Abstract
Background:
Metastasis is the primary cause of lung cancer-related death. Primary cancer cells invade through the lymphatic or blood vessels to distant sites. Recently, it was proposed that lymphatic metastasis was more a hallmark of tumor aggressiveness or metastatic potential than a gateway to metastases. Therefore, the underlying molecular mechanism of metastasis is not entirely clear.
Objectives:
This study aimed to explore the genetic mechanisms underlying liver metastases from lung cancer and to evaluate the efficacy of different therapies in these patients.
Design:
We retrospectively analyzed the mutation spectrum of different biopsy samples including primary lung tumors, liver, lymph node metastasis, and circulating tumor DNA (ctDNA) from 1090 non-small-cell lung cancer (NSCLC) patients with liver metastasis between the years 2017 and 2022.
Methods:
Demographic and disease characteristics were summarized using descriptive parameters. Time to treatment discontinuation was used to analyze the clinical outcome.
Results:
More liquid biopsies were performed than tissue biopsies, especially in the treated advanced NSCLC patients. Liver metastasis before treatment was associated with poor response to immune checkpoint inhibitors and targeted therapy. Liver and lymph node metastasis had higher levels of single nucleotide variants and copy number variants than primary lung tumors. In paired lung and liver, lymph nodes, and simultaneous ctDNA, we found actionable mutations were always shared, while metastasis samples had multiple private mutations. Serial ctDNA analysis identifies potential resistant mutations and describes the evolution of tumor cells.
Conclusion:
Liver and lymph node metastasis in NSCLC showed shared actionable mutations. Of note, the discrepancy of private mutations in liver and lymph node metastases indicated that liver metastases are mainly seeded by the primary tumor rather than the earlier colonized lymph node metastases.
Introduction
Primary lung cancer is the leading cause of cancer-related mortality worldwide. 1 Metastasis to distant organs remains the leading cause of cancer-related death. 2 For non-small-cell lung cancer (NSCLC) patients, the most common extra-pulmonary sites of distant metastasis are the brain, bone, adrenal gland, and liver.3–5 Approximately 4% of NSCLC patients have liver metastasis at first diagnosis, and more than 95% of patients have multiple metastases. 6 While surgical resection of metastases is sometimes curative, most patients with liver metastases are not considered resectable because of the number or location of the metastases.7–10 The presence of liver metastasis is an independent prognostic factor for shorter survival in NSCLC patients,11–13 and a recent study has shown that liver and bone metastases are associated with inferior outcomes compared to bone metastases. 4 Moreover, liver metastasis is associated with poor therapeutic response to targeted therapy14–16 and immunotherapy. 17
Next-generation sequencing (NGS) has been widely used to identify genetic variation in patients with lung cancer before targeted therapy and immunotherapy.18–20 In general, metastatic cancers carry mutations similar to those of the primary cancer, but additional mutations occur after transformation or treatments.21,22 Small cohort studies have shown that 66.3% of genetic mutations identified in paired primary and metastatic liver were shared with high consistency in common driver mutations such as TP53 and EGFR. 23 Nonetheless, genetic analysis of liver metastatic lesions was still limited, though a better understanding of mutational profiling might guide therapeutic strategies and improve patient outcomes. Liquid biopsy with plasma circulating tumor DNA (ctDNA), pleural effusions, or cerebrospinal fluid is increasingly adopted in clinical practice. 24 The concordance of genetic alteration between matched liquid biopsy and tissue was associated with tumor burden and previous therapies.25–28
In the recently published TRACERx studies, it was noted that approximately 75% of metastases diverge late in the primary tumor and the majority of clonal driver mutations persisted in the metastases as previous results. 29 Interestingly, it was reported that less than 20% of primary lymph node metastases seed progressive disease, suggesting that lymphatic metastasis was more a hallmark of tumor aggressiveness or metastatic potential than a gateway to metastases.30–32 Most patients with liver metastasis also had lymph node metastasis, whether cancer cells in lymph nodes could seed liver metastases has been a subject of considerable debate, though in mouse models, lymph node metastases can be a source of cancer cells for distant metastases. 33
Here, we conducted this retrospective study to examine a large cohort of patients with liver metastasis from NSCLC to show the real-world choice of samples for mutation analysis. We also analyzed the concordance of genetic alterations among different samples collected at advanced stages to provide information on metastatic evolution and for future decision-making for NSCLC patients with liver metastasis.
Methods
Patient recruitment
The cohort of 1090 NSCLC patients with confirmed liver metastasis from January 2017 to August 2022 was retrospectively recruited from five hospitals. Clinical information was collected and analyzed with mutation profiling. This study was conducted in accordance with the principles of the revised Declaration of Helsinki. In addition, the reporting of this study conforms to the STROBE statement (Supplemental File 1).
DNA extraction, library preparation, and target enrichment
NGS-based somatic mutation was performed in a College of American Pathologists-accredited laboratory, Geneplus-Beijing (Beijing, China). All tissue samples included in this study underwent pathology review onsite to confirm histologic classification and the adequacy of the tumor tissues, which required a minimum of 20% of tumor cells. Tumor DNA was extracted from formalin-fixed, paraffin-embedded (FFPE) tumor tissue specimens using the ReliaPrep™ FFPE gDNA Miniprep System (Promega, Madison, WI, USA). Genomic DNA was extracted from white blood cells as a germline control using the DNeasy Blood Kit (Qiagen, Valencia, CA, USA). cell-free DNA (cfDNA) was isolated from 4 to 5 ml of isolated plasma, pleural effusion, ascites, cerebrospinal fluid, and pericardial effusion supernatant using the QIAamp Circulating Nucleic Acid Kit (Qiagen, Hilden, Germany). DNA concentration was measured using a Qubit fluorometer and the Qubit dsDNA HS (High Sensitivity) Assay Kit (Invitrogen, Carlsbad, CA, USA). The size distribution of the cfDNA was assessed using an Agilent 2100 BioAnalyzer and a DNA HS kit (Agilent Technologies, Santa Clara, CA, USA). Sequencing libraries were prepared using the KAPA Library Preparation Kit (Kapa Biosystems, Wilmington, MA, USA). Libraries were hybridized to custom-designed biotinylated oligonucleotide probes (Roche NimbleGen, Madison, WI, USA), covering ~1.4 Mbp genomic regions of 1021 cancer-related genes (Supplemental Table S1) using Gene + Seq 2000 instrument (Gene + technology). 34
Sequencing and data analysis
Sequencing data were analyzed using default parameters. Adaptor sequences and low-quality reads were removed. The clean reads were aligned to the reference human genome (hg19) using the Burrows-Wheeler Aligner (BWA; version 0.7.12-r1039 from the sourceforge download). GATK (v3.6-0-g89b7209 from the Broad Institute) was employed for realignment and recalibration. Single-nucleotide variants (SNV) were called using MuTect (version 1.1.4, a broad tool included in GATK) and NChot, an in-house software developed for reviewing hotspot variants. Small insertions and deletions (InDel) were determined by GATK. CONTRA (v2.0.8 from http://contra-cnv.sourceforge.net.) was used to identify somatic copy number alterations. All final candidate variants were manually verified with Integrative Genomics Viewer. Targeted capture sequencing required a minimal mean effective depth of coverage of 300× and 1000× in tissues and plasma samples, respectively. Targetable genomic alterations simultaneously detected by this assay included base substitutions, short insertions and deletions, focal gene amplifications and deletions (copy number alterations), and selected gene fusions and rearrangements. Depending on whether the plasma variant occurs in matched tumor tissue, the plasma variants were classified into tissue-derived and ctDNA-private mutations.
Subclonal analysis
Samples with more than four somatic substitution/small insertions and deletions were applied to PyClone by default to analyze the clonal structure using a Bayesian clustering method. 35 Cancer cell fraction was calculated with the mean of predicted cellular frequencies. The cluster with the highest mean Variant Allele Frequency (VAF) was identified as the clonal cluster, and mutations in this cluster were defined as clonal mutations, otherwise subclonal.
Phylogenetic tree construction
Phylogenetic tree reconstructions were performed using Dendrogram Plot tools in Hiplot Pro (https://hiplot.com.cn/), a comprehensive web service for biomedical data analysis and visualization.
Statistical analyses
Due to the retrospective nature of the cohort, time to treatment discontinuation (TTD) was used instead of progression-free survival (PFS) for analyzing the clinical outcome. TTD was defined here as the time from the start of treatment to the end of therapy. TTD distribution was analyzed by the Kaplan–Meier method with a log-rank test. Patients with missing values were not included in the statistical analysis. Associations between any two categorical variables were analyzed with Fisher’s exact test. A two-sided p value of <0.05 represented statistical significance. All the statistical analyses were performed using GraphPad Prism (v. 8.0; GraphPad Software, La Jolla, CA, USA) software.
Results
Patient characteristics
A total of 1090 NSCLC patients with liver metastasis were investigated in this study (Figure 1). Among them, 185 patients were treatment naïve while 905 patients were treated with targeted therapy and chemotherapy with/without immunotherapy (Table 1). The median age of diagnosis was 59, ranging from 23 to 85. Most patients were adenocarcinoma (n = 965), followed by squamous cell carcinoma (n = 98), adenosquamous carcinoma (n = 14), and other rare types such as spindle cell carcinoma, sarcomatoid carcinoma, etc. (n = 13). Besides liver metastasis, a large number of patients also had bone, lung, brain, and adrenal gland metastasis (Table 1). Compared with treatment-naïve patients, treated patients had significantly more patients with bone (94/185 vs 554/905, p = 0.01) or brain (40/185 vs 291/905, p = 0.005) metastasis, but comparable lung (51/185 vs 309/905, p = 0.09) or adrenal metastasis (18/185 vs 108/905, p = 0.45) (Figure 2(a)–(d)).
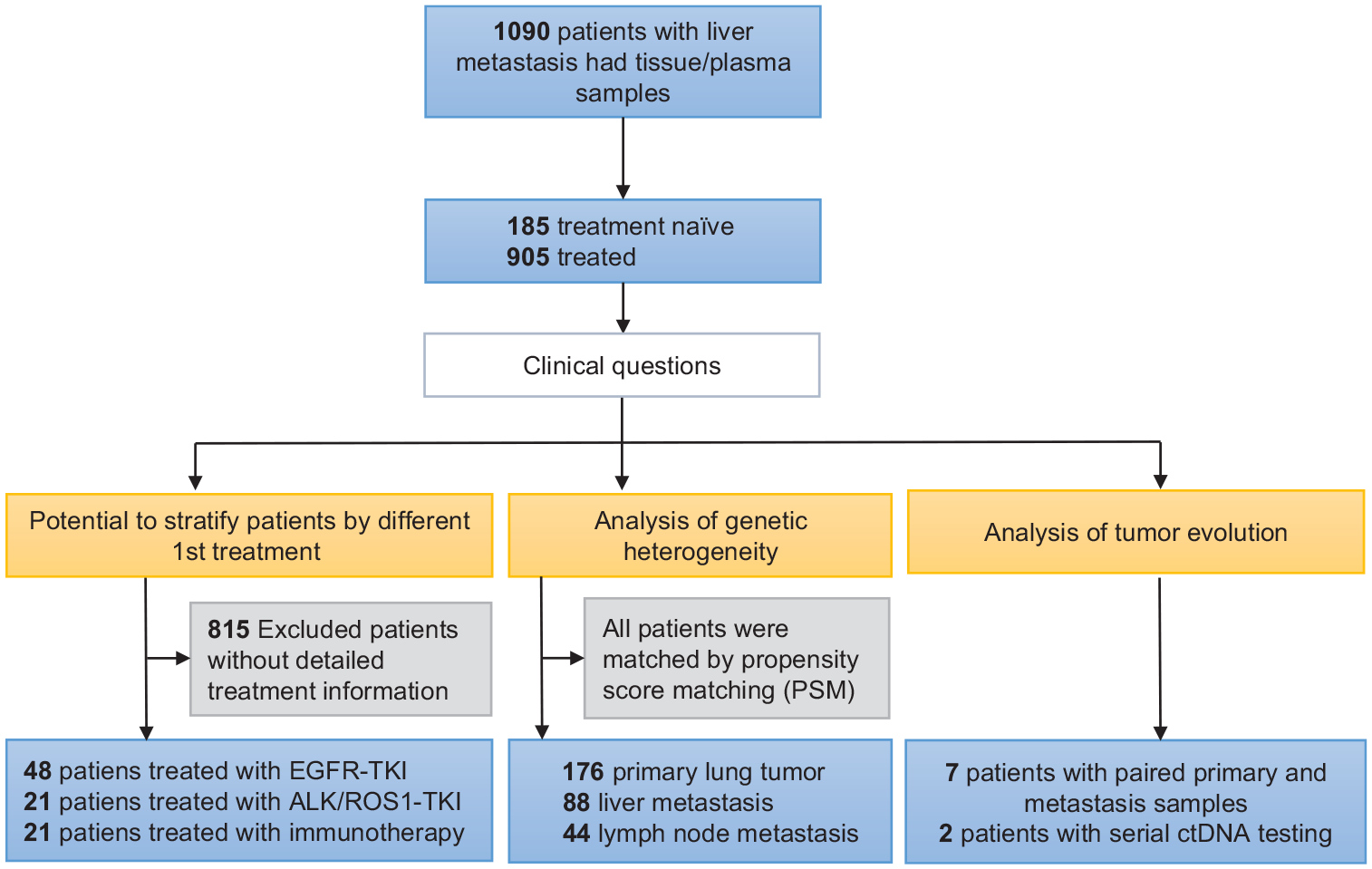
Study design.
Patient characteristics.
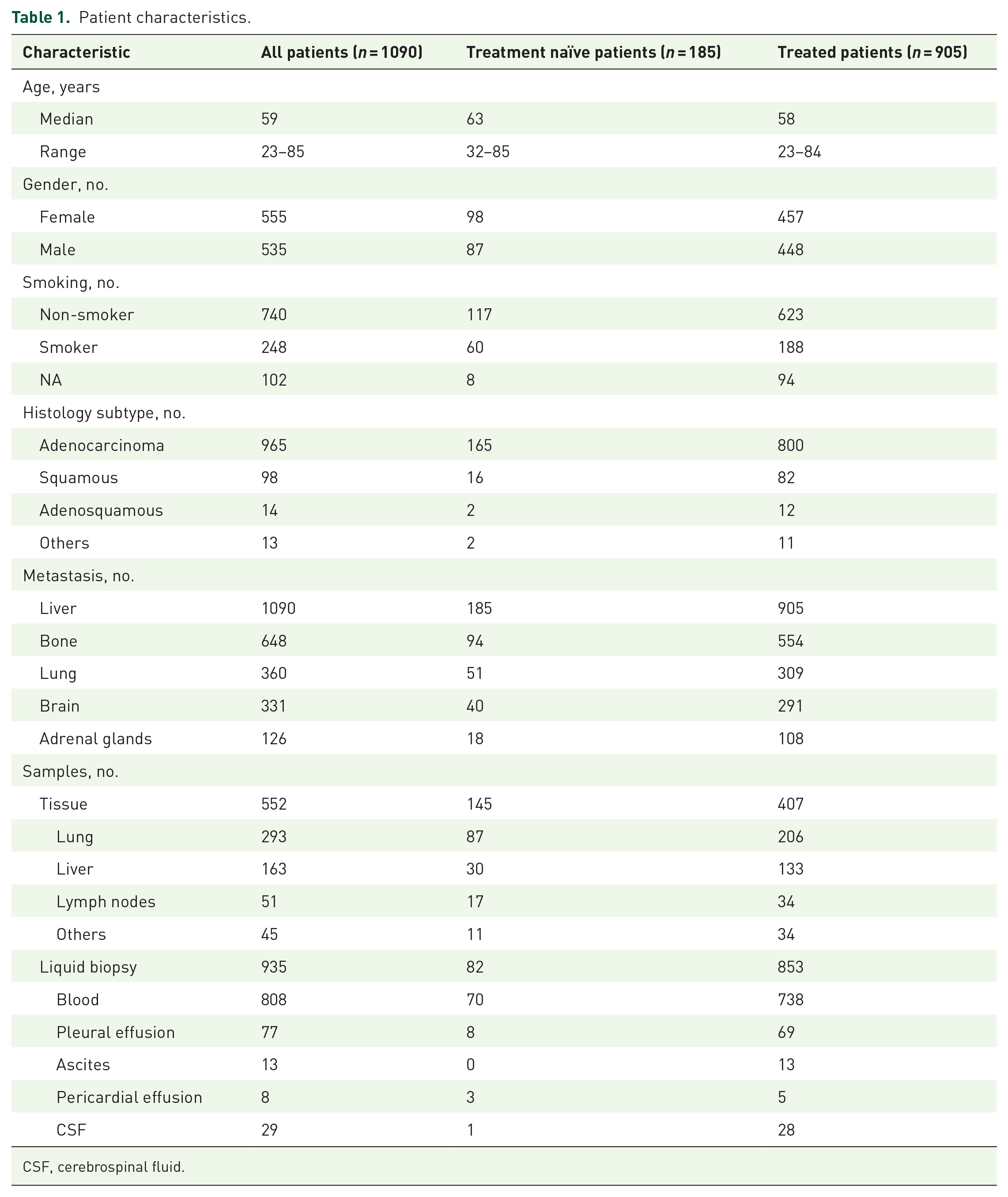
CSF, cerebrospinal fluid.
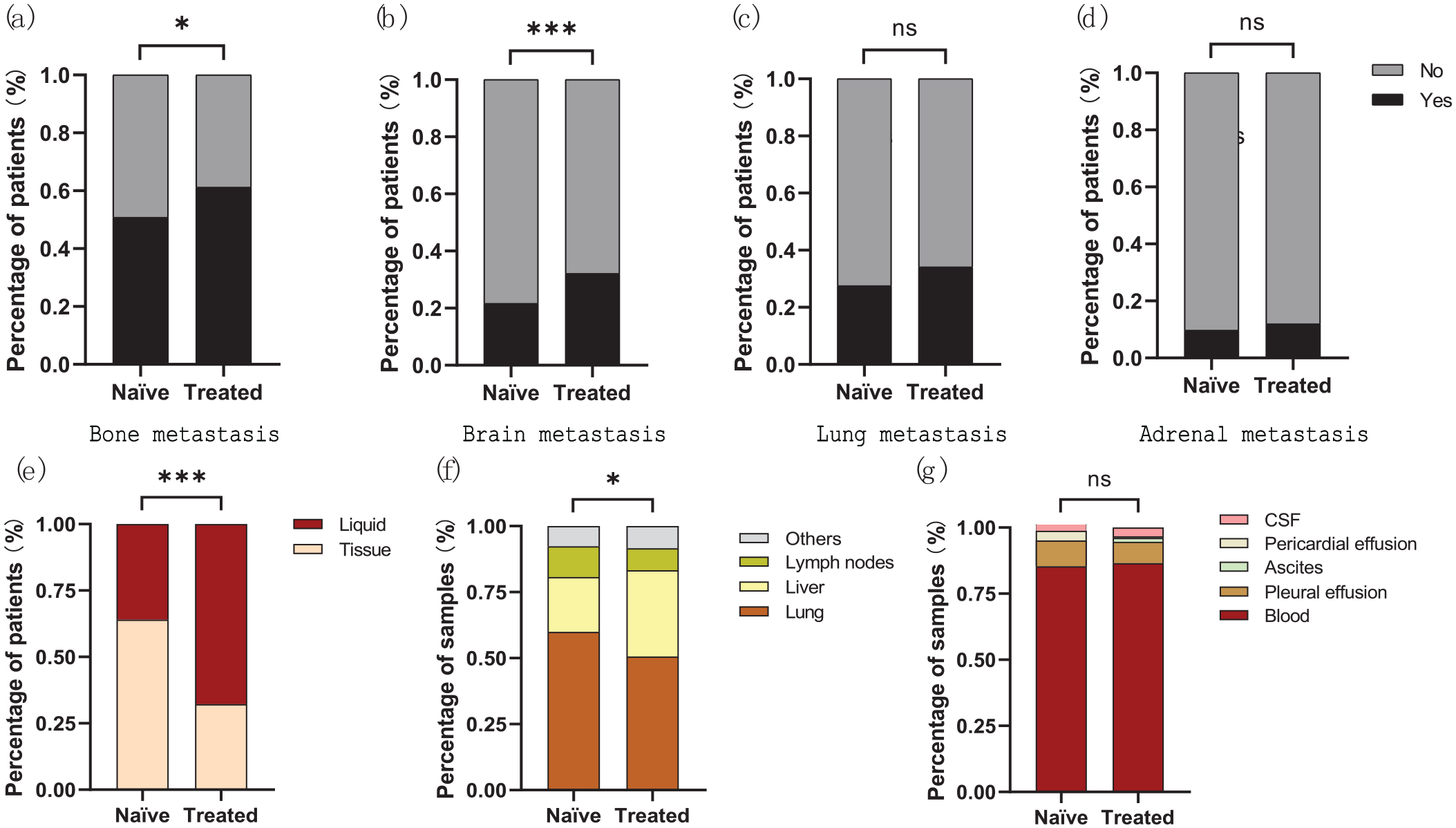
Metastases and choice of biopsy for mutation testing in the patient cohort (n = 1090). ((a)–(d)) Bone, brain, lung, and adrenal metastasis in treatment-naïve patients (n = 185) versus treated patients (n = 905); ((e)–(g)) Choice of liquid or tissue biopsy, specific tissue biopsy including lung, liver, lymph nodes and others (f), specific liquid biopsy including blood, pleural effusion, ascites, pericardial effusion, and CSF in treatment-naïve patients versus treated patients.
In total, there were 1487 samples from the 1090 patients who underwent mutation analysis. The 552 tissue biopsy samples included 293 from primary lung lesions, 163 from liver metastasis, 51 from lymph nodes, and 45 from other metastases such as pleura and bone. The 935 liquid biopsy samples were composed of 808 blood ctDNA, 77 pleural effusion, 13 ascites, 8 pericardial effusions, and 29 cerebrospinal fluid. Treatment-naïve patients used tissue biopsy more often than liquid biopsy, while treated patients used liquid biopsy more commonly (145/227 vs 407/1260, p < 0.001, Figure 2(e)). Moreover, treatment-naïve patients had more lung biopsy (87/145 vs 206/407, p = 0.05) and less liver biopsy (30/145 vs 133/407, p < 0.001) compared with treated patients (Figure 2(f)), but similar choice of liquid biopsy (Figure 2(g)).
Liver metastasis is associated with poor therapeutic response to targeted therapy of EGFR/ALK/ROS1 and immune checkpoint inhibitors
In our cohort, there were 560 patients underwent EGFR-TKI therapy, 246 patients got chemotherapy, 79 patients took ALK-TKI, 12 patients had other targeted therapies including ROS1-TKI or RET-TKI, and finally, 8 patients were treated with immune checkpoint inhibitor monotherapy (Figure 3(a)). We further compared the 158 samples with programmed death-ligand 1(PD-L1) expression data available and found that there was a numerical but not significantly higher percentage of PD-L1 expression in the lung samples (n = 95) than in liver samples (n = 47) or lymph nodes samples (n = 16, Figure 3(b)).

Liver metastasis is associated with poor therapeutic response. (a) Therapeutic history of the treated patients (n = 905); (b) PD-L1 expression in lung (n = 95), liver (n = 47), LN (n = 16); ((c)–(e)) Time to treatment discontinuation (TTD) of first-line EGFR-TKI (n = 48 (c)), ALK/ROS1-TKI (n = 21 (d)), and immunotherapy (n = 21 (e)).
We then compared the mutation spectrum of liver metastasis samples with or without treatment and found there was no difference in either SNV or copy number variant (CNV). Considering the complex treatment strategies, we separated patients into EGFR-TKI, ALK/ROS1-TKI, and immunotherapy (with or without chemotherapy) groups and analyzed the time to treatment discontinuation (TTD) of first-line EGFR-TKI (n = 48), ALK/ROS1-TKI (n = 21), and immunotherapy (n = 21). There were a few patients treated with anti-PD-1/PD-L1 after resistance to EGFR-TKI (n = 22), or ALK/ROS1-TKI (n = 6). Most patients treated with anti-PD-1/PD-L1 were combined with chemotherapy (n = 53). By reviewing previous therapeutic history, we were able to identify patients who had no liver metastasis when initiating the treatment, thus defined as having no liver metastasis before treatment. Consistent with previous reports,14–17 patients without liver metastasis when initiated the targeted therapy or immunotherapy had longer treatment, with mTTD of 15 months versus 8 months, HR 2.69, 95% CI 1.42–5.09, p < 0.001 for EGFR-TKI (Figure 3(c)); mTTD of 11 months versus 5 months, HR 2.77, 95% CI 0.97–7.86, p < 0.001 for ALK/ROS1-TKI (Figure 3(d)); and mTTD of 11.5 months versus 5 months, HR 2.62, 95% CI 1.09–6.32, p = 0.01 for immunotherapy (Figure 3(e)).
The mutational landscape in primary tumors and liver or lymph node metastases
A total of 4249 mutations were found in the primary lung tumor (n = 224), liver (n = 115), and lymph nodes (n = 44) samples, including 3134 SNV, 1021 CNV, and 94 fusions/rearrangements. EGFR and TP53 were the top two most frequently mutated genes, among which EGFR accounted for 53% of all samples and TP53 accounted for 67% (Supplemental Figure S1).
To understand the effect of metastasis on genetic heterogeneity, we then compared the mutation spectrum between primary tumor, liver metastasis, and lymph node metastasis. After matching, there were no statistically significant differences in baseline characteristics between primary tumor, liver metastasis, and lymph node metastasis (Supplemental Table S2). In comparison with primary tumors, liver metastatic samples had a higher proportion of ALK (13.64% vs 5.11%, p = 0.03), CTNNB1 (10.23% vs 2.27%, p = 0.11), FLCN (4.55% vs 0.57%, p = 0.04), MAX (4.55% vs 0.57%, p = 0.03), PIK3CG (7.95% vs 1.70%, p = 0.03), NRXN1 (3.41% vs 0.00%, p = 0.04), PBRM1 (3.41% vs 0.00%, p = 0.04), and TOP1 (3.41% vs 0.00%, p = 0.04) mutations, but a lower proportion of CDKN2A (1.14% vs 7.95%, p = 0.02) and LRP1B (3.41% vs 11.93%, p = 0.04) mutations (Figure 4(a)). And lymph node metastatic samples had a higher prevalence of ACIN1 (9.09% vs 1.70%, p = 0.03), PIK3CG (9.09% vs 1.70%, p = 0.03), CBL (4.55% vs 0.00%, p = 0.04), FOXP1 (4.55% vs 0.00%, p = 0.04), JAK3 (4.55% vs 0.00%, p = 0.04), MEN1 (6.82% vs 0.00%, p = 0.007), SLX4 (4.55% vs 0.00%, p = 0.04), SOX9 (4.55% vs 0.00%, p = 0.04), TLR4 (4.55% vs 0.00%, p = 0.04) mutations, as well as a lower proportion of KRAS (0.00% vs 10.80%, p = 0.02) mutations than primary tumors (Figure 4(a)). In addition, lymph node metastatic samples had a higher prevalence of ACIN1 (0.09% vs 0.00%, p = 0.01) and LRP1B (15.09% vs 3.41%, p = 0.02) mutations than liver metastatic samples (Figure 4(a)). In terms of CNV, there was higher percentage of C11orf30 (4.55% vs 0.57%, p = 0.04), CD274 (5.68% vs 0.57%, p = 0.02), CDK6 (5.68% vs 1.14%, p = 0.02), CDKN2A (22.73% vs 9.66%, p = 0.007), CDKN2B (20.45% vs 9.09%, p = 0.02), EGFR (30.68% vs 18.75%, p = 0.04), EXT1 (4.55% vs 0.57%, p = 0.04), FANCD2 (4.55% vs 0.57%, p = 0.04), FGF3 (6.82% vs 1.14%, p = 0.02), FGF4 (6.82% vs 0.57%, p = 0.006), FGFR1 (5.89% vs 0.68%, p = 0.02), IL7R (4.55% vs 0.57%, p = 0.04), MET (14.77% vs 3.98%, p = 0.002), PDCD1LG2 (6.82% vs 1.14%, p = 0.02), RAC1 (6.82% vs 1.14%, p = 0.02), WRN (9.09% vs 1.70%, p = 0.007), AKT1 (3.41% vs 0.00%, p = 0.04), FLCN (3.41% vs 0.00%, p = 0.04), MLH1 (4.55% vs 0.00%, p = 0.01), and SMAD4 (4.55% vs 0.00%, p = 0.01), but lower proportion of FGFR1 (0.00% vs 5.11%, p = 0.03) mutations in the liver metastatic samples than that of primary tumor of lung (Figure 4(b)). However, only EXT1 (6.82% vs 0.57%, p = 0.03), CCND3 (4.55% vs 0.00%, p = 0.04), EZH2 (4.55% vs 0.00%, p = 0.04) and WT1 (4.55% vs 0.00%, p = 0.04) mutations showed a higher incidence of CNV in the lymph nodes compared with lung lesion (Figure 4(b)). There were no significant differences between CNVs in liver metastatic and lymph node metastatic samples (Figure 4(b)). In addition, liver metastatic samples had a higher number of somatic cell mutations and higher tumor mutational burden (TMB) compared to primary tumor and lymph node metastasis samples, although there was no difference in tumor cell content among them (Supplemental Figure S2).

The genomic landscape of NSCLC patients with liver metastasis. (a) SNV in primary lung tumor versus liver metastasis or LN and (b) CNV in primary lung tumor versus liver metastasis or LN.
Subgroup analysis was performed according to the presence or absence of actionable alterations after all matched patients had balanced baseline clinical characteristics (Supplemental Tables S3 and S4). For NSCLCs with actionable drivers, liver metastatic samples had a higher proportion of SMARCA4 (7.53% vs 1.97%, p = 0.04) mutations, but lower proportion of CDKN2A (0.00% vs 5.23%, p = 0.03), ERBB2 (1.08% vs 7.19%, p = 0.03) mutations than primary tumors (Supplemental Figure S3A). And lymph node metastatic samples had a higher prevalence of CBL (5.71% vs 0.00%, p = 0.03), FOXP1 (5.71% vs 0.00%, p = 0.03), HDAC4 (5.71% vs 0.00%, p = 0.03), INPP4B (5.71% vs 0.00%, p = 0.03), MEN1 (5.71% vs 0.00%, p = 0.03), MYCN (5.71% vs 0.00%, p = 0.03), and SLX4 (5.71% vs 0.00%, p = 0.03) mutations than primary tumors (Supplemental Figure S3A). In addition, lymph node metastatic samples had a higher prevalence of ACIN1 (5.71% vs 0.00%, p = 0.049), CBL (5.71% vs 0.00%, p = 0.049), FAM123B (5.71% vs 0.00%, p = 0.049), FOXP1 (5.71% vs 0.00%, p = 0.049), MEN1 (5.71% vs 0.00%, p = 0.049), MLH1 (5.71% vs 0.00%, p = 0.049), MS4A1 (5.71% vs 0.00%, p = 0.049), and MYCN (5.71% vs 0.00%, p = 0.049) mutations than liver metastatic samples (Supplemental Figure S3A). Also, there was a higher percentage of C11orf30 (5.38% vs 0.65%, p = 0.03), CD274 (6.45% vs 0.00%, p < 0.001), CDKN2A (26.88% vs 9.15%, p = 0.001), CDKN2B (22.58% vs 7.84%, p = 0.04), FGF3 (7.53% vs 1.31%, p = 0.005), FGF4 (7.53% vs 0.65%, p = 0.006), FGFR19 (7.53% vs 1.96%, p = 0.03), MET (16.13% vs 5.23%, p = 0.02), MLH1 (5.38% vs 0.00%, p = 0.002), PDCD1LG2 (6.45% vs 0.00%, p = 0.007), SMAD4 (4.30% vs 0.00%, p = 0.002), and WRN (8.60% vs 1.96%, p = 0.02) mutations in the liver metastatic samples than that of primary tumor of lung (Supplemental Figure S3B). However, only EZH2 (5.71% vs 0.00%, p = 0.03) and WT1 (5.71% vs 0.00%, p = 0.03) mutations showed a higher incidence of CNV in the lymph nodes compared with lung lesions (Supplemental Figure S3B). Compared to lymph node metastatic samples, liver metastatic samples had a higher proportion of MYC (24.73% vs 8.57%, p = 0.049) mutation (Supplemental Figure S3B). For NSCLCs without actionable drivers, liver metastatic samples had a higher proportion of FLCN (13.64% vs 0.00%, p = 0.01) and PIK3CG (13.64% vs 1.41%, p = 0.04) mutations, but a lower proportion of MLL2 (0.00% vs 22.54%, p = 0.04) mutations than primary tumors (Supplemental Figure S3C). And lymph node metastatic samples had a higher prevalence of POLE (33.33% vs 1.41%, p = 0.003), SOX9 (22.22% vs 1.41%, p = 0.03), and SYK (22.22% vs 1.41%, p = 0.03) mutations than primary tumors (Supplemental Figure S3C). There were no significant differences between CNVs in primary lung tumors, liver metastatic, and lymph node metastatic samples (Supplemental Figure S3D).
As seven patients (P0134, P0172, P0699, P0862, P0914, P1012, and P0308) had paired primary lung tumor and liver metastasis samples available, we compared the mutation spectrums of them individually (Table 1). As expected, all seven patients had shared driver mutations between lung and liver lesions, including TP53, EGFR, EML4-ALK, and CD74-ROS1. Six of the seven liver metastatic samples also had additional CNVs which also indicated the potential role of CNVs in the development of liver metastasis (Figure 5(a)). By contrast, no patient had paired primary lung tumor and lymph nodes available. To use these liquid biopsy samples for mutational profiling and clinical decision-making, it is necessary to compare the mutational spectrum of liquid biopsy samples and tissue samples. In our cohort, there were 11 patients had paired ctDNA and lymph nodes samples collected simultaneously. As expected, CNVs were less common in ctDNA; however, only six ctDNA samples had ctDNA private mutations, while all the lymph node samples had private mutations including CNV and SNV (Figure 5(b)). Moreover, both shared and private mutations were also present between paired tissue and other liquid biopsy samples (Supplemental Figure S4).

Comparison of mutations in patients with paired samples. (a) Primary lung tumor versus liver metastasis; (b) LN versus ctDNA; and (c) the phylogenetic tree of somatic mutations for three tumors from a patient.
For further analysis of the evolutionary relationship between liver metastasis and lymph node metastasis, we conducted specific research on a phylogenetic tree (Figure 5(c)). The numbers of all somatic mutations in P0949 are labeled above the tree. The phylogenetic tree constructed from these data located the two tumors in two independent evolutionary branches.
Mutation evolution analysis with ctDNA
Liquid biopsy with plasma ctDNA is increasingly adopted in clinical practice, it is informative in the study of mutation evolution. Thus, we retrospectively analyzed two patients with four serial mutation tests (Figure 6(a) and (b)).

ctDNA analysis of mutation evolution. (a) Gain of EGFR-resistant mutations in P0318; (b) gain of combined BRAF, KRAS-resistant mutations in P0031; (c) ctDNA indicated early relapse of cancer in P0244, the gain of BRAF-resistant mutations during the EGFR-targeted therapy.
The patient P0318 was an 80-year-old male, and he was diagnosed with lung adenocarcinoma with bone and brain metastasis. The brain metastasis was treated with radiotherapy and icotinib was given as the first-line therapy as EGFR 19indel mutation was detected. At M10, the lung lesion progressed and an EGFR T790M was identified in ctDNA, so the patient switched to osimertinib. A partial response (PR) was achieved with clearance of ctDNA at M15; however, liver metastasis occurred at M22 and 2 EGFR C797S mutations (c.2389T>A, c.2390G>C) were detected. Then the patient was treated with combined osimertinib and chemotherapy/antiangiogenetic therapy. With the best response of stable disease (SD), the patient progressed at M31 with an additional EGFR CNG identified in the ctDNA (Figure 6(a)).
P031 was a 59-year-old male, and he was diagnosed with lung adenocarcinoma with bone, liver, and spleen metastasis. As both lung biopsy and ctDNA detected EGFR 19indel mutation, the patient was treated with afatinib. At M3, the patient reached PR in the lung lesion, and ctDNA showed no mutation detected. However, at M16, a BRAF V600E was detected though the patient was still on SD. At M20, an additional KRAS G12D mutation was detected and the patient switched to combined afatinib and chemotherapy (Figure 6(b)).
P0244 was a 41-year-old female, and she was diagnosed with lung adenosquamous carcinoma (pT1cN0M0, stage I). As vessel carcinoma embolus, tumor cells spread through air spaces, and a high Ki67 index (70%) was noticed in the surgically removed tumor, the patient underwent ctDNA after surgery. An NF2 mutation was detected in the ctDNA and thus the patient was treated with combined osimertinib and chemotherapy. At M1 after surgery, a liver lesion was noticed, and the patient kept on combined osimertinib and chemotherapy with the best response of PR. At M11, bone metastasis occurred and radiotherapy of the bone metastasis was added. At M15, intrapulmonary metastasis occurred, the liver metastasis enlarged, and a BRAF V600E mutation was detected in cfDNA (Figure 6(c)).
Discussion
In this large cohort of NSCLC patients with liver metastasis, we retrospectively analyzed the real-world choice of biopsy for mutation testing, compared the therapeutic efficacy of EGFR/ALK/ROS1-targeted therapy and anti-PD1/PD-L1 immunotherapy in patients with primary or secondary liver metastasis. We also described the mutation spectrum of primary lung tissue, liver biopsy, and lymph node biopsy, and finally tried to decipher the evolution of tumor mutations in patients with serial tissue or ctDNA testing.
In our cohort, we found there were more liquid biopsies than tissue biopsies, especially in the treated patients (Table 1 and Figure 2(e)). This indicated that more and more advanced NSCLC patients would accept liquid biopsy since repeated tissue biopsy was not always applicable. In terms of tissue biopsy, the majority of patients in our cohort would choose the primary lung tumor as the tissue biopsy, with liver and lymph nodes as the second and third common choice (Figure 2(f)).
Among the 905 treated patients, there were 560 patients treated with EGFR-TKI. The high incidence was probably due to the prevalence of EGFR actionable mutation in Asia and more patients with EGFR-TKI therapy would have additional mutation testing when the disease progressed. As expected, most patients treated with anti-PD-1/PD-L1 were combined with chemotherapy, and only a few patients treated with anti-PD-1/PD-L1 after resistance to EGFR-TKI. Consistent with previous reports, He et al., 14 Wu et al., 15 Jiang et al., 16 and Funazo et al. 17 report that liver metastasis was associated with poor therapeutic response to targeted therapy of EGFR and immune checkpoint inhibitors (Figure 3(c) and (e)). We also found that the time to treatment discontinuation (TTD) of ALK/ROS1-TKI (n = 21) in patients with liver metastasis before starting ALK/ROS1-TKI treatment was shorter than that without liver metastasis (Figure 3(d)).
By comparing the 88 liver biopsies and 44 lymph node biopsies with the primary lung tumor (n = 176), liver and lymph node metastatic samples were found to have a generally higher mutation level of SNV and CNV than the primary tumor (Figure 4(a) and (b)). This finding was further confirmed in the seven patients with paired primary lung tumor and liver metastasis samples (Figure 5(a)). Given that CNV is an important indicator of chromosomal instability (CIN) and previous publications suggested that CIN is associated with metastasis,36–38 it is worthwhile to investigate the significant role of CIN in the formation and process of liver, and lymph node metastasis in the future. In tumors with actionable alterations, the mutation level of SNV and CNV was generally higher in liver and lymph node metastasis samples than in primary tumors (Supplemental Figure S3A and S3B). For different NSCLCs without actionable drivers, liver and lymph node metastasis samples had higher mutation levels of SNV than primary tumors, but comparable CNV (Supplemental Figure S3C and S3D). Therefore, it is likely that the differences in actionable alterations are the main driver of the observed differences in primary lung tumor, liver, and lymph node metastatic samples. When comparing the 11 paired ctDNA and lymph node samples collected simultaneously, all the lymph node samples had private mutations including CNV and SNV, while only six ctDNA samples had ctDNA private mutations (Figure 5(b)). As reported previously, ctDNA-based sequencing was of sufficient sensitivity and specificity in SNVs but inferior performance in CNVs due to a high level of biases and artifacts from limited ctDNA fraction in the circulating free DNA (cfDNA).39–41 The discrepancy of SNV in paired ctDNA and lymph nodes is probably due to the following reasons: (1) not all lymph node metastatic tumor cells shed DNA into the peripheral blood and (2) lymph node metastatic tumor cells might not further metastasize to other organs as suggested by a previous study that the lymph node dissemination time was relatively early, and non-lymph node metastases were mainly seeded by the primary tumor rather than the earlier colonized lymph node metastases. 42 This notion was also consistent with the finding in the TRACERX study that perhaps less than 20% of primary lymph node metastases seed progressive disease.30–32
In both the paired samples (Figure 5) and mutation evolution analysis (Figure 6), we noticed the consistency of functional driver mutations in different samples of the same patients. It was also consistent with the previous study that liver metastasis presented with a far less discrepant mutational landscape than brain metastasis when compared with their primary tumors. The authors suggested that the evolutionary trajectory in lung adenocarcinoma of liver metastasis is more likely to have a linear model, while brain metastasis is more likely to have a parallel evolutionary model. 23 Collectively, our study also suggests that liver metastases are, to a certain extent, the relative early product of the evolution of primary tumors. 43
There are several limitations in this study. First, it was retrospective, and the patient population might be biased; thus, prospective studies are required to verify our findings. Second, as it is really hard to find a large cohort of patients who have matched primary tumor samples and multiple metastatic samples, we could only compare the mutation spectrum between different groups of patients and in patients whenever possible with paired samples. Nevertheless, the number of liver biopsy samples (n = 115) and lymph nodes (n = 44) was relatively high in the literature. Besides, there were no other matched lymph nodes and liver biopsy comparisons presented to provide further evidence of genetic divergence except for patient P0949. Third, all the patients used the NGS panel covering 1021 cancer-related genes instead of whole exome sequencing (WES), which is not commonly used in the clinic, to characterize the genetic alterations of the primary and metastatic samples. This might miss some other uncommon variants. Finally, due to its retrospective nature, we could not collect all the information on treatments and their efficacy, so TTD was used instead of PFS for analyzing the clinical outcome.
Conclusion
In summary, we explored the clinical choices of biopsy samples in this retrospective large cohort of NSCLC patients with liver metastasis and found that the mutation spectrum, especially the driver mutations of liver and lymph nodes was pretty much similar to the primary lung tumor. Moreover, liver metastases might be mainly seeded by the primary tumor rather than the earlier colonized lymph node metastases. Moreover, patients with liver metastasis are often accompanied by poor long-term prognosis. Therefore, more research is also needed to find better treatment options. The findings of this study provide new insights into liver metastasis and advocate for the development of clinical intervention strategies against advanced NSCLC patients.
Supplemental Material
sj-docx-1-tam-10.1177_17588359241275421 – Supplemental material for Molecular features of NSCLC patients with liver metastasis
Supplemental material, sj-docx-1-tam-10.1177_17588359241275421 for Molecular features of NSCLC patients with liver metastasis by Jun Zhao, Jia Zhong, Yujie Chen, Zipei Chen, Huan Yin, Yuange He, Rongrong Chen and Renhua Guo in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-docx-2-tam-10.1177_17588359241275421 – Supplemental material for Molecular features of NSCLC patients with liver metastasis
Supplemental material, sj-docx-2-tam-10.1177_17588359241275421 for Molecular features of NSCLC patients with liver metastasis by Jun Zhao, Jia Zhong, Yujie Chen, Zipei Chen, Huan Yin, Yuange He, Rongrong Chen and Renhua Guo in Therapeutic Advances in Medical Oncology
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
