Abstract
Background:
KRAS, TP53, CDKN2A, and SMAD4 have been the main driver mutations in pancreatic ductal adenocarcinoma (PDAC). Studies on the clinical significance and treatment response to 5-fluorouracil, leucovorin, irinotecan, and oxaliplatin (FOLFIRINOX) regimen in terms of the presence of these mutations remain inconclusive.
Objectives:
This study aimed to compare the survival outcome and response to FOLFIRINOX chemotherapy based on the presence of four driver mutation genes.
Design:
A multi-center retrospective study conducted at two tertiary medical centers.
Methods:
This study analyzed PDAC patients who were treated with FOLFIRINOX chemotherapy as the initial treatment. Tumor specimens were analyzed by a targeted next-generation sequencing platform at two tertiary referral hospitals from January 2016 to March 2022. Patients’ demographics, survival outcomes, and chemotherapeutic response were investigated and compared according to the presence of driver mutations.
Results:
The analysis included 100 patients. KRAS mutation was identified in 92 (92.0%) patients, followed by TP53, CDKN2A, and SMAD4 in 63 (63.0%), 18 (18.0%), and 17 (17.0%) patients, respectively. The TP53 wild-type group demonstrated longer overall survival (OS) than the TP53 mutated group (median OS: 29 vs 19 months, p = 0.03), and TP53 served as a prognostic factor for survival (hazard ratio = 1.74, 95% confidence interval: 1.00–3.00, p = 0.048). The difference in OS according to TP53 mutation was intensified in localized pancreatic adenocarcinoma (37 vs 19 months, p = 0.01). The TP53 wild-type group demonstrated a higher objective response rate to FOLFIRINOX chemotherapy than the TP53 mutation group in localized pancreatic adenocarcinoma (50.0% vs 17.6%, p = 0.024).
Conclusion:
PDAC patients with wild-type TP53 demonstrated longer OS than those with TP53 mutation, and this trend was intensified in patients with localized disease. This result may be due to an impaired response to FOLFIRINOX chemotherapy in patients with TP53 mutation.
Introduction
Pancreatic ductal adenocarcinoma (PDAC) is the fourth leading cause of death from cancer in the United States, and approximately 62,000 patients are diagnosed annually. 1 Only 10%–20% of PDAC cases are resectable upon diagnosis, which explains the poor prognosis of PDAC. 2 However, gradual improvement in PDAC treatment has been reported in the past decade with the introduction of 5-fluorouracil (5-FU), leucovorin, irinotecan, and oxaliplatin (FOLFIRINOX) regimen. FOLFIRINOX has demonstrated notable efficacy in treating metastatic PDAC, with an objective response rate (ORR) of 32%. 3 Currently, it is a preferred treatment option for patients with a tolerable performance status in both neoadjuvant and palliative settings.
The identification of molecular mutation profiles has spread and is commercially available at numerous centers globally since the introduction of next-generation sequencing (NGS). Studies that aimed to associate PDAC mutation profiles with treatment options have revealed limited results. Some options have been effective against specific mutations, but their applicability is limited to a small subset of patients with PDAC. 4 Mutations found with high frequency include the oncogene KRAS and tumor suppressor genes TP53, CDKN2A, and SMAD4.5,6 Several studies have been conducted on these main driver mutation genes and their clinical relevance. However, their relevance and clinical implications remain inconclusive. In addition, no studies focused on the response to FOLFIRINOX chemotherapy as an outcome of interest. Thus, our study aimed to compare the survival outcome and response to FOLFIRINOX chemotherapy based on the presence of four driver mutation genes in patients with PDAC.
Methods
Patient and study design
Patients diagnosed with PDAC whose tumor specimens underwent NGS testing from January 2016 to March 2022 and from March 2018 to August 2020 at Seoul National University Hospital (SNUH) and Seoul National University Bundang Hospital (SNUBH), respectively, were investigated. This study included those who were treated with FOLFIRINOX chemotherapy as the initial treatment among the investigated patients. Our analysis excluded patients (1) who underwent upfront resection, (2) whose initial chemotherapy regimen was not FOLFIRINOX, (3) assessed as having resectable PDAC, (4) whose histological diagnosis was not ductal adenocarcinoma, and (5) diagnosed with intraductal papillary mucinous carcinoma. Data from the study patients were retrospectively collected from electronic medical records. Demographics, Eastern Cooperative Oncology Group (ECOG) performance status, tumor location and size, pathologic reports, and follow-up data, including survival and progression, were collected. Subgroup analysis was conducted for localized and metastatic PDAC. Localized disease was defined as borderline resectable (BR) and locally advanced (LA) PDAC according to the National Comprehensive Cancer Network Clinical Practice Guidelines. 7 The institutional review boards of SNUH (IRB no. 2207-121-1342) and SNUBH (IRB no. B-2305-827-402) approved the study protocol and the informed consent was waived because of the retrospective nature of the study. The reporting of this study conforms to the Strengthening the Reporting of Observational Studies in Epidemiology statement. 8
NGS analysis
We used the NGS report of the SNUH pan-cancer panel (version 3.3), which included 184 genes (Supplemental Table 1), and the SNUBH-Macrogen panel (version 2.0), which targeted 544 genes (Supplemental Table 2). Both analyses extracted genomic DNA from formalin-fixed, paraffin-embedded tissue. The NextSeq 550Dx platform (Illumina Inc., San Diego, CA, USA) was used for paired-end sequencing. Sequenced reads were aligned to the human reference genome hg19 using the Burrows–Wheeler Aligner 9 and GATK Best Practice. Single-nucleotide variants and small insertions and deletions (indels) were detected using an in-house developed pipeline, GATK HaplotypeCaller, 10 SNVer, 11 and LoFreq 12 in the SNUH panel, and Mutect2 in the SNUBH panel. Copy number alterations were performed using an in-house developed pipeline and CNVkit. Mutations were annotated using ANNOtate VARiation. 13
Variants included in an in-house Panel of Normals were excluded to identify and remove germline variants and recurrent sequencing artifacts. Furthermore, variants with a population frequency of >0.1% on the Genome Aggregation Consortium (gnomAD) East Asian database, Korean Variant Archive, or Korean Reference Genome Database were filtered out. However, variants with a frequency of >1% of gnomAD in the SNUBH panel were filtered out. 14 We focused on the presence of four driver mutations (KRAS, TP53, CDKN2A, and SMAD4) for each patient. The analysis excluded synonymous variants among single-nucleotide polymorphisms. R package “maftools” was used to draw lollipop plots to visualize variant locations on TP53. 15
Assessment and definition
Overall survival (OS) was defined from the date of diagnosis to the date of death or last follow-up. The patient’s response to FOLFIRINOX was evaluated according to the Response Evaluation Criteria in Solid Tumors (RECIST) 1.1 criteria. 16 The best response, which was defined as the most favorable outcome observed throughout the treatment period of FOLFIRINOX, was investigated. Four authors (M.K.K., I.R.C., K.J., and J.K.) retrospectively reviewed computed tomography images to assess the response to FOLFIRINOX. In cases of disagreement, the corresponding author was involved in the discussion to conclude. The resectability of PDAC was defined following the National Comprehensive Cancer Network criteria. 7 The ORR was the percentage of people who had a partial response (PR) or complete response (CR).
Statistical analysis
Continuous variables were provided as median values with an interquartile range (IQR), and categorical variables were provided as numbers and proportions (%). χ2 test or Fisher’s exact test and the Kruskal–Wallis test were utilized to compare categorical variables between the two and three groups, respectively. Student’s t-test was used to compare continuous variables between groups. Kaplan–Meier survival analysis and log-rank test were used to compare OS between groups. A Cox proportional hazards analysis was conducted to evaluate prognostic factors related to survival. The multivariate Cox analysis included variables that were effective in the univariate Cox analysis (p < 0.05) or clinically meaningful. A p-value of 0.05 indicated statistical significance. R version 4.2.0 (R Foundation for Statistical Computing, Vienna, Austria) was used for all statistical analyses.
Results
Study population and baseline characteristics
A total of 207 patients were diagnosed with pancreatic cancer and underwent NGS at SNUH and SNUBH. Our study included 100 patients diagnosed with PDAC who underwent NGS panel-based tests and received FOLFIRINOX as their initial treatment after excluding patients who were not adequate for analysis. Of these patients, 61 were from SNUH and 39 were from SNUBH (Figure 1). Table 1 summarizes the baseline characteristics of the included patients. A total of 54 (54.0%) patients were male, the median age was 62 years (IQR: 57–67), and the median follow-up was 22 months. All patients demonstrated a good ECOG performance status of 0 or 1. PDAC consisted of BR (17.0%), LA (35.0%), and metastatic (48.0%) PDAC. KRAS mutation was determined in 92 (92.0%) patients, TP53 mutation in 63.0%, followed by CDKN2A (18.0%) and SMAD4 (17.0%). Among the patients, 62.0% had both KRAS and TP53 mutations, 30.0% had mutated KRAS and wild-type TP53, 1.0% had wild-type KRAS and mutated TP53, and 7.0% had both KRAS and TP53 wild type. The best response to FOLFIRINOX by the RECIST 1.1 criteria was as follows: PR (32, 32.0%), stable disease (SD) (49, 49.0%), and PD (19, 19.0%). CR was not reported. The median progression-free survival and OS of the entire cohort were 10 and 23 months, respectively. The median number of FOLFIRINOX cycles at the point of best response was 7 (IQR: 4–10).

Flow chart of patient selection in this study.
Baseline characteristics of patients.
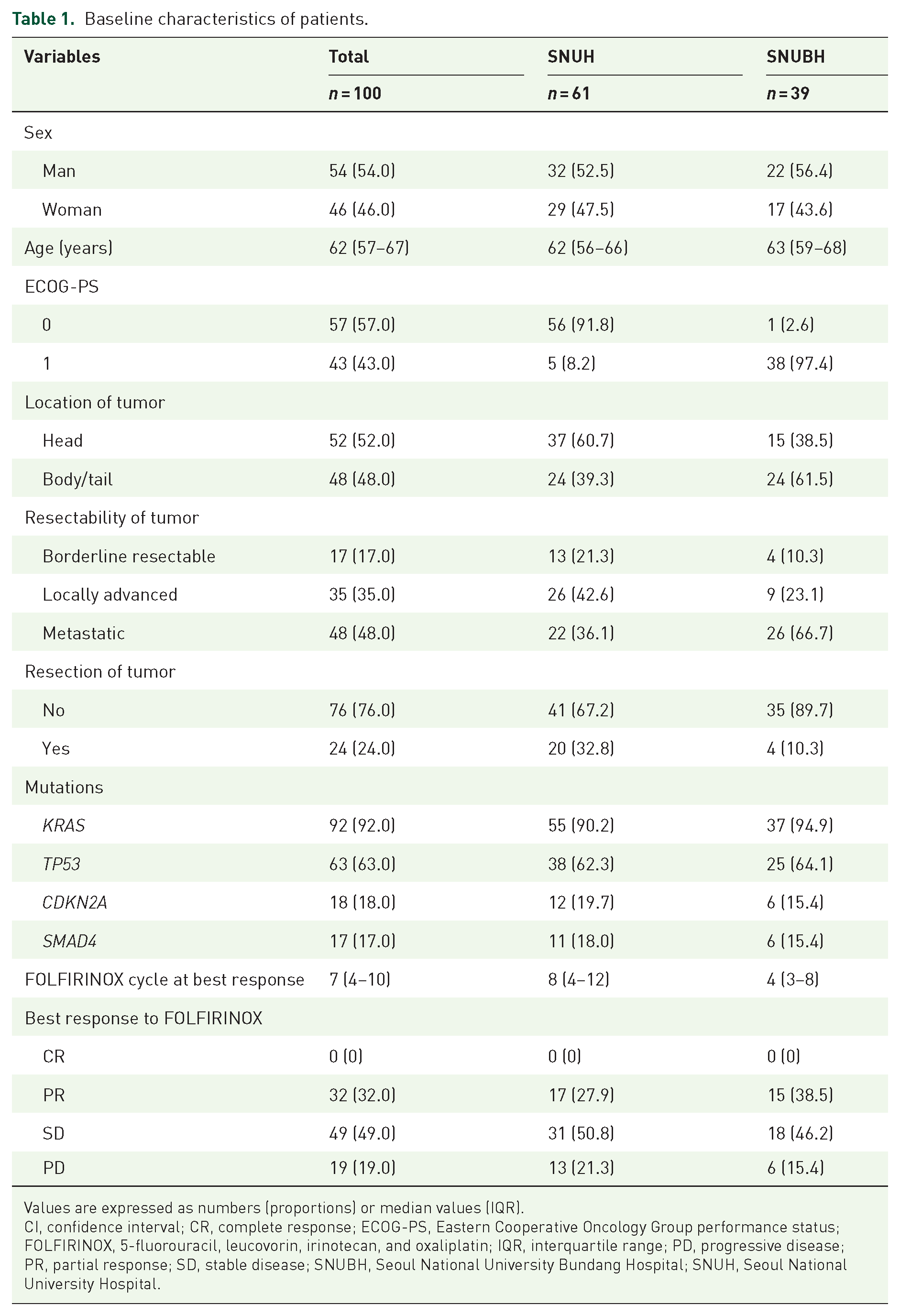
Values are expressed as numbers (proportions) or median values (IQR).
CI, confidence interval; CR, complete response; ECOG-PS, Eastern Cooperative Oncology Group performance status; FOLFIRINOX, 5-fluorouracil, leucovorin, irinotecan, and oxaliplatin; IQR, interquartile range; PD, progressive disease; PR, partial response; SD, stable disease; SNUBH, Seoul National University Bundang Hospital; SNUH, Seoul National University Hospital.
Result of the survival outcomes
We analyzed and compared the OS according to the presence of KRAS, TP53, CDKN2A, and SMAD4 mutations in the entire cohort (Figure 2). The TP53 wild type demonstrated a longer median OS compared with the mutated TP53 group (29 vs 19 months, p = 0.03). Subgroup analysis was performed on patients with localized and metastatic disease. The difference in OS according to TP53 mutation was intensified in localized PDAC (37 vs 19 months, p = 0.01, Figure 3). However, metastatic PDAC demonstrated no significant difference according to the presence of TP53 mutation (25 vs 19 months, p = 0.7, Supplemental Figure 1). No differences were observed in OS for other mutations, both in the entire cohort and subgroup analysis (Figures 2 and 3, and Supplemental Figure 1). The median OS for patients with both KRAS and TP53 mutations, KRAS mutation and TP53 wild type, and both KRAS and TP53 wild type were 19, 28, and 31 months, respectively. Only one patient had wild-type KRAS and TP53 mutation, with an OS of 15 months. However, these differences were not statistically significant (Supplemental Figure 2).
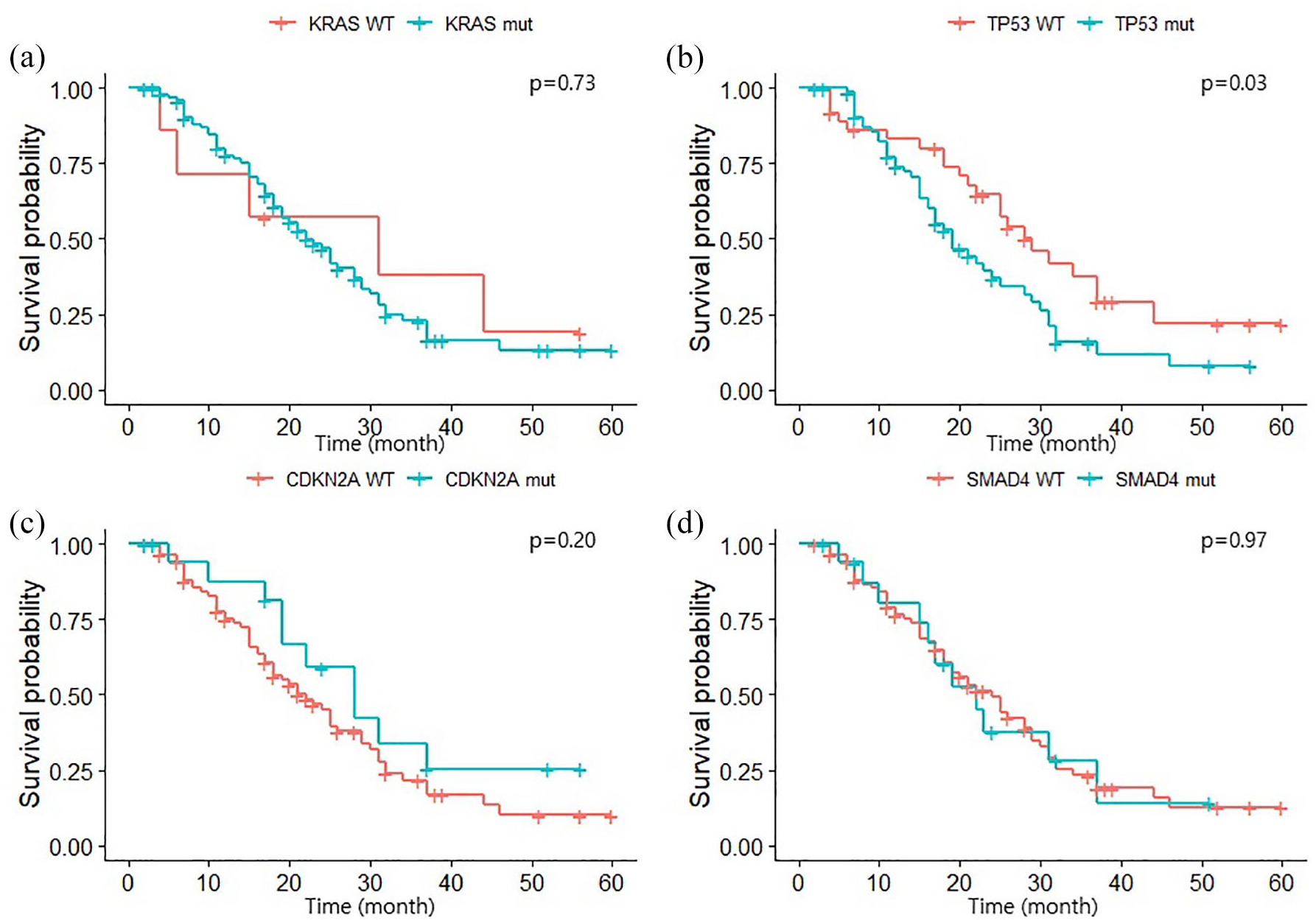
Kaplan–Meier curve of overall survival according to the presence of mutated (a) KRAS, (b) TP53, (c) CDKN2A, and (d) SMAD4 genes.

Kaplan–Meier curve of overall survival according to the presence of mutated (a) KRAS, (b) TP53, (c) CDKN2A, and (d) SMAD4 genes in localized disease.
Prognostic factors associated with survival
Cox proportional hazards analysis was conducted on various prognostic factors, including the presence of each mutation, to investigate their prognostic impact on survival outcomes. Multivariate analysis indicated that the presence of TP53 mutation (hazard ratio (HR) = 1.74, 95% confidence interval (CI): 1.00–3.00, p = 0.048) was a significant prognostic factor (Table 2). Subgroup analysis of patients with localized disease identified TP53 mutation as a significant prognostic factor for OS (HR: 4.49, 95% CI: 1.67–10.98; p = 0.002, Table 3). In localized disease, the SNUBH center was a negative prognostic factor for survival compared with the SNUH center in multivariate analysis as well (HR: 4.37, 95% CI: 1.87–10.21; p = 0.001).
Prognostic factors for survival of total cohort.

BR, borderline resectable; CI, confidence interval; ECOG-PS, Eastern Cooperative Oncology Group performance status; HR, hazard ratio; LA, locally advanced; M, metastatic; SNUBH, Seoul National University Bundang Hospital; SNUH, Seoul National University Hospital; WT, wild type.
Prognostic factors for survival in patients with localized disease.

BR, borderline resectable; CI, confidence interval; ECOG-PS, Eastern Cooperative Oncology Group performance status; HR, hazard ratio; LA, locally advanced; SNUBH, Seoul National University Bundang Hospital; SNUH, Seoul National University Hospital; WT, wild type.
Treatment response to FOLFIRINOX
The response to FOLFIRINOX according to the presence of mutations was analyzed and compared. No significant difference was observed in the distribution of PR, SD, and PD (Supplemental Table 3). The same analysis was conducted on a subgroup of patients with localized disease. The presence of mutations in KRAS and TP53 significantly affected the treatment outcome of FOLFIRINOX (Table 4). The post hoc analysis revealed that ORR (proportion of CR and PR) was different according to the presence of KRAS and TP53. ORR was higher in the KRAS wild type than in the mutated KRAS (100.0% vs 24.5%, p = 0.020) and in the TP53 wild type than in the mutated TP53 (50.0% vs 17.6%, p = 0.024, Table 5).
Comparison of treatment response to FOLFIRINOX according to the presence of main driver mutations in patients with localized disease.

FOLFIRINOX, 5-fluorouracil, leucovorin, irinotecan, and oxaliplatin; PD, progressive disease; PR, partial response; SD, stable disease; WT, wild type.
Post hoc analysis of treatment response to FOLFIRINOX based on the presence of KRAS and TP53 mutations in patients with localized disease.

FOLFIRINOX, 5-fluorouracil, leucovorin, irinotecan, and oxaliplatin; PD, progressive disease; PR, partial response; SD, stable disease; WT, wild type.
In-depth NGS analysis of the SNUH cohort
We conducted a deeper analysis of the mutational profiles of the genes for which status differed significantly among patient groups based on the best response to FOLFIRINOX. We focused on analyzing the assessment of vcf files that involve information on variants’ specific locations because it was limited to the SNUH cohort. Variants located at TP53 in each subgroup were further inspected, considering that TP53 was more frequently mutated in the SD and PD groups than in the PR group (p = 0.024). Missense mutations at amino acid position 33 were prevalently observed in all subgroups, all of which indicated the conversion of proline to arginine. Notably, the p53 transactivation domain was exclusively mutated in the SD and PD groups (Figure 4).

Lollipop plots of TP53 variants on each subgroup divided by response to FOLFIRINOX. The Y-axis indicates the number of variants detected in a certain position.
Discussion
This study focused on investigating the four most frequent mutations (KRAS, TP53, CDKN2A, and SMAD4) in PDAC and analyzing their association with survival and response to FOLFIRINOX chemotherapy, which is a largely used treatment option in PDAC. Patients who had wild-type TP53 demonstrated longer OS compared with the mutated TP53 group, and the trend was more prominent in PDAC with localized disease in the subgroup analysis. In addition, a higher ORR to FOLFIRINOX was observed in the TP53 wild-type group in patients with localized disease.
Previous studies have investigated the association between frequent driver mutations and clinical significance, including the prognosis of pancreatic cancer. In general, patients with pancreatic cancer with mutations in the main driver gene appeared to have a poor prognosis. Based on data obtained from 283 patients with resected pancreatic cancer, KRAS and TP53 gene mutations were associated with poor OS. Other studies revealed that among the four driver mutation genes (KRAS, TP53, CDKN2A, and SMAD4), patients with fewer mutated genes demonstrated better survival outcomes.17,18 Furthermore, a meta-analysis of 17 studies revealed that TP53 mutation overexpression was associated with poorer OS, along with other driver mutations. 6 Overall, the presence of main driver mutations has been associated with a poorer prognosis compared with the wild type. However, to the best of our knowledge, no other studies have determined these driver mutations in response to cytotoxic chemotherapy.
Tumor heterogeneity is clear in pancreatic cancer. Intratumor heterogeneity, including heterogeneity between the primary lesion and its metastatic part, plays a key role in tumor progression and drug resistance. 19 Two large studies involving 1080 and 718 patients have revealed a higher frequency of TP53 mutations in the metastatic sites compared with the primary sites despite the lack of direct studies that compare the main mutation profiles of matched primary and metastatic lesions in pancreatic cancer.20,21 Our results have revealed that the negative predictive role of mutated TP53 is more evident when excluding patients with distant metastasis. Intratumor heterogeneity observed in pancreatic cancer, in addition to the higher frequency of TP53 mutations in metastatic lesions, may explain this view.
Over the past decade, the spread of NGS and advancements in bioinformatics have resulted in the emergence of novel treatment strategies that target specific subgroups of PDAC based on their genomic profile. Golan et al. 22 revealed the effectiveness of using olaparib, a poly(adenosine diphosphate-ribose) polymerase inhibitor, as maintenance therapy in patients with metastatic pancreatic cancer who have germline BRCA mutation. Sotorasib confirmed anticancer effects against patients with KRAS G12C in phase I and II trials. 23 The KEYNOTE 158 study revealed that immune checkpoint inhibitor pembrolizumab was effective in mismatch repair deficiency/ microsatellite instability (MSI)-high and tumor mutatnioal burden (TMB)-high pancreatic cancer.24,25 The incidence of PDAC indicated for these options remains exceedingly low despite the introduction of these innovative treatments. Currently, no specific option is recommended for patients with main driver mutations. FOLFIRINOX remains the treatment of choice for most patients with PDAC with adequate performance status.
TP53 is a tumor suppressor gene that encodes the p53 protein genetic information. Furthermore, it is one of the most mutated genes in cancer. Wild-type p53 is known for its proapoptotic effects. It detects DNA damage, activates cell cycle checkpoints, and subsequently induces cell death.26,27 Our in-depth NGS analysis revealed that the p53 transactivation domain was exclusively mutated in the SD and PD groups, inferring the influence of the transactivation domain on protein function causing a poor treatment outcome. FOLFIRINOX is known to act on tumor cells by inducing DNA damage followed by apoptosis. 28 Full transcriptional p53 activity assisted by the transactivation domain is essential for DNA damage-induced apoptosis 29 ; thus these mutations may interfere with the cancer-fighting mechanism of FOLFIRINOX. Other studies confirm this view of FOLFIRINOX resistance and 5-FU mutation. A study that focused on colon cancer cell lines revealed that oxaliplatin was effective in inhibiting the growth of all p53 wild-type cell lines, whereas most of the p53 mutated cell lines demonstrated innate treatment resistance. 30 The association between 5-FU and p53 protein is more evident. As a DNA-damaging reagent, 5-FU effectively induces cell cycle arrest, preventing cancer cells from proliferating and triggering apoptosis. 31 TP53 plays a crucial role in regulating the cell cycle; therefore, the efficacy of 5-FU as a therapeutic agent is partially contingent on the TP53 status of cancer cells. One study revealed that the Ca2+-calmodulin-p53 axis is crucial to extrinsic apoptosis induced by 5-FU. Inhibiting this pathway eliminated the ability of 5-FU to induce caspase activity, indicating the role of p53 in 5-FU-induced cell death. Moreover, the apoptotic response to 5-FU was reduced by >50% in cells expressing mutant p53 compared with cells expressing exogenous wild-type p53.32,33 Furthermore, p53 may be involved in downstream signaling pathways in response to 5-FU. 34 Overall, TP53 mutations help develop resistance to FOLFIRINOX, although conclusive clinical evidence to confirm this remains unknown.
The evidence associated with FOLFIRINOX resistance is not as extensive for the other mutated genes included in our study. However, one study demonstrated improved survival outcomes in patients with pancreatic cancer with wild-type KRAS compared with those with mutated KRAS. Interestingly, this survival advantage was more prominent in the subgroup that received 5-FU and oxaliplatin treatment. 35
TMB and MSI were not included in our analysis from the perspective of the NGS data analysis. However, we confirmed that none of the included patients were classified as TMB-high or MSI-high. The SNUH-pan-cancer panel and SNUBH-Macrogen panel used a DNA-based targeted panel, considering targeted sequencing analysis is sufficient to analyze the presence of main driver mutations. Whole-genome sequencing and whole-exome sequencing are frequently used in contemporary studies investigating the mutational landscape of pancreatic cancer.36–39 A constraint was found in generalizing the findings of other studies in our analysis because our study used targeted sequencing analysis for categorizing mutations in pancreatic cancer. However, identifying the predominant driver mutations that are commonly occurring and easily identifiable in pancreatic cancer remains to be accomplished using cost-effective targeted sequencing analysis. Moreover, these key driver mutations serve as potential targets for treating pancreatic cancer. 4 The targeted gene panel was based on the understanding that only approximately 500, out of the vast pool of over 20,000 human genes, are true driver genes in cancer. 40 The molecular traits of tumor tissues can be analyzed simultaneously using NGS-based cancer gene panels, providing comprehensive coverage and allowing for the detection of minor allele frequencies cost-effectively. 41
Along with the aspect of NGS data analysis, our research has several limitations that should be addressed. First, this study included a relatively small number of patients, and the patient characteristics differed between the two institutions (SNUH and SNUBH). The SNUH group exhibited a higher proportion of localized patients, which could have affected the results. Second, the collection of specimens for the NGS test was not well organized. Specimens in certain patients were collected after FOLFIRINOX initiation, whereas specimens from nonprimary lesions were used in other patients. Finally, conducting a transcriptomic analysis is essential to gain a better understanding of the precise role played by p53 in FOLFIRINOX resistance, which was not performed in our analysis.
Conclusion
We investigated the relationship between the most frequently found driver mutation genes of PDAC and their clinical significance, including survival and FOLFIRINOX response. The TP53 wild-type group demonstrated better survival outcomes than the group with TP53 mutation, possibly due to improved response to FOLFIRINOX. In addition, our results revealed that TP53 could serve as a predictive marker for survival. However, the association between variants of TP53 and their impact on clinical features needs to be evaluated in future studies. Further in-depth analysis of the NGS panel data is required to obtain a more comprehensive understanding of this subject.
Supplemental Material
sj-docx-1-tam-10.1177_17588359241290482 – Supplemental material for Prognostic value of the TP53 mutation in patients with pancreatic ductal adenocarcinoma receiving FOLFIRINOX
Supplemental material, sj-docx-1-tam-10.1177_17588359241290482 for Prognostic value of the TP53 mutation in patients with pancreatic ductal adenocarcinoma receiving FOLFIRINOX by Min Kyu Kim, In Rae Cho, Yooeun Kim, Jin Ho Choi, Kwangrok Jung, Jaihwan Kim, Sheehyun Kim, Hongseok Yun, Jeesun Yoon, Do-Youn Oh, Kwangsoo Kim and Sang Hyub Lee in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-jpg-2-tam-10.1177_17588359241290482 – Supplemental material for Prognostic value of the TP53 mutation in patients with pancreatic ductal adenocarcinoma receiving FOLFIRINOX
Supplemental material, sj-jpg-2-tam-10.1177_17588359241290482 for Prognostic value of the TP53 mutation in patients with pancreatic ductal adenocarcinoma receiving FOLFIRINOX by Min Kyu Kim, In Rae Cho, Yooeun Kim, Jin Ho Choi, Kwangrok Jung, Jaihwan Kim, Sheehyun Kim, Hongseok Yun, Jeesun Yoon, Do-Youn Oh, Kwangsoo Kim and Sang Hyub Lee in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-jpg-3-tam-10.1177_17588359241290482 – Supplemental material for Prognostic value of the TP53 mutation in patients with pancreatic ductal adenocarcinoma receiving FOLFIRINOX
Supplemental material, sj-jpg-3-tam-10.1177_17588359241290482 for Prognostic value of the TP53 mutation in patients with pancreatic ductal adenocarcinoma receiving FOLFIRINOX by Min Kyu Kim, In Rae Cho, Yooeun Kim, Jin Ho Choi, Kwangrok Jung, Jaihwan Kim, Sheehyun Kim, Hongseok Yun, Jeesun Yoon, Do-Youn Oh, Kwangsoo Kim and Sang Hyub Lee in Therapeutic Advances in Medical Oncology
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
