Abstract
Although prostate cancer (PCa) is the most commonly diagnosed cancer in men, most patients do not die from the disease. Prostate specific antigen (PSA), the most widely used oncologic biomarker, has revolutionized screening and early detection, resulting in reduced proportion of patients presenting with advanced disease. However, given the inherent limitations of PSA, additional diagnostic and prognostic tools are needed to facilitate early detection and accurate risk stratification of disease. Serum, urine, and tissue-based biomarkers are increasingly being incorporated into the clinical care paradigm, but there is still a limited understanding of how to use them most effectively. In the current article, we review test characteristics and clinical performance data for both serum [4 K score, prostate health index (phi)] and urine [SelectMDx, ExoDx Prostate Intelliscore, MyProstateScore (MPS), and PCa antigen 3 (PCA3)] biomarkers to aid decisions regarding initial or repeat biopsies as well as tissue-based biomarkers (Confirm MDx, Decipher, Oncotype Dx, and Polaris) aimed at risk stratifying patients and identifying those patients most likely to benefit from treatment versus surveillance or monotherapy versus multi-modal therapy.
Introduction
Prostate cancer (PCa) is the most common solid organ cancer in men in the United States, with 191,930 new cases and 33,330 deaths in 2020. 1 Although the lifetime risk of developing PCa is 1 in 9, the risk of death from the disease is only 2%. 1 Screening and early detection of PCa is performed through a digital rectal exam (DRE), prostate specific antigen (PSA) and its derivatives. Those with abnormal DRE and elevated PSA would then proceed to a prostate biopsy. Prostate biopsy, however, is associated with the risk of infectious hospitalization, bleeding, or obstructive complications in ~1–3% of cases.2–5 To decrease unnecessary biopsies, multiparametric MRI (mpMRI) can be used to identify those at risk of clinically significant disease for biopsy. 6 But mpMRI has its shortcomings, including a wide range of diagnostic variability among centers, 7 a positive predictive value (PPV) of 35% for clinically significant PCa (csPCa) with PI-RADS ⩾ 3 lesions, 7 and up to 35% of csPCa in men with negative mpMRI.7,8 Molecular biomarkers may supplement PSA and mpMRI for early detection and risk stratification of csPCa. Here, we review test characteristics and clinical performance data for both serum [4 K score, prostate health index (phi)] and urine [SelectMDx, ExoDx Prostate Intelliscore, MyProstateScore (MPS), and PCa antigen 3 (PCA3)] biomarkers to aid decisions regarding initial or repeat biopsies as well as tissue-based biomarkers (Confirm MDx, Decipher, Oncotype Dx, and Polaris) for risk stratifying patients and identifying those patients most likely to benefit from treatment versus surveillance or monotherapy versus multi-modal therapy.
Biomarker characteristics and performance evaluation
A biomarker is ‘a defined characteristic that is measured as an indicator of normal biological processes, pathologic processes, or responses to an exposure or intervention’. 9 The performance of a biomarker can be evaluated using several metrics summarized in Table 1. Unlike sensitivity and specificity, positive predictive value (PPV) and negative predictive value (NPV) depend on the prevalence of the disease in the population of interest. In general, area under curve (AUC) is preferred for comparing biomarker performance since sensitivity and specificity are unique to a decision threshold. The AUC is more reflective of accuracy independent of the cut point. AUC values range from 0 (perfectly inaccurate) to 1 (perfectly accurate).10–12
Definitions and examples of test performance metrics mentioned in this review.
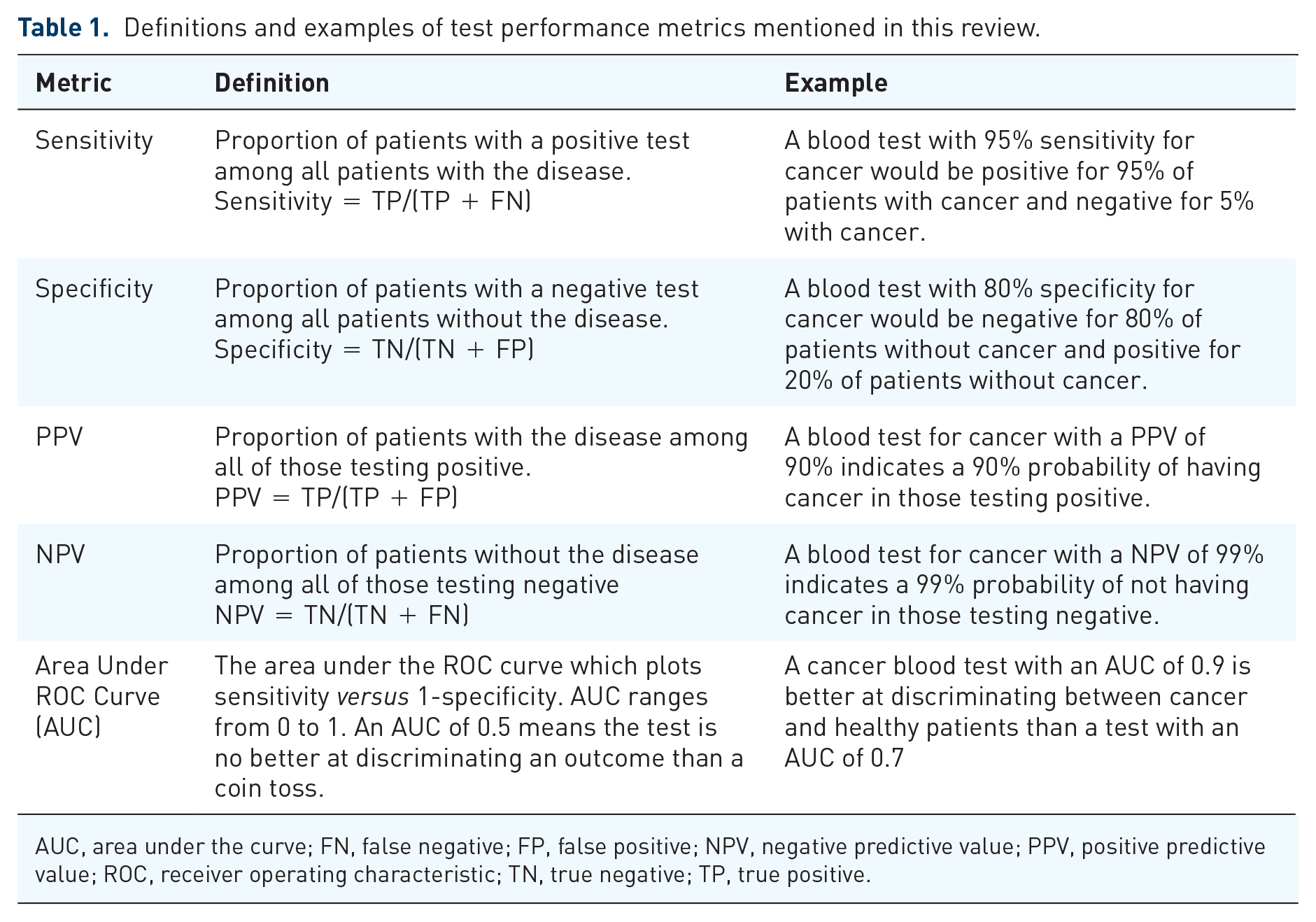
AUC, area under the curve; FN, false negative; FP, false positive; NPV, negative predictive value; PPV, positive predictive value; ROC, receiver operating characteristic; TN, true negative; TP, true positive.
Prostate Specific Antigen (PSA)
A member of the kallikrein family of serine proteases, PSA (human kallikrein 3) is the most widely used oncologic biomarker. 13 It is produced by prostatic luminal epithelial cells and plays a role in the regulation of semen coagulation. 14 Elevations of PSA are thought to occur due to disruptions of cellular architecture, and it circulates in both free and complex forms. PSA is FDA approved for PCa screening. Implementation of PSA testing at scale has been estimated to account for 45–70% of the decrease in PCa mortality observed in the 1990s. 15 However, while PSA is organ-specific, it is not cancer-specific, and PSA can be elevated in benign conditions like benign prostatic hyperplasia (BPH) and prostatitis. Thus, PSA screening can lead to unnecessary biopsies of benign prostates, over diagnosis and over treatment of indolent diseases, as well as missed diagnoses in some men with apparently normal PSA. 16 In the PCa Prevention Trial, the AUC for PSA to discriminate any PCa versus no cancer and Gleason ⩾ 7 versus Gleason < 7 PCa was 0.68 and 0.78 respectively. For detection of any PCa, PSA cutoff values of 1.1, 2.1, 3.1, and 4.1 yielded sensitivities of 83.4%, 52.6%, 32.2%, and 20.5%, respectively, and specificities of 38.9%, 72.5%, 86.7%, and 93.8%, respectively. 17 While the introduction of PSA certainly revolutionized the field of PCa detection and treatment, newer biomarkers may improve decision making along the complex and nuanced PCa clinical care pathway.
Overview of existing biomarkers in prostate cancer
The clinical care pathway for those with suspected PCa, based on elevation in PSA, abnormal DRE, or both, involves consideration of prostate biopsy. Per clinical guidelines, the decision to proceed with biopsy (initial or repeat) can be supported by multiparametric MRI (mpMRI) or molecular biomarkers (Table 2). Once prostate biopsy reveals cancer, biomarkers can also facilitate appropriate risk stratification for treatment versus surveillance, intensity of treatment, and the necessity for multi-modal therapy (Table 3). This review is focused on available molecular biomarkers in PCa, starting with serum and urine markers to inform prostate biopsy and concluding with tissue-based biomarkers for PCa risk stratification. A summary of the care pathway and available biomarkers in each clinical scenario is presented in Figure 1. For each biomarker, we will discuss the source (serum, urine, or tissue), what is measured, the performance of the test (including in racially diverse populations, specifically focusing on Blacks and/or African Americans), how it is interpreted, and, where available, approval status, insurance coverage, and inclusion in clinical guidelines from various societies. To facilitate direct comparison of biomarker performance, we will utilize AUC. In addition, we will include information on the number of biopsies avoided and the percentage of csPCa missed, if available.
Summary of early detection biomarkers usage, performance, and guideline inclusion.
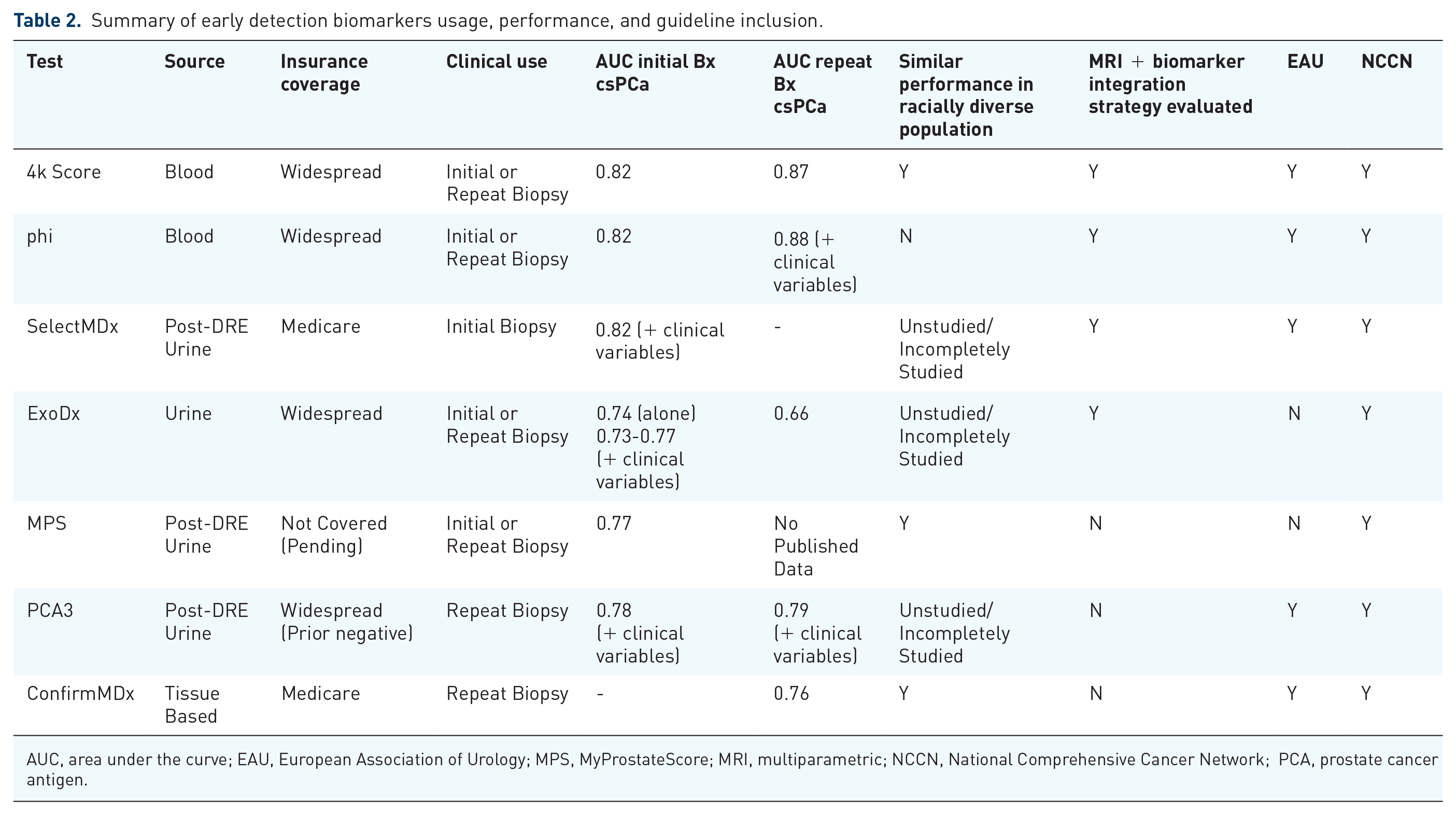
AUC, area under the curve; EAU, European Association of Urology; MPS, MyProstateScore; MRI, multiparametric; NCCN, National Comprehensive Cancer Network; PCA, prostate cancer antigen.
Summary of risk stratification biomarkers clinical applications, performance, and impact on patient management.

AUC, area under the curve; PCa, prostate cancer; RP, radical prostatectomy.

Graphical summary of the decision points prior to and following prostate cancer diagnosis and the biomarkers that are relevant to each phase.
Article selection process
Studies were selected for inclusion in the review based on the following criteria: (1) Patients have a suspicion for or a diagnosis of non-metastatic prostate cancer AND being evaluated for initial or repeat biopsy, placement on active surveillance, treatment with radical prostatectomy (RP) or radiation, OR post-RP treatment by one or more of the relevant commercially available biomarkers, (2) Original research, (3) Publication in the English language, (4) Reporting on outcomes that enable quantification and cross comparison of diagnostic or prognostic performance, namely sensitivity, specificity, area under the ROC curve (AUC). Articles were identified using the PubMed database. Search terms included a combination of the disease state (e.g. ‘prostate cancer’), stage of the clinical care pathway (e.g. ‘initial biopsy’), desired study characteristics (e.g. ‘African American Patients’, ‘mpMRI’), and the biomarker name. Sample test reports were sourced from the manufacturer’s websites.
Biomarkers for consideration of initial biopsy
4kscore
The 4 K score is a CLIA-certified test, included in the NCCN guidelines, that combines serum levels of four kallikrein proteins, including PSA, fPSA, intact PSA, and human kallikrein 2 (hK2), with DRE findings, age, and history of prior biopsy. The algorithm was developed based on data from the European Randomized Study of Screening for PCa (ESRPC) studies and the Prostate Testing for Cancer and Treatment (ProtecT) study. 18 The 4Kscore provides a probability (0–100%) that a patient will have csPCa on biopsy, defined as Gleason ⩾ 7 (Figure S1). The direct bill cost of the test is $500. 19
In a large, multi-center study of 1012 men, the 4 K score outperformed a modified Prostate Cancer Prevention Trial Risk Calculator (PCPTRC) 2.0 score in the prediction of csPCa (AUC 0.82 versus 0.74, p < 0.0001). At a cutoff of ⩾ 6%, 30% of biopsies can be avoided while delaying diagnosis in 1.3% of those with csPCa. 18 In a multi-institutional validation study done at the VA Health System in which most of the subjects were African American (56%), the AUC of the 4Kscore test was 0.81 compared to 0.74 in a base model consisting of age, DRE, and total PSA (p < 0.01). 20 Using the official score cutoff of ⩾ 7.5%, there would be a 22% biopsy avoidance rate with a 1% rate of missed csPCa. 20
When studied in conjunction with MRI, the highest NPV for PI-RADS 1-2 lesions on mpMRI were for patients with a low (96.9%) or intermediate (97.1%) 4 K score. In assessing the sequence of MRI and biomarkers, the initial 4 K score followed by mpMRI if 4 K score was ⩾7.5% followed by subsequent biopsy if mpMRI was positive, or 4 K score was ⩾ 18% would miss 2.7% of csPCa and avoid 34.2% of biopsies. This was postulated to be the most viable of seven strategies evaluated by decision curve analysis. 21 In a cohort of 925 men with a previous negative biopsy, the 4 K score had a higher discriminative accuracy for PCa than PSA and DRE, with an AUC of 0.68 compared to 0.58 (p < 0.001). For high grade PCa (Gleason ⩾ 7), AUC improved from 0.76 to 0.87 (p = 0.003). At a 15% risk threshold, the number of biopsies would be reduced by 71% while missing < 6% of high-grade cancers (3/53). 22
Prostate Health Index (phi)
The Prostate Health Index (phi) is an FDA-approved test that combines three PSA isoforms in the blood, including [-2] proPSA (p2PSA), free PSA (fPSA), and total PSA (tPSA), to create a composite score that predicts the likelihood of PCa on biopsy (Figure S2). 23 The test is included in the NCCN guidelines and is the least expensive of commercially available PCa biomarkers in the USA, at a cost to the patient with insurance coverage of $80. 19 Notably, samples should be centrifuged within 1–3 h of collection as [-2]proPSA increases significantly after blood draw at room temperature.24,25
In a multicenter study evaluating the ability of phi to detect aggressive PCa in biopsy naïve men, de la Calle et al. 26 reported an AUC of 0.815 and 0.783 in the discovery and validation cohorts, respectively. In another multicenter cohort of 658 men with a PSA between 4 and 10 ng/ml, Loeb et al. 27 reported that phi outperformed each of the three individual PSA isoforms for predicting Gleason ⩾ 7 PCa (AUC of phi 0.70, fPSA 0.65, [-2]proPSA 0.55, PSA 0.55). Using a phi cutoff of 28.6, 30.1% of patients could have been spared an unnecessary biopsy. 27 In a head-to-head comparison with PCA3 and PSA derivatives in 300 patients undergoing initial biopsy, phi had an AUC of 0.77 for the prediction of PCa on biopsy, which was not significantly different than PCA3 (0.73), despite a significant outperformance of PSA derivatives (fPSA (0.60), %fPSA (0.62), and p2PSA (0.63)). On decision curve analysis, there was a very close net benefit profile between phi and PCA3. 28 Further evidence for the utility of phi in detecting csPCa was provided by Nordström et al., 29 who found that phi outperforms base clinical models of PSA and age in discriminating between benign versus GG1 as well as ⩾ GG2 PCa. The performance of phi in biopsy naïve African American men to predict ⩾ GG2 PCa was not significantly different from PSA (AUC = 0.63 versus 0.57, respectively, p = 0.27). 30 Phi has also been shown to be an effective predictor of higher T stage (T3 versus T2, AUC = 0.85) and high-volume disease (tumor volume > 0.5 ml, AUC = 0.94). 31
An expanding body of literature has evaluated strategies for combining phi and mpMRI. In a study of 196 biopsy-naïve men undergoing mpMRI, phi outperformed PI-RADS (difference in AUC of 0.14, p < 0.001) in predicting positive biopsy. A subset of patients went on to undergo RP, and it was found that phi (cutoff of 61.68) and PI-RADS of 4 or greater had a comparable performance in identifying csPCa at RP. 32 In another single center study, Tosoian et al. 33 demonstrated that there were no GG ⩾ 2 cancers in those with a phi < 27 and a PI-RADS ⩽ 3, and phi can reduce the rate of prostate biopsy by 9% (39% versus 48%, p < 0.001). A small prospective study of 102 Asian men found that the AUC for prediction of Gleason ⩾ 7 PCa was greater with the combination of phi and mpMRI (0.87) than with phi alone (0.74, p = 0.002) or mpMRI alone (0.830, p = 0.04). Restricting biopsy to patients with either PI-RADS 5 or PI-RADS 3 or 4 with a phi ⩾ 30% would have avoided 50% of biopsies with only one csPCa being missed. 34 In patients undergoing repeat biopsy, the incorporation of phi into a clinical prediction model demonstrated an AUC = 0.85 for all PCa and 0.88 for Gleason ⩾ 7 PCa. 35
SelectMDX
SelectMDx combines clinical parameters with the mRNA expression levels of DLX1 and HOXC6 to estimate the likelihood of any PCa and Gleason ⩾ 7 PCa on biopsy (Figure S3). It is a non-FDA approved test performed on post-DRE urine in a CLIA certified lab and is included in the NCCN guidelines. The test has limited coverage by Medicare, and the direct bill cost is $500. 36 Utilization of SelectMDx resulted in a cost savings of $1694 per patient over an 18-year horizon. 36
In a prospective multicenter study, Van Neste et al. 37 found that a combined model using both HOXC6 and DLX1 as well as clinical risk factors (age, DRE, PSAD, previous biopsy, PSA, and family history) yielded an AUC of 0.9 (95% CI, 0.85–0.96) in the training cohort and 0.86 (95% CI, 0.80–0.92) in the validation cohort for detection of csPCa, with a potential to decrease total biopsies by 42%. Notably, PSA was higher in both the discovery (Mean: 15.8, Median: 7.4, IQR 5.5–11.1) and validation (Mean: 11.9, Median: 7.3, IQR: 5.2–10.9) cohorts used in this study than the contemporary biopsy referral population. In a follow up large multicenter study of 1,955 men in Europe using a model encompassing urinary HOXC6 and DLX1 mRNA levels, patient age, DRE, and PSA density, AUC was 0.82 (95% CI, 0.79–0.86) in the validation cohort for detecting GG ⩾ 2 PCa, compared to 0.70 (95% CI, 0.65–0.74, p < 0.001) for the PCPTrc. 38 Notably, the diagnostic and prognostic accuracy of the test in a racially diverse study population is unknown.
SelectMDx testing may contribute to the stratification of patients for mpMRI. A study of 310 patients who underwent SelectMDX testing and mpMRI prior to prostate biopsy revealed that SelectMDx was more sensitive (86.5% versus 51.9%) but less specific (73.8% versus 88.3%) than mpMRI for the detection of PCa on biopsy. The same trend was observed for sensitivity (87.1% versus 61.3%) and specificity (63.7% versus 83.9%) for the detection of csPCa. 39 SelectMDx scores (AUC = 0.83) has been shown to outperform PSA (AUC = 0.66) and PCA3 (AUC = 0.65) for the prediction of suspicious lesions (PIRADS 4 and 5) on mpMRI. It could be of additional value when a PI-RADS 3 lesion is found, but developing a formalized strategy would require further evaluation. 40
ExoDx prostate intelliscore
The ExoDx Prostate Intelliscore measures urinary exosomal mRNA of PCA3 and ERG normalized with SPDEF. It is reported on a scale of 0–100 (Figure S4). 41 The test is performed in a CLIA certified lab, is non-FDA approved, and is included in the NCCN guidelines. The test is covered by most insurance plans, including Medicare and private payers, with a direct bill cost of $795. The ExoDx test has a distinct advantage of not requiring a DRE prior to sample collection and does not incorporate any clinical variables as part of the score.
In a multicenter study, the ExoDx Prostate IntelliScore improved discrimination between Gleason ⩾ 7 PCa and benign disease, with an AUC of 0.77 (95% CI, 0.71–0.83) when combined with clinical variables versus clinical variables such as PSA level, age, race, and family history of PCa alone (AUC = 0.66, 95% CI, 0.58–0.72) (p < 0.001). The ExoDx Prostate IntelliScore alone had an AUC of 0.74. Using a cutoff of > 15.6 to prompt a biopsy avoids 20% of all biopsies while only missing 2% of Gleason ⩾ 7 PCa. In an independent validation cohort of 519 patients, the assay plus clinical variables (AUC = 0.73, 95% CI, 0.68–0.77) outperformed clinical variables alone (AUC = 0.63, 95% CI, 0.58–0.68, p < 0.001). With the official cut point of 15.6, 27% of biopsies could have been avoided while only 5% of patients with primary pattern 4 Gleason ⩾ 7 PCa would have been missed. 41 Using pooled data from three studies (n = 1212) of biopsy naïve patients with a median age and PSA of 63 and 5.2 ng/ml respectively, the Prostate Intelliscore AUC (0.70) was superior to PCPT-RC (0.62), ESRPC (0.59) and PSA alone (0.56) (p < 0.001) for discriminating ⩾ GG2 from GG1 and benign histology. In this combined cohort, 17% identified as African American, but no distinct studies have yet been done to specifically evaluate the performance of the test in a racially diverse population. 42
In the study by de la Calle et al., 43 a strategy consisting of up front ExoDx, mpMRI in those with a ExoDx score of ⩾ 15.6, and biopsy in those with ExoDx score ⩾ 19 or those with ExoDx score ⩾15.6 and a PI-RADS 3-5 lesion on mpMRI resulted in a biopsy avoidance rate of 43% while missing 4.8% of GG ⩾2 cancers. In 229 men with a prior negative biopsy, ExoDx had an AUC of 0.66 (95% CI: 0.55–0.78), which was superior to PSA alone (0.54, 95% CI: 0.43–0.66), and the ERSPC Risk Calculator (AUC = 0.47, 95% CI = 0.36–0.58) in predicting GG ⩾ 2 PCa. 44
MyProstateScore (MPS)
Previously referred to as the Mi-Prostate Score (MiPS), the MyProstateScore (MPS) is a non-FDA approved test, not yet specifically endorsed by the NCCN, that combines urinary PCA3 and TMPRSS2:ERG fusion gene expression with PSA to provide a score, ranging from 0 to 100. Scores increase with the likelihood of detecting Gleason ⩾ 7 PCa, and they are grouped into low, intermediate, or high-risk (Figure S5). The urinary gene expression assay requires a post-DRE urine specimen. The test is not yet covered by public or private payers, and the direct bill cost is $760. Using MPS as a tool to guide biopsy decisions was found to be more cost-effective than MRI or ordering biopsies for all men with indeterminate PSA levels. 45
In a multicenter trial consisting of a training cohort of 677 men and a validation cohort of 1225 men, MPS was able to discriminate GG ⩾ 7 PCa with better performance (AUC = 0.77) than PSA (AUC = 0.65, p < 0.001), PSA with TMPRSS2:ERG (AUC = 0.73, p < 0.001), and PSA with PCA3 (AUC = 0.75, p < 0.001). 46 MPS-based models had significantly greater AUC (p < 0.001) than models incorporating PCPTRC with PCA3 or TMPRSS2:ERG. 46 In a separate study that investigated the use of a MPS cutoff of 10 to proceed with biopsy in a combined validation cohort consisting of 1,525 patients from community and academic practices, MPS testing would have led to biopsy avoidance in 26% of men while missing GG ⩾ 2 PCa in 3%. Among African Americans (n = 150), sensitivity was similar (97.6% versus 97% overall), and 15% of men were able to avoid biopsy with 2.4% of GG ⩾ 2 PCa missed. 47
Tosoian et al. have reported that MPS score increases with increasing PI-RADS score (p < 0.001). Furthermore, in the PI-RADS 3 (43.2 versus 28.4, p = 0.003), 4 (40.6 versus 30.1, p < 0.001), and 5 (48.0 versus 28.2, p = 0.04) populations, MPS was significantly higher in those with GG ⩾ 2 PCa. In the equivocal PI-RADS 3 population, MPS outperformed PSA (AUC = 0.55), PSA Density (AUC = 0.62) for prediction of GG ⩾ 2 PCa. A MPS threshold of ⩽ 25 would prevent 39% of biopsies in the PI-RADS 3 population while missing 6% of GG ⩾ 2 PCa. 48 MPS has also been evaluated in men with a prior negative biopsy. Using a slightly higher threshold of MPS ⩽ 15 to rule out biopsy would avoid 21% of biopsies with a 100% NPV. 49
Biomarkers for consideration of repeat biopsy
There are an estimated 1 million prostate biopsy procedures in the United States annually, the majority of which reveal benign pathology. 1 In a serial biopsy study that included 2500 men, the authors found that nearly 25% of cancers are missed on initial biopsy, with detection rates of 17%, 14%, and 11% on the second, third, and fourth biopsies, respectively. 50 Implementation of molecular testing to reduce unnecessary repeat biopsies may result in cost savings. 51
PCA3
The prostate cancer antigen 3 (PCA3) gene is a non-coding mRNA with expression that is 60-100-fold greater in cancerous than in benign tissues. 52 The PCA3 test is FDA-approved and requires the collection of the first 20–30 ml of post-DRE urine. The quantitative result provides the ratio of PCA3 mRNA transcripts in the sample to serum PSA (Figure S6). The test costs roughly $230. 53
In an early study of 570 men undergoing biopsy, the performance of PCA3 for detection of PCa was found to be equivalent in biopsy naïve (AUC = 0.70) and prior negative biopsy (AUC = 0.68). A model using PCA3 score, serum PSA, prostate volume and DRE results increased the AUC in the combined cohort (biopsy naïve and repeat biopsy) from 0.69 (PCA3 alone) to 0.75 (p = 0.0002). 54 For the detection of high-grade PCa, AUC improvements for initial and repeat biopsies were 0.74 to 0.78 and 0.74 to 0.79, respectively. 55 In another prospective multi-center study published by Wei et al., 55 adding PCA3 to the PCPT risk calculator increased the AUC from 0.64 to 0.69 (p < 0.001) for detection of any cancer and from 0.74 to 0.79 (p < 0.003) for high grade cancers, in the repeat biopsy setting. In a separate, prospective multi-center study in Europe (n = 463), the AUC for PCA3 score in predicting a positive repeat biopsy was 0.66 compared to 0.578 for %fPSA (p = 0.08) and using a PCA3 cutoff of 20 was estimated to reduce the repeat biopsy rate by 44% while missing 9% of >/= GG2 disease. 56 PCA3 has also been shown to be an accurate predictor of high stage (T3 versus T2, AUC = 0.74) and high tumor volume (>0.5 ml, AUC = 0.86) disease. 31 In a multi-center cohort study, O’Malley et al. demonstrated that PCA3 did not provide any significant increase in predictive value for csPCa in AAM. However, the study was limited by the small number of AA patients (n = 72, 10% of the cohort). 57 PCA3 has been previously demonstrated in combination with mpMRI in the repeat biopsy setting to improve the NPV for detection of PCa on biopsy (95%) in comparison to PCA3 (40%) and mpMRI (83%) alone. 58
ConfirmMDX
ConfirmMDX is a Medicare-covered tissue-based assay included in the NCCN guidelines that tests for hypermethylation of three PCa associated genes – GSTP1, APC, and RASSF1 (Figure S7). 59 The assay capitalizes on the field effect principle – detectable changes can occur in non-malignant tissue that is contiguous in space with cancerous tissue, thus mitigating the sampling error issue that is inherent in traditional prostate biopsy. 60 The specificity of GSTP1 hypermethylation for PCa (present in 82% of cancer cases, 5% of benign controls), 61 coupled with the strong field effect of APC and RASSF1A, provides the basis for the ConfirmMDX test. The retail price for the assay is $206 per core or $2060 for a 10-core biopsy. 51
In the MATLOC study (n = 483 men), ConfirmMDx was able to identify 68% of cases (sensitivity) while confirming the benign diagnosis (specificity) in 64% of men, with a NPV of 90%. 59 The DOCUMENT multi-center study in the US, Partin et al. 62 found a similarly high NPV (88%), supporting the conclusion that the epigenetic assay can decrease unnecessary repeat prostate biopsies. In a combined study, the AUC for the identification of men with high grade PCa on repeat biopsy was 0.76, which was significantly higher than PSA (0.57, p = 0.004) and PCPTRC (0.62, p = 0.029). Analyses of the performance of the assay in an African American population consisting of 211 men from 7 centers across the US revealed a sensitivity of 78% and a specificity of 53% for the detection of Gleason ⩾ 7 PCa. The NPV in this study for the detection of Gleason ⩾ 7 PCa was 94.2% consistent with the performance of assays in other large multicenter studies. 63 Notably, in a small study of 113 patients undergoing ConfirmMDX testing and MRI, 71% of ConfirmMDX negative patients also had a negative MRI (PI-RADS1-2), and most of the other cases were PI-RADS3 transition zone lesions in large glands. 64
Guideline statements – early detection
The NCCN states that while biomarkers are not yet first-line screening tests in conjunction with serum PSA, the probability of high-grade cancer can be further defined using the PHI, SelectMDX, 4 K score, and ExoDX prostate tests. PHI, 4KScore, ConfirmMDX, and PCA3 are also listed as options to improve biopsy specificity in individuals who have previously undergone a negative prostate biopsy. It is not yet known how these tests can be applied in optimal combination with MRI. The AUA does not mention specific tests but provides a similar statement on the applicability of urine and serum biomarkers to inform biopsy decisions. Finally, the EAU concluded that further studies are required to validate the efficacy of biomarkers, and at the moment, the data is too limited to implement these markers into routine screening programs. The EAU guidelines consider MPS and ExoDx as investigational while PCA3, SelectMDx, 4 K score, PHI, and Confirm MDX are all listed as options with weak evidence to guide biopsy decision making in those with PSA between 2 and 10 ng/dl and a normal DRE.65–67
Biomarkers for risk stratification
Tissue-based molecular biomarkers may facilitate accurate risk stratification for appropriate treatment in the clinical continuum of PCa. For example, since more than half of patients on active surveillance eventually proceed to treatment, biomarkers could help stratify patients with favorable risk disease for appropriate management.68–70 Another distinct clinical challenge is determining the optimal treatment approach for men with NCCN high-risk disease, especially with the high rate of recurrence (50–95%) in this group of patients. 71 Furthermore, in the event of adverse pathology at RP or BCR, only a fraction of men receives adjuvant and salvage radiation, respectively. 72 Molecular biomarkers may play an integral role in identifying patients for additional treatment following initial therapy.
Decipher
The Decipher tissue-based gene expression classifier is an insurance reimbursed test included in NCCN and ASCO guideline statements. The Decipher score is based on the microarray-based expression of 22 genes involved in androgen receptor signaling, cell proliferation, differentiation, motility, and immune modulation. 73 Notably, even though a small number of genes are used to calculate the score, the microarray measures the expression data of 46,000 genes. The reported score is on a scale of 0–1 and is independent of clinical or demographic data. Those with scores ⩽ 0.6 are classified as low risk, while those with scores above this threshold are classified as high risk. The test can be performed on either a positive biopsy specimen or a post-RP specimen. The report provides several prognostic variables, including percentage risk of distant metastasis in 5- and 10-years, 15-year disease specific mortality, and the risk of adverse pathology at RP (Figure S8). With over 130 studies across the spectrum of disease after positive biopsy, Decipher is the most broadly validated of all the tissue-based biomarkers.
Prognostic value of Decipher
For patients with very-low, low, and favorable intermediate risk disease, Decipher alone (AUC = 0.65, 95% CI, 0.56–0.74) and Decipher in conjunction with Cancer of the Prostate Risk Assessment (CAPRA) groups (AUC = 0.65, 95% CI, 0.58–0.70) improved prediction of adverse pathology at RP.74,75 In a prospective statewide registry study by Vince et al., a high risk Decipher score was associated with shorter time to treatment in men on active surveillance (AS) [hazard ratio (HR) = 2.51, 95% CI = 1.52–4.13, p < 0.001], 76 suggesting that this test may play a role in identifying patients for AS.
In patients undergoing primary radiation therapy, Decipher testing on biopsy specimens has a higher AUC (0.76–0.86) for predicting 5-year metastasis free survival (MFS) compared to NCCN risk groups (0.54–0.63). 77 Similar findings were observed in a cohort of 100 intermediate and high-risk patients in a single center receiving definitive radiation and androgen deprivation therapy (ADT). 78
The decipher has been extensively studied in post-RP samples for its ability to predict future recurrence and metastasis in patients undergoing RP. 79 In a multi-center cohort of 561 patients with high-risk PCa (PSA > 20, Gleason score 8–10, stage pT3b or pN1), the AUC for biochemical recurrence (BCR) was 0.72 (0.03 greater than CAPRA-S) and the AUC for metastasis was 0.64 (0.03 greater than the CAPRA-S). 80 In a multicenter study by Spratt et al., 81 a clinical-genomic model consisting of 6 tiers combining NCCN risk groupings and Decipher scores demonstrated improved performance (AUC = 0.84, 95% CI = 0.61–0.93) in comparison to NCCN (AUC = 0.73, 95% CI = 0.6–0.86) and CAPRA (AUC = 0.74, 95% CI, 0.65–0.84) risk groupings in prediction of 10-year distant metastasis.
In patients with BCR after surgery, Decipher has also been shown to predict later development of metastasis (AUC = 0.82, 95% CI, 0.76–0.86), more so than Gleason score (AUC = 0.64, 95% CI, 0.58–0.70) and PSA doubling time (AUC = 0.69, 95% CI, 0.61–0.77). 82 Spratt et al. demonstrated similar discrimination ability in a multi-center study of 477 patients, 150 of whom had a detectable PSA postoperatively. In this subset, the 5-year metastasis rate was 0.90% for Decipher low or intermediate risk, and 18% for Decipher high risk (p < 0.001). 83 In a study within the VA system of 548 patients deemed to be high risk after surgery, 301 of whom were African American (AAM), Decipher had a higher AUC than CAPRA-S for 5-year metastasis in AAM versus non-AAM (0.86 versus 0.69) as well as 10-year PCa specific mortality in AAM versus non-AAM (0.91 versus 0.78). 84
In multiple studies by Den et al. analyzing the optimal timing of post-operative radiation therapy, the authors suggest that those with low Decipher scores are best treated with salvage RT while those with higher scores may benefit from adjuvant therapy. The AUC for prediction of metastasis 5 years after radiation therapy was 0.66 for the CAPRA-S model (95% CI, 0.56–0.78) and 0.83 for the Decipher score (0.83, 95% CI, 0.27–0.89). 85 A retrospective multi-center and multi-ethnic cohort study by Freedland et al. 86 also demonstrated that Decipher provides a high predictive accuracy (AUC = 0.85, 95% CI = 0.73–0.88) for metastasis after salvage RT. In an analysis of Decipher data from patients in the NRG/RTOG 9601 trial investigating salvage RT +/- 2 years of bicalutamide, Decipher scores were independently associated with distant metastasis (HR = 1.19, 95% CI = 1.06–1.35, p = 0.003), PCa specific mortality (HR = 1.37, 95% CI = 1.18–1.61, p < 0.001), and OS (HR = 1.16, 95% CI = 1.06–1.28, p = 0.002) on a multivariable analysis adjusting for clinical parameters. The absolute impact of ADT was less in patients with a lower versus higher Decipher score. 87
Impact of Decipher on treatment decisions
In assessing the role of the Decipher GC in influencing treatment decision making, Michalopoulos et al. 88 demonstrated in the PRO-ACT study that review of Decipher increased the proportion of patients recommended treatment who were deemed Decipher high risk and an increase in the number of patients recommended observation who were deemed Decipher low risk (p < 0.001). Similar conclusions have been made by Badani et al., 89 who showed a 20% increase in observation recommendations for Decipher low risk patients and a 16% increase in treatment recommendations for Decipher high risk patients, and Marascio et al., 90 who demonstrated a ‘number needed to test’ of 3 to change one treatment decision, with Decipher testing changing recommendations for 39% of patients.
Oncotype Dx
The OncotypeDx Gene Expression classifier is a quantitative RT-PCR assay specifically mentioned in both the ASCO and NCCN guidelines that measures the expression of 17 genes (12 cancer specific and 5 housekeeping genes). The numeric Oncotype score, on a scale of 0 to 100, is combined with NCCN risk groupings on the test report to provide a comprehensive risk score. Furthermore, the report provides a percentage risk of PCa death within 10 years, metastasis within 10 years, and risk of adverse pathology (defined as Gleason ⩾ 4 + 3 or pT3 +) on RP (Figures S9-10). The cost of the test is $4520, but studies have shown an average savings of $2286 per patient (including Oncotype test cost) in the first 180 days after diagnosis for very low-risk and low-risk patients. 91
Prognostic value of Oncotype
In a single institutional cohort of 514 prostatectomy patients (91% white) with NCCN low or intermediate-risk disease, every 20-point increase in Oncotype score was associated with high grade (OR = 2.3, 95% CI = 1.5–3.7) and non-organ confined pathology (OR = 1.9, 95% CI = 1.3–3.0). Adding Oncotype to CAPRA resulted in less treatment of patients with favorable pathology. 92 In a separate study of 279 men with low, intermediate, and high-risk PCa, the addition of Oncotype to CAPRA improved the AUC for prediction of PCa specific death in a 10-year interval from 0.78 to 0.84 (p < 0.001). 93 A larger meta-analysis consisting of 732 patients revealed improved prediction ability for favorable pathology when Oncotype was added to CAPRA (AUC = 0.68–0.73) and NCCN (AUC = 0.64–0.70) risk groups. 94 In two separate cohorts that included NCCN unfavorable intermediate risk patients, incorporation of Oncotype improved the AUC for prediction of BCR to 0.76 from 0.63 and distant metastasis to 0.76 from 0.69 when incorporating only clinical variables. In addition, the AUC improved to 0.78 from 0.69 for prediction of PCa specific death in the unfavorable intermediate patients within the cohorts. 95 In a racially diverse cohort of 431 men (20% AA) with very low, low or intermediate-risk PCa, incorporation of Oncotype improved the AUC for prediction of adverse pathology from 0.63 (NCCN risk groups alone) to 0.72, and from 0.60 to 0.69 in the subset with Gleason 3 + 3 and low volume Gleason 3 + 4 (p = 0.001). For prediction of BCR, Oncotype improved the AUC from 0.59 (NCCN risk group alone) to 0.68. The association between Oncotype and recurrence free interval and adverse pathology was statistically significant in both white patients alone and African American patients alone. 96
Impact of Oncotype testing on treatment decisions
In a 297-patient cohort of men with low-risk PCa eligible for AS, AS utilization was higher in the Oncotype tested cohort than the baseline cohort, and the percentage of men remaining on AS at 1 year was 55% and 34%, respectively. Decision conflict scores fell from an average of 29 to 16 after Oncotype testing and the fraction of men expressing low decisional conflict increased from 36% to 59%. 97 Dall’Era et al. 98 showed that in a low-risk or very low-risk cohort of 211 men, the use of Oncotype testing, increased the recommendation and use of AS by 22% and 56%, respectively. Similarly, Lynch et al. 99 showed a 12% increase in AS utilization in Oncotype tested patients within a VA cohort. The impact of Oncotype testing on clinical decision-making in high-risk patients after prostatectomy was evaluated by Badani et al. They found that knowledge of the OncotypeDx score changed treatment recommendations in 31% of patients (95% CI, 27–35), with a 20% increase in observation recommendations in low-risk patients and a 16% increase in treatment escalation (e.g. adjuvant radiation) in those deemed to be high risk by the genomic classifier. Oncotype predicted probability of metastasis was the dominant risk factor driving decision making in a multivariable analysis consisting of clinical variables (OR = 8.6, 95% CI = 5.3–14.3, p < 0.001). 89
Use of Oncotype in the context of MRI
In a study of the relationship between Oncotype results and MRI, Oncotype scores were significantly higher in those with positive MRI (28, IQR 21–34) than indeterminate (23, IQR 14–27) and negative MRI (16, IQR 13–21). While the authors found stratification among MRI findings, additional study is required to recommend definitive clinical sequencing of MRI and tissue-based biomarkers. 100
Polaris
The Polaris test, which is mentioned in the NCCN and ASCO guidelines, consists of measurement of mRNA levels of 46 genes, comprising 31 genes involved in cell cycle progression and 15 housekeeping genes by RT-PCR. 101 The score is reported on a 10-point scale and gives the score percentile within the patient’s NCCN risk group (Figures S11 and S12). The report also provides a percentage risk of 10-year disease specific mortality and distant metastasis. Although the test costs $3400, the estimated cost savings per patient in a commercial health plan using Polaris testing is estimated to be ~$1900. 102
Prognostic value of Polaris score
The Prolaris score has been shown in several studies to predict BCR, metastasis, and PCa-specific death. In a multivariate analysis, the Polaris score was independently associated with a higher risk of BCR (HR = 1.77, 95% CI = 1.40–2.22, p = 4.3 × 10−6), but there was no difference in the AUC for BCR in the RP cohort (AUC = 0.825 for clinical variables alone, AUC = 0.84 for a combined model including the Polaris score). 101 Following primary radiotherapy, Polaris’s score was associated with 5-year BCR, with a HR per score unit of 2.55 (95% CI = 1.43–4.55, p = 0.002). In addition, the Polaris score was associated with 10-year disease-specific mortality, with a HR per score unit of 3.77, 95% CI = 1.37–10.4, p = 0.013. 103 In a group of 582 biopsy patients from multiple centers, Polaris was a significant predictor of BCR on multivariable analysis (HR = 1.47, 95% CI = 1.23–1.76, p = 4.7 × 10−5). Although the cohort included only 12 metastatic cases, the Polaris score was a highly significant predictor of metastatic disease (HR = 4.19, 95% CI = 2.08–8.45, p = 8.2 × 10−6). 104 In a conservatively managed TURP cohort, the Polaris score was also independently associated with PCa specific death risk on multivariate analysis (HR = 2.56, 95% CI = 1.85–3.53, p = 1.3 × 10−8). Furthermore, AUC for prediction of PCa death increased from 0.806 to 0.878 with the addition of Polaris to clinical variables. 101 A combined model consisting of CAPRA-S and Polaris was better able to predict BCR after prostatectomy as compared to CAPRA-S alone (AUC 0.77 versus 0.73) in a post-RP, low-risk cohort of 413 men. 105
Impact of Polaris testing on treatment decisions
In 305 cases from a national registry, Prolaris testing de-escalated treatment from interventional to non-interventional and vice versa in low and intermediate risk patients. There were no significant changes, however, in high-risk patients. Overall, clinicians felt that Polaris testing was useful in clinical decision making in 88% of cases. 106 In the larger PROCEDE-1000 prospective registry (n = 1206), 47.8% had a change in treatment, 72.1% of which were de-escalations; 26.9% were intensifications in treatment. 107 With respect to active surveillance, Kaul et al. showed in a multicenter study of 547 men after prostate biopsy that the rate of selection of AS in low-risk patients was 84.2% when patients had both Polaris score and NCCN risk criteria. This is significantly higher than the 20–40% utilization of AS in men with low-risk disease that was observed in other studies. In addition, there was a durability of AS utilization observed, with a 4-year durability rate of 69% in the academic setting and 63% in the community setting among patients in this study. 108
Guideline statements
Per NCCN guidelines, Decipher is covered post-biopsy for NCCN very-low, low-risk, favorable-intermediate, and unfavorable intermediate-risk PCa in patients with at least 10 years life expectancy who have not received treatment and are candidates for active surveillance or definitive therapy. Post-RP, Decipher is recommended and covered for those with pT2 disease with positive margins, any pT3 disease, and rising PSA. Oncotype and Polaris are covered post biopsy for very-low, low, and favorable intermediate risk patients. These tests are primarily validated in low and intermediate-risk patients, and no explicit recommendation is made for their use post-RP or in higher risk patients. 109
Per ASCO guidelines, Oncotype, Polaris, and Decipher are to be offered to patients who are likely to benefit from active surveillance and in whom the assay result is likely to change management. Decipher is specifically mentioned to help guide the decision of post-prostatectomy adjuvant or salvage radiation. 110
Other biomarkers in clinical development
Although this review focuses on biomarkers that are commercially available in the United States at present, there are several on the horizon with promising results. The miR Sentinel platform is certainly one of these tests. It consists of three modules that rely on exosomal miRNAs isolated from urine – the Sentinel PCa test, which distinguishes PCa from benign prostatic tissue; the Sentinel CS test, which distinguishes GG1 from GG2-5 disease; and the Sentinel HG test, which delineates patients with GG3-5 cancer who are candidates for therapeutic intervention rather than active surveillance. The AUCs of Sentinel PCa for detection of cancer and Sentinel CS for detection of GG2-5 disease were 0.98, and the AUC of Sentinel HG for detection of GG3-5 disease was 0.99. 111
Another emerging biomarker for early detection is Stockholm 3 (S3M), a test developed at the Karolinska Institute. It combines clinical variables with 232 SNPs and 6 protein markers (total PSA, fPSA, intact PSA, hK2, MSMB, and MIC1). In a multicenter study conducted in Norway and Sweden, S3M had an AUC significantly higher than PSA and PSA density in predicting GG2+ cancer (85.9 versus 74.8).112,113 The major limitations of the S3M are a validation population consisting of mainly Northern Europeans and reliance on 232 SNPs. It is currently only commercially available in Sweden. Proclarix is another novel test that combines thrombospondin-1 and cathepsin D with age, PSA and fPSA and has been shown to predict csPCa on biopsy with an AUC of 0.83, which is significantly higher than fPSA alone (AUC = 0.64). However, this was a retrospective, single-center study.113,114
Circulating immune cells have also been proposed as biomarkers across the PCa clinical care continuum, including predicting adverse pathology at RP and risk stratification of patients being considered for AS. While none of these tests have been implemented at scale clinically, circulating neutrophil, platelet, eosinophil, and lymphocyte content have shown early promise.115,116
Conclusion
The selection of the appropriate biomarker is highly nuanced and dependent on the specific clinical scenario. Here, we have provided foundational information to guide decision making on the use of these biomarkers and suggest opportunities for additional data collection. In the early detection space, such areas of study include more robust studies of test performance in racially diverse populations and how these biomarkers can be integrated with MRI. The risk stratification biomarkers all demonstrate similar prognostic performance and utility for low-risk patients considering active surveillance. Decipher is unique in that it has been extensively studied in high-risk patients and in the guidance of multi-modal therapy in post-RP patients with high-risk features or treatment failure. Future opportunities exist to further evaluate how these biomarkers can be used in the management of higher-risk patients requiring multi-modal therapy and in the guidance of surgical approaches.
Supplemental Material
sj-docx-1-tau-10.1177_17562872221103988 – Supplemental material for Biomarkers for prostate cancer detection and risk stratification
Supplemental material, sj-docx-1-tau-10.1177_17562872221103988 for Biomarkers for prostate cancer detection and risk stratification by Mark W. Farha and Simpa S. Salami in Therapeutic Advances in Urology
Footnotes
Author contributions
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Conflict of interest statement
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: M.W.F. has a contractor engagement providing consulting services to LynxDx. S.S.S. is on a Study Advisory committee for Bayer Pharma.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
