Abstract
Background:
Triple-negative breast cancer (TNBC) includes approximately 20% of all breast cancer and is characterized by its aggressive nature, high recurrence rates, and visceral metastasis. Pathological complete response (pCR) is an established surrogate endpoint for survival. The window of opportunity studies provide valuable information on the disease biology prior to definitive treatment.
Objectives:
To study the association of dynamic change in pathological, imagining, and genomic biomarkers that can prognosticate pCR. The study aims to develop a composite prognostic score.
Design:
Clinical, interventional, and prognostic biomarker study using the novel window of opportunity design.
Methods:
The study aims to enroll 80 treatment-naïve, pathologically confirmed TNBC patients, administering a single dose of paclitaxel and carboplatin during the window period before neoadjuvant chemotherapy (NACT). Tumor tissue will be obtained through a tru-cut biopsy, and positron emission tomography and computed tomography scans will be performed for each patient at two time points aiming to evaluate biomarker alterations. This will be followed by the administration of standard dose-dense NACT containing anthracyclines and taxanes, with the study culminating in surgery to assess pCR.
Results:
The study would develop a composite prognostic risk score derived from the dynamic change in the Ki-67, tumor-infiltrating lymphocytes, Standardized Uptake Value (SUV max), Standardized Uptake Value for lean body mass (SUL max), and gene expression level pre- and post-intervention during the window period prior to the start of definitive treatment. This outcome will aid in categorizing the disease biology into risk categories.
Trial registration:
The current study is approved by the Institutional Ethics Committee [Ethics: Protocol. no. JIP/IEC/2020/019]. This study was registered with ClinicalTrials.gov [CTRI Registration: CTRI/2022/06/043109].
Conclusion:
The validated biomarker score will help to personalize NACT protocols in patients in TNBC planned for definitive treatment.
Plain language summary
We are investigating new ways to predict how well a particular treatment will work in patients with a specific type of breast cancer called triple-negative breast cancer. The study goal is to find biomarkers that change in response to drugs to predict the complete elimination of cancer in patients before it spreads to other parts of the body. To do this, we are using a special research approach called a ‘window of opportunity design.’ This information could be valuable in personalizing and improving cancer treatments.
Keywords
Introduction
Triple-negative breast cancer (TNBC) includes approximately 20–15% of all breast cancer and is characterized by its aggressive nature, high recurrence rates, and tendency to metastasize to the brain. 1 Although targeted therapies for TNBC are currently limited, around one-third of patients with TNBC can achieve a pCR with standard taxane or anthracycline chemotherapy, which is considered the standard of care. 2 pCR changes significantly among various breast cancer subtypes and is an established surrogate endpoint for survival in triple-negative and her2neu-positive breast cancer individuals. 3 The pCR is identified only after the neoadjuvant therapy is given and surgery performed. 1 The pCR rates in TNBC depend on the drugs used varying from 30% with anthracyclines and taxanes to 45% with the addition of platinum and 60% with the addition of immunotherapy. 2 Ki-67, tumor-infiltrating lymphocytes (TILs), and gene expression studies have been validated in different studies as prognostic markers and their association with pCR. Noninvasive biomarkers like positron emission tomography (PET) scan (SUV max) which exploits the metabolic activity of the tumor have recently been used in breast cancer to predict response to chemotherapy. 4
Several studies stated that patients reaching a pathological complete response (pCR) post-NACT have shown greater outcomes in comparison with those who failed to achieve it. 5 Thus, the convenience of NACT is the rapid determination of tumor response to treatment. 6 The discovery of the taxanes such as paclitaxel and docetaxel and understanding of their cytotoxicity action through cell cycle arrest by tubulin stabilization have reformed breast cancer treatment. 7 In the window of opportunity study design, newly diagnosed cancer patients awaiting neoadjuvant chemotherapy (NACT) followed by surgery receive an interventional drug during the Window of Opportunity (WoO) period which is between diagnosis and before definitive treatment. 8 The WoO studies can give information about the therapeutic efficacy and biological effects of new therapeutic techniques. 9
In the context of NACT, achieving a pCR is a significant indicator of improved survival. However, the identification of reliable biomarkers prior to NACT that can predict which patients are likely to achieve pCR remains a challenge. This proposed study seeks to bridge this by investigating a comprehensive set of pathological, imaging, and molecular biomarkers early in the treatment protocol. By utilizing the window of opportunity design, the study aims to assess how the change in these biomarkers correlates with pCR in TNBC patients and the ultimate goal is to tailor personalized treatment strategies based on a patient’s individual biomarkers profile, thereby enhancing therapeutic efficacy and patient outcome.
Review of literature
The commonly studied baseline and dynamic pathological, molecular, and imaging prognostic biomarkers in terms of response and survival are described in Table 1.
Prognostic biomarkers in conventional and WoO breast cancer trials.
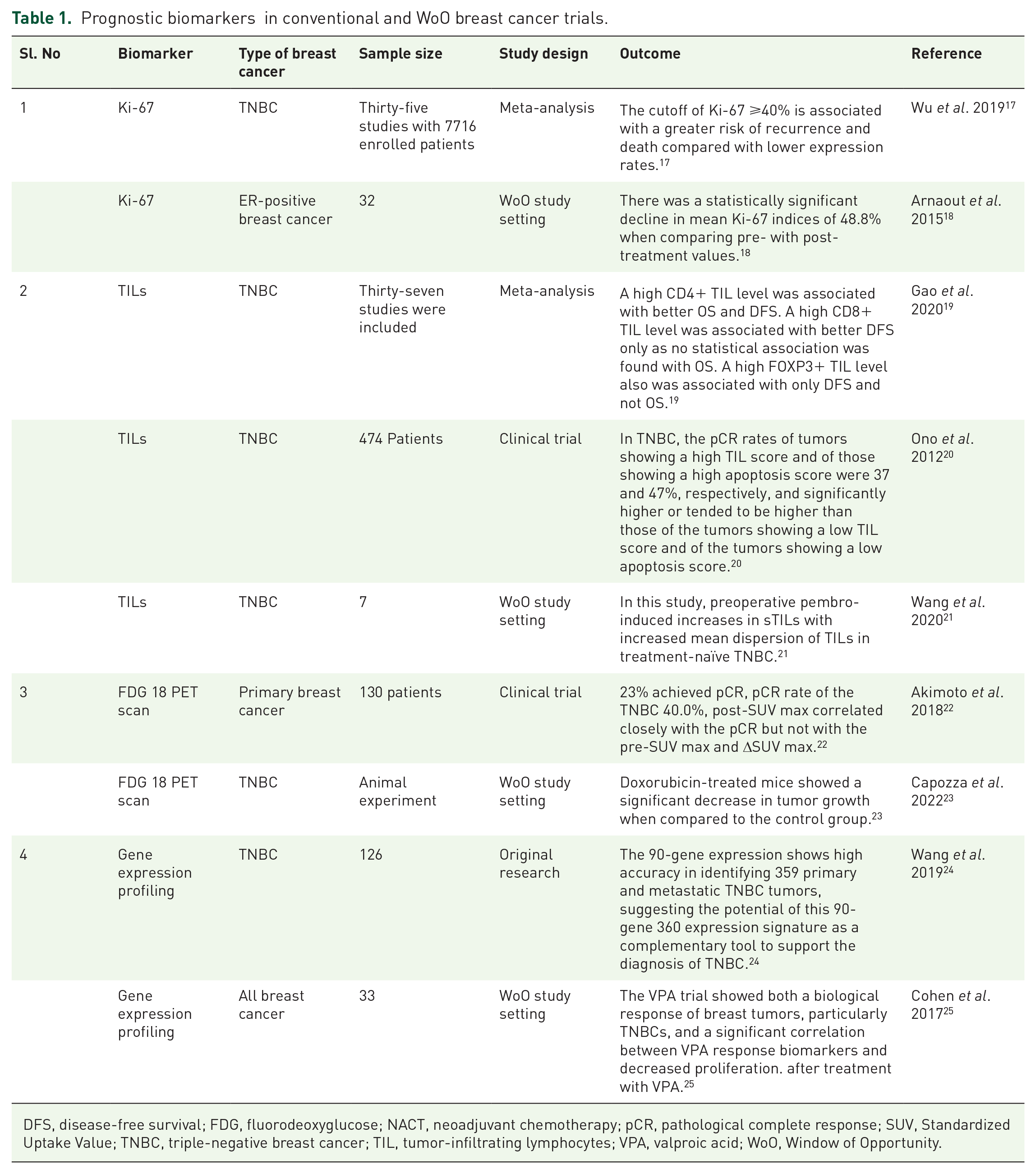
DFS, disease-free survival; FDG, fluorodeoxyglucose; NACT, neoadjuvant chemotherapy; pCR, pathological complete response; SUV, Standardized Uptake Value; TNBC, triple-negative breast cancer; TIL, tumor-infiltrating lymphocytes; VPA, valproic acid; WoO, Window of Opportunity.
Ki-67
Ki-67 expression is strongly associated with aggressive tumor biology and tumor proliferation, and recognition has grown for Ki-67 as an excellent prognostic biomarker. 10 High Ki67 would predict the increased proliferation of breast cancer cells and could be considered a prognostic marker. 11 Penault-Llorca et al. 12 and Breast Cancer International Research Group 001 trial recently reported that high levels of Ki-67 were predictive of benefit from adding docetaxel to fluorouracil and epirubicin chemotherapy as adjuvant treatment for patients with Estrogen receptor (ER)-positive tumors in the PACS01 randomized trial.
Thus, the data on the identification of patients benefiting from chemotherapy require confirmation before the use of Ki-67 reaches clinical utility.
Tumor-infiltrating lymphocytes
Several studies showed that the tumor microenvironment is one of the driving factors of tumor progression and invasion. 13 High TIL counts at baseline and a significant reduction in TIL counts after neoadjuvant therapy are associated with higher pCR rates. 14
Gene expression profiling
Early gene expression studies are crucial for targeting TNBC due to its molecular heterogeneity.The common genes and pathways regulated in breast cancer are described in Table 2. They identify subtypes and predict treatment response. Early gene expression profiling facilitates the selection of treatments based on individual genetic makeup and tumor characteristics, leading to improved outcomes and reduced side effects. Studies on gene expression offer promising avenues for better patient stratification, advanced targeted therapies, and significant advancements in breast cancer treatment. Molecular profiling is a promising diagnostic approach that has the potential to provide an objective classification of metastatic cancers with an uncertain or unknown tissue of origin and to facilitate more time- and cost-effective diagnostic work-up of cancer patients. 15
Common genes and their regulation in breast cancer.

PET-computed tomography scan
A phase II study by Connolly et al. correlating SUV with pCR to pertuzumab and trastuzumab has demonstrated that early changes in SUV max by D15 of the first cycle in Her2neu-positive breast cancer correlated with pCR. 16
Experimental design
Hypothesis:
Reduction in Ki67 expression & SUV value of the tumor and alteration gene expression profile (GEP) following single dose of paclitaxel and carboplatin prior to neoadjuvant chemotherapy during window of opportunity period may predict pCR in triple negative breast cancer.
Objective
To study the association of change in prognostic biomarkers Ki-67, TILs, FDG PET-computed tomography (CT) scan, and gene expression profiling with pCR in patients with non-metastatic TNBC following single-dose paclitaxel and carboplatin during the window period before NACT.
To develop a composite prognostic score using the above parameters for the assessment of pCR in TNBC before NACT.
Study design
Clinical, interventional, and prognostic biomarker study using the novel window of opportunity design.
Intervention
A single weekly dose of carboplatin (AUC-2) and paclitaxel (80 mg/m2) will be given to patients as an intervention during the WoO period (diagnosis and starting of definite treatment which is usually injection paclitaxel 80 mg/m2 in 1 unit NS in Glass bottle IV infusion over 1 h using Di(2-ethylhexyl) phthalate (DEHP) free set and 0.2-µ filter. Injection carboplatin AUC 2 was calculated using Calverts’ formula in 1 unit 5% dextrose intravenous (IV) infusion over 1 h. Dose (mg) = (GFR + 25) * 2, GFR is calculated using the Cockcroft–Gault formula.
Study setting
This is an exploratory study that uses the window of opportunity design to identify the chemo sensitiveness of the tumor before starting NACT to assess biomarkers early which can predict pCR post-NACT, thereby tailoring treatment in TNBC. Triple negativity is defined as ER and PR (0 or <1%) or Alred score of 0 and 2 HER 2 neu 0, 1+, if 2+ FISH, HER 2 no amplification. All patients fulfilling the inclusion, and exclusion criteria after obtaining consent will be included in the study. Baseline assessment of Ki-67, TILs, gene expression profiling and will be done on the initial biopsy specimen. A baseline PET scan will be performed. Patients will be receiving a single weekly dose of paclitaxel and carboplatin. On D15 of the chemotherapy, a second biopsy will be obtained to study the Ki-67, TILs, and gene expression. Before the biopsy, a hemogram is obtained the absolute neutrophil count of more than 1500 and platelets of more than 1 lakh would be the prerequisite before the second biopsy. A PET-CT scan will also be performed to study the changes in the SUV and SUL max of the tumor. Patients will then receive NACT with a standard anthracycline and taxane regimen (EC × 3 cycles followed by Doce × 4 cycles), later patients will undergo surgery. The post-surgery pathological response will be studied. pCR is a condition of a complete absence of invasive components in the axilla and breast. The study duration would be 3 years Figure 1.

Study design.
Study duration and registration: 3 years
Outcomes
Primary endpoint: pCR will be defined as an absence of invasive components in the axilla and breast (yp T0 N0).
Secondary endpoint: Disease-free survival (DFS) will be defined as the duration from cancer diagnosis to relapse or death due to any cause.
The expected outcome from this project is to identify pathological, imaging, and molecular biomarkers that would predict pCR in TNBC patients using the window period before NACT. We would develop a composite prognostic score that will enable us to identify patients who will achieve pCR. This risk score needs to be validated in a larger population. Once validated, we will prospectively use this score in a clinical trial setting to decide on chemotherapy intensity in TNBC patients planned for NACT.
Study population
Eligibility criteria
Inclusion criteria:
Age 18–65 years.
Newly diagnosed TNBC patients, as triple negative defined by - ER and PR score 0, HER2- negative (HER2 0 or 1+ on immunohistochemistry, or 2+ on immunohistochemistry and negative on fluorescence in situ hybridization FISH).
Early and locally advanced breast cancer (T2/T3/T4b, with or without node-positive).
ECOG PS 0/1.
Planned for neo-adjuvant chemotherapy with anthracycline and taxanes.
Bone marrow function-Hb >9 g%, absolute neutrophil count >1.5/mic L, platelet >1.5 lakhs/mic L.
Normal renal function with serum creatinine <1.5 mg%.
Normal liver function with serum bilirubin >1.5 mg%.
Normal cardiac function LV ejection fraction <50%.
Exclusion criteria:
Patient is not willing for the second biopsy.
Previously allergic to paclitaxel and carboplatin.
Pregnant women.
Breastfeeding mother.
Prior history of any other malignancy.
Bilateral breast cancer patients.
Patient has active local site infection.
Who will take informed consent?
Each patient diagnosed with TNBC will undergo preoperative screening to determine their eligibility based on specific inclusion and exclusion criteria. An authorized investigator will verbally convey to the patient the objectives of the study, procedures, and potential risks involved, as well as the process of treatment during the window of opportunity period. The patients receive a written informed consent form, the sheet would be provided and sufficient time would be given to the patients/guardians to go through the sheet and ask questions and clarify, which follows ICH guidelines in Good Clinical Practice.
Sample size and sampling strategy
The minimum expected AUC for establishing prognostic accuracy of the biomarkers Ki-67, PET SUV max, and the gene expression profile for estimating the sample size was 0.80 at a 5% level of significance (Software used-SYSTAT 13.2). The null hypothesis value was taken as 0.6. The ratio of non-pCR (negative outcome):pCR (positive outcome) is 70:30. The estimated sample size is 72, considering sample loss an additional 10% is added to the sample size. The ultimate estimated sample size is 80 patients.
Participant timeline
All patients with breast lumps reporting to Surgery OPD at JIPMER Pondicherry and planned for tru-cut biopsy will be screened. Baseline biopsy tissue will be collected in the RNA later. Patients who have been pathologically confirmed to have TNBC are provided with an explanation about the project and asked for their consent. All the baseline investigations will be done for the patient. Based on the report intervention, the date will be planned for the patient on the 14th or 15th day 2nd PET scan will be done in the Department of Nuclear Medicine and 2nd biopsy will be done in the Department of Surgery. Three bits of tissue are collected during 2nd biopsy; two bits are collected in a formalin container for the making FFPE, and one bit will be collected in the RNA later solution for RNA extraction fresh tissues collected from the tru-cut biopsy at two different time points are stored in the RNA later container in the −80 deep freezer FFPE blocks will be made and stored for the histopathological assessment, followed by standard of care and assessment of endpoint pCR-(ypT0N0M0) and non-pCR after obtaining the histopathology report (Figure 2).

Participant timeline.
Recruitment
This study will be conducted among newly diagnosed non-metastatic TNBC patients at the healthcare center. The patients would be solicited to participate in the study. All patients will be screened as per the inclusion–exclusion criteria. If they agree to provide written informed consent, they will be considered for the biomarker study.
Methods followed in biomarker analysis
Pathological biomarkers
Ki-67
The immunohistochemistry (IHC) assessment of mouse antihuman Ki-67 will be quantified using a visual scoring system. Stained cells will be counted and expressed as a percentage. Nuclear staining will be incorporated into the Ki-67 score, which is defined as the percentage of positively stained cells among the total number of malignant cells scored. The patients will be divided into two groups according to changes in the Ki-67 score namely decreased group who demonstrate a Ki-67 score of at least 1% less in the second biopsy. The no decrease group will be defined as an increase or no change in Ki-67 expression.
Tumor-infiltrating lymphocytes
TIL counts will be determined on hematoxylin–eosin stained full sections using the scoring guidelines of the International Immuno-Oncology Biomarker Working Group on Breast Cancer. 32 Stromal TIL scores will be defined as the percentage of tumor stroma area that is occupied by mononuclear inflammatory cells. Inflammatory infiltrates in the stroma of noninvasive lesions and normal breast structures will be excluded from TIL counts. 33
Genomic biomarkers
Fresh tissue will be collected and RNA will be extracted using Qiagen kits.
Phase I: RNA sequencing of 35 patients (70 samples) at two different time points.
Phase II: Selecting the target gene (top 10 genes) based on phase I results considering the frequency of gene expression and pathways related to breast cancer followed by RT-PCR for analysis of gene expression in the 80 samples.
RNA extraction
RNA purification involves extracting high-quality RNA from biological samples while eliminating contaminants. The RNeasy technology achieves this by combining a silica-based membrane’s selective binding with microspin efficiency. The process starts with sample lysis in a denaturing guanidine–thiocyanate buffer, deactivating RNases. Ethanol is added for optimal binding, and the sample is applied to an RNeasy Mini spin column. Here, RNA (>200 bases) binds to the membrane, and impurities are washed away, followed by elution in water. Notably, RNAs <200 nucleotides are excluded, enriching mRNA. The method’s versatility is evident in tailored protocols for different samples, primarily differing in lysis, homogenization, and binding conditions, while the purification steps remain consistent, ensuring reliable results.
The Dynabeads® mRNA purification kit streamlines the isolation of mRNA from total RNA samples through the following steps
Preparation of RNA
Begin with 75 μg of total RNA. Adjust its volume to 100 μl using distilled Diethyl pyrocarbonate (DEPC)-treated water or 10 mM Tris–HCl, pH 7.5. Heat the mixture to 65°C for 2 min to disrupt secondary structures, and then cool it on ice.
Preparation of Dynabeads®
Take 200 μl (1 mg) of well-resuspended Dynabeads® and place them on a magnet for 30 s to collect them against the tube wall. Remove the supernatant and calibrate the beads with 100 μl binding buffer. Afterward, add 100 μl of binding buffer to the Dynabeads®. If the RNA is more dilute than 75 μg/100 μl, adjust the volume with binding buffer.
Isolation of mRNA
Mix the total RNA with the Dynabeads®/binding buffer suspension to allow mRNA to anneal to the oligo(dT)25 on the beads. Rotate the mixture for 3–5 min at room temperature. Place the tube on the magnet until the solution becomes clear and remove the supernatant. Wash the mRNA-bead complex twice with 200 μl washing buffer B. Ensure efficient removal of supernatant using the magnet.
Elution of mRNA
If elution is needed, add 10–20 μl of 10 mM Tris–HCl, pH 7.5. Heat the mixture to 65–80°C for 2 min and immediately place the tube on the magnet. Transfer the eluted mRNA to a new RNase-free tube for downstream applications.
Regeneration and reuse of Dynabeads® Oligo(dT)25
Resuspend used Dynabeads® (original volume 200 μl) in 200 μl reconditioning solution and incubate at 65°C for 2 min. Place the tube in a magnetic field to remove the supernatant and repeat the wash with the reconditioning solution twice. Resuspend the Dynabeads® in storage buffer oligo(dT)25 through a series of washes and magnetic separations. The Dynabeads® are now ready for reuse in mRNA isolation.
Quality control of RNA sample
✓ RNA samples will be subjected to qualification and quantification using 1% agarose gel electrophoresis and Qubit/Nanodrop spectrophotometer, respectively.
✓ If received samples fail to meet initial QC parameters, re-sampling will be required.
Phase I : RNA sequencing of 35 patients (70 samples) at two different time points.
Workflow for mRNA sequencing on the Illumina platform
Wet lab:
RNA Seq. library construction will be carried out using an Illumina-specific library preparation kit. mRNA will be enriched from total RNA and will be converted to cDNA as per the manufacturer’s protocol. cDNA will be used for library preparation. Fragmentation of cDNA will be carried out using the enzymatic method followed by adapter ligation. Enrichment of adapter-modified DNA fragments by PCR will be performed and quantification of the enriched library will be performed using a qubit fluorometer. Samples will be pooled before sequencing to generate defined data. Sequencing prepared libraries will be sequenced on Illumina HiSeqX/NovaSeq to generate 30 M, 2 × 150 bp reads/sample. Up to 75% of the sequenced bases will be of Q30 value. Sequenced data will be processed to generate FASTQ files and uploaded on the FTP server for download.
Data generation
Data: 4.5 GB/sample.
Platform: Illumina platform.
File format: FASTA.
Bioinformatics workflow
Quality check for raw data- Data will be checked for the bad-quality reads, bases and adapter sequences. If any adapter sequences found in the reads will be removed by appropriately trimming/removing the reads. The analysis will provide an illustrated summary of the quality of the data generated.
Preprocessing of raw data includes adapter sequence removal and contamination sequence removal (tRNA, rRNA): data will be mapped to the human genome, identify reads generated from structural RNA molecules (rRNA/tRNA), and filter them. This is the contamination removal step.
Alignment of preprocessed data to the human reference genome (hg19) using HiSAT2: the quality filtered/contamination/adapter removed data will be mapped to the human genome reference sequence (HG19) using HiSAT. The statistics for data loss at each of the three steps (quality filtering, contamination filtering, and mapping) will be recorded.
Genes expression estimation (raw) using feature counts: feature counts will be used to derive raw read counts mapping to known genes. These read counts will normalized in DESeq2, to assess gene expression levels.
Differential expression analysis will be performed using DESeq2: we use DESeq2 for identifying differentially expressed genes. Table 3 includes the Illumina stranded mRNA preparation.
Illumina stranded mRNA Prep.

FFPE, Formalin-Fixed Paraffin-Embedded; SNP, single nucleotide polymorphisms; UDI, unique dual indexes.
Phase II: Selecting the target gene (top 10 genes) based on phase I results considering the frequency of gene expression and pathways related to breast cancer results followed by RT-PCR for analysis of gene expression in the 80 samples.
Reverse Transcription Quantitative Polymerase Chain Reaction (RT-qPCR)
mRNA expression will be assessed using Qiagen mRNA primer sets. All qPCR reactions will be conducted on the Quant Studio™ 5 Real-Time PCR System (Applied Biosystems, ThermoFisher Scientific, USA). The synthesized cDNA will serve as the template for qPCR, employing the appropriate mRNA-specific primers and probes (QuantiTect RT-PCR Kits, Cat. No. 204443). Cycling conditions will comprise an initial heat activation step at 95°C for 2 minutes, followed by 40 cycles of denaturation at 95°C for 5 seconds and annealing/extension at 56°C for 30 seconds. Relative mRNA concentration will be determined using the relative cycle threshold method. Expression differences will be evaluated through relative quantification, normalizing the mean Ct values of the target genes to the mean Ct values of suitable endogenous controls.
Imaging biomarker
PET-CT scan
Participants will undergo a PET scan at baseline (post-biopsy) and D14 before the (second biopsy). All patients undergoing whole-body fluorine-18 fluorodeoxyglucose (PET/CT) (F-18 FDG PET/CT) study will be instructed to be on fasting for 4–5 h and avoid any strenuous exercises 24 h before the study. Patients will be checked for fasting blood sugar to be less than 180 mg/dl before F-18 FDG administration. F-18 FDG dose of 0.1 mCi per kg body weight is administered intravenously and image acquisition is performed after 45 min in the ‘Discovery MI digital-ready PET/CT’ model from Wipro GE Healthcare Pvt limited. A whole-body F-18 FDG PET/CT study will be performed from the base of the skull to the mid-thigh level. The images will be processed and displayed in AW 4.7 workstation (Advanced workstation from GE Healthcare Private Limited). SUV max and SUL max of the primary breast tumor will be recorded.
Biomarker analysis – Proposed method of analysis is mentioned in the Table 4.
Biomarker analysis.

IHC, immunohistochemistry; PET-CT, positron emission tomography-computed tomography; PCR, polymerase chain reaction; TIL, tumor-infiltrating lymphocyte.
The Table 5 provides a structured timeline for when each parameter is assessed throughout the screening, intervention, and post-intervention phases, as well as during neoadjuvant chemotherapy and surgery.
Assessment schedule.

CBC, complete blood count; pCR, pathological complete response; PET-CT, positron emission tomography-computed tomography; TIL, tumor-infiltrating lymphocyte.
Project implementation
The developed composite prognostic score can be used for grouping patients based on their level of risk for the assessment of pCR in TNBC which would also help to decide chemotherapy intensity in patients planned for NACT thus, personalizing therapy.
Composite prognostic score
The composite prognostic score is a combination of multiple biomarkers and clinical parameters that have been identified as predictive factors for pCR. These could include gene expression profiles, TIL count, Ki-67 levels, and other relevant molecular and clinical features.
Grouping patients by risk
The composite prognostic score allows the grouping of TNBC patients into different risk categories based on their likelihood of achieving pCR after NACT. Patients with higher scores are likely to achieve pCR, while those with lower scores may have a lower chance of complete response.
Tailoring chemotherapy intensity
With the knowledge of the patient’s risk category, oncologists can tailor the intensity of chemotherapy accordingly. Patients at higher risk of not achieving pCR may require more aggressive and personalized chemotherapy regimens to improve their chances of a complete response.
Personalizing therapy
The composite prognostic score enables personalized therapy by guiding treatment decisions based on individual patient characteristics. Instead of a one-size-fits-all approach, patients receive treatments that are specifically designed to maximize effectiveness and minimize adverse effects.
Clinical significance
Achieving pCR in TNBC is associated with better long-term outcomes and overall survival. Thus, identifying patients likely to achieve pCR allows for the optimization of treatment strategies and potentially improves patient prognosis.
Reducing overtreatment
By stratifying patients based on their risk of pCR, the composite prognostic score can help avoid unnecessary overtreatment in patients with a high likelihood of achieving pCR. This reduces the risk of drug toxicity and associated costs.
Research and clinical implementation
The development of the composite prognostic score involves rigorous scientific research, including validation in clinical trials. If the score proves to be reliable and robust, it can be integrated into routine clinical practice to aid treatment decisions. Figure 3 depicts the project implementation plan.

Project implementation plan.
Data collection
Patient details will be entered in a standard proforma.
Data entry: Data will be entered into Microsoft Excel as a spreadsheet.
Data extraction: The data extraction process for this exploratory study employing the window of opportunity design entails a comprehensive collection of various data points at distinct junctures throughout the investigation. Initially, baseline data is gathered, encompassing patient demographics, confirmation of triple-negative breast cancer (TNBC) status based on estrogen receptor (ER), progesterone receptor (PR), and HER2/neu expression, along with adherence to inclusion and exclusion criteria. Baseline biomarker assessments, including Ki67 expression levels, tumor-infiltrating lymphocytes (TILs) assessment, and gene expression profiling from the initial biopsy specimen, are meticulously recorded alongside baseline PET scan results. During the chemotherapy phase, meticulous monitoring of treatment administration, patient clinical status, and adverse events occurs, while hemogram results and second biopsy findings, such as Ki67 expression levels, TILs assessment, and gene expression changes, are documented. Additionally, PET CT scan results are analyzed to discern changes in Standardized Uptake Value (SUV) and Standardized Uptake Value of lean body mass (SUL) max of the tumor. Post-surgery, data includes completion of the neoadjuvant chemotherapy regimen, surgical outcomes detailing the type and extent of tumor removal, lymph node involvement, and post-surgery pathological response assessment, particularly focusing on the presence or absence of invasive components in the axilla and breast, as well as evaluation of pathological complete response (pCR) status. Long-term follow-up extends over a duration of 3 years, monitoring for disease recurrence, metastasis, and survival outcomes, all meticulously documented to ensure comprehensive analysis and interpretation while upholding ethical standards and patient confidentiality throughout.
Privacy and confidentiality
Participants in this trial will receive unique identification codes, and all data collected will be de-identified. Identifiable participant information will be stored separately, and patient-reported data will only be used for this study. This approach ensures privacy and confidentiality. During the trial, strict confidentiality measures will be implemented to safeguard all collected information. Personal sensitive data, including full names, contact information, and identification details, will not be included in the clinical research Performa. All data will be processed confidentially to ensure privacy and protect participants’ identities.
Statistical analysis
Statistical methods for primary and secondary outcomes
The categorical variables such as clinical characteristics, histopathological profile, comorbidity, etc. will be expressed as percentages and frequency. The continuous data such as Ki-67, gene expression, SUV, and age, will be expressed as median with range or mean with standard deviation. The change in the Ki-67, gene expression, and SUV over time will be explored by paired t-test otherwise Wilcoxon signed-rank sum test. The comparison of the change in Ki-67, gene expression, and SUV concerning pCR status will be carried out using a Mann–Whitney U test or independent student’s t-test. The association of the pCR status with different categorical variables mentioned above will be carried out using Fisher’s exact test or chi-square test. Kaplan–Meier curve will be utilized to estimate the survival function and a log-rank test will be done to compare the same between different characteristics. All statistical analysis will be estimated at a 5% level of significance and p value.
Composite prognostic score
The multiple logistic regression analysis will be done to estimate the odds ratio with a 95% confidence interval to investigate the correlation between candidate prognostic markers and pCR. 34 The final selection of prognostic variables will be based on the change in Ki-67, TILs, SUV max and SUL max value, and gene expression profiling. The selected predictors will be given a weightage or a score by multiplying their corresponding regression coefficients (β) by a factor of 10. Further categorization of the derived score (low, moderate, high) will be based on dividing the range of scores into tertiles. Receiver Operating Characteristic (ROC) curve will be plotted and the resulting Area Under the Curve (AUC) value will be used to compare the pCR outcomes based on the derived prognostic score model.
Discussion
In this study, preoperative setup emphasizes the chance to assess biomarkers change and surrogate endpoints of cancer in vivo in response to the interventional drug. Currently, a potent approach to personalize chemotherapy in the early stages of TNBC does not exist. Regardless of significant response rates in a subgroup of patients, mostly all patients will receive the same number of cycles of chemotherapy. NACT decreases tumor size, improves surgical outcomes, and evaluates chemotherapy response. However, tailored therapies based on the pathological response have not been well established. One of the reasons to lag in tailoring personalized therapy is there are no established biomarkers that have been identified before NACT to identify patients likely to achieve pCR. These biomarkers may be clinical, biological, or imaging. Clinical and demographic features, such as menstrual status, family history, racial disparity, patient’s age, and mammographic breast density, have been associated with pCR in various studies.
Ki-67 expression was considerably associated with tumor proliferation and has been recognized Ki-67 as an excellent biomarker. 35 In meta-analysis study carried out by Wu et al. in the TNBC population stated that the pooled results of 35 studies showed that highly expressing Ki-67 was correlated with overall survival of poor DFS. 17 Arnaout et al. carried out a study in a window of opportunity set up in ER-positive breast cancer patients, where the patient received anastrozole as an interventional drug during the window period and found a statistically significant fall of 48.8% in mean Ki-67 indices while comparing pre-treatment Ki-67 indices with post-treatment. 18
Several studies showed that the tumor microenvironment can be one of the driving factors of tumor invasion and progression. High TIL counts at baseline and a significant reduction in TIL counts after neoadjuvant therapy are associated with higher pCR rates. 36 Elevated expression of CD4+ TIL count significantly correlates with better DFS and OS. 14 Studies reported that a high CD8+ TIL level was associated with better DFS only as no statistical association was found with OS. 19 The increased FOXP3+ TIL level also was correlated only with DFS but not with OS. 37 In TNBC, tumors having the highest TIL score achieved 37% of pCR, and those exhibiting high apoptosis showed 47%. 20 Therapeutic drugs such as pembrolizumab can elevate sTILs, thereby increasing the mean dispersion of TILs in naïve TNBC tumors. 38 A phase II study carried out by Connolly et al. 39 in the breast cancer population to know the correlation of SUV with pCR to trastuzumab and pertuzumab, thereby, found that post-SUL max value and delta SUL max value correlated with pCR. A study conducted in the primary breast cancer population stated that pCR significantly correlates with post-SUV max value. 40
Mostly, primary tumors will be identified based on IHC and morphological assessment; however, these biomarkers exhibit significantly lesser sensitivities in TNBC comparatively ER-positive breast cancer. Molecular profiling is a promising diagnostic approach that is believed to aid in the classification of metastatic cancers with an unknown primary tissue origin. 41 Wang et al. 24 reported a small subgroup of genes from the 90 panel of genes aiding diagnostic utility in the TNBC subgroup, where they found 17 genes were upregulated, 15 genes were downregulated, and recognized as candidate genes to differentiate TNBC from other types of malignancy. Interesting research conducted in a primary breast cancer population of 25 patients found that 125 genes were upregulated, and 116 genes were downregulated at 24 h of NACT followed by 193 genes upregulated, 238 genes downregulated at 48 h of NACT, surprisingly, significant transcriptional response occurred in all patients during therapy which is supporting the hypothesis that genes respond to drugs within the short period. 42
Strength
Utilization of a window of opportunity design to assess biomarker changes before NACT and also to check the chemotherapy tolerance in patients. During the waiting period (window of opportunity), patients are administered a single dose of chemotherapy to offset any potential delays in receiving dose-dense chemotherapy. Due to the study intervention involving a single shot and a low dose, the expected toxicity is anticipated to be minimal. Potential to enhance treatment outcomes by predicting pCR and guiding therapy based on individual biomarker profiles. Addressing the need for reliable biomarkers to predict pCR in TNBC patients undergoing NACT contributes to the understanding of biomarkers associated with treatment response in TNBC and may inform future clinical trials.
Personalized treatment: Aim to develop a predictive score for identifying TNBC patients likely to achieve pCR, leading to tailored treatment strategies.
Limitations
Transcriptomic analysis in the current study will be done for only a small sample size which may or may not represent the large population. The primary endpoint of the study is pCR which is only a surrogate endpoint to overall survival which would be ideal.
Supplemental Material
sj-docx-1-tam-10.1177_17588359241248329 – Supplemental material for Assessment of novel prognostic biomarkers to predict pathological complete response in patients with non-metastatic triple-negative breast cancer using a window of opportunity design
Supplemental material, sj-docx-1-tam-10.1177_17588359241248329 for Assessment of novel prognostic biomarkers to predict pathological complete response in patients with non-metastatic triple-negative breast cancer using a window of opportunity design by Chitradurga Rajashekhar Akshatha, Dhanapathi Halanaik, Rajesh Nachiappa, Nanda Kishore, Prasanth Ganesan, Smita Kayal, Harichandra Kumar and Biswajit Dubashi in Therapeutic Advances in Medical Oncology
Footnotes
Acknowledgements
We thank Dr Vijai Joseph, Associate Attending Geneticist, Department of Medicine, MSKCC, Associate Professor of Genetics Research in Medicine, Weill Cornell Medical Center, Cornell University, Associate Director, Bioinformatics, Niehaus Center for Inherited Cancer Genomics, MSKCC and Dr. P. S. Suresh, Assistant Professor, School of Biotechnology, National Institute of Technology, Calicut, Kerala, India for their valuable suggestions in designing the protocol.
Declarations
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
