Abstract
Background:
Different RAS/BRAF allele mutations imply distinct biological properties in various solid tumors. Recently, several studies have focused on the predictive and prognostic roles of various RAS/BRAF allele mutations in colorectal cancer (CRC) but the results remain controversial.
Methods:
Between March 2017 and September 2022, the patients diagnosed as stages I–IV CRC with detailed medical records including next-generation sequencing (NGS) data and clinicopathological follow-up information available at our center were enrolled. Survival data were estimated using the Kaplan–Meier method, and the difference was tested in a log-rank test. Multivariate tests were carried out using Cox models.
Results:
A total of 1029 CRC patients were included, and the incidence of RAS/BRAF mutation was 58.4%. The hypermutated cohort was defined as patients with microsatellite instability-H or POLE/D mutation. In the non-hypermutational cohort, only KRAS G13D mutation was associated with a higher incidence and inferior disease-free survival in patients with stage I-III CRC. In the cohort of patients with non-hypermutated metastatic colorectal cancer (mCRC), we assessed the risk of various RAS/BRAF allele mutations and subsequently reclassified patients into four groups based on first-line median progression-free survival: wild type (group 1), low-risk RAS/BRAF mutation (group 2, RAS/BRAF mutations other than KRAS G13D/G12V/G12C or BRAF V600E), high-risk RAS mutation (group 3, KRAS G13D/G12V/G12C), and BRAF V600E mutation (group 4). mCRC patients with high-risk RAS mutation could significantly benefit from intensive triplet chemotherapy (hazard ratio, 2.54; 95% confidence interval, 1.36–5.12; p = 0.0091).
Conclusion:
In the non-hypermutated CRC cohort, the prognostic risk of various RAS/BRAF allele mutations varied between local and metastatic CRC. KRAS G13D mutation tended to be the only prognostic marker for stages I–III CRC; however, KRAS G13D/G12V/G12C mutations collectively defined a high-risk subgroup of mCRC patients with poor prognosis, who would benefit from intensive triplet chemotherapy.
Introduction
Predominant oncogene RAS/BRAF mutations occur in 50–60% of colorectal cancer (CRC) patients1,2 and are considered negative biological markers due to anti-epidermal growth factor receptor (anti-EGFR) therapy resistance and poor prognosis.3,4 The frequency of the RAS/BRAF gene and their allele mutations was highly variable across tumor types. 5 For most cancers, such as CRC and lung cancer, KRAS is the dominant mutant but in melanoma and thyroid cancer, NRAS mutation is more common.6,7 In lung cancer, KRAS G12C is the most popular mutated codon, but in CRC, the top five prevalent RAS/BRAF allelic mutations are KRAS G12D, KRAS G12V, KRAS G13D, BRAF V600E, and KRAS G12C.5,8 Therefore, with the popularization of NGS in precision medicine, our attention to RAS/BRAF mutations should not be limited to the gene level but should be refined to the allele level.
Recently, an increasing number of studies have shown that different RAS/BRAF allele mutations have distinct biological functions and lead to disparate clinical outcomes in cancers. KRAS mutations at different codons showed different RAF affinities and GTP intrinsic hydrolytic activities. G12A/G12V/G12R/Q61 showed high RAF affinity, while the G12R/G12D/G12V mutation showed low affinity. According to the intrinsic hydrolytic activity of GTP, the mutant sites could be divided into a high-activity group (G12C, G12D, and G13D) and a low-activity group (G12A, G12R, G12V, Q61L, and Q61H). Studies have shown that high RAF affinity and low intrinsic Guanosine Triphosphatase (GTPase) activity will exhibit more intense and sustained RAF activation. 9 Meanwhile, the genetic commutation network of oncogenic KRAS mutations is allele- and tissue-specific. In CRC, KRAS allele-specific derived networks are weaker than those in lung cancer, with only a few genes involved in integral KRAS signaling pathways linking the alleles together, including an increased co-mutation interaction between KRAS G12A and MAP2K3, a reduced co-mutation interaction between KRAS G12D and ERBB4, and a very strong increased rate of co-mutation between KRAS G12C and STK11. 10 There are discrepancies in the main downstream activation pathways and activation intensities at different KRAS site mutations.11,12 In non-small-cell lung cancer, the KRAS G12D mutation activated PI3K/AKT, whereas the KRAS G12C mutation decreased growth-factor-dependent Protein Kinase B (AKT) activation. 13 Among various NRAS allelic mutations, discrepancies in biochemical and signaling properties were also observed. 14 Approximately 200 BRAF mutant alleles have been identified in human tumors. Based on their mechanisms of activation, oncogenic BRAF mutants may be divided into three categories. 15 The heterogeneity of the biological function of RAS/BRAF allele mutations means that site-specific considerations are required for their value in tumor prognosis and treatment decision-making.
Previous research has shown that the KRAS codon 12 mutation, but not the codon 13 mutation, is associated with a worse prognosis in CRC 16 ; however, other studies have shown opposite results: the KRAS G13D mutation exhibited the worst prognosis.17,18 Some heterogeneous retrospective series have evaluated the prognostic value of KRAS G12D with inconsistent results.19,20 The prognostic value of G12C for CRC is also controversial.21–23 By contrast, the conclusion that the KRAS G12V mutation is associated with greater aggressiveness and poorer outcomes was relatively consistent24–26 as well as BRAF-V600E. 27 In addition to prognosis, different RAS/BRAF mutation alleles lead to different drug susceptibilities. While KRAS/NRAS mutations have been considered to be nonresponsive to anti-EGFR therapy, it was shown that patients with KRAS G13D mutations may benefit from cetuximab. 28 The classification of BRAF mutant alleles determines their sensitivity to inhibitors; class 1 BRAF mutations (BRAF V600 mutations) are RAS-independent, signal as monomers, and are sensitive to current RAF ‘monomer’ inhibitors; class 2 BRAF mutants are RAS-independent, signal as constitutive dimers, and are resistant to vemurafenib but may be sensitive to novel RAF dimer inhibitors or MEK inhibitors; class 3 BRAF mutants have impaired kinase activity or are very sensitive to ERK-dependent feedback of RAS and are also resistant to vemurafenib and may be effectively treated with combinations that include inhibitors of the RTKs responsible for driving RAS activation. 15
FOLFOXIRI plus targeted therapy has become one of the standard treatment options for mCRC recommended by the European Society of Medical Oncology and American Society of Clinical Oncology clinical practice guidelines.29,30 Patients receiving the intensive triplet regimen have a better objective response rate, progression-free survival (PFS), and overall survival (OS). 31 However, strong chemotherapy-related grades 3–4 toxicities limit its clinical use. Therefore, we need to accurately identify patients who can benefit from a triplet regimen. The BRAF V600E mutation accounts for approximately 95% of all BRAF mutations in CRC and is responsible for its poor prognosis.8,32 BRAF V600E is a reliable predictor for triplet chemotherapy but studies have been inconsistent on whether mCRCs with RAS mutations benefit from this regimen. The TRIBE 2 study showed that the combination of triplet chemotherapy and bevacizumab was a better choice for patients with RAS mutation or right-sided colon cancer.33–35 A subsequent large meta-analysis confirmed the above conclusions. 36 However, in another study, multivariate analysis showed that RAS mutation status was not a predictive factor for triplet chemotherapy. 37 Given the poor prognosis of KRAS G12C, a recent study suggests that this subgroup of mCRCs may benefit from intensive triplet chemotherapy compared to standard doublet chemotherapy. 38 We speculate that the reason for this inconsistency is that mCRC with RAS mutation is a very heterogeneous population due to various alleles.
Therefore, we conducted a large retrospective study to estimate the prognostic and predictive roles of various RAS/BRAF-mutated alleles in CRC. Based on the first-line median progression-free survival (mPFS) of metastatic CRC patients, we reclassified RAS/BRAF allele mutations and explored valid treatment regimens for different groups.
Methods
Patients and study design
From March 2017 to September 2022, patients who were diagnosed with CRC and had available genomic data as well as clinicopathological follow-up information at the First Affiliated Hospital of Zhejiang University were included in our study. All data were retrospectively collected from electronic medical records, and the inclusion criteria were as follows: (a) histologically confirmed CRC, (b) patients with available genomic status based on NGS, using Onco-Screen Plus panel or the Colon Core panel, (c) baseline as well as post-treatment imaging was accessible at our center for efficacy evaluation, and (d) adequate organ function. Exclusion criteria were as follows: (a) multiple primary malignancies, (b) RAS/BRAF multisite mutations, (c) patients have available genomic status but not based on NGS or not using Onco-Screen Plus panel or the Colon Core panel, (d) less than 3 months of follow-up as of September 2022, and (e) irregular monitoring rhythmicity. The study was approved and supervised by the Ethics Committee of the First Affiliated Hospital, Zhejiang University School of Medicine (No. IIT20210185B), and carried out in accordance with the Declaration of Helsinki. Due to the retrospective nature of our study, written informed consent was exempted. We have followed the relevant Equator guidelines and the reporting of this study conforms to STROBE guidelines (https://www.equator-network.org/) 39 (Supplemental Table S1).
Genomic status
RAS/BRAF mutation and hypermutation status were identified using the Onco-Screen Plus panel or the Colon Core panel that targeted 520/41 cancer-related genes, based on surgically resected or punctured tissue of cancer, provided by Burning Rock, a clinical laboratory based in Guangzhou, China. The hypermutated cohort was defined as those harboring either microsatellite instability (MSI)-H or pathogenic POLE/D mutations. The DNA isolation and targeted sequencing procedures were conducted at Burning Rock Biotech, a commercial clinical laboratory that holds accreditation from the College of American Pathologists and certification from the Clinical Laboratory Improvement Amendments. The process of target capture involved the utilization of a commercially available panel comprising 520/41 genes associated with cancer, which collectively spanned 1.64 megabases of the human genome. The obtained sequence data were aligned to the reference human genome using Burrows-Wheeler Aligner version 0.7.10. Subsequently, local alignment optimization, duplication marking, and variant calling were conducted using Genome Analysis Tool Kit version 3.2 (Manufacturer: Broad Institute) and VarScan version 2.4.3 (Manufacturer: Koboldt Laboratory at Washington University School of Medicine). To identify somatic variants, the tissue samples were compared against their respective white blood cell controls. In addition, an in-house algorithm called markSV was employed for the analysis of structural rearrangements.
Outcomes
The response was assessed according to the Response Evaluation Criteria version 1.1. Disease-free survival (DFS) was defined as the interval between surgery and the emergence of locoregional failure, distant metastasis, or death from any cause. PFS was defined as the time from first-line chemotherapy initiation to disease progression or death from any cause. Our follow-up protocol is based on National Comprehensive Cancer Network (NCCN) guidelines: for stage I CRC patients, CEA was tested every 6–12 months for a total of 5 years, and chest/abdominal/pelvic computed tomography (CT) was performed when CEA is abnormal or clinical symptoms are present. For stages II–III CRC patients, CEA was examined every 3–6 months for 2 years and every 6 months thereafter for a total of 5 years; chest/abdominal/pelvic CT scans were performed every 6–12 months for a total of 5 years. For stage IV CRC patients, who received first-line therapy are evaluated for efficacy every 2 months based on CEA and chest/abdominal/pelvic CT scans. For all patients, colonoscopy was performed in 1 year after surgery, if advanced adenoma, repeat in 1 year, if no advanced adenoma, repeat in 3 years, then every 5 years.
Statistical analysis
We used the χ2 test or Fisher exact test to estimate differences between categorical values, and the χ2 test, t-test, and Mann–Whitney to assess differences between wild-type and RAS/BRAF allele mutations. Survival estimates for the study population were generated using the Kaplan–Meier method. The association of several variables with DFS was assessed using the COX proportional risk regression model, and clinicopathological variables of known prognostic significance, such as age, sex, T-stage, primary tumor location (colon or rectum), regional lymph node status, CEA level, and metastatic organs, were tested for stepwise exclusion, and those that were statistically significant in univariate analysis (p > 0.05) were excluded. Variables were retained in the multivariate model. The independence of independent factors was assessed by calculating the previously described β coefficients. All analyses were performed using Statistical Package for the Social Sciences (SPSS) statistical software (version 26; IBM) and R (version 3.5.3). All tests were two-sided, with p < 0.05 defined as statistically significant.
Results
RAS/BRAF allele landscape
Between March 2017 and September 2022, a total of 1029 CRC patients were identified, and 59 patients were excluded due to multiple primary tumors (n = 54) and no baseline data (n = 5) [Figure 1(a)]. Finally, 970 patients met the criteria and were enrolled in the subsequent analysis. Baseline characteristics of the population are listed in Table 1. The incidence of MSI was 57 (5.8%), and the incidence of POLD/E was 11 [1.1%, 7 MSI-H, 4 microsatellite stable (MSS)]. KRAS G13D (20.3% versus 9.6%, p = 0.008) and BRAF V600E (11.86% versus 6.08%, p = 0.040) occurred more frequently in the hypermutated cohort than in the non-hypermutated cohort, while KRAS G12V (0% versus 10.6%, p = 0.008) showed the opposite tendency (Supplemental Table S2). Considering the distinct biological characteristics of the hypermutated cohort, we excluded this cohort and further explored the roles of RAS/BRAF mutations in the non-hypermutated cohort. In the 880 non-hypermutated CRCs, the incidence of KRAS, NRAS, and BRAF mutations was 47.6%, 3.3%, and 7.1%, respectively. The most prevalent KRAS-mutated alleles were KRAS G12D (14.7%), KRAS G12V (10.7%), KRAS G13D (9.8%), KRAS G12C (3.2%), and KRAS A146T (2.0%). The most prevalent NRAS-mutated alleles were NRAS Q61K (0.8%) and NRAS G12C (0.6%). The most prevalent BRAF-mutated allele was BRAF V600E (6.1%). The remaining mutated RAS/BRAF alleles were classified separately in the ‘KRAS/NRAS/BRAF-Other’ group [Figure 1(b)].

(a) Flowchart of 1029 patients with CRC receiving NGS and (b) frequency of RAS/BRAF site mutation variant in 880 patients with CRC (only mutation sites which were found in more than five patients were listed in the figure).
Baseline clinical characteristics of patients.

AC, adenocarcinoma; IRI, irinotecan; MC, mucinous adenocarcinoma; MSI, microsatellite instable; MSS, microsatellite stable; OX, oxaliplatin; SRCC, signet-ring cell carcinoma; POLE, polymerase epsilon; KRAS, kirsten rat sarcoma viral oncogene homolog; NRAS, neuroblastoma ras viral oncogene homolog; BRAF, b-raf proto-oncogene.
KRAS G13D was the only mutated allele related to higher incidence and inferior DFS in stages I–III CRC patients
There are different frequencies of RAS/BRAF site mutations between stages I–III and stage IV CRCs. The incidence of KRAS G13D was significantly higher in stages I–III CRCs than in stage IV CRCs (13.1% versus 6.5%, p = 0.001) [Figure 2(a) and (b), Supplemental Table S3]. By September 2022, disease progression had occurred in 184 patients (43.8%) in the stages I–III population. Among the patients with stages I–III disease, KRAS G13D variant was significantly associated with inferior DFS [median 18.0 (95% CI, 14.0–22.0) months versus 23.0 (95% CI, 19.3–26.7) months versus 21.0 (95% CI, 15.0–25.0) months (p = 0.0034) compared with wild-type and non-KRAS G13D variant [Figure 2(c) and (e)]. However, no statistically significant differences were observed for BRAF V600E or other RAS/BRAF allele mutations [Figure 2(d), Supplemental Figure S1]. In the multivariate analysis, KRAS G13D variants remained significantly prognostic for DFS [hazard ratio (HR), 3.735, 95% confidence interval (CI), 1.468–9.505, p = 0.006] [Table 2(a)].

Frequencies of RAS/BRAF mutation subtypes in I–III (a)/IV stage (b) CRC. DFS between KRAS G13D (c)/BRAF V600E (d)/KRAS G13D (e), and wild type in I–III stage CRC.
Univariate and multivariate analyses of prognostic factors in non-hypermutated I–III/IV stage CRC.
(a) DFS of non-hypermutated I–III CRC.

(b) PFS of non-hypermutated IV CRC.

AC, adenocarcinoma; CA, carbohydrate antigen; CEA, carcinoembryonic antigen; TNM, tumor-node-metastasis; CI, confidence interval; DFS, disease-free survival; DLN, distant lymph node metastasis; HR, hazard ratio; M, metastatic; MC, mucinous adenocarcinoma; PFS, progression-free survival; SRCC, signet-ring cell carcinoma; WT, wild type; RAS, rat sarcoma viral oncogene homolog.
Reclassification of RAS/BRAF-mutated alleles based on first-line mPFS in metastatic CRC patients
Multivariate analysis identified the following factors as significantly associated with mPFS of synchronous mCRC: CEA, peritoneum metastasis, number of distal metastatic organs, RAS mutation, and BRAF mutation. Further analysis based on mutated alleles found that only BRAF V600E, KRAS G13D, KRAS G12V, and KRAS G12C mutants were significantly associated with worse mPFS than the wild-type state [Table 2(b)]. The mPFS ranked BRAF V600E at 6 months (95% CI, 4.8–7.1, p < 0.0001), KRAS G13D at 9.0 months (95% 7.6–10.3, p = 0.0004), KRAS G12V at 10.0 months (95% CI, 8.2–11.7, p < 0.001), and KRAS G12C at 11.0 months (95% CI, 5.5–16.5, p = 0.0094) [Figure 3(a)–(d)]. Based on the above results and multivariate analysis results of mPFS, we regrouped RAS/BRAF mutations: BRAF V600E was still defined as the group with the worst prognosis, KRAS G13D/G12V/G12C were classified as high-risk RAS mutations with the second worst prognosis, and KRAS G12D/A146T, NRAS, and RAS/BRAF-Other were classified as low-risk RAS/BARF mutations [Table 2(b), Supplemental Table S4]. The results were also consistent when all patients with simultaneous or heterogeneous metastases were included (Supplemental Figure S3). Therefore, all patients with mCRC were reclassified into four groups: wild type (group 1), low-risk RAS/BRAF mutation (group 2), high-risk RAS mutation (group 3), and BRAF V600E mutation (group 4) (Supplemental Table S4). After regrouping, there was a statistically significant difference in mPFS between the four groups [p < 0.0001, Figure 3(e)], as well as between the high-risk RAS and low-risk RAS/BRAF groups [p < 0.0019, Figure 3(f)]. Moreover, there was no statistically significant difference in mPFS between alleles in the high-risk RAS or low-risk RAS/BRAF group [Figure 3(g) and (h)]. All of the above results prove that our grouping is reasonable. The results were also consistent when all patients with simultaneous or heterogeneous metastases were included.
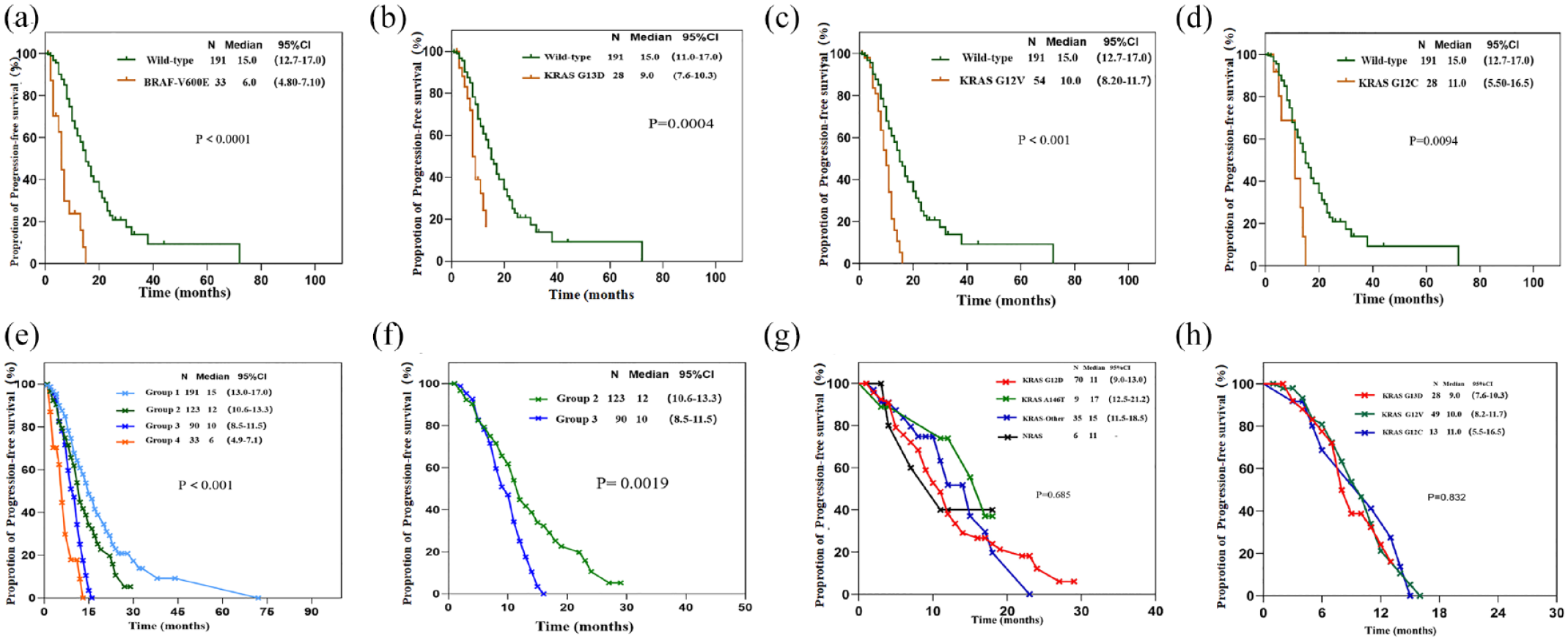
Kaplan–Meier survival curves in RAS/BRAF gene locus mutation in IV stage CRC. (a) PFS according to BRAF V600E/wild type; (b) PFS according to KRAS G13D/wild type; (c) PFS according to KRAS G12C/wild type; (d) PFS according to KRAS G12V/wild type; (e) PFS between group1/2/3/4; (f) PFS between group2/3; (g) PFS in group 2 (KRAS G12D/A146T/other KRAS/NRAS locus mutations); and (h) PFS in group 3 (KRAS G13D/G12V/G12C locus mutations).
Then, we further analyzed whether the reclassification of RAS/BRAF allele mutations was associated with the clinical and pathological characteristics of CRC patients. In this study, the primary tumor located on the right colon, aggressive histopathology (mucinous adenocarcinoma; signet-ring cell carcinoma), peritoneal metastasis, and distant lymph nodes were the variables different among the four groups (Table 3). However, no difference was found between group 2 and group 3, so we speculate that group 4 (BRAF V600E) may be responsible for these differences (Supplemental Table S5).
Baseline clinical characteristics of patients after regroup.

AC, adenocarcinoma; DLN, distance lymph nodes; IRI, irinotecan; MC, mucinous adenocarcinoma; OX, oxaliplatin; SRCC, signet-ring cell carcinoma.
Intensive triplet regimens significantly improved the mPFS of mCRC patients in group 3
A total of 363 mCRC patients received first-line oxaliplatin- or irinotecan-based chemotherapy combined with targeted therapy. Among them, 289 were treated with a first-line doublet regimen (FOLFOX/FOLFIRI + Cet/Bev), and 74 were treated with a first-line triplet regimen (mFOLFIRINOX + Bev). Disease progression occurred in 191 (52%) of 363 patients (44 in the triple-regimen group and 147 in the double-regimen group). Based on the new reclassification of RAS/BRAF allele mutations, we found that only patients in group 3 (with high-risk RAS mutation) benefited from an intense triplet regimen (HR 2.54, 95% CI, 1.36–5.12; p = 0.014) [Figure 4(a)–(c), Supplemental Figure S4]. For mCRC patients with KRAS G13D/G12V/G12C mutations, the mFOLFIRINOX group reported a median PFS of 12 months compared with 9 months in the doublet chemotherapy group (p = 0.0091) [Figure 4(b)].

Intensive triplet regimens significantly improved the mPFS of mCRC patients in regroup (a): Forest plots of double drug regime versus triple-drug regime within four groups; (b): PFS in KRAS group 3 compared double drug regime with the triple-drug regime; and (c): PFS in KRAS group 2 compared double drug regime with the triple-drug regime.
Discussion
We retrospectively analyzed RAS/BRAF allele variants and their association with clinicopathologic characteristics and prognosis in CRC patients. The frequencies of KRAS, NRAS, and BRAF mutations in our cohort were 47.6, 3.3, and 7.1%, respectively. The top five most prevalent RAS allelic mutations were KRAS G12D (14.7%), KRAS G12V (10.7%), KRAS G13D (9.8%), KRAS G12C (3.2%), and KRAS A146T (2.0%), which is in accordance with previous studies.20,40 Furthermore, BRAF V600E and KRAS G13D were significantly more frequent in hypermutated CRCs, while G12V appeared more frequently in non-hypermutated CRCs but never in hypermutated populations, which is also in concordance with other reports.41,42
In stages I–III CRC patients, the prognostic value of RAS/BRAF is controversial. Roth et al. 43 showed that KRAS mutation status does not have prognostic value based on a study of 1404 stages II–III CRC patients. In addition, a study concluded that KRAS mutation was not associated with DFS in stage III colon cancer patients. 44 However, some studies,45,46 as well as a large retrospective analysis of stage III MSS CRC by Taieb et al., 47 have yielded opposite results. The prognostic value of BRAF is also controversial.48,49 We suspect that the reason for this controversy is that most previous studies have included hypermutated cohorts with MSI and POLE/POLD1 mutations. However, the prognosis of the hypermutated population is excellent, and previous studies and our study have clearly shown that many RAS/BRAF sites differ significantly between populations with and without hypermutation. Considering the distinct biological characteristics of the hypermutated cohort, we further explored the roles of RAS/BRAF allele mutations only in the CRCs with non-hypermutation. In our univariate analysis, prognosis was worse in CRCs with RAS/KRAS/NRAS mutations than in those with wild type but similar trends were not observed in the multivariate analysis. Further multivariate analysis found that patients with KRAS G13D mutation showed a worse DFS than those without KRAS G13D mutation, and the KRAS G13D mutation was the most significant independent prognostic factor associated with DFS. Based on our study, for patients with stages I–III CRC, we recommend that not only MMR status but also RAS/BRAF allele mutations be determined as early as possible. Patients with KRAS G13D but not BRAF V600E or other allele mutations in the non-hypermutated cohort may need more intensive adjuvant therapy and a more rigorous follow-up schedule.
It is controversial whether mCRC patients with RAS mutations have a poor prognosis, both at the gene and the exon levels.50–53 For example, one study showed a similar prognosis for KRAS exons 2 and 3 mutation, 54 while another study showed a better prognosis for KRAS exon 2. 55 Some studies focusing on RAS/BRAF codon mutations have been more controversial and even come to the opposite conclusions.16,18,56,57 Therefore, the classification of RAS/BRAF mutations in terms of genes, exons, and alleles is not accurate, and we need to overcome the limitations of these traditional classifications. In our study, the heterogeneity of mPFS of the first-line treatment in mCRCs between RAS/BRAF variants was well demonstrated at the allele level. The mPFS for each allele mutation was generally consistent with that previously reported, 58 indicating that our cohort could be representative of the entire mCRC population. We performed an innovative reclassification of RAS/BRAF mutations based on first-line mPFS and identified a high-risk RAS-mutated population with KRAS G13D/G12V/G12C, whose survival was much worse than that of the low-risk RAS/BRAF-mutated population, close to BRAF V600E.
Reclassification of RAS/BRAF mutations is to better stratify treatment. Except for a few patients with targetable mutations, most mCRC patients need to receive doublet or triplet chemotherapy plus targeted drugs (anti-EGFR or anti-VEGF). Previous studies have shown that patients with poorer biological behavior are more likely to benefit from aggressive triplet chemotherapy. For mCRCs with BRAF V600E, a first-line intensive triplet regimen is necessary 32 but for RAS/BRAF wild type, doublet chemotherapy combined with anti-EGFR is sufficient because of the benign biological behavior. 59 However, there has been controversy over patients with RAS mutations. More than 100 RAS mutant alleles have been identified in CRC, which has different biological functions and leads to different clinical outcomes or drug responses. As a result, gene-level analysis is no longer enough to accurately stratify treatments. At the same time, it should be taken into account that the mutation frequency of some RAS/BRAF alleles is very low, and it is difficult to obtain exact conclusions from allelic analysis alone. Regrouping RAS/BRAF allele mutations with similar prognoses is an effective strategy for stratified therapy. To verify the significance of regrouping for treatment decision-making, we analyzed which strategy of doublet or triplet chemotherapy was better in groups 1–4 and found that only group 3 (the high-risk RAS mutation) benefited from an intense triplet regimen. This indicates that for patients with high-risk RAS mutations (KRAS G13D/G12V/G12C), more intensive triplet chemotherapy plus targeted drug is needed, while for patients with low-risk RAS mutations in group 2, doublet regimen is sufficient. The reason for the negative result of group 4 was that the well-documented benefit of first-line triplet chemotherapy for BRAF V600E mCRCs influenced real-world treatment decisions at our center, resulting in a control sample size that was too small to analyze. To our knowledge, this is the first study to define a population with high-risk RAS mutations (KRAS G13D/G12V/G12C) who can benefit from an intensive first-line triplet regimen, and its implication for real-world treatment decision warrants further prospective randomized controlled clinical studies.
This study has some limitations. The main limitation is the insufficient follow-up time. Unable to provide accurate OS Kaplan–Meier curves, more than 60% of the patients survived at the end of the follow-up, and the shortest follow-up time was only approximately 10 months. Thus, the outcomes seem less rigorous. In addition, due to the retrospective nature of the study, a degree of selection bias was unavoidable.
Conclusion
The frequencies of various RAS/BRAF allele mutations varied between hypermutated and non-hypermutated cohorts or locally advanced and metastatic CRC. In the non-hypermutated cohort, the KRAS G13D mutation tended to be more common and aggressive for patients with stages I–III CRC. Our study revealed a significant association between high-risk RAS allele mutations and shorter PFS in non-hypermutated mCRC patients undergoing standard first-line therapy. Meanwhile, we found that intensive first-line therapy with triplet chemotherapy in combination with targeted therapy may be considered a viable and effective treatment option for high-risk RAS allele mutations.
Supplemental Material
sj-doc-1-tam-10.1177_17588359231225035 – Supplemental material for Reclassification of RAS/BRAF allele mutations predicts the survival benefit of triplet chemotherapy in metastatic colorectal cancer
Supplemental material, sj-doc-1-tam-10.1177_17588359231225035 for Reclassification of RAS/BRAF allele mutations predicts the survival benefit of triplet chemotherapy in metastatic colorectal cancer by Xiang Zhang, Haizhong Ma, Yinjun He, Wenguang He, Nan Chen, Yandong Li, Weixiang Zhong, Guosheng Wu, Xile Zhou, Hanju Hua, Feng Ye, Hui Cai and Weiqin Jiang in Therapeutic Advances in Medical Oncology
Supplemental Material
sj-doc-2-tam-10.1177_17588359231225035 – Supplemental material for Reclassification of RAS/BRAF allele mutations predicts the survival benefit of triplet chemotherapy in metastatic colorectal cancer
Supplemental material, sj-doc-2-tam-10.1177_17588359231225035 for Reclassification of RAS/BRAF allele mutations predicts the survival benefit of triplet chemotherapy in metastatic colorectal cancer by Xiang Zhang, Haizhong Ma, Yinjun He, Wenguang He, Nan Chen, Yandong Li, Weixiang Zhong, Guosheng Wu, Xile Zhou, Hanju Hua, Feng Ye, Hui Cai and Weiqin Jiang in Therapeutic Advances in Medical Oncology
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
