Abstract
Background:
Platinum derivatives are important treatment options for patients with esophageal carcinoma (EC), and a predictive marker for platinum-based therapy is needed for precision medicine.
Patients and methods:
This study contained two cohorts consisting of EC patients treated using platinum-based chemoradiation therapy (CRT) as the first-line and another external cohort of nationwide clinicogenomic data from the BioBank Japan (BBJ).
Results:
Genome-wide association study (GWAS) of therapeutic outcomes, refractory disease or not, following platinum-based CRT as first-line in 94 patients in the first cohort suggested the association of 89 SNPs using p < 0.0001. The top 10 SNPs selected from each chromosomal region by odds ratio were evaluated for progression-free survival (PFS) and overall survival (OS) hazard ratios in the first cohort, resulting in four candidates (p < 0.0025). The four selected candidates were re-evaluated in another cohort of 24 EC patients, which included patients prospectively enrolled in this study to fulfill the sample size statistically suggested by the results of the first cohort, and of the four, only rs3815544 was replicated (p < 0.0125). Furthermore, this candidate genotype of rs3815544 proceeded to the re-evaluation study in an external cohort consisting of EC patients treated with platinum derivatives and/or by radiation therapy as the first-line treatment in BBJ, which confirmed that the alternative allele (G) of rs3815544 was statistically associated with non-response (SD or PD) to platinum-based therapy in EC patients (odds ratio = 1.801, p = 0.048). The methylation QTL database as well as online clinicogenomic databases suggested that the region including rs3815544 may regulate MSX1 expression through CpG methylation, and this down-regulation was statistically associated with poor prognosis after platinum-based therapies for EC.
Conclusion:
rs3815544 is a novel candidate predictive marker for platinum-based EC therapy.
Keywords
Introduction
Esophageal carcinoma (EC) is one of the leading causes of neoplastic death worldwide.1,2 Currently, multimodalities consisting of neoadjuvant chemotherapy (ChT) or chemoradiation therapy (CRT) followed by surgery have become the gold standard treatment for EC.3,4 Alternatively, definitive CRT has become another powerful treatment for curability because some patients treated with neoadjuvant CRT followed by surgery achieved pathological complete remission (pCR). 5 In fact, comparable prognoses after definitive CRT to surgery alone have been reported by a couple of independent prospective studies.6,7 Thus, CRT is now the most important treatment option for EC, regardless of whether it is combined with surgery or not. Platinum derivatives such as cisplatin have played important roles in CRT.5,8 Even for patients with distant metastases, who are commonly encountered clinically, although radiation therapy is often omitted in those, platinum derivatives are still important treatment options. 9 Therefore, it is necessary to identify predictive markers for platinum derivatives.
Quite a few studies have attempted to identify factors predictive of platinum-based ChT/CRT efficiency, mostly by molecular-profiling endoscopic biopsies before treatment, but have not been as successful. 10 Actors such as intratumoral heterogeneity cause difficulties in identifying predictive markers using biopsy samples. 11 The subclonal nature of genomic driver mutations in tumor contributes to important clinical implications that may not necessarily be observed in a single biopsy. 12 Therefore, some studies have suggested several genetic polymorphisms as predictive markers for platinum-based therapies using candidate-gene or candidate-polymorphism approaches.13–17 However, the whole genome approach may be better because responsible genes or precise molecular pathways with anti-tumor effects have not yet been completely unveiled. 11
Therefore, we assessed the effectiveness of genome-wide association studies (GWAS) to unveil noble single-nucleotide polymorphisms (SNPs) as predictive markers for platinum-based therapy. For this purpose, we recently developed a population-optimized SNP array ‘Japonica SNP-array’ for a GWAS using resequencing data from 2,000 Japanese individuals. 18 This Japonica SNP array allowed us to catalog more comprehensive genetic polymorphisms in terms of predictive markers for platinum-based therapy, especially in the Japanese population. This study used the whole-genome approach in the first cohort, followed by candidate-SNP approach in the second cohort including prospective recruitment of patients, before re-evaluating a candidate-polymorphism in an external cohort obtained from the nationwide clinicogenomic database, the BioBank Japan (BBJ).19,20 We selected EC patients treated with platinum-based CRT for the two independent cohorts to make the treatment background as uniform as possible; the second cohort was designed and prepared based on the survival hazard ratios of the first cohort. On the other hand, the cohorts from BBJ consisted of clinicogenomic data from a number of hospitals from all over Japan, in which EC patients were treated by various therapeutic combinations with platinum derivatives and/or radiation therapies and reflected real-world clinical practice. The candidate SNP suggested by the two aforementioned cohorts was re-evaluated for platinum-based therapies as well as for radiation therapy, using this nationwide clinicogenomic data. We also tried to unveil the molecular and pathogenic functions induced by this candidate SNP by referring to online published multiomics databases. The present study followed the guidelines of the reporting recommendations for tumor marker prognostic studies (REMARK).21,22
Patients and methods
Ethical approval and consent to participate in two cohorts
This study contained two cohorts. The first cohort consisted of all of 94 patients who provided written informed consent to participate in this study between April 2003 and October 2005 at Tohoku University Hospital. The second cohort consisted of 24 patients, 12 of whom were enrolled at Tohoku University Hospital between December 2005 and June 2016, whereas the other half were enrolled at National Hospital Organization Mito Medical Center between May 2006 and February 2008. This study was approved by the Tohoku University Graduate School of Medicine IRB (permission number 2003-003, 2005-124, 2016-1-331), National Hospital Organization Mito Medical Center IRB (permission 20060529), National Hospital Organization Sagamihara National Hospital (permission 2018-051) and National Center for Global Health and Medicine (permission NCGM-A-003267). Written informed consent was obtained from all enrolled patients by a board-certified esophageal surgeon (TM).
All the authors confirmed that this study was performed in accordance with the Declaration of Helsinki.
Patient characteristics of the first and the second cohorts
Both cohorts consisted of patients with EC who were primarily treated with definitive CRT (Figure 1(a), Table 1). We selected the second cohort after the whole genome SNP association study was conducted, which eventually enrolled 24 patients with EC primarily treated using CRT, including 6 patients who were prospectively recruited to fulfill the requirements of the study design and the sample size as calculated based on the results obtained by the first cohort; the number of enrolled patients, 24, in the second cohort was suggested by ‘Two Arm Survival’ of The Cancer Research and Biostatics (CRAB) statistic (https://stattools.crab.org/Calculators/twoArmSurvival.html) using the following: α = 0.05, PFS hazard ratio = 5, proportion in standard group = 0.15, power = 0.8 for differences (β = 0.2), follow-up years = 10, and survival years of standard group = 8 years, based on the results for PFS by the first cohort. Both cohorts were Japanese and differed only in the dates of patient enrolment, as described above. The clinical and pathologic data of the enrolled patients are summarized in Table 1. We did not observe any familial relationships between the 118 patients.
Patients’ characteristics of the two cohorts in this study.
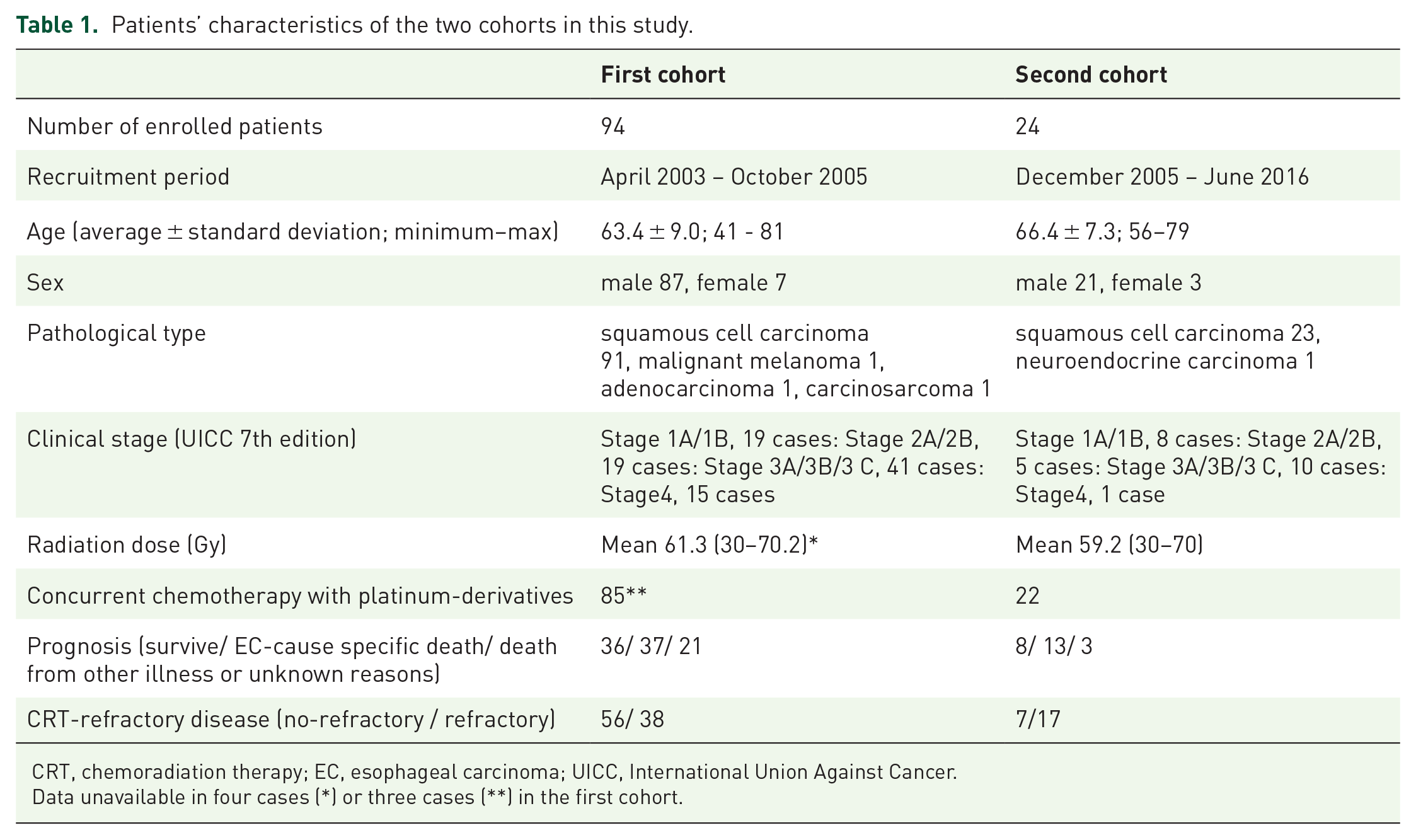
CRT, chemoradiation therapy; EC, esophageal carcinoma; UICC, International Union Against Cancer.
Data unavailable in four cases (*) or three cases (**) in the first cohort.

The flow diagrams of this study. (a) The flow diagram from two cohorts to the BBJ cohort. This study contained two cohorts. The first cohort consisted of 94 patients treated using platinum-based CRT as first-line. GWAS for screening of potential candidate SNPs for prognosis outcome, CRT-refractory disease or not, was applied to the first cohort, followed by OS/PFS analyses. The four candidate SNPs identified in the first cohort were examined in the second cohort recruited after the first cohort. Only rs3815544 was replicated by the candidate-SNP approach. (b) The flow diagram of the preparation of the external BBJ cohorts from nationwide clinicogenomic data. The clinicogenomic data from all EC patients enrolled in the BBJ were used. There were multiple registrations for chemotherapy data sheets or for radiation data sheets from a single patient because each registration was required for each chemotherapy regimen or for irradiation treatment for each lesion, as indicated by asterisks.
All patients were followed up until the date of death or for as long as possible. Clinical stages were determined based on the International Union Against Cancer (UICC) TNM classification of malignant tumors, 7th edition.23
SNP genotyping and imputation
Genomic DNA was isolated from the peripheral white blood cells of each patient by using a publicly known procedure after obtaining written informed consent. All samples were genotyped using the Japonica Array v1 (Toshiba Corp, Tokyo, Japan). We conducted genotype calling using Affymetrix Power Tools (version 1.18.2; Thermo Fisher Scientific Inc., Waltham, MA). All samples met the manufacturer’s quality control criteria (dish QC ⩾ 0.82 and sample call rate ⩾ 97%). SNPs that were categorized as ‘Recommended’ by the SNPolisher package (version 1.5.2; Thermo Fisher Scientific Inc.) were used for subsequent analyses. No cryptic relatives were found among the samples using the Maximum Unrelated Set Identification (IMUS) method implemented in PRIMUS v1.8.0 24 with default settings. Haplotype phasing was conducted using SHAPEIT (v2.r644) 25 as a pre-phasing for the genotype imputation after filtering SNPs with a call rate < 97.0%, and a Hardy–Weinberg equilibrium test result of p < 10−6, or minor allele frequency (MAF) < 0.5%. Genotype imputation was performed using IMPUTE2 (ver. 2.3.1) 26 with the options -Ne 2000, -k_hap 1000, -k 120, -burnin 15, and -iter 50 using a phased reference panel of 1,070 Japanese individuals.27,28 Each genotype was accepted if the highest genotype probability was higher or greater than 0.9. Otherwise, the genotype was treated as missing. Finally, SNPs in Hardy–Weinberg equilibrium with a p-value < 0.0001, call rate < 0.990, or MAFs < 0.01 were filtered out, and 4,095,500 SNPs were evaluated in the downstream analysis.
Statistical analysis
A genome-wide association test for the first screening of potential candidate SNP markers of prognosis outcome, CRT-refractory disease (n = 38) or not (n = 56), was applied to the imputed SNPs. We conducted the logistic regression analysis with sex as a covariate using Plink (version 1.9). 29 SNPs with p-values < 0.0001 were used in subsequent analyses as candidate SNPs to avoid losing potential candidates that might have an estimated P-value no less than GWAS significance (5.0E-8), since the sample size of this study was considered.
After a GWAS, we examined the hazard ratios (HRs) for PFS and OS by Cox univariate regression analysis including age at CRT, sex, radiation doses and clinical stages in the first cohort, and only age at CRT was statistically associated with HRs for PFS and OS (data not shown). We then incorporated age at CRT and each SNP into a Cox proportional hazard model to determine HRs for PFS and OS.
Survival analysis using the Kaplan–Meier method and hazard ratio evaluation using a Cox proportional hazard model were performed using SPSS (version27) (IBM, Armonk NY, USA).
The EC cohorts from the BioBank Japan (BBJ)
We selected all EC patients that were treated with platinum-based ChT/CRT as first-line therapy and evaluated by the Response Evaluation Criteria In Solid Tumor (RECIST) version 1.1 of post-treatment imaging (platinum-based therapy cohort as first-line) or those that were treated by radiation therapy with radiation doses no less than 40 Gy (radiation therapy cohort as first-line), from all of 1,338 EC patients in the BBJ project, as shown in Figure 1(b). We adopted tumor response estimated by RECIST version 1.1 as the endpoint of clinical outcome for the platinum-based cohort because there are missing values in long-term prognosis; however, therapeutic efficiency was precisely evaluated by post-treatment imaging according to RECIST in this BBJ-EC cohort. In contrast, the therapeutic outcome of radiation therapy was estimated by the 1-year, 2-year and 3-year survivals due to unavailability of RECIST evaluations in clinical data sheets for radiation therapy in BBJ.
Association between rs3815544 and methylation status of MSX1 gene CpG sites
We used the iMETHYL database published online (http://imethyl.iwate-megabank.org) to evaluate the contribution of the candidate SNP, rs3815544, to the methylation status of CpG sites around the MSX1 gene. The database provides whole-DNA methylation data (almost 24 million autosomal CpG sites) obtained from whole-genome bisulfite sequencing of CD4 + T lymphocytes and monocytes collected from a cohort of healthy subjects of Japanese ethnicity. 30
Evaluation of MSX1 gene features using online published databases
The data used for the analyses were obtained from the following four portals: Wanderer (http://www.maplab.cat/wanderer), 31 a tool that helps to visualize genomic and epigenomic data from The Cancer Genome Atlas (TCGA) Research Network (http://cancergenome.nih.gov/) was used for CpG methylation around the MSX1 gene in ECs and normal esophageal mucosae. The Gene Expression Omnibus (GEO) portal site (https://www.ncbi.nlm.nih.gov/geo/) was used for H3K27ac peaks in EC cell-lines from results using ChIP sequencing assays (GSE106433) and expression data in ECs and normal esophageal mucosae before neo-adjuvant CRT (GSE45670) or neo-adjuvant chemotherapies (GSE104958) using microarray and pathologic diagnoses of therapeutic effects on EC tumors after esophagectomies. Shiny Methylation Analysis Resource Tool (SMART) (http://www.bioinfo-zs.com/smartapp/#tab-8724-4) was used to assess the correlation between CpG-site methylation and MSX1 expression in ECs. The cBio Cancer Genomics Portal (https://www.cbioportal.org),32,33 from which 17 cBioPortal.org Esophagus/Stomach studies representing 3,791 patients were identified and filtered in June 2021 and prognostic outcomes such as progression-free survival (PFS) and disease-free survival (DFS) were compared between patients with (n = 50) and without (n = 3,241) MSX1 alterations among the 3, 291 patients examined for MSX1.
Summary of clinical endpoints of the first, the second and BBJ cohorts and neoadjuvant CRT/ChT cohorts in online published clinicogenomic database (GSE studies)
The clinical endpoints of each analysis in this study were as follows: CRT-refractory disease for a GWAS in the first cohort, hazard ratios for OS and PFS by a Cox proportional hazard model for selected 10 SNPs in the first cohort, hazard ratios for PFS by a Cox proportional hazard model for four selected SNPs in the second cohort, tumor response estimated by RECIST version 1.1 for the BBJ platinum-based cohort, and pCR induction for neoadjuvant CRT (GSE45670 study) or ChT (GSE104958 study). OS was not included in the second cohort because the mean follow-up period was not sufficient in the second cohort compared to the first cohort (28 months in the second cohort vs 79 months in the first cohort). OS and PFS of the first and the second cohorts were re-evaluated by Kaplan–Meier method. The reason that we adopted tumor response estimated by RECIST as an endpoint of clinical outcome in the BBJ platinum-based cohort is described above.
Summary of biological functions of rs3815544 and MSX1 in ECs examined by online omics databases
We used five online-published omics databases as follows: iMETHYL database to assess the association between rs3815544 and CpG methylation, Wanderer tool to assess CpG methylation status around the MSX1 gene in ECs and normal esophageal mucosae, GEO portal site to assess H3K27ac peaks in ECs (GSE106433) and to assess the association between pCR induction and MSX1 expression status before platinum-based neoadjuvant CRT (GSE45670) or ChT (GSE104958), SMART portal site to assess the correlation between CpG-site methylation and MSX1 expression in EC, and cBio Cancer Genomics Portal to assess MSX1 genetic alterations and prognosis of the EC/Stomach cancer cohort. The details of each online multiomics database are described above in the ‘Patients and Methods’ section.
Results
The first and the second cohorts’ characteristics
The characteristics of the first and the second cohorts of this study are summarized in Table 1. Mean follow-up months, from the initiation of the primary therapy with CRT to patients’ death or the end of follow-up, were 79.3 months and 27.9 months in the first and the second cohorts, respectively.
Pathologic diagnosis of squamous cell carcinoma, male patients, and mean age at initiation of primary treatment accounted for 97% (n = 91), 93% (n = 87) and 63.4 years, respectively, in the first cohort, and for 96% (n = 23), 83% (n = 21), and 66.4 years, respectively, in the second cohort, which were generally representative of typical Japanese patients with EC. We included histological types other than squamous cell carcinoma, such as malignant melanoma or neuroendocrine carcinoma, as long as they had been treated with platinum-based CRT as first-line treatment.
The mean radiation doses were 61.3 Gy in the first cohort (data unavailable in four cases) and 59.2 Gy in the second cohort, which were almost equal to the mean radiation doses of definitive CRT previously reported. 5 A total of 93 % (n = 85) of the first cohort (data unavailable in three cases) and 92 % (n = 22) of the second cohort received CRT with concurrent use of platinum derivatives.
The overall survival (OS) and progression-free survival (PFS) in both cohorts were analyzed using the Kaplan–Meier method. In the first cohort, median OS and median PFS were 88.10 months (48.7—127.5 months; 95% confidence interval (CI)) and 88.10 months (40.1—136.1 months; 95% CI), respectively. In the second cohort, median OS and median PFS were 13.40 months (7.7—19.1 months; 95% CI) and 5.00 months (2.6—7.4 months; 95% CI).
Potential candidate SNPs for poor prognosis outcomes
All SNPs identified in the first cohort are summarized as potential candidates for prognosis-outcome prediction after CRT for EC in Figure 2(a) and Table 2. Eighty-nine SNPs were identified as risk factor for CRT-refractory diseases (Table 2).
SNPs selected in the first cohort by GWAS with P-value < 0.0001.
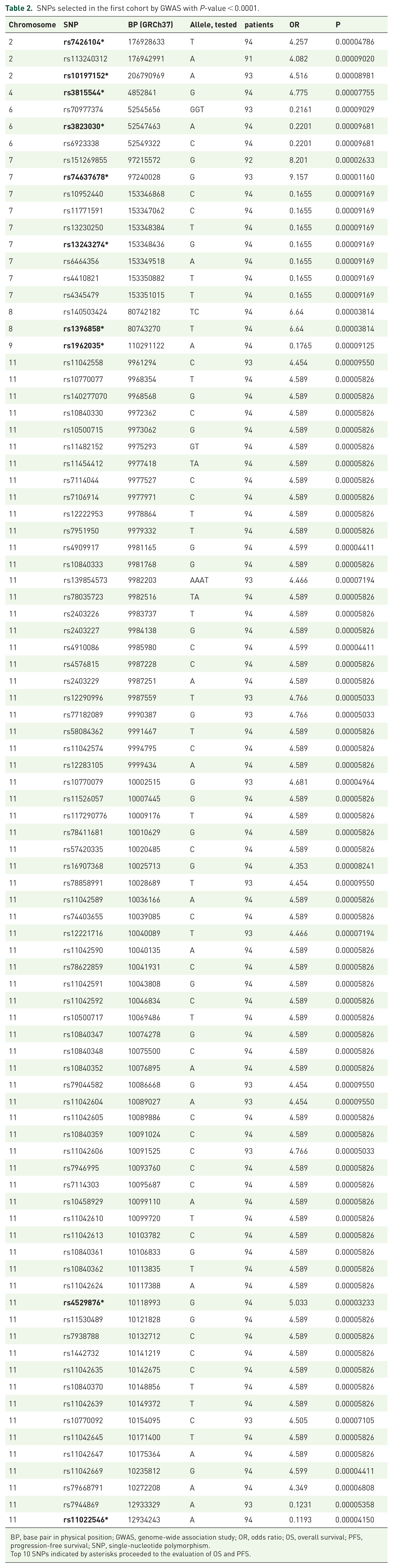
BP, base pair in physical position; GWAS, genome-wide association study; OR, odds ratio; OS, overall survival; PFS, progression-free survival; SNP, single-nucleotide polymorphism.
Top 10 SNPs indicated by asterisks proceeded to the evaluation of OS and PFS.

(a) Manhattan plot of association of SNPs with CRT-refractory disease in 94 EC patients; the results from 38 EC patients with CRT-refractory disease and 56 EC patients of control (non-CRT-refractory). (b) An example of the detailed view of GWAS; results around rs3815544 in the region of chromosome 4. rs3815544 was selected as a candidate predictive marker from this chromosomal region.
We selected the leading 10 SNPs filtered by chromosomal location (Table 2), odds ratio (OR), and the number of genotyped patients, and examined the hazard ratios for PFS and OS using a Cox proportional hazard model incorporating age and each SNP, because only age at CRT was statistically associated with prognosis by Cox univariate regression analysis among age, sex, radiation dose, and clinical stage (data not shown). The results of the multivariate analyses are summarized in Table 3 (upper panel). Four SNPs were significant in PFS or OS, with a p-value < 0.0025 after Bonferroni correction. Each of these four was included in the candidate-polymorphism approach in the second cohort in terms of PFS hazard ratio by multivariate analysis using a Cox proportional hazard model (Table 3, lower). Using this candidate-polymorphism approach, only rs3815544, located close to MSX1, was found to be statistically significant (p < 0.0125, after Bonferroni correction) among the four SNPs in the second cohort.
PFS and OS hazard ratios by Cox proportional hazard analyses, for each of 10 selected SNPs in the first cohort (upper) and four selected SNPs in the second cohort (lower).

CRT, chemoradiation therapy; OS, overall survival; PFS, progression-free survival; SNP, single-nucleotide polymorphism.
Four SNPs with significance, which proceeded to analysis by the second cohort, are marked with asterisks in the first cohort.
In the BBJ cohort for platinum-based therapy, we adopted tumor response estimated by RECIST version 1.1 34 as an endpoint of clinical outcome because there are missing values in long-term prognosis and follow-up; however, therapeutic efficiency was precisely evaluated by the post-treatment imaging according to RECIST in this BBJ-EC cohort (Figure 1(b)). The alternative allele (G) of rs3815544 was significantly associated with the risk of non-response (SD or PD) to platinum-based ChT/CRT (OR = 1.801, p = 0.0480) (Table 4, right), despite the fact that there were no statistical differences among the patients with each genotype of rs3815544 in terms of age, sex, clinical stage and details of treatment procedures (Table 4, left). In contrast, we could not find any statistical significance between rs3815544 genotype and the therapeutic effect of radiation therapy, although the number of patients might not be enough and the therapeutic effect by radiation therapy was only estimated by the 1-year, 2-year, and 3-year survivals due to unavailability of RECIST evaluations in clinical data-sheets for radiation therapy in BBJ (Figure 1(b)); the ORs of the 1-year, 2-year, and 3-year survivals were 1.71 (p = 0.2339), 1.28 (p = 0.5617) and 1.42 (p = 0.4331), respectively.
The association between genotypes of rs3815544 and RECIST-based evaluations by post-treatment imaging in the BBJ-EC cohort obtained from BioBank Japan.
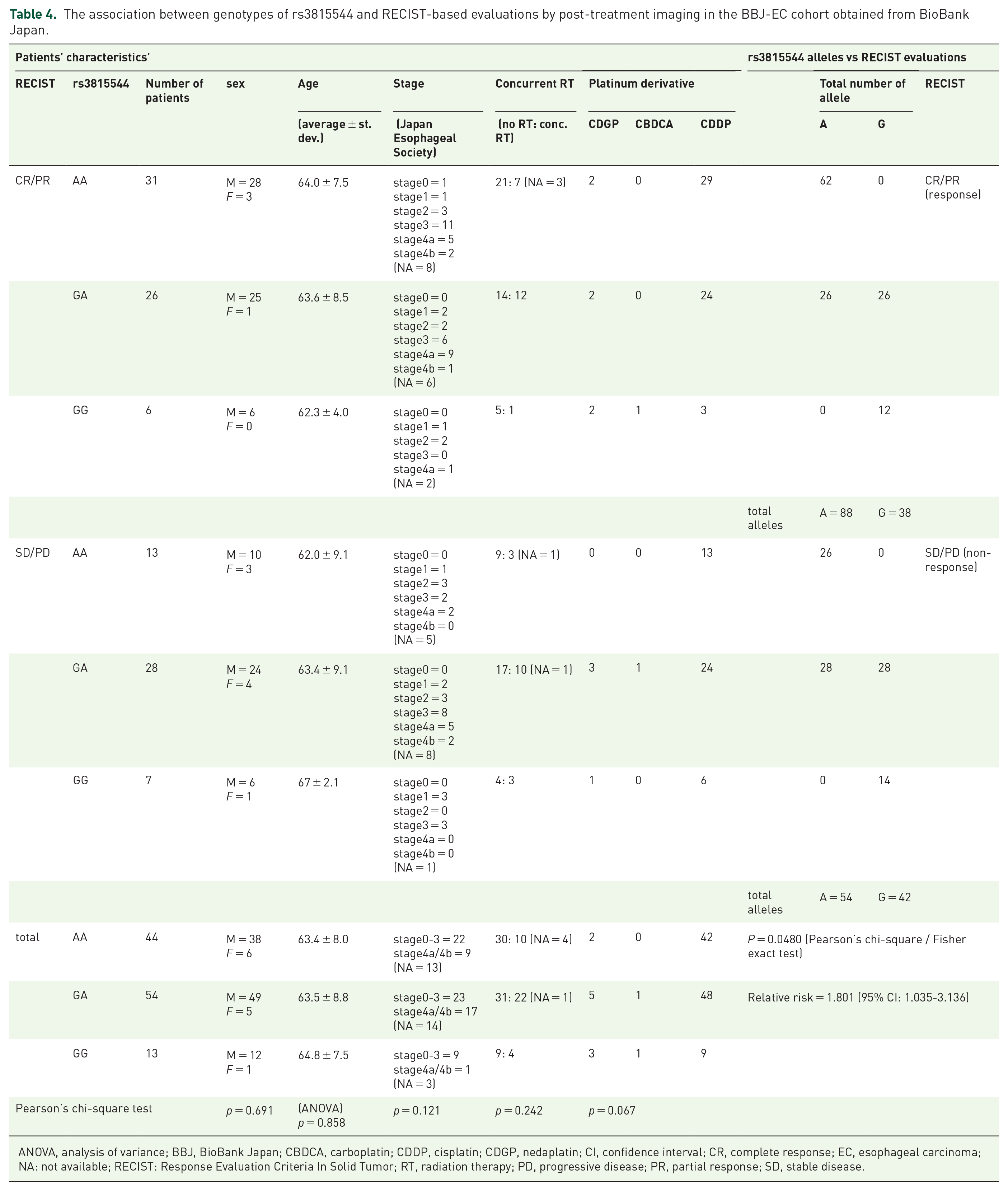
ANOVA, analysis of variance; BBJ, BioBank Japan; CBDCA, carboplatin; CDDP, cisplatin; CDGP, nedaplatin; CI, confidence interval; CR, complete response; EC, esophageal carcinoma; NA: not available; RECIST: Response Evaluation Criteria In Solid Tumor; RT, radiation therapy; PD, progressive disease; PR, partial response; SD, stable disease.
Attribution of rs3815544 to methylation of CpG sites within the 5′-flanking region of the MSX1 gene
We used the Wanderer portal site 31 containing original methylation data from TCGA to examine CpG methylation levels around MSX1 gene in ECs and normal esophageal mucosae. As shown in Figure 3 (upper), the CpG methylation probe near rs3815544 was mostly methylated in both the normal mucosa and EC. In addition, there were two regions that were more methylated in ECs than in normal mucosae, including the CpG site that was located at a transcription start site (TSS) and contained an H3K27 acetylation peak. Meanwhile, the contribution of rs3815544 to epigenetic modification of the promoter or enhancer regions of MSX1 gene was examined using iMETHYL, which revealed that rs3815544 contributed to the methylation of CpG sites not only around rs3815544 but also other CpG sites, including a CpG (chr4: 4861443), which was close to the TSS of MSX1 and H3K27 acetylation peak (Figure 3, middle). Specifically, the CpG site at chr4: 4861443 is located within the CpG island that contains other CpG sites, such as chr4: 4861101, chr4: 4861330, and chr4: 4862240. Among them, methylation of chr4: 4861330 and the region between chr4: 4861675 and chr4: 4861880 was associated with MSX1 suppression by previous reports,35,36 and the methylation of 13 CpGs between chr4: 4861101 and chr4: 4862240 was statistically correlated with the suppression of MSX1 within ECs by the SMART (Figure 3, lower).
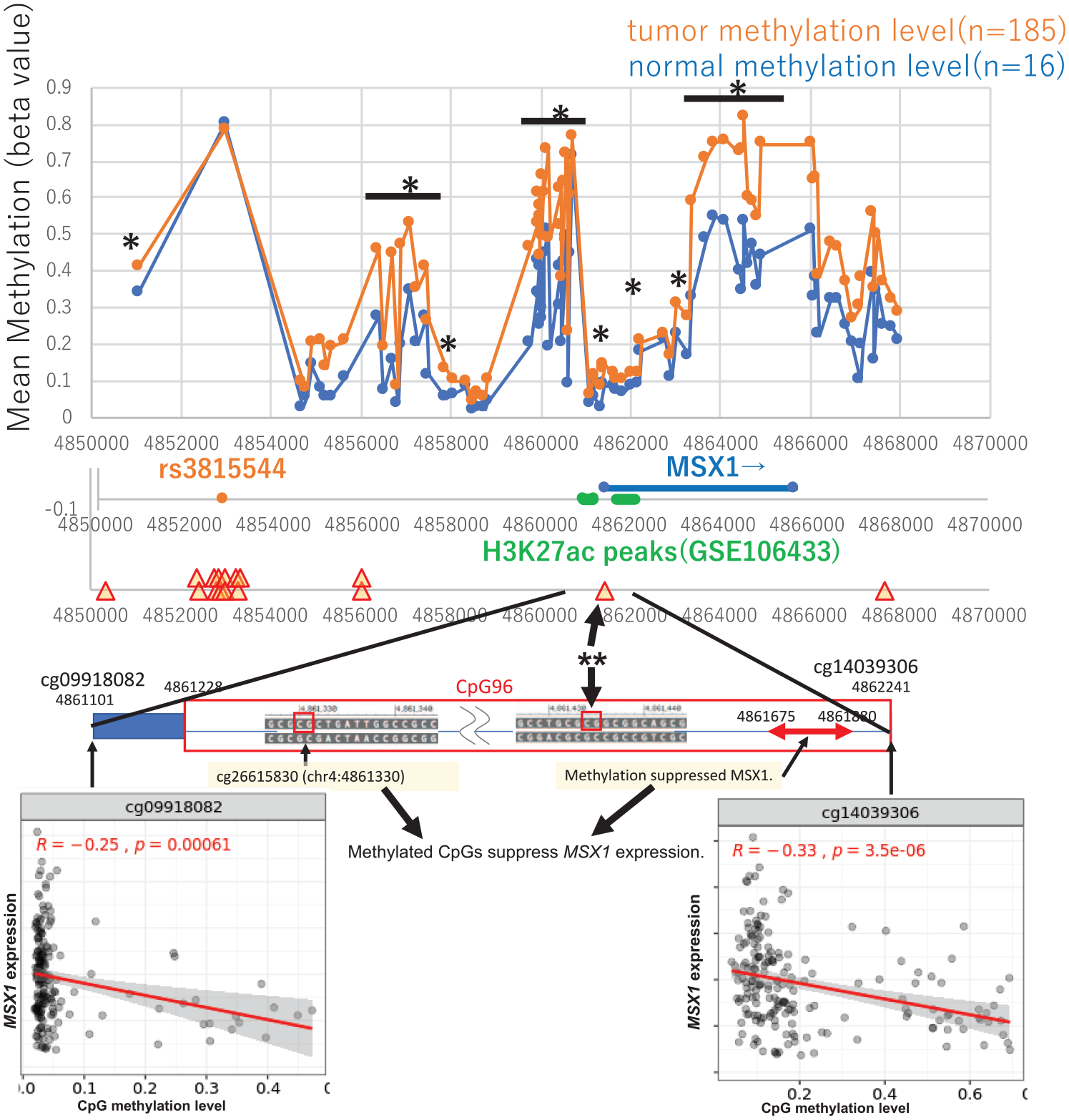
The mean methylation of MSX1 and its promoter region in normal esophageal mucosa (blue) and tumor tissues of esophageal carcinoma (EC, red) were adopted from ‘Wanderer’ (http://www.maplab.cat/wanderer) in the upper row. Mean methylation as the beta-value is indicated along the Y-axis, and chromosomal locations are indicated on the X-axis (GRCh37). Each asterisk indicates a CpG site with a statistically significant methylation status difference between normal mucosae and tumor tissues (p < 0.05). An asterisk with an under-bar indicates a region significantly more methylated by all probes within the region. Attributions of rs3815544 to CpG sites are based on the iMETHYL database, and H3K27ac peaks in EC cell lines are derived from ChIP assay results (GSE106433), as shown in the middle of this figure. rs3815544 is indicated by an orange dot. Triangles indicate rs3815544-attributed CpG methylations. H3K27ac peaks around the MSX1 gene in the EC cell-lines KYSE510, KTSE70, and TT, suggested by ChIP sequencing, are indicated by green bold lines. The online database, SMART, suggested that CpG site indicated by an arrow with double asterisks locates within CpG island (indicated as CpG96) containing other CpG sites (named as cg09918082 and cg14039306) that suppress MSX1 within ECs through methylation, as shown at the bottom. The other two CpG sites were also reportedly associated with MSX1 suppression.
MSX1 gene expression and EC prognoses after platinum-based chemotherapy or chemoradiation therapy in online published data
Comprehensive gene expression data from pretreatment biopsies and pathologic diagnoses for neoadjuvant platinum-based ChT or CRT efficiencies are available in GSE45670 and GSE104958.37,38 We downloaded the data from the GEO portal site. The total number of ECs examined by microarray was 68 (28 in GSE45670 and 40 in GSE104958), accompanied by 15 normal mucosae (10 in GSE45670 and five in GSE104958). We determined the standard value of MSX1 expression using the average and standard deviation (STDV) calculated from all values in normal mucosae in each GSE study. We summarized the suppression of MSX1, which we determined below one STDV from the average in each GSE study, compared to pathologic diagnoses after platinum-based ChT or CRT (Table 5). The results showed that none of the cases with suppression of MSX1 showed pathological complete remission (pCR), and the relative risk of negative MSX1 for non-pCR was estimated as 1.488 with 95% confidence interval 1.1254—1.766 (p < 0.05).
MSX1 expression levels before CRT or neoadjuvant chemotherapy and pathologic assessment of EC after neoadjuvant chemotherapy or CRT, by the online published data.

CRT, chemoradiation therapy; EC, esophageal carcinoma; pCR, pathological complete remission; SD, standard deviation.
Inactivating alterations such as homozygous deletions of MSX1 as prognostic-marker candidates of EC suggested by online published data
DFS and PFS were compared between the two groups, with or without MSX1 alterations, and Kaplan–Meier curves were generated using the cBio Cancer Genomics Portal. EC patients, including gastric cancer patients in some studies, with altered MSX1 had significantly worse prognoses both in DFS and PFS, with p-values of 1.082 × 10–5 and 0.0225, respectively (Figure 4(a) and (b)). Of EC patients with altered MSX1, 50% (n = 25) were homozygous deletions (Figure 4(c)). Although there were many missing values in the details of treatment procedures in this clinicogenomic data, platinum derivatives such as oxaliplatin and cisplatin were used in all cases for whom treatment data were available, excluding gastric acid suppressants (Figure 4(d)).

In the current study, 17 cBioPortal.org Esophagus/Stomach studies representing 3,791 patients were identified and filtered in June 2021, from the cBio Cancer Genomics portal (https://www.cbioportal.org). Prognostic outcomes such as PFS and DFS between the two subgroups, that is, EC/GC patients with or without MSX1 alterations, were compared by Kaplan–Meier method. EC/GC patients with altered MSX1 significantly had worse prognoses both in DFS (a) and PFS (b) with p-values of 1.082 × 10-5 and 0.0225, respectively. (c) Of those with altered MSX1, 50% (n = 25) were homozygous deletions. (d) Although there were many missing values in details of treatment procedures in this clinicogenomic data, platinum derivatives such as oxaliplatin or cisplatin were used in all cases available for treatment information, excluding gastric acid suppressants. Each figure above was created and adopted from cBio Cancer Genomics (https://www.cbioportal.org).
Discussion
This study involved GWAS screening for CRT-refractory diseases, followed by OS and PFS hazard ratio evaluation using a Cox proportional hazard model in the first cohort. This implicated the four candidate SNPs, as well as the sample size and follow-up periods of the second cohort for validation. Among these four, only rs3815544 was statistically confirmed in the validation study by the second cohort. This result was also re-evaluated using other external nationwide clinicogenomic data from the BioBank Japan (BBJ). Furthermore, the molecular and pathogenic functioning of this SNP was assessed using online published multiomics databases. The Wonderer portal site was used to study the methylation status around MSX1 gene within ECs and corresponding normal mucosae; the iMETHYL portal was used to investigate the relation between candidate SNP and CpG methylations around MSX1 gene. Finally, the SMART portal was used to assess the relation between CpG methylations and expression status of MSX1 within ECs; in addition, data from two GSE studies were analyzed to identify the relations between MSX1 suppression and pCR induction in ECs.
To our knowledge, this study is the first to suggest a predictive SNP candidate utilizing the population-optimized whole-genome SNP array for patients with EC that had been treated primarily using platinum-based therapy. For patients with EC primarily treated using platinum-based therapy as first-line treatment, the number of enrolled patients expected from previous studies and the incidence of this disease suggested that the current study design must be similar to that of rare and lethal cancer studies, 39 rather than previous GWASs that suggested predictive markers of more common malignant diseases such as breast cancer, lung cancer, and cervical cancer.40–42 In fact, the total number of enrolled patients, 118 in this study, and the length of observation after CRT – an average of 80 months of the first cohort – have been rarely observed in previous studies. For example, two representative large-scale long-term cohort studies on CRT for ECs consisted of 55 EC patients with a median survival time of 16 months and 56 EC patients with 14 months of median survival time.5,43
We re-evaluated GWAS results using a Cox hazard proportional model for OS and PFS, incorporating age at CRT and each of 10 SNPs as potential candidates from GWAS filtered by chromosomal location, ORs and the number of genotyped patients (Table 2). From the above, we selected four SNPs (Table 3), which were used in the candidate-polymorphism study in the second cohort consisting of 24 patients that were suggested by CRAB statistics, including patients who were prospectively recruited to fulfill the study design. Eventually, only one SNP, rs3815544, was found to be significant hazard ratios for PFS (Table 3, lower).
Kaplan–Meier analyses showed that PFS and OS were significantly associated with each of alleles of rs3815544, suggesting the alternative allele (G) as a risk allele following first-line platinum-based CRT (Figure 5). For the first and the second cohorts, we selected EC patients treated with platinum-based CRT to make the treatment background uniform, but we predicted that rs3815544 might be predictive not only for CRT but also for platinum-based chemotherapy for the reasons suggested by preceding studies, as described later. Therefore, we decided that this result should be validated in another external cohort of EC patients treated with platinum-based therapy obtained from BBJ.

Kaplan–Meier survival curves of 118 EC patients from the two cohorts in this study by the genotypes of rs3815544. Survival ratios were significantly dependent on the genotypes of rs3815544 by Kaplan–Meier analyses, both in PFS with p = 0.000058 (logrank) (a) and OS with p = 0.000151 (log rank) (b); median survival time in each genotype was 5.0 months (GG), 26.0 months (GA), 162.6 months (AA) in PFS, and 22.4 months (GG), 53.6 months (GA), 162.6 months (AA) in OS.
BBJ is one of the largest disease biobanks in the world and included 1,338 patients with EC.19,20 We prepared the external cohort to investigate the association between platinum derivatives and objective evaluations of therapeutic effects such as RECIST evaluation (Figure 1(b)). The reasons that we adopted RECIST evaluation of post-treatment imaging as an endpoint of the prognosis outcome were as follows: first, patients with EC of the BBJ cohort included those who were not available for medium- or long-term follow-up data for survival analyses; second, precise, objective, and widely shared methods to evaluate therapeutic effects were needed because this nationwide cohort consisted of clinicogenomic data from a number of hospitals all over Japan. The results indicated that the alternative allele (G) of rs3815544 was statistically associated with tumor non-response (SD or PD) (Table 4, right). It should be noted that the result was replicated even in an external cohort that consisted of patients treated with various combinations of platinum derivatives with or without concurrent radiation therapy. There were no statistical differences in patient background, such as age, sex, and clinical factors such as clinical stages and the use of concurrent radiation therapy, between genotypes of rs3815544 in this BBJ-EC cohort (Table 4, left). In fact, patients with relatively early stage ECs might account for a substantial portion of those with homozygous alternative alleles (GG) (p = 0.121), which might rather support the view that the alternative allele (G) of rs3815544 is associated with poor prognosis. We also investigated the association of the rs3815544 genotype and the therapeutic effect of radiation therapy but did not find any statistical significance.
MSX1 reportedly plays an important role as a functional tumor suppressor in the genesis of various types of human cancers.44–48 A germline variant in MSX1 was identified in a Dutch family with clustering of Barrett’s esophagus and esophageal adenocarcinoma, suggesting that the loss of MSX1 plays an important role in esophageal carcinogenesis. 49 In fact, MSX1 was listed among candidate driver genes for esophageal carcinogenesis, as suggested by copy number variants from the TCGA database. 50 Furthermore, a promoter hypermethylation signature of MSX1 has been reported in various types of human malignancies by genome-wide screening for promoter methylation, 51 which has also been confirmed in EC (Figure 3). Consistently, MSX1 alterations in EC were mostly homozygous deletions, as suggested by the cBio Cancer Genomics Portal database (Figure 4(c)). Taking these factors into consideration, MSX1 was suspected to play an important role in esophageal carcinogenesis, suggesting the presence of an EC subpopulation driven by deleterious MSX1 either from homozygous deletion or promoter methylation.
The suppression of MSX1 has previously been reported to be associated with platinum-resistant diseases in high-grade epithelial ovarian cancers, in which the expression of MSX1 was dependent on CpG-methylation status. 52 High MSX1 expression was listed among the genes highly expressed in the intestinal type of gastric carcinoma, which is more sensitive to platinum-based chemotherapy than the diffuse type. 53 These previous studies strongly suggest that the expression/suppression of MSX1 contributes to sensitivity/resistance to platinum derivatives. We examined the association between MSX1 expression and pathological CR (pCR) induction in EC patients using online published data; comprehensive gene expression data by microarray before neo-adjuvant CRT/ChT and pathologic diagnoses after treatments have been published online as GSE45670 study (platinum-based CRT) and GSE104958 study (platinum-based ChT: a combination of cisplatin, docetaxel, and 5-fluorouracil).37,38 This result was compatible with the current study; all cases with reduced MSX1 showed non-pCR. The estimated relative risk of reduced MSX1 for non-pCR was 1.488 (p < 0.05, 95% confidence interval 1.1254—1.766) (Table 5). In addition, the cBio Cancer Genomics Portal database suggested that EC patients with MSX1 alterations that mostly consisted of homozygous deletions had poor prognoses in both DFS (p = 1.082 × 10-5) and PFS (p = 0.0225) compared to those without MSX1 alterations (Figure 4(a) and (b)). Platinum derivatives, such as oxaliplatin and cisplatin, were used in the all patients for whom treatment data were available. Taken together, it is plausible that the suppression of MSX1 might confer resistance to platinum-based chemotherapy in patients with EC.
To date, there is no direct evidence of an association between rs3815544 and MSX1 expression in EC. However, rs3815544 could be attributed to CpG methylation around the 5′-flanking region of MSX1 (Figure 3, upper panel). Using online multiomics databases such as Wonderer, iMETHYL and SMART, we found that rs3815544 contributed to the methylation of some CpG sites, including one at chr4: 4861443. This CpG site at chr4: 4861443 was close to the transcription start site (TSS) of MSX1 and the H3K27 acetylation peak and located within the CpG island that contains other CpG sites, such as chr4: 4861101, chr4: 4861330, and chr4: 486224 (Figure 3, middle). These three CpG sites were all reportedly associated with MSX1 suppression in ECs35,36 (Figure 3, bottom). Therefore, the G allele of rs3815544 induces the methylation of CpGs around the H3K27 acetylation sites and downregulates MSX1, which was supported by a phenomenon of gene expression control by DNA methylation and histone modification. 54
In conclusion, the current study suggests a novel predictive marker candidate of platinum derivatives for EC patients, but further studies are needed to validate the findings in another large-scale clinical trial as well as by molecular research to unveil the underlying mechanisms of MSX1 suppression through CpG methylation around the 5′-promoter/enhancer region.
Footnotes
Acknowledgements
The authors are grateful to Dr. Akira Ono (National Center for Global Health and Medicine) for his technical support and helpful advice. The authors also express sincere thanks to all physicians and laboratory members who have contributed to the foundation and development of the BBJ, which has been added to the coauthors’ list as a banner.
Author contributions
Conflict of interest statement
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: TM received personal fees from OncoTherapy Science outside the submitted work in 2017 and 2018. MN holds a concurrent post at the Department of Cohort Genome Information Analysis endowed by Toshiba Corporation. MN received research funding from Toshiba Corporation.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by a Grant-in-Aid from the Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan [grant numbers 18591451 and 16K10488], and was partially supported by the Japan Agency for Medical Research and Development (AMED) [grant number JP18km0405205], the Center of Innovation Program from the Japan Science and Technology Agency, JST, and the research fund from Repertoire Genesis.
