NY) with a Retiga 2000R camera (QImaging, Surrey, BC, Canada) controlled by QCapture Pro 7.0 (QImaging). Confocal images were acquired through the 40x (UPLFL, 1.3 n.a.) oil immersion objective of an inverted Olympus Fluoview FV1000 microscope (Olympus, Center Valley, PA). Images were processed by MetaMorph 7.0 software (Molecular Devices, LLC, San Jose, CA, United States).
Isolation of Myenteric plexus Cells
Whole-mount sheets of myenteric plexus from the mouse colon were prepared as previously described (Grubisic et al., 2020). Briefly, colons were flushed with ice-cold DMEM/Nutrient Mixture F-12 (Life Technologies) supplemented with L-glutamine and HEPES, the remaining mesentery was removed, and colons were cut in half to yield two 3–4 cm segments. The segments were individually placed on plastic rods (≈ 2 mm in diameter) and cotton swabs wetted with ice-cold DMEM were used to remove the myenteric plexus and longitudinal muscle layers. Preparations were rinsed twice with HBSS substituted with HEPES (10 mM) and digested in 2.5 ml of HBSS/HEPES buffer substituted with Liberase TM (0.13 Wünsch units/ml) and DNAse I (100 Kuntz units/ml) for 30 min at 37°C with occasional mixing. A gentleMACS Dissociator and gentleMACS C Tubes were used for cell dissociation. The enzyme reaction was stopped by an ice-cold complete medium composed of DMEM/Nutrient Mixture F-12 supplemented with L-glutamine and HEPES, penicillin, and streptomycin (100 U/mL and 100 µg/mL), and 10% fetal bovine serum (FBS, Denville Scientific, Inc). Cell suspensions were sequentially filtered through 100 µm and 40 µm cell strainers before flow cytometry.
Flow Cytometry
Filtered cell suspensions were spun down (10 min at 350 × g and 4°C), resuspended, and aliquoted in 1.5 ml centrifuge tubes containing 1 ml of flow cytometry buffer (FCB) containing 1% bovine serum albumin in PBS. Cells were resuspended in 100 µl FCB containing TruStain FcX™ (anti-mouse CD16/32, dilution1:50) for 15 min on ice, and then APC-conjugated anti-ACSA2 antibody was added and incubated for 15 min at 4°C. Cells were washed twice with FCB and viability dye 4′,6-diamidino-2-phenylindole (DAPI, final concentration 1 µM) was added 5 min before analysis on a BD Influx [Becton Dickinson (BD) Biosciences, Franklin Lakes, NJ] run by BD Software, version 1.0.0.650. Unstained, single stained, and fluorescence minus one stained controls were used for proper gating. Post-hoc analysis was performed using FCS Express 7 Research Edition (Win64) v7.01.0018.
Statistics
Data were analyzed by GraphPad Prism 9.1.2 (GraphPad Software, San Diego, CA). Normality was tested by the Shapiro-Wilk test with P < 0.01 as a statistically significant difference. We used multiple unpaired t tests or Mann-Whitney U tests setting a level of P < 0.05 as the cutoff for statistical significance.
Results
Astrocyte Cell Surface Antigen 2 (ACSA2), Cluster of Differentiation 9 (CD9), Lysophosphatidic acid receptor 1 (LPAR1), and ProteoLipid Protein 1 (PLP1) were selected as candidates for cell surface markers of enteric glia based on available genomic data showing that enteric glia express Cd9 (Delvalle et al., 2018) and Plp1 (Rao et al., 2015), functional data showing that enteric glia respond to LPA (Segura et al., 2004; Ahmadzai et al., 2022), and their known similarities with astrocytes which are labeled by ACSA2 (Figure 1) (Batiuk et al., 2017). We began by screening enteric glia-specific transcript expression of the genes encoding these proteins using publicly available data sets at a single-cell/-nuclei resolution (Drokhlyansky et al., 2020; Zeisel et al., 2018) and glia-specific translating mRNA expression in healthy intestines and after dinitrobenzene sulfonic acid (DNBS)-induced colitis (Delvalle et al., 2018). Sox10 was used as a control based on its known expression in adult enteric glia. The Atp1b2 gene was used to assess potential ACSA2 expression because suitable ACSA2 antibodies target the ATPase Na+/K+ Transporting Subunit Beta 2 (ATP1B2) (Batiuk et al., 2017). Gene expression of Sox10, Atp1b2, Cd9, Lpar1, and Plp1 were observed in the enteric nervous system (ENS) of the small intestine and colon in all data sets (Figure 2, S1-2). Single-cell/-nuclei transcriptional data show that Sox10, Lpar1, and Plp1 have significantly higher expression in glia than neurons (P < 0.05, unpaired t or Mann-Whitney U tests) (Figure 2A-C), and neuronal expression of Sox10 and Plp1 is virtually zero in datasets where single nuclei sequencing was used (Figure 2A-B). Cd9 is highly expressed in the ENS by neurons and glia with significantly higher neuronal expression observed in the large intestine [1.05 to 9.5 neuronal interquartile range (IQR) versus 0.65 to 0.88 IQR (neurons vs. glia), P = 0.011, Mann-Whitney U test] (Figure 2B). Atp1b2 expression was also detected at low levels in both neurons and glia (Figure 2A-C); however, some enrichment may be present in glia (small intestine samples; 0.00 vs. 0.015 medians, neurons vs. glia, P = 0.055, Mann-Whitney U test) (Figure 2A). Data obtained by bulk sequencing of translating glial mRNA detected Sox10, Atp1b2, Cd9, Lpar1, and Plp1 with Cd9 exhibiting the highest expression levels (Figure 2D). Comparisons of expression in samples from healthy and inflamed mice suggested that expression levels of Cd9 and Lpar1 are modulated during acute colitis (P = 0.039 and 0.021, unpaired t-tests, Figure 2D) with lessor observed effects on Sox10, Atp1b2, and Plp1. Recent single-cell transcriptomics of mouse large intestine (Progatzky et al., 2021) showed that Sox10, Atp1b2, Cd9, Lpar1, and Plp1 are expressed by enteric glia of naïve mice and in an animal model of parasitic infection (Figure S3).
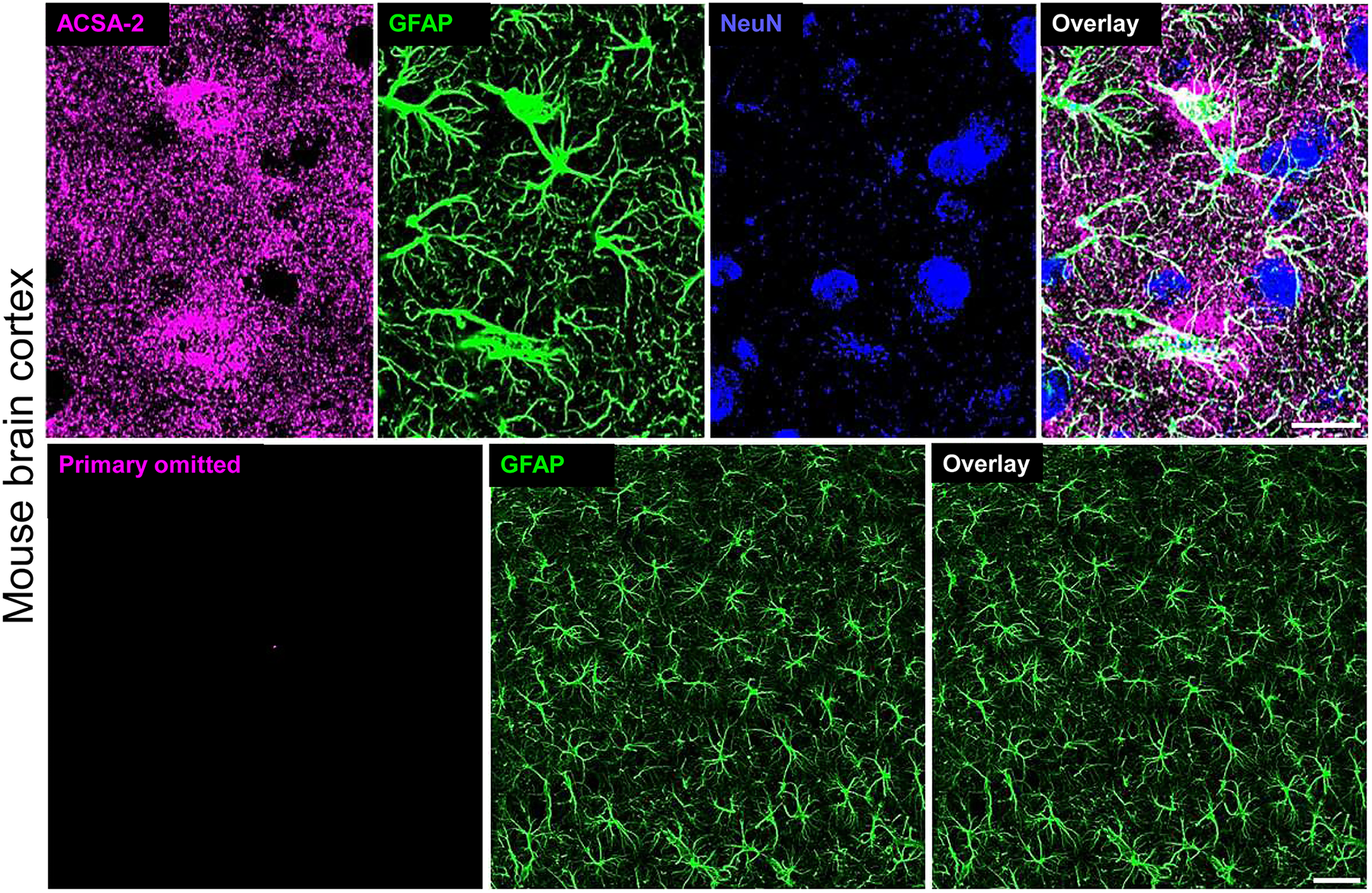
Characterization of the anti-ACSA2 antibody. Mouse brain slices were co-stained (top row) with ACSA2-APC (magenta), the glial marker GFAP (Green), and the neuronal marker NeuN (blue) or stained with only glial marker GFAP (bottom row). Scale bars, 20 and 40 µm (top and bottom).
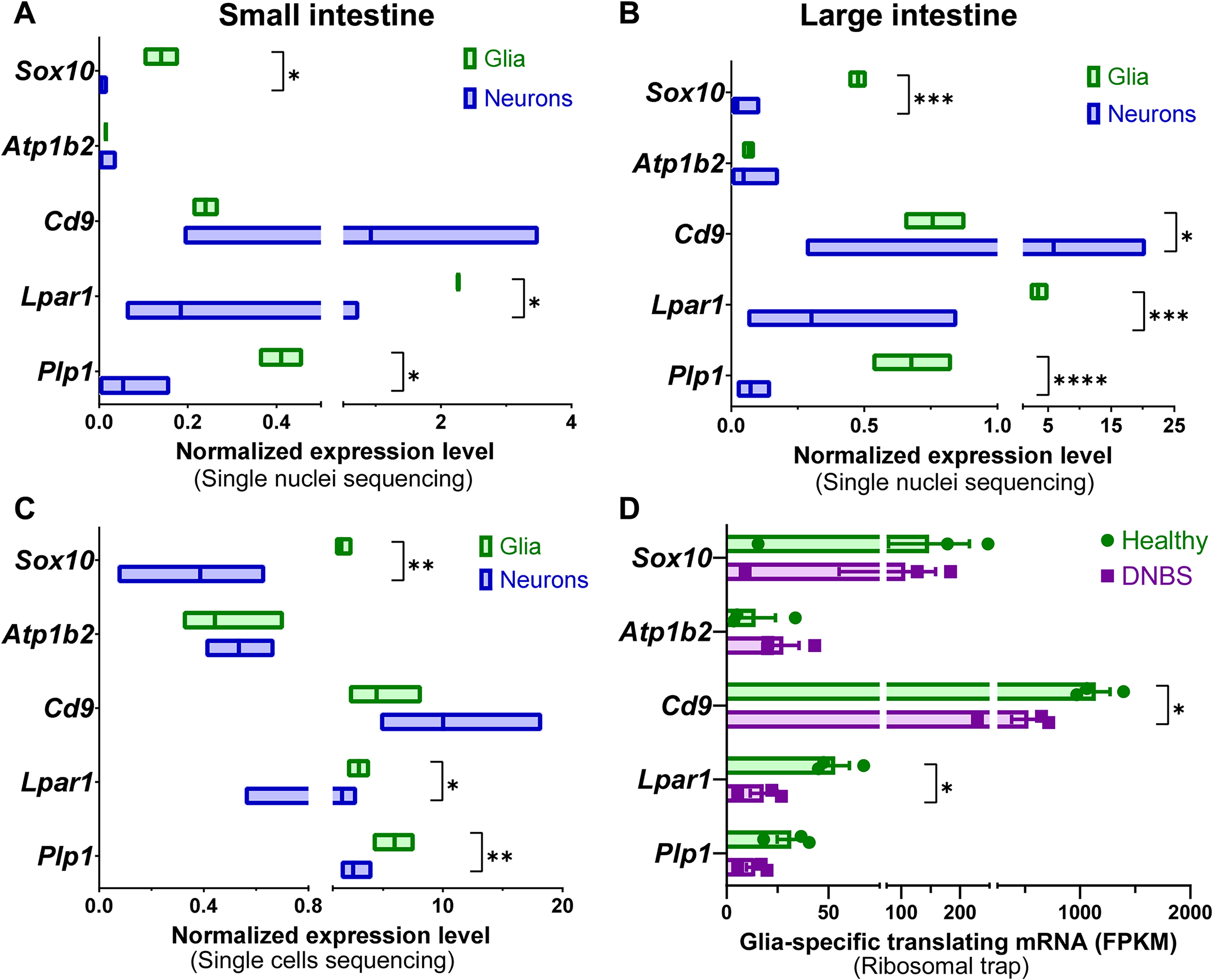
Cell-specific transcript expression of potential surface antigens. Graphs are prepared from published data sets using single nuclei (A-B), single-cell (C), and translating glial mRNA (D) transcriptomics from mouse small (A, C) and large (B, D) intestines. A-B) Single nuclei transcriptome from mouse small (A) and large (B) intestines (Drokhlyansky et al., 2020). Data show normalized expression levels (median ± range) of combined neuronal or glial subpopulations (more detailed summary in Figure S1). *, P < 0.05, ***, P < 0.001, ****, P < 0.0001, multiple Mann-Whitney U tests. C) Single-cell sequencing from adolescent mouse small intestine (Zeisel et al., 2018). Data show normalized expression levels (median ± range) of combined neuronal or glial subpopulations (more detailed summary in Figure S2). *, P < 0.05, **, P < 0.01, multiple Mann-Whitney U tests. D) Ribosomes were isolated from dinitrobenzene sulfonic acid (DNBS) treated glial RiboTag mouse colons (Sox10CreERT2; Rpl22fl/fl) and their healthy controls to perform sequencing of the translating glial mRNA (Delvalle et al., 2018). FPKM, fragments per kilobase per million mapped fragments. *, P < 0.05, multiple unpaired t tests.
The genomic data above show that genes for each of the candidate cell surface markers are expressed by enteric glia at varying levels and with varying overlap with expression in neurons. To assess whether the genomic data reflect protein expression, we performed immunolabeling in whole-mount preparations of the myenteric plexus and mucosa from the mouse colon. Whole-mount preparations were labeled with antibodies against the candidate cell surface markers and co-labeled with GFAP or s100b as known markers of enteric glia (Figures 3 and 4). Positive glial labeling was observed with antibodies directed against ACSA2, LPAR1, and CD9 at the level of the myenteric plexus (Figure 3). Labeling with the ACSA2 and LPAR1 antibodies was specific for enteric glia while labeling for CD9 localized to glia and some presumed immune cells. Labeling for ACSA2 was confined to intra- and inter-ganglionic glial cells within and between the myenteric ganglia while labeling for LPAR1 detected both myenteric and intramuscular glia. Labeling with antibodies against PLP1 did not detect enteric glia despite producing good labeling of oligodendrocytes in the brain (Figure S4). This result is expected because enteric glia are non-myelinating glia and do not express the full PLP1 protein.
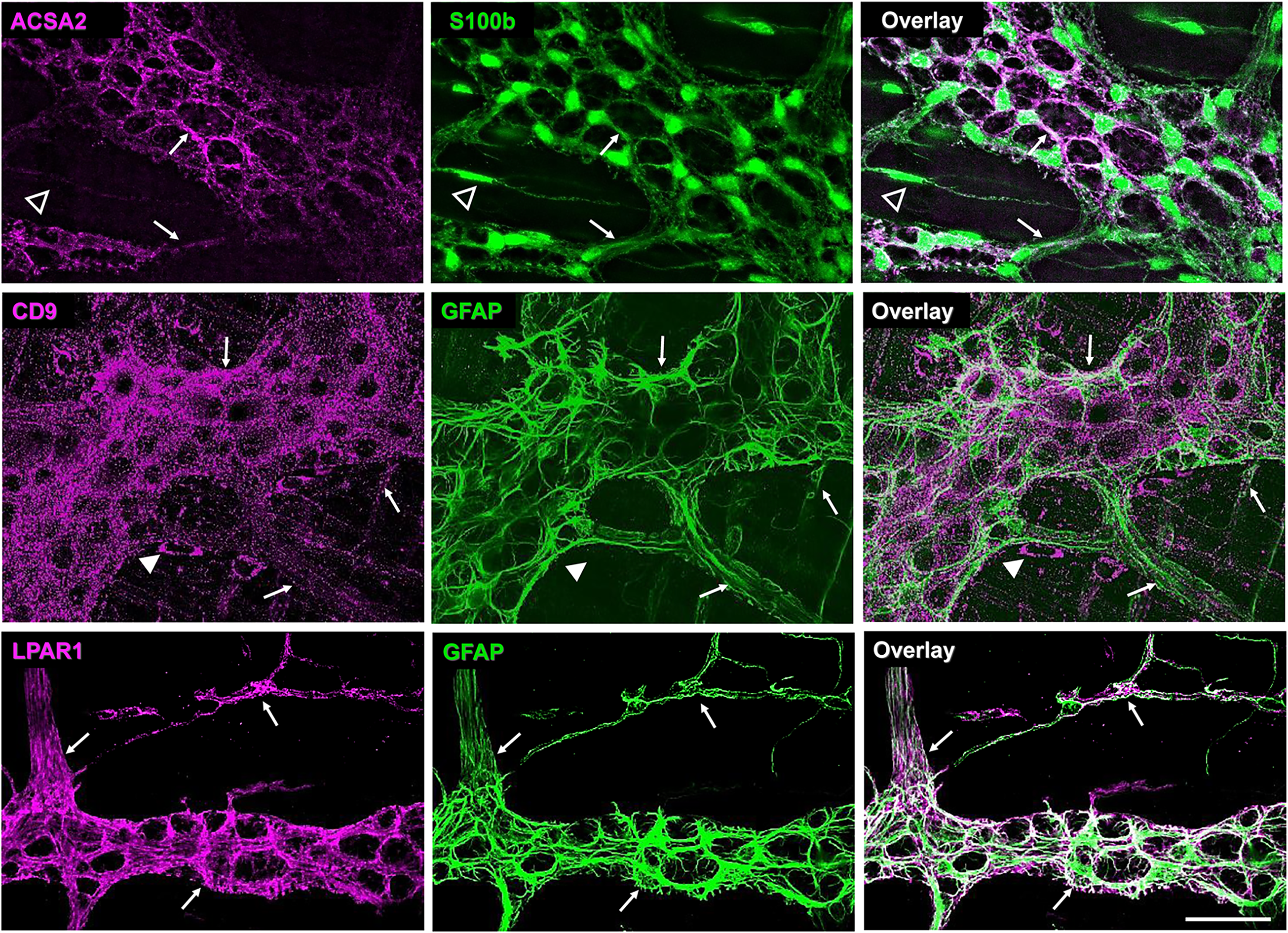
Antigen expression of ACSA2, CD9, and LPAR1 in the mouse colon muscularis. Whole mounts of colon longitudinal muscle-myenteric plexus preparations were stained with one of the anti-antigen antibodies (left column, magenta) and one of the glial markers anti-GFAP or anti-s100b (middle column, green). Overlay images are in the right column. Arrows, closed arrowheads, and open arrowheads point to the co-expressed immunoreactivity (ir), only antigen ir, and only glial marker ir, respectively. Note that ACSA2 ir colocalizes with inta- and inter-ganglionic glia, while CD9 and LPAR1 antibodies label intraganglionic and extraganglionic glia. CD9 antibodies also label GFAP- cells outside the myenteric plexus. Scale bar = 50 µm.
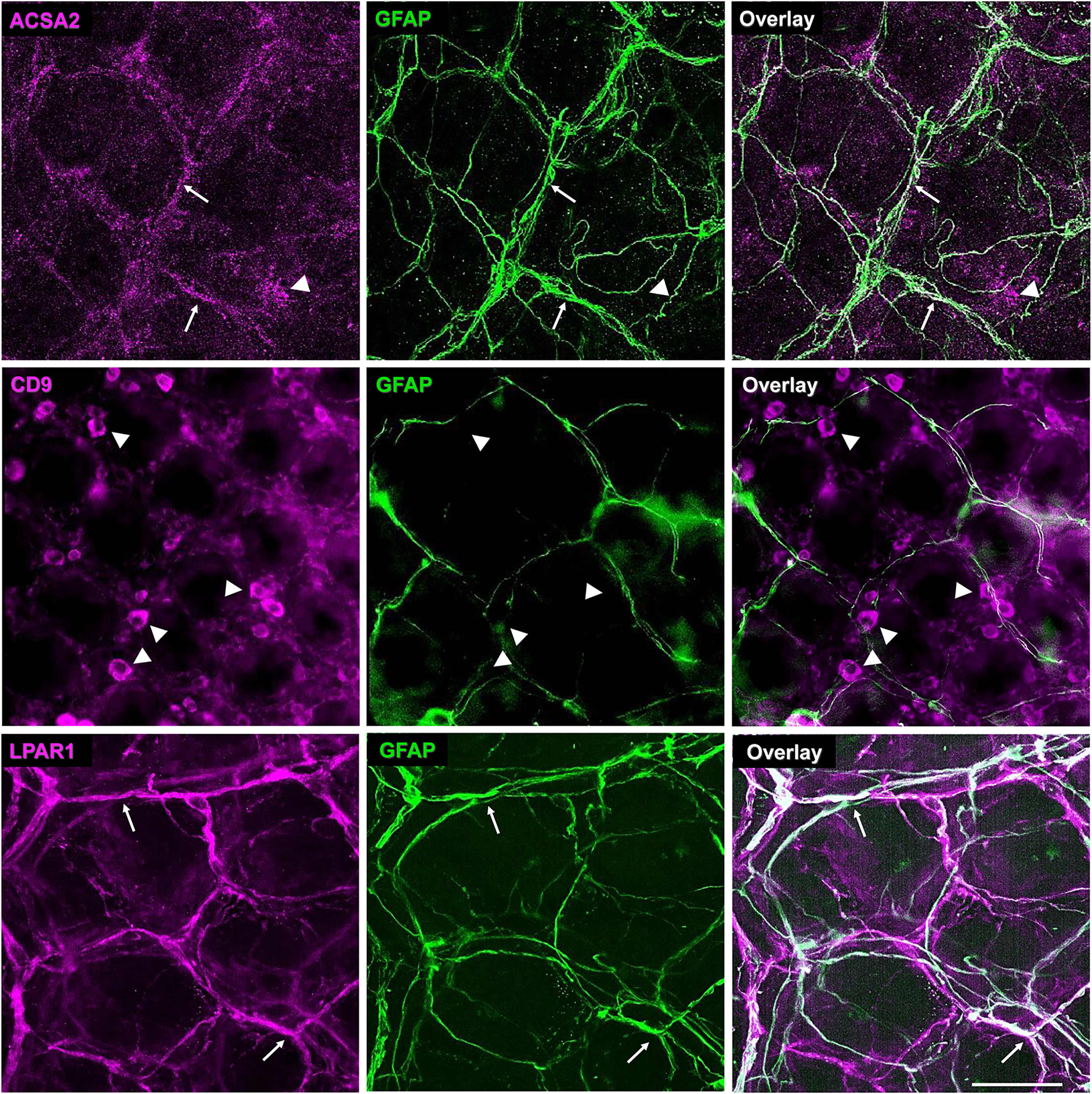
The antigen expression in the mouse colon mucosa. Whole mounts of colon mucosa were stained with one of the anti-antigen antibodies (left column, magenta) and the glial marker anti-GFAP (middle column, green). Overlay images are in the right column. Arrows and closed arrowheads point to the co-expressed ir and only antigen ir, respectively. Note that ACSA2 and LPAR1 ir co-localize with mucosal glia, while CD9 antibody labels GFAP- cells; ACSA2 antibody also labels GFAP- structures due to non-antigen specific reactions (see Figure S4 showing staining with the isotype control). Scale bar = 50 µm.
Labeling in whole-mount preparations of the colonic mucosa showed colocalization between immunoreactivities for LPAR1 and GFAP (Figure 4). ACSA2 immunoreactivity in the mucosa also exhibited co-localization with GFAP but additionally stains some structures that were not GFAP positive and did not resemble the morphology of mucosal glia. These GFAP-negative structures were also labeled by an APC-conjugated isotype control (Figure S5) suggesting that ACSA2 is also specific for mucosal glial cells and that labeling in the other structures was due to non-specific binding of the antibody. Labeling for CD9 in the mucosa produced good labeling of immune cells with no observed co-localization with GFAP. Overall, the rank order of the antibodies tested in the colonic mucosa is anti-LPAR1 > anti-ACSA2 >>> anti-CD9 in terms of glial specificity (Figure 4).
Low gene expression of Atp1b2 in the mouse colon (Figure 2B and D) corresponded with low ACSA2 immunoreactivity in the myenteric plexus (Figure 3), while CD9 and LPAR1 exhibited both high transcript abundance (Figure 2B and D) and high antibody immunoreactivity (Figure 3). At this juncture, it is unclear if the high CD9 and LPAR1 immunoreactivity within the myenteric ganglia also comes from myenteric neurons. Thus, we investigated the co-localization of these antigens with the glial and neuronal markers GFAP and HuC/D (Figure 5). We found that the antigens only colocalize with the glial marker indicating glial specificity of CD9 and LPAR1 antibodies within the myenteric plexus.
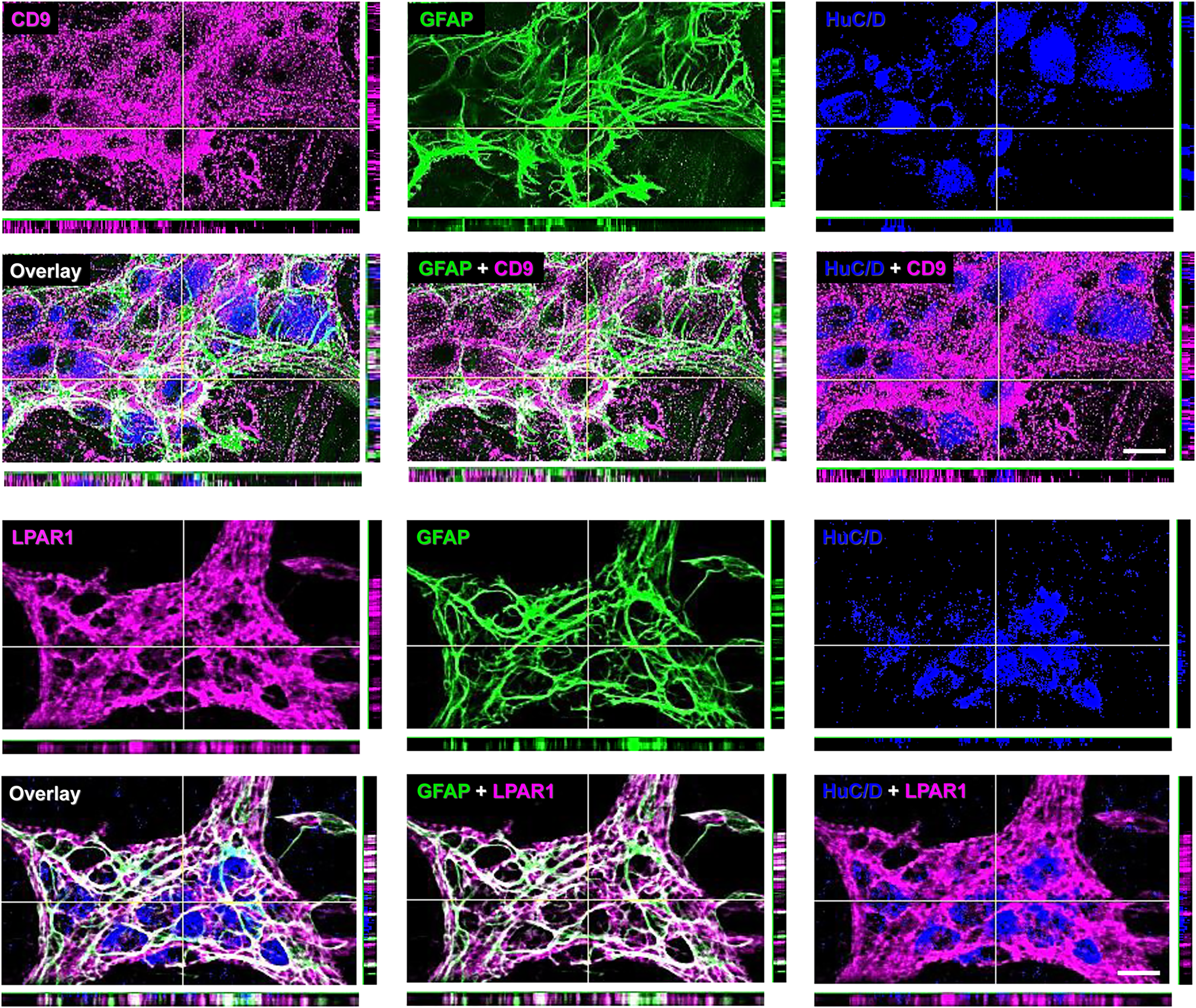
Colocalization of CD9 and LPAR1 immunoreactivity with glial and neuronal markers within myenteric ganglia. Whole mounts of colon longitudinal muscle-myenteric plexus preparations were stained with CD9 (top panels) or LPAR1 (bottom panels) antibodies and co-stained with the anti-GFAP glial marker (middle column) and anti-Hu neuronal marker (right column). Overlay images are in the second row of each panel. Optical sections are 5 µm thick. Vertical and horizontal yellow lines in z-projections (large images) mark the optical sections of x- and y-projections placed adjacently at the bottom and the right side of the respective images. Note that CD9 and LPAR1 colocalize with GFAP but not HuC/D. Scale bars, 20 µm.
The data to this point show that LPAR1 exhibits the best glial specificity in terms of RNA and protein expression but exhibited reduced transcription during acute intestinal inflammation. ACSA2 showed stable transcript expression during inflammation and protein expression is preferentially expressed by mucosal and myenteric glia. We were intrigued by the relative selectivity of ACSA2 as a marker of myenteric glia and further investigated the antibody specificity using flow cytometry. We used flow cytometry on cells suspensions derived from the colon muscularis of Sox10CreERT2; tdTomato mice (Figure 6) in which tdTomato expression is present in virtually all enteric glia of adult mice. After gating for single live cells (Figure 6A), we found that ACSA2 stained 12–16% (interquartile range, IQR), and tdTomato is expressed in 32–36% (IQR) of the gated cells (Figure 6B). More importantly, 96 ± 4% (mean ± SD) of the ACSA2+ cells also expressed tdTomato indicating that ACSA2 exclusively labels glial cells (P = 0.07, one-sample t-test from theoretical mean 100, N = 5 mice). To extrapolate the relative size of the ACSA2+ glial subpopulation, we normalized raw data to the maximal tdTomato+ population (Figure 6C). We found that 33–45% (IQR) of tdTomato+ cells are co-labeled by ACSA2 antibody while 60–72% (IQR) of the tdTomato+ cells remain unstained. This outcome indicates that only a subpopulation of glial cells in the muscular layers express ACSA2 which is consistent with the immunolabeling data showing ACSA2 labeling only in glia within myenteric ganglia (Figure 3). Together, these data suggest that ACSA2 is a suitable cell surface marker of a subpopulation of enteric glia.
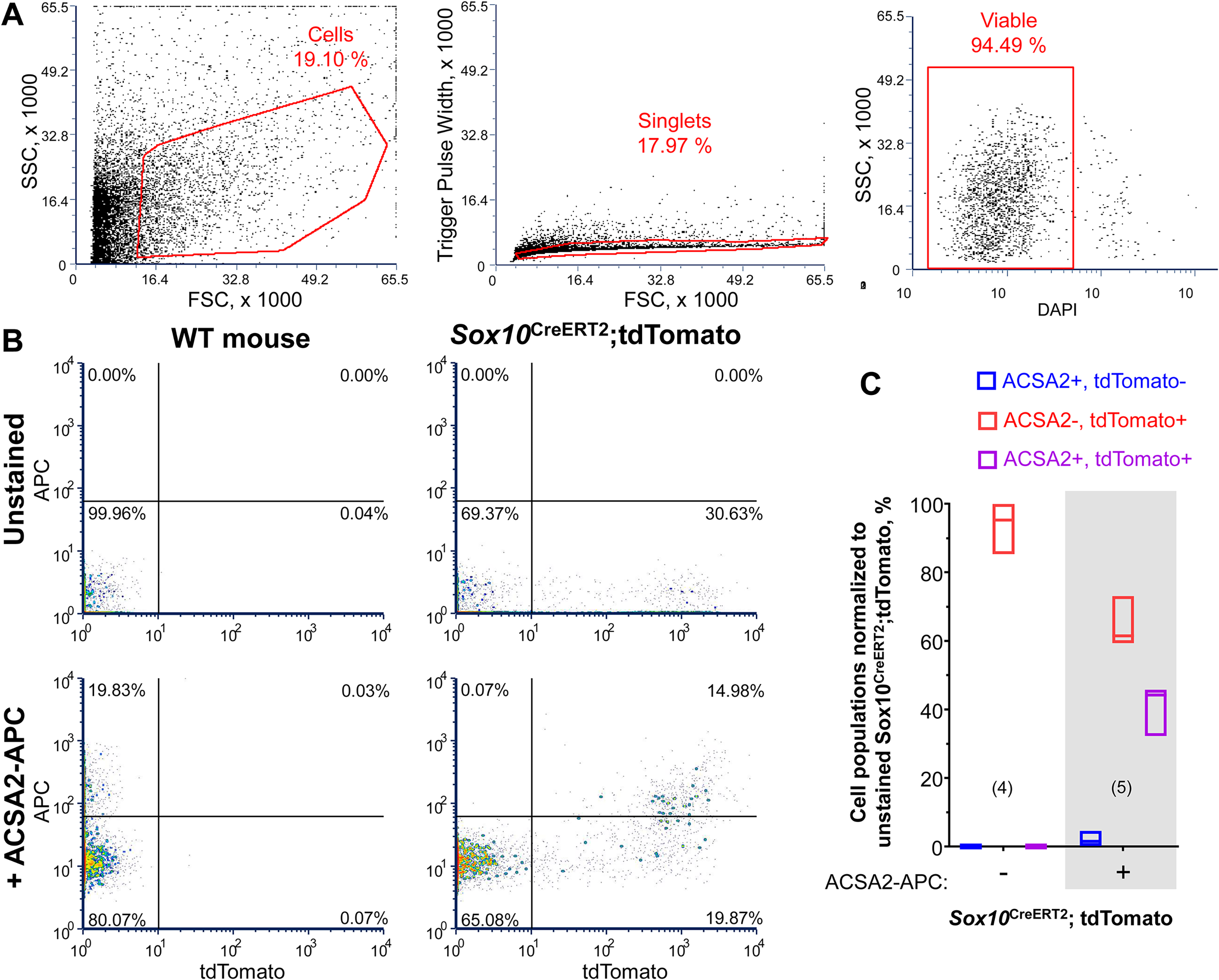
Flow cytometry using the anti-ACSA2 antibody. Live mouse longitudinal muscle-myenteric plexus cell suspensions were stained with APC-conjugated anti-ACSA2 antibody while DAPI was used as a viability dye. (A) Gating strategy for cells (left), single events (middle), and live cells (right). (B) APC vs tdTomato fluorescence from the unstained (top) and stained (bottom) cell suspensions derived from WT (left) and Sox10CreERT2; tdTomato (right) mice. The gating was adjusted using fluorescence minus one controls. (C) Summary graph showing unstained (left, unshaded) and APC-ACSA2 stained (right, shaded) cells from the Sox10CreERT2; tdTomoto mice that were normalized to the maximal tdTomato+ population. The graph shows three groups of cells: ACSA2+ tdTomato- (blue), ACSA2- tdTomato+ (red), and ASCA-2+ tdTomato+ cells (purple). Note that ACSA2 exclusively labels about 40% of the tdTomato+ cells. N = 4–5 mice.
Discussion
In this study, we characterized several potential cell surface markers of enteric glia (Table 2). LPAR1 labels all subpopulations of enteric glia in the large intestine, but its transcription is reduced in acute intestinal inflammation. In contrast, the gene encoding ACSA2 has stable transcription during inflammation and ACSA2 labeling identifies a subpopulation of myenteric glia and mucosal glia. CD9 is highly expressed by myenteric glia, but its expression is not glial-specific and is also modulated by inflammation. The Plp1 gene is broadly expressed by enteric glia and seems relatively stable in inflammation, but it does not appear that enteric glia express the full-length protein.
RNA and Protein Expression Stability and Specificity of the Selected Surface Antigens.
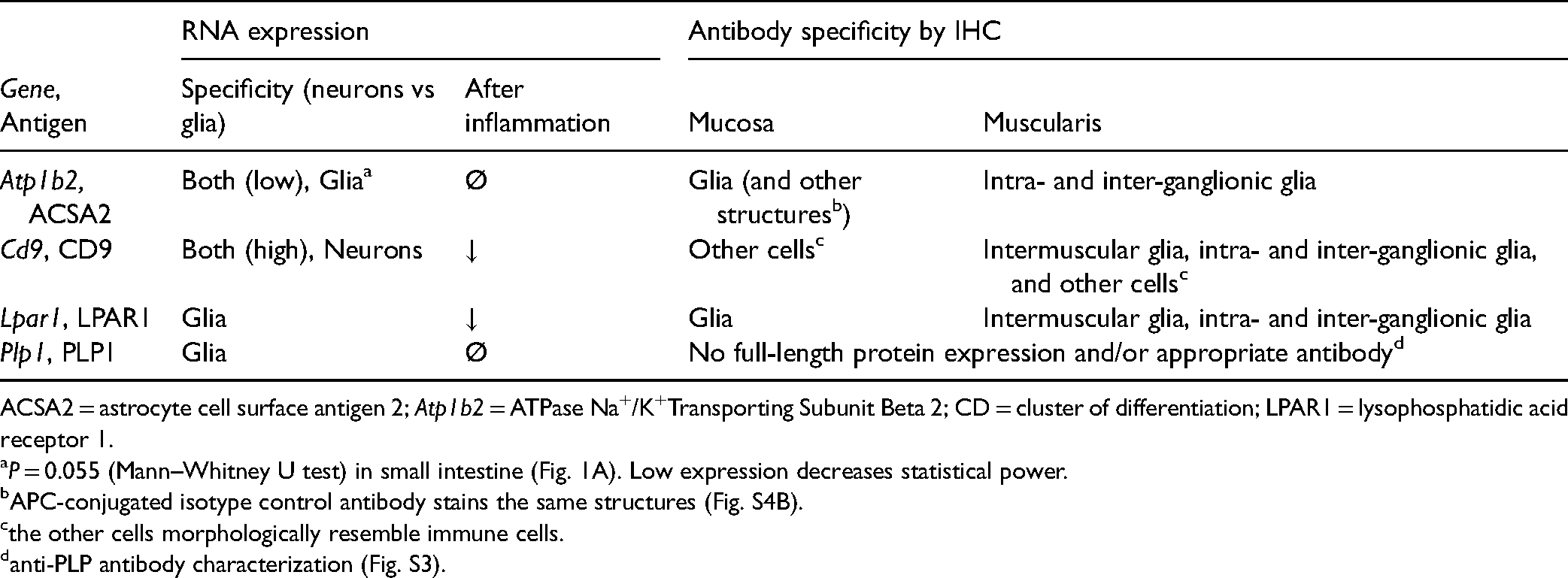

ACSA2 = astrocyte cell surface antigen 2; Atp1b2 = ATPase Na+/K+Transporting Subunit Beta 2; CD = cluster of differentiation; LPAR1 = lysophosphatidic acid receptor 1.
P = 0.055 (Mann–Whitney U test) in small intestine (Fig. 1A). Low expression decreases statistical power.
APC-conjugated isotype control antibody stains the same structures (Fig. S4B).
the other cells morphologically resemble immune cells.
anti-PLP antibody characterization (Fig. S3).
The main outcome of this study is the identification of several candidate cell surface markers for enteric glia; however, the expression patterns of these markers also provide additional insight into the heterogeneity of enteric glia. There are at least 4 different subpopulations of enteric glia that can be differentiated based on their anatomical locations and morphologies (Boesmans et al., 2015; Gulbransen & Sharkey, 2012). Intra- and inter-ganglionic glia are present in enteric ganglia with the enteric neurons and nerve fibers that form functional neurocircuits in the gut wall. In addition, extraganglionic enteric glia are present in the intestinal mucosa (mucosal glia) and with nerve fibers in the smooth muscle coats of the intestine (intramuscular glia). Single-cell transcriptional profiling (Drokhlyansky et al., 2020; Progatzky et al., 2021; Zeisel et al., 2018), immunolabeling (Boesmans et al., 2015), and functional studies (Ahmadzai et al., 2021) suggests additional complexity to glial heterogeneity in the gut. It is currently unclear whether the subtypes of enteric glia display distinct genetic profiles since data from single-cell transcriptional profiling studies are inconsistent in terms of cell clusters identified (Drokhlyansky et al., 2020; Progatzky et al., 2021; Zeisel et al., 2018).
Intraganglionic glia display morphological and functional similarities to astrocytes in the CNS and the data presented here show that ACSA2, a marker of CNS astrocytes, also labels this subpopulation of enteric glia. The microenvironment within enteric ganglia is distinct from that in the mucosa or muscle and differences in the glia residing in these locations would be expected. Myenteric ganglia are encapsulated by a connective tissue barrier (Dora et al., 2021; Gabella, 1972; Gershon & Bursztajn, 1978; Kiernan, 1996) that protects the rest of the intestinal environment and makes the environment within myenteric ganglia CNS-like. Our flow cytometry data indicates that ACSA2 labels about 40% of the glial cells in the mouse colon muscularis and our IHC labeling demonstrated that these ACSA2+ cells are glia within the myenteric plexus. Taken together, these outcomes suggest that about 60% of the glial cells in the muscle coats that are ACSA2- belong to the extraganglionic glia. This is an exciting new idea because the current opinion is that majority of the glial cells outside mucosa are found in the enteric ganglia. Indeed, myenteric ganglia show the high intensity of glial markers but this could be due to the high packing density of glial cells within the myenteric ganglia while the intramuscular glia are interspersed among smooth muscle cells and their signal gets diluted. This hypothesis should be further investigated by determining the numbers of myenteric and intermuscular glia. An alternative explanation is that ACSA2 labels a subpopulation of glial cells within the myenteric ganglia. ACSA2 is also expressed by mucosal glia, albeit at lower apparent levels, and this should be further investigated. Enteric glia in intestinal lamina propria are replenished throughout adulthood by migration from the myenteric plexus (Kabouridis et al., 2015), so glial ACSA2 expression may be suppressed once outside the myenteric plexus.
The outcomes of this study also point to critical differences between gene and protein expression, and the importance of investigating both. Outcomes of single-cell/-nuclei transcriptomics revealed very low Atp1b2 transcription. The abundance of Atp1b2 transcripts was at the level of neuronal expression of glial markers Sox10 and Plp1, and therefore, could be interpreted as non-expressing and dismissed for more detailed investigation. However, the ribosomal trap-enriched glial transcriptome and our antibody labeling data in IHC and flow cytometry show reliable Atp1b2 translation and ACSA2 expression in enteric glia. Perhaps the discrepancy between the RNA and protein expression lies in their molecular stability and complexity or lengthiness of the experimental procedures. On the other hand, Cd9 had the highest RNA expression. Although Cd9 transcript levels were comparable between glia and neurons, we found mostly glial CD9 expression in the myenteric plexus of the mouse colon. Due to the proximity and dense packing of neurons and glia in the myenteric ganglia, additional studies using cell-specific knockouts are needed to properly investigate the cell specificity of the Cd9 RNA and protein expression. Lastly, enteric glia show strong Plp1 transcription but we were unable to reliably label Plp1-expressing cells with antibodies. Enteric glia are non-myelinating peripheral glial cells but retain Plp1 promoter activity (Rao et al., 2015). We were unable to produce positive labeling of enteric glia using commercially available anti-PLP antibodies. Similar discordances between the Plp1 gene and protein expression have been reported in CNS neurons, which express the Plp1 gene but not the functional protein (Miller et al., 2009). Furthermore, the Plp1 gene encodes two proteins, PLP and DM20 (Griffiths et al., 1995), and unpublished research shows that Plp1 expression in the intestine predominantly occurs during early postnatal development, that the main product is the DM20 isoform, and that the gene is expressed in enteric neurons in addition to glia (Patyal et al., 2021).
In summary, our findings indicate two good candidates for cell-specific surface labeling of enteric glia. While LPAR1 labels enteric glia in general, ACSA2 is more specific for a glial subpopulation within myenteric ganglia and mucosal glia. These new tools will open new avenues for basic and translational research and, in turn, generate discoveries that will expand our fundamental knowledge of glial biology/pathophysiology and new ways for the treatment of human intestinal diseases.
Supplemental Material
sj-docx-1-asn-10.1177_17590914221083203 - Supplemental material for Astrocyte Cell Surface Antigen 2 and Other Potential Cell Surface Markers of Enteric glia in the Mouse Colon
Supplemental material, sj-docx-1-asn-10.1177_17590914221083203 for Astrocyte Cell Surface Antigen 2 and Other Potential Cell Surface Markers of Enteric glia in the Mouse Colon by Vladimir Grubišić and Brian D. Gulbransen in ASN Neuro
Footnotes
List of Abbreviations Used
Acknowledgments
The data presented herein were obtained using instrumentation in the MSU Flow Cytometry Core Facility with the support of Core staff. The facility is funded through user fees and the financial support of Michigan State University's Office of Research & Innovation, College of Osteopathic Medicine, and College of Human Medicine. We thank Vedrana Bali for preparing the brain slices and Jonathon McClain for animal care.
Author Contribution
Project conceptualization and design were developed by V.G. and B.D.G. Experiments and analyses were performed by V.G. The manuscript was written and edited by V.G. and B.D.G.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Crohn’s and Colitis Foundation, Research Fellowship Award [577598] to VG and National Institute of Diabetes and Digestive and Kidney Diseases grants [R01DK103723 and R01DK120862] to BDG.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
