Abstract
Background:
The early detection of digestive cancers and precancerous diseases remains a significant challenge. This study aimed to investigate the performance of the blood methylated SEPT9 (mSEPT9) assay, and the combination of this assay with serum protein markers, in hospital-based opportunistic screening strategies for digestive cancers.
Methods:
Opportunistic screening was performed in the participating hospitals on outpatients and inpatients who met specific inclusion criteria. We recruited a total of 2030 subjects, including 764 cancer patients [291 colorectal cancer (CRC), 239 gastric cancer (GC), 106 esophageal cancer (EC), and 128 hepatocellular carcinoma (HCC)], 423 subjects with precancerous diseases, and 843 normal subjects. All samples were transported to an authenticated clinical laboratory where the mSEPT9 tests were performed.
Results:
When used separately, the mSEPT9 detected CRC, GC, EC, and HCC, with a sensitivity of 76.6% [area under the receiver operating characteristic curve (AUC) = 0.86)], 47.7% (AUC = 0.76), 42.6% (AUC = 0.69), and 76.7% (AUC = 0.85) and a specificity of 94.6%, 92.3%, 92.5%, and 87.7%, respectively. The mSEPT9 assay also had potent ability to discriminate cancer from non-cancer subjects. The combination of mSEPT9 with CEA, CA724, SNCG, or AFP significantly enhanced the sensitivity for CRC, GC, EC, and HCC to 86.4% (AUC = 0.99, specificity = 92.8%), 63.6% (AUC = 0.86, specificity = 91.1%), 71.3% (AUC = 0.81, specificity = 82.1%), and 83.3% (AUC = 0.93, specificity = 85.1%), respectively. The performance of the mSEPT9 assay was influenced by cancer stage, patient age, pathological types, and the location of cancer. We also identified that mSEPT9 was an independent risk factor and was a valuable predictor for the long-term survival of digestive cancer patients, with a hazard ratio of 2.84, 2.07, 1.88, and 2.45, for CRC, GC, EC, and HCC, respectively.
Conclusion:
The blood mSEPT9 assay, whether used alone or in combination with serum protein markers, is effective for the opportunistic screening of digestive cancers. Furthermore, mSEPT9 is an independent risk factor and a predictive marker for the long-term survival of digestive cancer patients.
Introduction
The methylated SEPT9 (mSEPT9) assay has been approved by the US Food and Drug Administration (FDA), the European Union, and the Chinese FDA as a test for colorectal cancer (CRC) screening or for the early detection of disease. This assay has been widely recognized as an effective method for the screening of CRC in average-risk populations, 1 high-risk populations, 2 or for the early detection of disease.3,4 The sensitivity of this assay for CRC detection ranges from 48.2% to 95.6%, with a specificity of 79.1–99.1%. 3 Although its efficacy for CRC screening has been proven by several studies, the utility of the test for the population screening of CRC still requires improved sensitivity for the detection of early-stage cancers and advanced adenomas. Opportunistic screening has been suggested as an effective method for cancer screening in a hospital environment. This strategy was designed to screen symptomatic subjects visiting hospitals for diagnosis and therapy, and therefore represents a screening strategy for high-risk populations. Both the fecal immunochemical test (FIT) and the mSEPT9 assay have been used for the opportunistic screening of CRC with satisfactory levels of performance.3–5
The development of effective blood methylation markers for the detection of cancer faces many challenges. One of the major issues is the balance between sensitivity and specificity. Several methylation markers exhibit acceptable levels of sensitivity across several types of cancer, although specificity was low due to an unsatisfactory capability for discriminating between cancer types or the discrimination of malignant from benign lesions.6–8 However, these markers can still be used for cancer screening since further imaging or pathological methods are available for diagnostic confirmation. In this scenario, pan-cancer screening by blood methylation markers may be a good choice for initial screening as this represents a cheap, convenient, and sensitive method. Although the performance of the mSEPT9 assay for CRC has been confirmed by many studies, 3 its ability to detect other cancers, especially digestive cancers, has not been fully investigated. Some preliminary reports suggested that the mSEPT9 assay can detect hepatocellular carcinoma (HCC), 9 indicating potential for the detection of digestive cancer beyond its current use for CRC screening.
In this study, we systematically investigated the performance of the mSEPT9 assay, and its combination with serum protein markers, for the detection of CRC, gastric cancer (GC), esophageal cancer (EC), and HCC, along with corresponding benign lesions and subjects with no evidence of diseases (NED) in an opportunistic screening program. We found that the mSEPT9 assay may be capable of detecting all four types of digestive cancers with acceptable levels of sensitivity and specificity, with better performance for CRC and HCC than for GC and EC. The assay can also effectively discriminate cancers from benign lesions and NED. The combination of the mSEPT9 assay with CEA, CA724, SNCG, or AFP further improved the detection performance for CRC, GC, EC, and HCC. Furthermore, the mSEPT9 assay was identified as an independent risk factor and survival predictor for all types of digestive cancers. For the first time, our study demonstrated the efficacy of mSEPT9 for the opportunistic screening and long-term survival prediction of digestive cancers, thus facilitating its future use for pan-cancer screening of the digestive system.
Materials and methods
Ethics
The detailed experiment plans and protocols for the study were submitted to the ethics/licensing committees of the named participating hospitals for review and approval before the start of the clinical study, and were approved by the corresponding committees of hospitals, including the Chinese PLA General Hospital, the Fifth Medical Center of the Chinese PLA General Hospital, and the Eighth Medical Center of the Chinese PLA General Hospital. Confirmation of approval for clinical studies was received from the ethics board of the Chinese PLA General Hospital (approval number: S2018-081-02) before the start of the clinical study, and was shared by all participating centers. Informed consent was obtained from all subjects involved in the study before the collection of blood samples, and all subjects were informed of the usage of plasma and the test results.
Study design, patients, and confirmation of diagnosis
The opportunistic screening study was designed and performed in four hospitals with the mSEPT9 assay [BioChain (Beijing) Science and Technology, Co., Ltd., Beijing, China], including the Chinese PLA General Hospital, the Fifth Medical Center of the Chinese PLA General Hospital, the Eighth Medical Center of the Chinese PLA General Hospital. Clinical status of all subjects was not determined before blood draw for SEPT9 assay, and blood samples were obtained from all outpatients and inpatients who met the selection criteria in the participating hospitals (Supplemental Figure 1). The main inclusion criteria were: adults over 18 years old with complete clinicopathological information and confirmed diagnosis of digestive cancer, digestive benign diseases, or no evidence of disease (normal subjects) by imaging examination (including endoscopy, ultrasound, magnetic resonance imaging, computed tomography, etc.) and/or subsequent pathological examination. The main exclusion criteria were: pregnant woman and history of any cancer. Information on patient age, sex, family history, Tumor/Lymph node/Metastasis (TNM) staging, cancer pathological types, primary tumor size, metastatic location, cancer differentiation and mSEPT9 test results were collected. Subjects with incomplete information were also excluded (Supplemental Figure 1).
As a result, a total of 2030 subjects were recruited (Table 1). The 764 cancer patients recruited comprised 291 CRC, 239 GC, 106 EC, and 128 HCC; 423 subjects with precancerous benign diseases were recruited, comprising 164 subjects with colorectal adenoma or hyperplastic polyps, 60 with atrophic gastritis, 114 with chronic esophagitis, esophageal achalasia or Barrett’s esophagus, and 85 with hepatic cirrhosis (HC). A total of 843 subjects with NED were recruited, comprising 295 in the colon and rectum group, 224 in the stomach group, 116 in the esophagus group and 208 in the liver group (Table 1). The cohorts of CRC patients and the corresponding normal subjects were from one of our previous studies 4 and were used here for comparison with GC, EC, and HCC.
Clinicopathological factors and the number of patients for four types of cancers, corresponding benign lesions, and normal subjects involved in this study.

Ade/HP, colorectal adenoma or hyperplastic polyps; AG, atrophic gastritis; CE/EA/BA, chronic esophagitis, esophageal achalasia or Barrett’s esophagus; CRC, colorectal cancer; EC, esophageal cancer; GC, gastric cancer; HC, hepatic cirrhosis; HCC, hepatocellular carcinoma.
Sample size estimation
Sample size estimation was based on the following equation for known detection sensitivity: N = Z 2 × [p (1–p)]/E 2 . The parameters are defined as follows: Z is a statistical parameter [Z = 1.96 for 95% confidence interval (CI)]; E represents the error (10% was chosen in this study), and p represents the putative positive detection rate). The p value for each cancer was obtained from previous reports or our pilot studies looking at the sensitivity of the mSEPT9 assay in digestive cancers. If the p value is 0.7, 3 0.5, 0.4, and 0.759 for CRC, GC, EC, and HCC, respectively, an estimated 81 CRC cases, 96 GC cases, 92 EC cases, and 72 HCC cases were required, making a total of at least 341 cancer cases. Considering that there may be 10% loss due to unqualified samples, the total number of cancer cases should be at least 375. The NED subjects were recruited in a ratio of at least 1:1 to that of the cancer cases (Table 1).
Sample collection and storage
Samples were collected from outpatients and inpatients, and the sample information was recorded in sample collection forms. A 10 ml peripheral blood sample was collected with 10 ml K2EDTA anticoagulant tubes (BD biosciences, Franklin Lakes, NJ, USA) to ensure the accuracy of the assay. Sample storage and transportation followed the instructions of the mSEPT9 assay kit. In brief, the K2EDTA anticoagulant tubes with collected blood samples were stored under 4°C after blood draw, and were centrifuged at 1350 × g for 10 min under 4°C within 4 h of blood collection. The supernatant (plasma) was centrifuged again at 1350 × g for 10 min under 4°C to remove the cell debris and avoid genomic DNA contamination. The plasma was stored under –80°C for future DNA extraction. All samples were transported in the presence of dry ice within 2 weeks from collection to a certified clinical laboratory (lab) for lab-developed testing, and the mSEPT9 tests were performed in the same lab.
DNA extraction and qualitative polymerase chain reaction analysis of SEPT9
DNA extraction from plasma samples and bisulfite conversion were performed according to the manufacturer’s instructions of the mSEPT9 assay [SensiColon, BioChain (Beijing) Science and Technology, Inc.; http://www.biochainbj.com/products_detail/productId = 25.html]. 4 In brief, the mSEPT9 assay [polymerase chain reaction (PCR) fluorescence probe method] is composed of two general steps. First, total cell-free DNA in plasma was extracted from 3.5 ml plasma samples (10 ml whole blood sample) using the plasma processing kit manufactured by BioChain (Beijing) Science and Technology, Inc. The DNA was then incubated with bisulfite, in which unmethylated cytosine is converted to uracil while methylated cytosines are not. In the second step, methylated target sequences in the bisulfite-converted DNA (bisDNA) template are amplified by real-time PCR on an ABI 7500 Fast Dx Real Time PCR device (Life Technologies). Instead of performing three PCR reactions containing 30 µl each as in the Epi proColon 2.0 assay, a single 60 µl PCR was performed for each sample in the new mSEPT9 assay, and therefore double the amount of extracted DNA was used in the single PCR of the modified assay compared with the Epi proColon 2.0 assay. PCR results for beta-actin (ACTB) and mSEPT9 for each reaction were recorded using the instrument software. The validity of each sample batch was determined on the basis of mSEPT9 and ACTB threshold count (Ct) values for the positive and negative controls. ACTB was used as an internal reference to assess the integrity of each sample. Positive control was regarded as valid when the Ct of SEPT9 ⩽41.1 and the Ct of ACTB ⩽29.8, and negative control was regarded as valid when no Ct is obtained and the Ct of ACTB ⩽37.2. The controls were tested in parallel with the samples for each plate at each PCR run. To minimize the plate-to-plate variation, the plasma cell-free circulating DNA was extracted from the same volume of plasma and the bisDNA was prepared using the identical procedure for each plate. The identical amount of bisDNA was mixed with the PCR master mix each time the reaction was established. The tests were performed by the same group of laboratory technicians using the same set of instruments. The Ct values of ACTB were assessed for all samples on each plate and compared with empiric values. If the Ct values of ACTB failed to pass the standard or exhibited huge difference from empiric values, the assay procedure would be examined and standard samples would be run to identify any variation.
Analysis of serum protein marker levels and interpretation
The levels of serum protein markers CEA, AFP, CA50, CA125, CA153, CA199, CA242, and CA724 were tested by the corresponding chemoluminescence kits [Roche Diagnostics(Shanghai) Co., Ltd.] and the hospital in-house automated chemoluminescence equipment (Roche Cobas 6000). The cut-off value for the above protein markers was established based on the recommendations from the instruction for use, and was 5 ng/ml for CEA, 20 ng/ml for AFP, 24 U/ml for CA50, 35 U/ml for CA125, 25 U/ml for CA153, 37 U/ml for CA199, 20 U/ml for CA242, and 6 U/ml for CA724. The cut-off value for gamma synuclein (SNCG) was set at 1.0 ng/ml based on the instruction for use from the manufacturer (Sino Biological Co., Ltd., Beijing, China).
Data analysis and interpretation
The data from the PCR reactions of the assay was analyzed using the 1/1 algorithm, which means that a sample was considered to be positive if the result of PCR was positive and was considered to be negative if the result of PCR was negative. For each sample, a relative methylation value was determined using the ΔΔCt method adapted for DNA methylation analyses as previously described. 10 In brief, ΔΔCt values were calculated as follows.

Calibrator here refers to the positive control. Analyses including the Student t-test, one-way analysis of variance, χ test, and receiver operating characteristic (ROC) curves, Kaplan–Meier survival curves were performed and figures were plotted with Graphpad PRISM 5.0 software (GraphPad Software, Inc., La Jolla, CA 92037, USA). ΔΔCt values, instead of Ct values, were used throughout this study to demonstrate the quantification of mSEPT9 levels in all statistics and analysis. The optimal cut-off in ΔΔCt for each type of cancer was determined based on the best balance between sensitivity and specificity by calculating the Youden’s index (sensitivity + specificity – 1). The bigger the Youden’s index was, the better the balance was between sensitivity and specificity. The corresponding cut-off value for the biggest Youden’s index was chosen as the optimal cut-off.
When both mSEPT9 and any one of the protein markers were combined in each cancer, the combined test was considered to be positive when either assay was positive and was considered negative when both assays were negative.
Results
Performance of the mSEPT9 assay for detecting cancers and precancerous diseases
In order to evaluate the performance of the blood mSEPT9 assay for four types of digestive cancers, we first defined the cut-off values for each type of cancer by examining blood samples from subjects with cancers, precancerous diseases, or NED. We then plotted ROC curves for cancer against NED, cancer against precancerous diseases, and precancerous diseases against NED. The cut-off values were defined based on the ROC curves, the best balance between sensitivity and specificity (the optimal cut-off for the best balance between sensitivity and specificity was determined by the Youden’s index), and the best division of ΔΔCt values for cancers and precancerous diseases against NED (Figure 1; top row). The final cut-off was determined as Ct values of 41.0 (ΔΔCt = –4.0) for CRC, 41.0 (ΔΔCt = –4.0) for GC, 41.1 (ΔΔCt = –4.1) for EC, and 40.9 (ΔΔCt = –3.9) for HCC.

The detection performance of the methylated SEPT9 assay in CRC, GC, EC, and HCC and corresponding precancerous diseases, and NED. The top row shows the ΔΔCt values for each type of cancer, precancerous diseases, and NED. The dashed lines indicate the cut-off values for distinguish positive from negative detection. The middle lane shows the receiver operating characteristic curves for cancer versus NED, cancer versus benign diseases, and benign diseases versus NED for colon and rectum, stomach, esophagus, and liver. The bottom lane shows positive detection rate for cancers, benign diseases and NED. The 95% confidence interval is shown for the positive detection rate of all groups in the bottom row.
The blood mSEPT9 assay showed the best performance for CRC and HCC when cancers were defined as positive and NED was defined as negative, with a sensitivity of 76.6% (AUC = 0.86, 95% CI: 71.3–81.4%) and 76.7% (AUC = 0.85, 95% CI: 68.1–83.4%) and a specificity of 94.6% and 92.3%, respectively. However, the assay was less potent for the detection of GC and EC, with a sensitivity of 47.7% (AUC = 0.76, 95% CI: 41.2–54.2%) and 42.6% (AUC = 0.69, 95% CI: 33.0–52.4%), and a sensitivity of 92.5% and 87.7%, respectively (Figure 1; middle and bottom rows). This level of detection remained effective when cancers were defined as positive, and benign diseases (precancerous diseases) were defined as negative, with an AUC of 0.78, 0.77, 0.70, and 0.67 for CRC, HCC, GC, and EC, respectively (Figure 1; middle row). However, the level of detection was less potent when precancerous diseases were defined as positive and NED was defined as negative; the AUC was 0.51, 0.66, 0.60, and 0.57, for colorectal, hepatic, gastric, and esophageal diseases, respectively (Figure 1; middle row). It was also apparent that the mSEPT9 assay may be a potent marker for distinguishing cancer from non-cancer conditions (precancerous diseases and NED), but was less potent in discriminating precancerous diseases from NED.
In order to characterize the profile of mSEPT9 detection in digestive cancers, we next investigated the detection sensitivity of this assay for different stages and locations of cancers. Figure 2 shows that the detection sensitivity was lower for stage I than stage II–IV for all types of cancers, and that there was no significant difference between stages II–IV (except for the much higher sensitivity of stage IV in EC), thus suggesting a higher detection sensitivity for more severe conditions. In terms of the sensitivity of this assay for different locations, we found no significant difference in sensitivity among CRC samples from the ascending, transverse, descending or sigmoid colon, and the rectum. Furthermore, there were no significant differences in sensitivity in terms of ECs from the upper, middle, or lower esophagus. No significant differences in sensitivity were observed in terms of HCCs from the left or right liver. However, significantly higher levels of sensitivity were observed in GC from the body of the stomach than those from the fundus and pylorus. These observations suggest that cancer location did not affect the detection sensitivity of mSEPT9, except for GC.
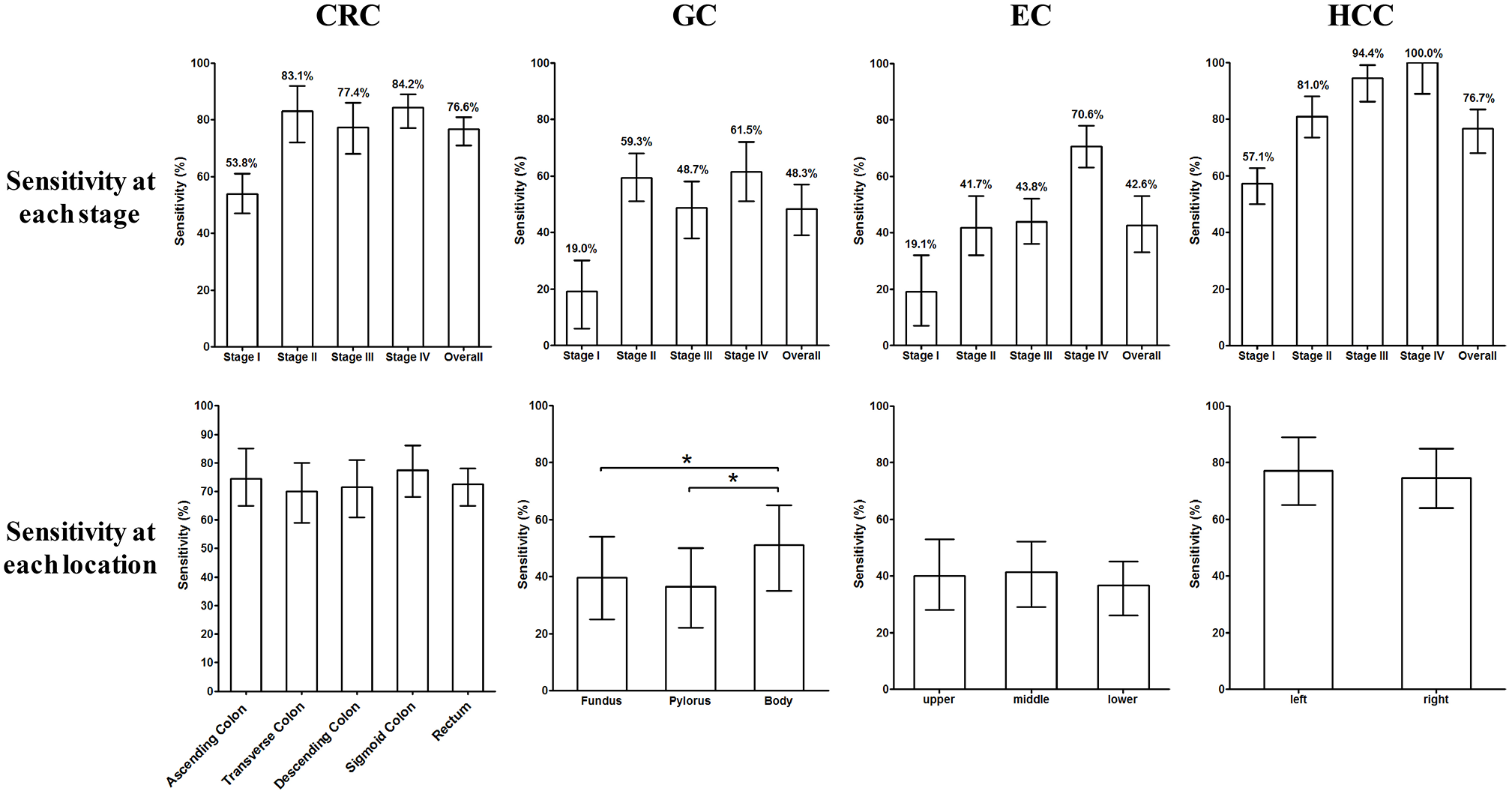
Stage-relevant and location-relevant sensitivity of methylated SEPT9 (mSEPT9) in the four types of cancers. The upper lane shows the sensitivity for each stage from stages I–IV for CRC, GC, EC, and HCC. The lower lane shows the sensitivity of mSEPT9 for each cancer location in CRC, GC, EC, and HCC. The 95% confidence interval is shown for the sensitivity of each group.
Next, we investigated the detection sensitivity in terms of different pathological types and levels of differentiation. In terms of gross classification, infiltrative CRC exhibited significantly higher positive detection rate (PDR) than ulcerative and protruding CRC, although there were no differences in sensitivity when compared among ulcerative/infiltrative, superficial/flat, and protruding cancers in GC and EC (Figure 3; top row). In terms of histological classification, we observed no differences in sensitivity among tubular adenocarcinoma, mucinous adenocarcinoma, and other types of CRC, among adenocarcinoma, mucinous carcinoma, and other types of GC, or among squamous cell carcinoma and adenocarcinoma in EC (Figure 3; middle row). The sensitivity of the assay for highly differentiated CRC was significantly lower than that for subjects with moderate and low differentiation, although this difference was not observed with GC and EC (Figure 3; bottom row). This comparison was not performed for liver cancer as HCC was the only pathological type investigated in this study.
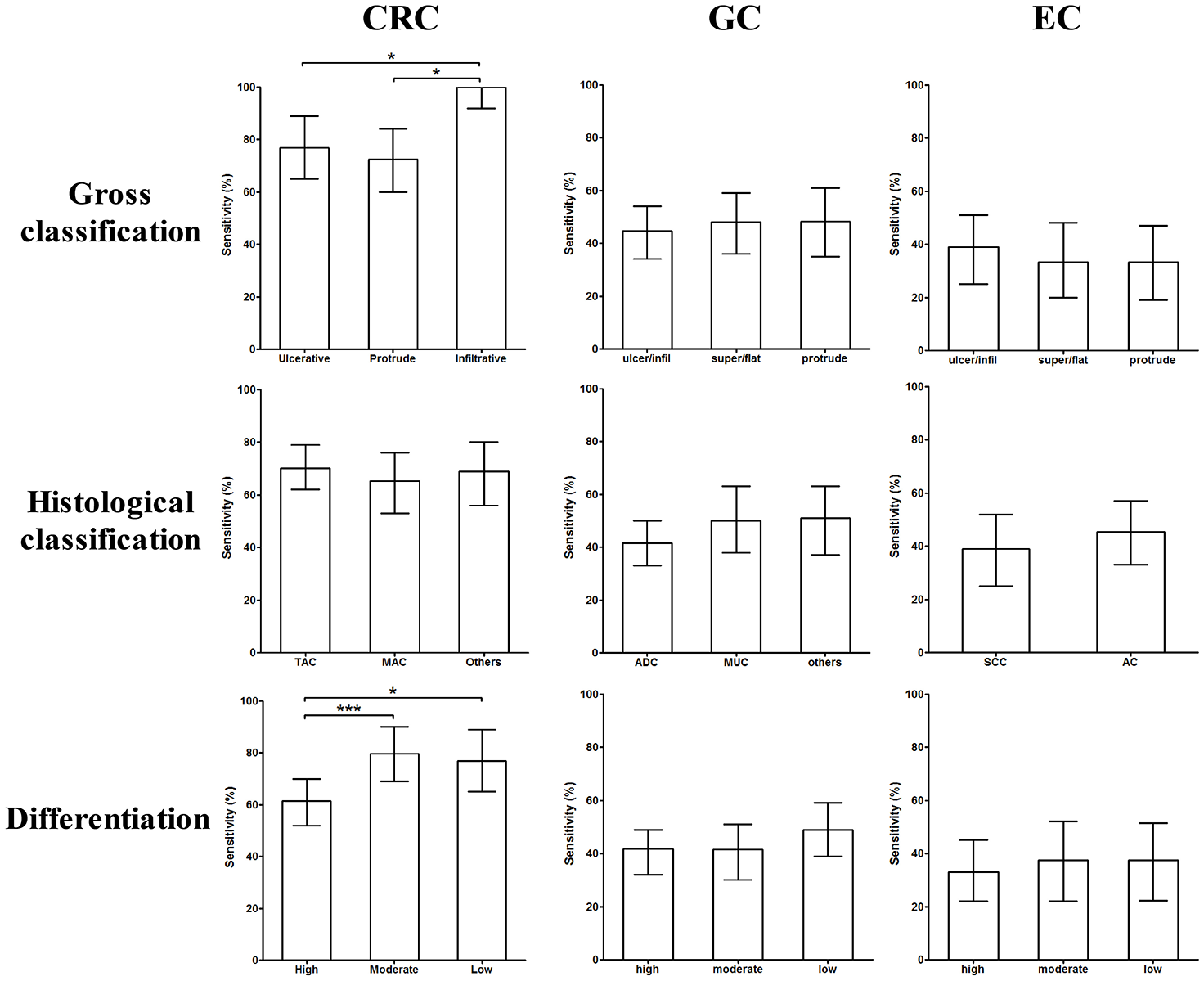
Comparison of detection sensitivity for various pathological types and differentiation of CRC, GC, and EC. Gross classification is compared in the top row, histological classification is compared in the middle row, and various degrees of differentiation are compared in the bottom row. The 95% confidence interval is shown for the sensitivity of each group.
Further analysis on patient gender revealed that males and females exhibited almost identical levels of sensitivity in the mSEPT9 assay for all four types of digestive cancers (Figure 4; upper row), thus suggesting that there were no gender preferences with regard to detection. In contrast, analysis by different age groups showed that the mSEPT9 assay exhibited clear age-dependent sensitivity for all types of digestive cancers. This was more obvious for GC, EC, and HCC than for CRC, in which there was a clear trend for increased sensitivity with increasing age (Figure 4; lower row).
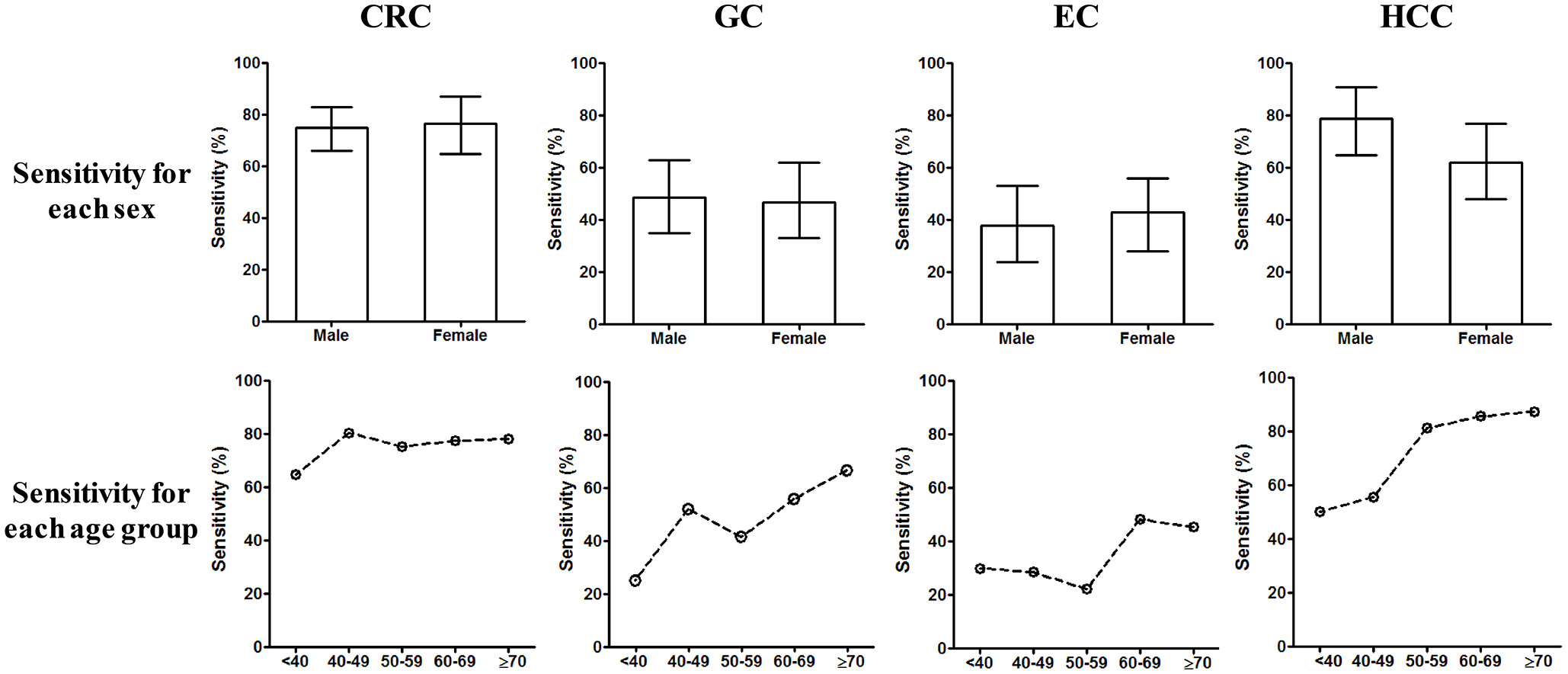
Comparison of detection sensitivity of methylated SEPT9 in different sexes and age groups. The upper lane shows the comparison of sensitivity between male and female in the four types of cancers. The lower lane shows the comparison of sensitivity at different age groups in the four types of cancers. The 95% confidence interval is shown for the sensitivity of each group.
The performance of the combination of mSEPT9 and protein markers for opportunistic screening
In order to improve the detection sensitivity of digestive cancers in opportunistic screening, we combined mSEPT9 assay with a range of protein markers and tested the diagnostic capability of this combined screen. The selection of protein markers was based on the assay packages available in hospitals and covered most of the commonly used markers for each type of cancer. First, we identified the most potent protein markers from the available serum protein assays by parallel examination of multiple markers. The protein marker with the highest levels of sensitivity for each type of cancer was identified and its performance in combination with the mSEPT9 assay was examined. Figure 5 shows that CEA (sensitivity: 40.3%), CA724 (sensitivity: 45.2%), SNCG (sensitivity: 41.5%), and AFP (sensitivity: 56.7%) were the markers with the highest levels of sensitivity for CRC, GC, EC, and HCC, respectively, at a detection specificity of 95% (Figure 5).

The detection sensitivity of protein markers in the four types of cancers. Comparison of several serum protein markers is shown for CRC, GC, EC, and HCC. The 95% confidence interval is shown for the sensitivity of each group.
Next, we studied the detection sensitivity of digestive cancers when the mSEPT9 assay and protein markers were combined (Figure 6; upper row). Figure 6 shows that this combination significantly enhanced the detection performance for all types of cancers compared with the mSEPT9 assay alone, or any protein marker alone. For CRC, the combination of the mSEPT9 assay and CEA increased the sensitivity to 86.4% (AUC = 0.99) at a specificity of 92.8%; this was significantly higher than the mSEPT9 assay alone (sensitivity = 76.6%, AUC = 0.86) or the CEA alone (sensitivity = 40.3%, AUC = 0.74). For GC, the combination of the mSEPT9 assay and CA724 increased the sensitivity to 63.6% (AUC = 0.86) at a specificity of 91.1%; this was significantly higher than the mSEPT9 assay alone (sensitivity = 47.7%, AUC = 0.76) or CA724 alone (sensitivity = 45.5%, AUC = 0.72). For EC, the combination of the mSEPT9 assay and SNCG increased the sensitivity to 71.3% (AUC = 0.81) at a specificity of 82.1%; this was significantly higher than the mSEPT9 assay alone (sensitivity = 42.6%, AUC = 0.79) or SNCG alone (sensitivity = 41.5%, AUC = 0.69). In HCC, the combination of mSEPT9 and AFP increased the sensitivity to 83.3% (AUC = 0.93) at a specificity of 85.1%; this was not significantly different from the mSEPT9 assay alone (sensitivity = 76.7%, AUC = 0.85), but was significantly higher than AFP alone (sensitivity = 56.7%, AUC = 0.80).

Detection sensitivity and ROC curves for mSEPT9, individual protein markers, and combined detection. The upper lane shows the detection sensitivity for mSEPT9 and CEA, CA724, SNCG, and AFP in CRC, GC, EC, and HCC, respectively. The lower lane shows the ROC curves for mSEPT9 alone, individual protein marker alone, or both combined in the four types of cancers. The 95% confidence interval is shown for the sensitivity of each group.
Pre-therapeutic mSEPT9 levels were good predictors for long-term survival
Next, we examined the predictive capability of mSEPT9 by following up all cancer patients and comparing survival data between patients with positive pre-therapeutic mSEPT9 detection and those with negative detection. Patients were followed up to 820 days for CRC, 610 days for GC, 798 days for EC, and 2012 days for HCC. Figure 7 shows that the pre-therapeutic mSEPT9 level was a predictive marker for the long-term survival of all four types of digestive cancers, regardless of cancer stage and the methods of treatment. Significant differences in survival were identified between the mSEPT9 positive and negative groups for CRC (p < 0.01), GC (p < 0.05), EC (p < 0.05), and HCC (p < 0.05). Interestingly, when survival data was stratified by cancer stage (Supplemental Figure 2), significant or almost significant differences were detected between mSEPT9-positive and -negative subjects in stage III of CRC (p = 0.15), GC (p = 0.047), EC (p = 0.071), and HCC (p = 0.265); and in stage IV of CRC (p = 0.052), but not in stage I and stage II. Further multivariate regression analysis showed that the mSEPT9 assay was an independent risk factor for predicting long-term survival, with a hazard ratio of 2.84 (95% CI: 1.38–5.82) for CRC, 2.07 (95% CI: 0.96–4.46) for GC, 1.88 (95% CI: 1.08–3.35) for EC, and 2.45 (95% CI: 0.95–6.32) for HCC. These observations confirmed the predictive role of mSEPT9 for all types of digestive cancers.

Kaplan–Meier survival analysis for CRC, GC, EC, and HCC patients. Patients were divided into two groups based on mSEPT9 detection, that is, positive or negative, and were followed up to 820 days in CRC, 610 days in GC, 798 days in EC, and 2012 days in HCC.
Discussion
Application of the mSEPT9 assay for the opportunistic screening of digestive cancers
The mSEPT9 assay was initially developed for the early detection of CRC 3 and was subsequently approved by the US FDA as a screening test for CRC 1 for average-risk populations above 50 years of age. Many studies have investigated the performance of the mSEPT9 assay and have demonstrated that the assay was effective for both the detection and screening of CRC. 3 Although some studies have evaluated the use of the mSEPT9 assay to detect other forms of cancer, including HCC 9 and lung cancer, 11 such studies are scarce and have yet to be fully addressed. Systematic studies of the performance of the mSEPT9 assay for digestive cancers have yet to be carried out. The present study represents the first systematic investigation of the performance of the mSEPT9 assay for pan-digestive cancers as part of an opportunistic screening strategy, and examined the capability of the assay for screening CRC, GC, EC, and HCC. By combining the mSEPT9 assay with serum protein markers, we found that the efficacy of the assay was significantly enhanced, rendering the assay very useful for clinical practice.
Opportunistic screening is quite different from screening an average-risk population. Opportunistic screening aims to screen symptomatic outpatients and inpatient populations who come to hospital for diagnosis or treatment. Consequently, the chances of cancer diagnosis are much higher than those in an average-risk population. As such, this is not a case of screening a well-defined population; patients could present with a variety of disease backgrounds and associated risk factors. However, this does represent a good opportunity to identify cancer patients and those with precancerous conditions, as patients are normally very compliant in a hospital environment, and it is much easier to organize such screening in hospital where subjects can be examined easily and sent for other tests if necessary. Therefore, it is more practical and easier to perform cancer screening in a hospital environment than in the community when the accessibility of screening is low for large average-risk populations.
Our group was the first to perform opportunistic screening for CRC using the mSEPT9 assay.2,4 In the present study, we expanded the range of this screening package for other digestive cancers, and enhanced the performance of the assay by combining it with protein assays. Our work showed that the mSEPT9 exhibited similar detection capability for HCC as for CRC. Although the detection of HCC using the mSEPT9 assay has been reported previously in a western population, 9 the current study represents the first study in a Chinese population and in patients with precancerous conditions, such as HC. Our study showed that the mSEPT9 assay was not only specific for CRC; it could also be used to detect HCC, thus suggesting that these two forms of cancer involve similar changes in mSEPT9. Furthermore, mSEPT9 detected HC with a sensitivity of 34.1%, thus suggesting that alterations in the methylation status of SEPT9 already happen before HC transforms into HCC. However, this also creates difficulties with regard to discriminating HC from HCC with the mSEPT9 assay. The diagnosis of HC is more dependent on imaging techniques, such as ultrasonic examinations. The detection sensitivity of mSEPT9 with regard to precancerous conditions of the colon, rectum, stomach, and esophagus was similar to the positive rate of detection for the NED population; thus, the blood-based mSEPT9 assay was not effective enough for detecting benign conditions. Since this assay was designed to detect cancer, it has proved to be a good assay with which to distinguish benign from malignant lesions in all of the digestive cancers studied thus far. Our study also showed that the detection of GC and EC was less potent than that of CRC and HCC, and that the sensitivities for the two types of cancer were very close. This observation suggests that alterations of SEPT9 methylation in GC and EC may be similar, perhaps due to anatomical similarities; these observations were different from those for CRC and HCC. These observations may also suggest that GC and EC exhibit growth patterns that differ from CRC, as the detection of mSEPT9 in blood is heavily dependent on the release of DNA from cancer cells into the blood. If the growth of cancer does not involve abundant angiogenesis and vessel invasion, the detection rate for mSEPT9 may be low. Further research should now investigate the lower levels of sensitivity for GC and EC.
Factors that could alter the performance of the mSEPT9 assay for the screening of digestive cancer
Previous reports highlight the fact that the performance of the mSEPT9 assay for CRC can be influenced by cancer stage, patient age, and pathological types, but not by cancer location or patient gender.12–20 Almost all reported blood methylation markers are stage-dependent, thus implying that the detection sensitivity of early-stage cancer is generally not as good as that of subsequent stages.12–20 This is more obvious for stage 0 or stage I cancer, in which the sensitivity was 50–60% or lower.12–20 However, this does not hinder the use of the assay for the early detection of cancer; it can still detect over half of cancer cases; this is far better than most other in vitro diagnostic tests. Furthermore, some studies suggest that the mSEPT9 assay can detect colorectal precancerous conditions, such as adenoma and polyps. Although this level of sensitivity appears to be low, the assay can still be used for screening, as evidence has shown that routine and repetitive screening can detect early stage precancerous or cancerous lesions.2,21 Therefore, stage-dependent detection should not be a problem for the mSEPT9 assay during screening. In the present study, we observed stage-dependent sensitivity for all types of cancer, clearly indicating the pan-cancer intrinsic properties of the assay.
In our study, differences in detection sensitivity that were relevant to specific pathological types were observed only with CRC; no such differences were observed for GC and EC. Interestingly, it appeared that CRC, which has a more severe phenotype (i.e. infiltrative type and low/moderate differentiation) exhibited higher levels of sensitivity. This suggests that the detection sensitivity of the mSEPT9 assay for CRC may be positively correlated with the severity of cancer. Previous observations suggested that the higher sensitivity for more severe phenotypes resulted from higher levels of SEPT9 methylation in the blood. 22 This is in accordance with the observation that higher stages of cancer exhibited higher levels of sensitivity. Furthermore, our data also suggested that there were no significant differences in blood mSEPT9 levels when compared across different pathological types in GC and EC; these findings were different from those for CRC.
Age-dependent detection is widely observed for methylation markers. 23 Evidence has shown that normal tissues undergo low-level de novo methylation of CpG islands, although this increases with age.24–26 Furthermore, aberrant methylation at some loci in tumors also took place in normal cells in vivo.27–29 Previous investigations of mSEPT9 suggested that the background methylation in normal subjects increased substantially over the age of 60 years. 12 In the present study, we also observed a significant increase in the detection sensitivity of the mSEPT9 assay in patients above 60 years of age for GC and EC, and above 50 years of age for HCC. This could be due to the additive effect of background methylation and cancer methylation. Although we did not observe such trends for CRC in the present study, previous studies have described this possibility. 12
In the present study, we also found that GC located in the stomach body exhibited significantly higher levels of sensitivity than those located in the fundus or pylorus, while the location of cancer did not affect the detection sensitivity of the mSEPT9 assay for CRC, EC, and HCC. This means that the assay can detect CRC, EC, and HCC in a non-biased manner. The reasons underlying this difference in sensitivity across different locations of GC are not clear but are worth investigating further.
The combination of the mSEPT9 assay with protein markers for the screening of digestive cancer
The combined detection of methylation and protein markers for clinical scenarios requires acceptable levels of sensitivity and specificity, good levels of complementation for the markers being used, and no substantial increase in costs. In the present study, we systematically investigated the combination of the mSEPT9 assay with the best-performance protein markers for CRC, GC, EC, and HCC. Although these protein markers have been widely used in clinical tests, we proved for the first time that the combination of these markers with the mSEPT9 assay further enhanced the detection performance of both types of markers in an opportunistic screening strategy. We previously showed that the combination of mSEPT9 and CEA could enhance the detection of CRC; 4 in the present study, we demonstrated that the same rationale can be applied to GC, EC, and HCC. The combined effect was more obvious for GC and EC, in which the detection sensitivity of mSEPT9, or protein marker alone, was not satisfactory. The combination of these factors increased the levels of sensitivity to an acceptable level that should benefit the screening of both cancers. In contrast, the sensitivity of the mSEPT9 assay alone for CRC and HCC was already high and the enhancement in sensitivity by combining this assay with other factors was statistically significant but not as large as those for GC and EC. We believe that this combined test is particularly useful for the future opportunistic screening of digestive cancers because it has high levels of sensitivity, is cost-effective, and is very convenient to carry out.
Prediction of long-term survival and therapeutic responses in clinical practice
The mSEPT9 has been demonstrated to be an independent risk factor and can predict the long-term survival of both CRC and HCC.9,30,31 In the present study, we demonstrated that this assay is an independent factor and predictive marker for long-term survival for all four types of digestive cancers. Predictions based on dichotomized pre-therapeutic mSEPT9 interpretation suggested that patients with digestive cancer who had undetectable levels of mSEPT9 in their blood prior to therapy would generally exhibit significantly better survival rates than those with detectable levels of mSEPT9 in their blood. Since the blood level of mSEPT9 is correlated with the severity of cancer, this observation suggests that cancers in patients who are negative for mSEPT9 prior to therapy may be less severe or self-constrained. These include some early-stage cancers and a minority of advanced cancers. This property of mSEPT9 may be used in the clinical prediction of survival and to facilitate the selection of therapeutic strategies. Indeed, this marker has been shown to be effective in a variety of applications for CRC, including the screening of average-risk populations, 1 opportunistic screening,3,4 the monitoring of therapeutic effects in surgery, 30 the monitoring of therapeutic effects during late-stage CRC, 31 and the prediction of therapeutic effects and long-term survival in patients with CRC.30,31 However, it should be noted that the prediction of survival by mSEPT9 was largely dependent on cancer stage, and that patients with advanced stages of cancer have higher levels of mSEPT9 in their blood; consequently, these patients are more likely to test positive and have a worse prognosis than those with early stages. Therefore, our stratification of survival by stage clearly showed that the marker may be predictable for the long-term survival of stage III–IV patients but not predictable for stage I–II patients under the current study design and follow-up duration. This could be due to the fact that the follow-up time may not be long enough for patients with early-stage cancers, and the fact that predictions based on mSEPT9 were not significant. Based on the findings of the present study, the mSEPT9 assay appears to have far broader implications for all types of digestive cancers and has significant potential for future clinical practice.
Limitations of this study
One of the major limitations of this study was that validation cohorts are still needed to confirm our results and conclusions. Independent samples should be collected prospectively to validate the performance of the new assay for all types of cancers, precancerous diseases, benign diseases, and in normal subjects. Although this study featured a training cohort, we believe that the data reported herein strongly supports the use of SEPT9 for pan-cancer screening. Future validation should focus on the new findings of this study, including the performance of blood SEPT9 for GC and EC, the combined performance of protein markers with SEPT9, and the detection efficacy of SEPT9 for precancerous conditions. Future research should also consider the economic implications of the assay.
Supplemental Material
SuppFig1 – Supplemental material for Opportunistic screening and survival prediction of digestive cancers by the combination of blood mSEPT9 with protein markers
Supplemental material, SuppFig1 for Opportunistic screening and survival prediction of digestive cancers by the combination of blood mSEPT9 with protein markers by Lele Song, Yan Chen, Yuan Gong, Jun Wan, Shaohua Guo, Hongyi Liu, Yuemin Li, Zhen Zeng and Yinying Lu in Therapeutic Advances in Medical Oncology
Supplemental Material
SuppFig2 – Supplemental material for Opportunistic screening and survival prediction of digestive cancers by the combination of blood mSEPT9 with protein markers
Supplemental material, SuppFig2 for Opportunistic screening and survival prediction of digestive cancers by the combination of blood mSEPT9 with protein markers by Lele Song, Yan Chen, Yuan Gong, Jun Wan, Shaohua Guo, Hongyi Liu, Yuemin Li, Zhen Zeng and Yinying Lu in Therapeutic Advances in Medical Oncology
Footnotes
Author contributions
LS, YL and ZZ designed the study. YC, YG, SG, HL and YLi recruited the subjects, collected the samples, and liaised with the laboratory. YCh and YG analyzed the data. LS made the figures and tables and wrote the manuscript. YL and ZZ proofread the manuscript. LS submitted the manuscript.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Conflict of interest statement
LS was a previous employee of BioChain, a collaborator of Epigenomics, Inc., who developed the first mSEPT9 assay for CRC screening and detection and obtained approval from the US FDA, the European Union, and the Chinese FDA. Others authors declare no conflict of interests.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the National Science and Technology Major Project ‘Prevention and Control of Major Infectious Diseases including AIDS and Viral Hepatitis’ Special Project ‘Development and Transformation of Hepatitis B-Related Hepatic Cancer Markers for Accurate Diagnosis and Treatment’ (2018ZX10302205) subproject ‘Translational Study of Tissue Microenvironment on Metastasis, Recurrence and Intervention of Liver Cancer’ (2018ZX10302205-001). This study was also supported by the National Key R&D Program of China (2017YFC0908401) and the National Science and Technology Major Project (2018ZX10723204 and 2018ZX10302205). This study was also supported by the Special health research projects of 2019 funded by the Chinese PLA General Hospital, under the project ‘Screening of early-stage CRC and adenoma by combined methylated SEPT9 and FIT assays in military aging population’ (NLBJ-2019003).
Supplemental material
Supplemental material for this article is available online.
