Abstract
In an era of powerful computing tools, radiogenomics provides a personalized, precise approach to the detection and diagnosis in patients with prostate cancer (PCa). Radiomics data are obtained through artificial intelligence (AI) and neural networks that analyze imaging, usually MRI, to assess statistical, geometrical, and textural features of images to provide quantitative data of shape, heterogeneity, and intensity of tumors. Genomics involves assessing the genomic markers that are present from tumor biopsies. In this article, we separately investigate the current landscape of radiomics and genomics within the realm of PCa and discuss the integration and validity of both into radiogenomics using the data from three papers on the topic. We also conducted a clinical trials search using the NIH’s database, where we found two relevant actively recruiting studies. Although there is more research needed to be done on radiogenomics to fully adopt it as a viable diagnosis tool, its potential by providing personalized data regarding each tumor cannot be overlooked as it may be the future of PCa risk-stratification techniques.
Introduction
In 2022, the American Cancer Society estimates prostate cancer (PCa) to lead new cancer incidences in men, accounting for nearly 27% of new cancer cases, which is more than double the estimated incidence of the second leading cause of new cases. 1 Unfortunately, PCa is also estimated to be the second leading cause of cancer deaths in male patients. Therefore, it is vital to adopt and implement new techniques that can improve the outcomes of those diagnosed with PCa.
Radiogenomics provides a personalized, precise approach to the detection and diagnosis in patients with PCa. Imaging plays an important role in PCa diagnosis. The use of magnetic resonance imaging (MRI) and target biopsy has been demonstrated to detect more clinically significant PCa (CSPCa). 2 Radiomics leverages a variety of image metrics using artificial intelligence (AI) to provide quantitative measurements of tumors, which may provide unique information in several pathologies. In addition, the robust research on gene expression levels has augmented the rapid development of genomics. Certain gene mutations and genomic markers have been found to be related to aggressiveness and prognosis of the tumor. 3 In recent years, the association between imaging and genomics information has become a new field, known as radiogenomics. It has the advantages of both radiomics and genomics, and can provide gene information on imaging, which serves as a promising tool in the diagnosis, classification, and prognosis of various malignancies. The objective of this review is to summarize the current status of the use of radiogenomics techniques in the evaluation for PCa and look into the future development.
Methods
We conducted a literature search in PubMed using the terms ‘radiomics and prostate cancer’, ‘genomics and prostate cancer’, and ‘radiogenomics and prostate cancer’. References from these published papers were reviewed as well for any potential studies. Based on the screening, we summarized the most relevant studies evaluating the use in PCa risk stratification in this narrative review. Considering the scarcity of studies in this field, we also looked at potential clinical trials data involving radiogenomics in PCa studies on clinicaltrials.gov.
Radiomics
The current landscape of PCa imaging and diagnosis involves the standard transrectal ultrasound-guided biopsy (TRUS Bx), via either transperineal or transrectal approach, to acquire 12–16 cores within pre-determined locations in the prostate of men and the recently adapted multiparametric magnetic resonance imaging (mpMRI) with MRI-ultrasound (US) fusion biopsies. Many of the radiomics in PCa research studies utilize the data obtained from mpMRI into newly developed machine learning models to predict outcomes.
In 2020, Bourbonne et al. developed an MRI-based radiomic model to predict the biochemical recurrence (BCR) and BCR-free survival of patients and compared it with another clinical prediction model which only used the patient’s age at surgery and postoperative prostate-specific antigen (PSA). In the model, they had a size-zone emphasis from apparent diffusion coefficient (ADC) maps extracted from 107 pre-therapeutic diffusion-weighted imaging (DWI) MRIs. 4 In the two cohorts tested, the model demonstrated a 78% BAcc (balanced accuracy) of BCR patients in the external cohort over the 56% BAcc of the initial testing cohort, highlighting the potential benefit of radiomic modeling. Similarly, Woźnicki et al. 5 created an ensemble model combining a radiomics model, with models for Prostate Imaging Reporting and Data System (PI-RADS), prostate-specific antigen density (PSAD), and digital rectal examination (DRE) where they achieved highly predictive results for the differentiation of malignant from benign prostatic lesions and of clinically significant PCa from clinically insignificant PCa.
Perhaps, the most important study to date regarding the potential of radiomics in PCa risk stratification is the Miami MAST trial, which aims to gather imaging features from mpMRI-guided MRI-US fusion biopsies to identify higher grade tumors earlier on in PCa workup so that it may lead to better risk-stratification patient outcomes for PCa (ClinicalTrials.gov: NCT02242773). In addition, it gives physicians better insight into underlying molecular pathways associated with imaging.
Radiomics in PCa detection poses a high upside; however, the aspect of image acquisition still remains a complex process as determining the regions of interest and choosing the right statistics to be analyzed and applied still remain difficult.
Genomics
There are three different commercially approved genomic panels that can be performed on biopsied tissue to risk-stratify patients between active surveillance (AS) and treating patients: Genomic Health’s Oncotype Dx test®, Myriad’s Prolaris test®, and Genome Dx’s Decipher test®. These are summarized in Figure 1.
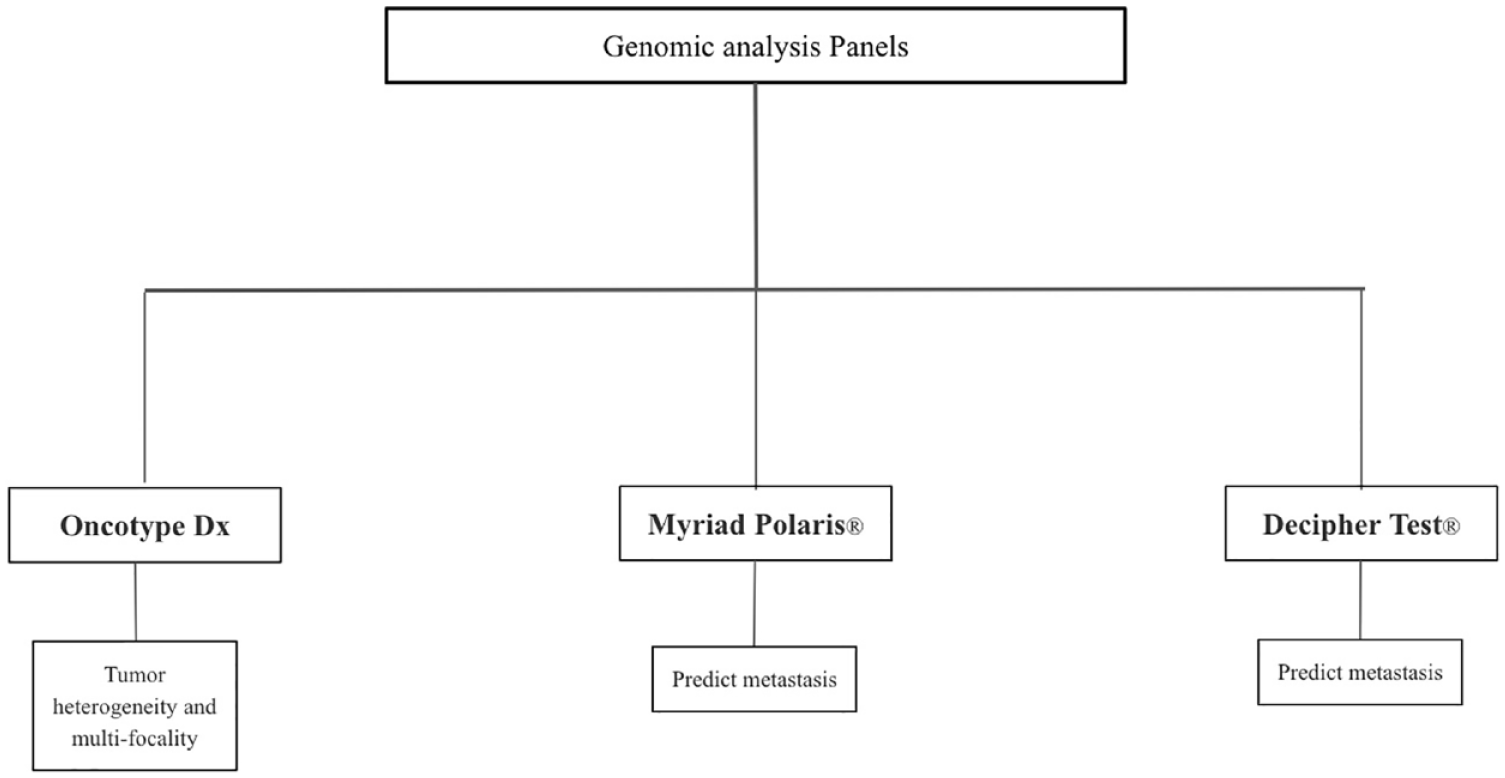
A summary of gene expression risk-stratification products for PCa.
Genomic Health’s Onctoype Dx is a 17-gene signature test that was mainly created to combat the issues of prostate tumor heterogeneity, which poses risks of underdiagnosis of clinically significant PCa. 6 Myriad’s Prolaris® uses a 31-gene cell cycle progression (CCP) signature created to predict metastasis in a radical prostatectomy (RP). 7 Cooperberg et al. 8 conducted a study using 413 patients who underwent RP and had a minimum follow-up of 5 years, later concluding that the CCP signature highly correlated with the likelihood of recurrence among low- and intermediate/high-risk patients. The Decipher Test®, developed by Genome Dx, is a 22-gene panel signature used to predict metastasis using RP tissue. 9 Its clinical use mainly focuses on the need for adjuvant radiotherapy or androgen deprivation therapy (ADT). 10 Although there is much promise with the use of genomics in PCa detection, the issue of tumor heterogeneity still exists as these genomic analyses are often conducted on the highest grade tumor sections. Similarly, there seems to be some hesitation about the usefulness of these biomarkers, as in 2018 a team investigated 176 tissue samples covering a wide range of PCa pathologies using three different tests and concluded that the gene signatures did not help determine the presence of sampled or unsampled high-grade cancer. 11
Radiogenomics
By combining the strengths of radiomics and genomics, radiogenomics offers a highly personalized approach to risk-stratify patients with PCa. The high utility of radiogenomics in PCa risk stratification stems from its inherent ability to determine the heterogeneity of PCa, which, many times, compromises diagnoses. More research is needed on specific biomarkers; however, the current three genomic panels serve as a good launch point for future studies. The growing role of mpMRI in PCa characterization has provided abundant metrics, such as statistical, geometrical, and textural data, to better inform machine learning models to provide tumor measurements regarding intensity, shape, and heterogeneity.
Wibmer et al. published a study in 2019 exploring the association between MRI features in PCa and Myriad’s Prolaris® test. Two board-certified radiologists were blinded to patient cell cycle risk (CCR) scores and tasked with analyzing prostate MRI and evaluating PI-RADS scores, the tumor’s extracapsular extension, and tumor length and volume on T2-weighted (T2w) images and ADC maps. They found a high association between patient CCR scores and extracapsular extension. Overall, 37 patients underwent standard TRUS Bx and the specimens were applied with the Prolaris® test. The results of the Prolaris® test were then used to calculate patient’s CCP scores, which were then combined with the patient’s CAPRA score (preoperative predictor of disease recurrence after radical prostatectomy) in a mathematical algorithm to determine CCR scores. Higher CCR scores were associated with higher mortality. The genetic and imaging results were further statistically evaluated using Fisher’s exact test, Wilcoxon rank-sum test, and Kruskal–Wallis test, indicating the strong association between CCR scores and extracapsular extension; however, there was no association between MRI features and genetic testing which classified the tumor’s pathology as more aggressive or less aggressive.12,13
Mccann et al.’s study, in 2015, looked at phosphate and tensin homolog (PTEN), a tumor suppressor gene on chromosome 10, and its correlation with mpMRI features. The mpMRI data were obtained using an endorectal coil to picture the whole prostate. Imaging features were then assessed from the initial MR images using T2w imaging signal intensity imaging features, DWI-based imaging features, and dynamic contrast-enhanced (DCE)-MRI-based image features. Pathologists looked at the peripheral zone cancer foci and outlined the regions of interest on MR images. PTEN of the biopsied tissues was graded by pathologists based on the presence of positive-staining tumor cells. A statistical analysis using the data from the imaging features and PTEN presence was then run using Spearman’s rank correlation coefficient and it was determined that there was a weak, but significant, association between peripheral zone PCa DCE-MRI features and Gleason score with PTEN expression. 14
Stoyanova et al. used current radiomic modules to implement their own radiogenomics models. The idea of this study is to create ‘radiophenotypes’ that are associated with and complement genomic risk-stratification biomarkers in PCa obtained from the Decipher gene panel test. First, they combined DCE-MRI and ADC to create maps indicating areas of high-, medium-, and low-risk cancer in the prostate. They created a software, named Insight Segmentation and Registration Toolkit (ITK), used for image analysis. Using MIM, a java plug-in, they were able to integrate the ITK data into the RedCap database. RNA extraction and microarray hybridization were conducted to analyze the genetics of the 17 biopsies obtained from six patients. The genetic analysis was then examined for its correlation with the Gleason score of the biopsies. The results showed a strong correlation between Gleason and Decipher scores (as demonstrated on Pearson’s correlation distance diagram in article). 15
Clinical trials search results
In our review, we also looked at potential clinical trials data involving radiogenomics in PCa studies. We used the National Institutes of Health’s (NIH) clinical trials database (clinicaltrials.gov) and inputted ‘prostate cancer’ in the ‘condition or disease’ search criteria and used the keywords ‘imaging genomics’ and ‘radiogenomics’ for the ‘other terms’ search. These are summarized in Figure 2. We were also able to obtain information about the Miami MAST trial which has been referenced in a few other papers.

A summary of clinical trials related to search results.
The PCa/imaging genomics search came back with 23 results, of which 2 studies were completed, 11 were recruiting, and 10 were terminated/not recruiting/unknown. To note, 11 out of the 13 studies we looked at did not meet our studies’ inclusion standard of dealing with imaging and genomic studies. The first study, ‘PSMA-PET Imaging for Detecting Early Metastatic Prostate Cancer in Men w/ High Decipher Test Scores’, is actively recruiting patients who have a higher Decipher genomic test score to consent to F-DCFPyL-prostate-specific membrane antigen (PSMA) positron emission tomography (PET) imaging. The goal of this study is to estimate the correlation between high Decipher scores and early metastatic disease using PSMA-PET imaging. The estimated study completion date is around June 2023. One of the focuses of the second study, ‘Collection and Measurement of Biomarkers in Prostate Cancer Radiotherapy Patients (COMBINE)’, is to evaluate how MRI and PET imaging changes relate to gene expression in tumor tissue before treatment from prostate biopsy or a prior prostatectomy. The estimated completion time for this study is in 2029.
In our clinical trials search using keywords ‘prostate cancer’ and ‘radiogenomics’, we obtained two results, of which the status of one study was ‘unknown’ and the other was ‘completed’. However, the completed study lacked the data of radiomics and imaging we sought after.
Discussion
The relationship between radiographic data and gene expression has provided great insight into the future of PCa risk stratification and diagnosis. As more data are fed into the different machine learning programs, they will be able to provide more accurate and predictive results about tumor characteristics. Radtke et al. looked at analyzing PI-RADS scores with genomics. In their study, they used standard techniques of image collection and biopsies to gather genomic information in prostate regions of interest. Using Pearson’s correlation, they were able to determine a strong association (r = 0.805, p < 0.001) between version 2 of PI-RADS and gene expression from the Decipher Genomic Resource Information Database (GRID); however, there were two outliers across all the models they used, so there is clearly room for improvement. The importance of precise prostate sampling was also required to obtain accurate genomic data as their study showed no genomic concordance between adjacent benign samples. 16 In regard to genomic data, there have been studies indicating that aggressive PCa has genomic alterations leading to prognostic significance; however, to create a more robust radiogenomics model, these specific genomic markers need to be further evaluated. 14
Nevertheless, there have been many studies examining the genetic profile of PCa. For example, Sun et al. looked at the hypoxia-related gene profiles in PCa, including a 15-gene universal hypoxia set, a 32-gene prostate-specific hypoxia gene set, and a 44-gene high-frequency hypoxia gene set. The purpose of this study was to understand the spatial distribution of hypoxia within the prostate because it creates great challenges in radiation therapy due to increased radioresistance within hypoxic tumor cells. Despite finding a weak association between mpMRI features and hypoxia-related gene expressions, the proof of the correlation is enough to continue research into finding specific genetic markers within PCa to direct future treatment options/risk stratification standards. 17 Similarly, Houlahan et al. discovered multiple small nucleolar RNAs (snoRNAs) and a noncoding RNA (SCHLAP1) in their study to determine the genetic markers involved leading to PCa visibility on mpMRI. The altered snoRNAs were pulled from the C/D box family, H/ACA box family, and a Cajal body-specific RNA family and synergized with a nimbosus hallmark (indicating increased metastatic potential). 18 SCHLAP1 was found in a previous study to be linked to PCa progression. 19 The study concluded that in tumor tissue visible on mpMRI, there was an overexpression of snoRNAs and SCHLAP1. More specifically, nimbosus hallmarks compounded with snoRNA levels to predict visible tumors with 87% accuracy compared with the 60% accuracy of the normal clinical characteristics that utilize just tumor and prostate volume. 20
The ability to match genome sequencing with imaging data poses as the main limitation due to the differences in imaging protocols and patient tumor characteristics. In addition, the retrospective nature of radiogenomic studies leads to less standardization of protocols. 12
Although radiogenomic analysis requires more data and research to increase its validity, its promise in improving patient outcomes cannot be dismissed.
In summary, radiogenomics holds a promising future in the growing era of personalized medicine. With more research being done into fine-tuning deep neural networks and explainable AI, only raw radiographic data and genomic data are needed to help these programs grow. However, for these AI-directed programs to hold any validity, it is necessary for standardization to occur in image collection/recognition and also in genomic analysis.
