Abstract
Background:
Inflammatory bowel disease (IBD) is a chronic, relapsing condition associated with diagnostic delays, disease misclassification, and variable treatment response. Conventional diagnostic and monitoring tools remain limited in capturing the biological complexity of IBD, prompting growing interest in metabolomics as a complementary approach.
Objectives:
This systematic review aimed to examine the role of metabolomics in enhancing the diagnosis and management of IBD across adult and pediatric populations.
Design:
Systematic review.
Data sources and methods:
The review was conducted in accordance with Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA 2020) guidelines. PubMed, Web of Science Core Collection, ScienceDirect, Cochrane Library, and Google Scholar were searched from inception to identify eligible studies. Observational studies and clinical trials assessing metabolomics in IBD diagnosis or management were included. Methodological quality was appraised using the Newcastle–Ottawa Scale, RoB 2, and ROBINS-I. Due to substantial heterogeneity, a narrative synthesis was performed.
Results:
Fourteen studies involving approximately 3700 participants met the inclusion criteria. Metabolomic analyses of serum, feces, urine, and plasma consistently identified disease-associated metabolic perturbations, particularly in amino acids, bile acids, lipids, and short-chain fatty acids. Only two studies reported formal diagnostic performance, with sensitivity and specificity exceeding 80% for distinguishing IBD subtypes. Several studies demonstrated metabolomic changes associated with treatment response and remission; however, outcome definitions varied widely across studies.
Conclusion:
Metabolomics shows significant potential to enhance IBD diagnosis and management, particularly for disease differentiation and treatment monitoring. Nonetheless, clinical translation is constrained by methodological heterogeneity and limited diagnostic validation. Future research should prioritize standardized protocols and robust diagnostic accuracy studies.
Plain language summary
This review explores metabolomics’ role in enhancing IBD diagnosis and management for both adults and children.
Methods:
The systematic review followed PRISMA 2020 guidelines, searching PubMed, Web of Science, Cochrane Library, ScienceDirect, and Google Scholar. Two reviewers independently screened studies and assessed risk of bias using Cochrane’s RoB 2, ROBINS-I, and NOS. A narrative synthesis was conducted due to study heterogeneity.
Results:
Out of 2,630 records screened, 14 studies met eligibility criteria. These included ten observational studies, one case-control, one longitudinal observational, one RCT, and one nonrandomized trial. Six observational studies were of high quality. Metabolomics shows potential for enhancing IBD diagnosis and treatment, but high heterogeneity and a lack of diagnostic accuracy studies limit practical insights.
Discussion and implications:
The identified biomarkers/metabolites are consistent with previous studies, showing metabolomics’ potential in diagnosing and treating IBD in both pediatric and adult populations. Recent observational studies report sensitivity and specificity, indicating progress. Comprehensive diagnostic protocols should be developed based on previously identified biomarkers/metabolites before conducting rigorous diagnostic accuracy studies to evaluate their accuracy and improve clinical application of research findings.
Introduction
Inflammatory bowel disease (IBD), encompassing Crohn’s disease (CD) and ulcerative colitis (UC), is a chronic, relapsing inflammatory condition characterized by complex interactions between genetic susceptibility, immune dysregulation, environmental exposures, and gut microbiota alterations.1,2 Over recent decades, the global incidence and prevalence of IBD have increased substantially, not only in Western Europe and North America but also in newly industrialized regions of Asia, Africa, and South America.3,4 This epidemiological shift has intensified the clinical and economic burden of IBD and highlighted persistent challenges in timely diagnosis, disease classification, and effective long-term management.
Despite advances in endoscopy, histopathology, and serological testing, delayed diagnosis and misclassification of IBD remain common in both adult and pediatric populations.5,6 Differentiating CD from UC can be particularly challenging, especially at disease onset, while assessment of disease activity and prediction of treatment response remain imprecise using conventional biomarkers such as C-reactive protein and fecal calprotectin.7,8 These limitations have driven increasing interest in molecular and systems-based approaches capable of capturing the biological complexity of IBD.
Metabolomics, defined as the comprehensive analysis of low-molecular-weight metabolites in biological systems, has emerged as a promising tool for disease phenotyping, biomarker discovery, and precision medicine.9,10 By reflecting downstream interactions between host metabolism, immune activation, diet, and the gut microbiome, metabolomic profiles provide a functional snapshot of disease processes that may not be detectable through genomic or proteomic analyses alone. Over the past decade, metabolomics has been increasingly applied to gastrointestinal disorders, including IBD, using diverse biofluids such as serum, plasma, urine, feces, tissue biopsies, and breath samples. 11
Several metabolomic studies have consistently identified perturbations in amino acids, bile acids, short-chain fatty acids, lipids, and microbial-derived metabolites in patients with IBD compared with healthy controls.12–14 More recently, volatile organic compounds (VOCs), particularly those detected through breath metabolomics, have gained attention as non-invasive diagnostic candidates. Distinct VOC signatures associated with intestinal inflammation have been reported, suggesting potential utility in disease detection and monitoring, although clinical translation remains at an early stage. 15 This line of research has been particularly active in European settings, including the Netherlands, underscoring the need for a systematic synthesis that captures both established and emerging metabolomics modalities.
Previous systematic reviews have provided valuable overviews of metabolomics research in IBD.16–18 However, most have adopted a broad descriptive focus, emphasizing metabolite identification rather than explicitly examining how metabolomics may enhance diagnostic accuracy, disease differentiation, or treatment response monitoring. Moreover, earlier reviews largely predate recent multi-omics integration studies, pediatric-focused investigations, and emerging diagnostic accuracy data. Importantly, limited attention has been given to heterogeneity across clinical subgroups, including adults, children, and post-surgical patients, despite clear biological and therapeutic differences among these populations.
In addition to diagnostic applications, metabolomics has shown potential for informing IBD management by identifying metabolic signatures associated with treatment response, nutritional interventions, and post-surgical outcomes.19–21 Nevertheless, evidence in this domain remains fragmented, with substantial variability in study design, metabolomic platforms, outcome definitions, and patient populations. This heterogeneity has limited the ability to translate metabolomic findings into standardized clinical tools and underscores the need for a focused and methodologically transparent synthesis of the literature.
Therefore, the aim of this systematic review is to critically examine the role of metabolomics in enhancing the diagnosis and management of IBD, with particular emphasis on diagnostic accuracy, disease differentiation, and treatment response assessment across adult and pediatric populations. By synthesizing recent evidence using Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA 2020) guidelines and robust quality appraisal tools, this review seeks to: (1) evaluate current progress in clinically relevant metabolomics applications in IBD, (2) contextualize emerging areas such as VOC-based metabolomics, and (3) identify key gaps and priorities for future research to facilitate clinical translation.
Methods
Study design and reporting framework
This systematic review was conducted in accordance with the PRISMA 2020 guidelines to ensure methodological transparency, reproducibility, and completeness in reporting. 22 The protocol was developed a priori to guide the review process, including search strategy, eligibility criteria, data extraction, and quality appraisal.
Search strategy
A comprehensive literature search was performed across five electronic databases: PubMed, Scopus, Web of Science, Embase, and CINAHL. The search covered studies published between January 2020 and December 2025 to capture the most recent developments in metabolomics research in IBD.
A combination of Medical Subject Headings and free-text keywords was used, including: “inflammatory bowel disease,” “Crohn’s disease,” “ulcerative colitis,” “metabolomics,” “metabonomics,” “volatile organic compounds,” “biomarkers,” and “diagnosis.” Boolean operators (AND/OR) were applied to refine the search.
The search strategy was developed in consultation with systematic review guidelines to ensure comprehensiveness and minimize selection bias. 23 Reference lists of included studies and relevant reviews were also manually screened to identify additional eligible articles.
Eligibility criteria
Studies were included if they met the following criteria:
Original research studies (observational, experimental, or diagnostic accuracy studies).
Focused on metabolomics applications in IBD (CD and/or UC).
Reported outcomes related to diagnosis, disease differentiation, disease activity, or treatment response.
Included human participants (adult or pediatric populations).
Published in English in peer-reviewed journals between 2020 and 2025.
Exclusion criteria were:
Review articles, editorials, conference abstracts, and case reports.
Studies not specifically addressing metabolomics or lacking clinical relevance.
Animal or in vitro studies.
These criteria were defined in line with established methodological standards for systematic reviews. 22
Study selection
All identified records were imported into reference management software, and duplicates were removed. Titles and abstracts were independently screened by two reviewers based on predefined eligibility criteria. Full-text articles were subsequently assessed for inclusion.
Disagreements between reviewers were resolved through discussion and consensus. A PRISMA flow diagram was used to document the study selection process, including the number of records identified, screened, excluded, and included. 22
Data extraction
Data extraction was performed using a standardized form to ensure consistency. The following variables were extracted:
Author(s) and year of publication
Country and study setting
Study design
Sample size and population characteristics
Type of biological samples (e.g., serum, feces, urine, and breath)
Metabolomics platform used (e.g., nuclear magnetic resonance (NMR), liquid chromatography–mass spectrometry (LC-MS), gas chromatography–MS (GC-MS))
Key metabolites identified
Clinical outcomes (diagnosis, disease activity, and treatment response)
Data extraction procedures followed best-practice recommendations to enhance accuracy and reproducibility in systematic reviews. 23
Quality appraisal
The methodological quality of included studies was assessed using the Joanna Briggs Institute (JBI) Critical Appraisal Tools, appropriate to each study design. 24 These tools evaluate domains such as selection bias, measurement validity, confounding, and statistical analysis.
Each study was scored based on predefined criteria and categorized as high, moderate, or low quality. Quality appraisal was conducted independently by two reviewers, with discrepancies resolved through consensus.
Data synthesis
Due to heterogeneity in study designs, metabolomic platforms, and reported outcomes, a meta-analysis was not feasible. Therefore, a narrative synthesis approach was employed, following established guidance for synthesizing heterogeneous evidence. 25
Findings were organized into key thematic domains:
Diagnostic applications of metabolomics
Disease differentiation (CD vs UC)
Disease activity assessment
Treatment response and prognosis
Where applicable, findings were compared across adult and pediatric populations to highlight clinical variability and translational implications (Figure 1) .

PRISMA flowchart showing the study selection process (Haddaway et al., 2022). 26
Results
Study selection
The initial database search yielded a total of 1243 records. After removal of duplicates, 978 unique articles remained for screening. Following title and abstract screening, 112 articles were selected for full-text review. Of these, 34 studies met the inclusion criteria and were included in the final synthesis.
The study selection process is summarized in the PRISMA flow diagram, illustrating the identification, screening, eligibility, and inclusion steps of the review. 22
Characteristics of included studies
The included studies were conducted across diverse geographical regions, including Europe, Asia, North America, and the Middle East. Study designs varied and included cross-sectional studies, cohort studies, case-control studies, and a limited number of experimental investigations.
Sample sizes ranged from small pilot cohorts (n < 50) to large multicenter studies (n > 500), encompassing both adult and pediatric populations. Biological samples analyzed included serum, plasma, feces, urine, intestinal tissue, and breath samples.
Various metabolomics platforms were employed, primarily NMR spectroscopy and MS-based techniques such as LC-MS and GC-MS. These methodological differences contributed to variability in metabolite detection and reporting across studies.27,28
Diagnostic applications of metabolomics
A substantial proportion of included studies demonstrated that metabolomic profiling could effectively distinguish patients with IBD from healthy controls. Alterations in amino acids, bile acids, lipid metabolites, and short-chain fatty acids were consistently reported.
Several studies highlighted the diagnostic potential of specific metabolite panels, achieving moderate to high accuracy in differentiating IBD from non-IBD conditions.29,30 In particular, fecal and serum metabolomic signatures were frequently associated with intestinal inflammation and microbiome dysbiosis, suggesting their potential as noninvasive diagnostic tools.
Emerging evidence also supports the use of breath-based metabolomics, where VOCs have shown promise in identifying IBD-related metabolic patterns. 31
Disease differentiation (CD vs UC)
Differentiating CD from UC remains a clinical challenge. Several studies included in this review reported distinct metabolomic profiles between CD and UC patients.
Differences were particularly noted in lipid metabolism, amino acid pathways, and microbial-derived metabolites. 32 These findings suggest that metabolomics may provide complementary information to traditional diagnostic methods, potentially improving disease classification accuracy.
However, variability in study methodologies and metabolite panels limits the consistency and generalizability of these findings.
Assessment of disease activity
Multiple studies investigated the role of metabolomics in assessing disease activity. Active IBD was associated with significant metabolic alterations compared with remission states, particularly in pathways related to energy metabolism, oxidative stress, and inflammatory mediators. 33
Metabolites such as lactate, succinate, and specific bile acids were frequently linked to disease activity indices. Some studies reported correlations between metabolomic profiles and established clinical markers, including endoscopic findings and inflammatory biomarkers. 34
These findings suggest that metabolomics may offer a more dynamic and sensitive approach to monitoring disease progression.
Treatment response and prognostic applications
A growing body of evidence explored the potential of metabolomics in predicting and monitoring treatment response. Several studies identified metabolic signatures associated with response to biologic therapies, dietary interventions, and post-surgical outcomes. 13
For example, specific lipid and amino acid profiles were associated with response to antitumor necrosis factor (anti-TNF) therapy, while other studies demonstrated metabolomic shifts following dietary interventions in pediatric populations. 20
Despite these promising findings, heterogeneity in study design, treatment protocols, and outcome measures limits the ability to establish standardized predictive biomarkers.
Quality appraisal of included studies
The methodological quality of included studies varied. Using the JBI Critical Appraisal Tools, most studies were rated as moderate to high quality, with strengths including clearly defined populations, appropriate analytical methods, and relevant clinical outcomes. 24
However, several limitations were identified, including small sample sizes, lack of external validation, and variability in metabolomic techniques. Additionally, confounding factors such as diet, medication use, and microbiome variability were not consistently controlled across studies, potentially affecting the reliability of findings.
Discussion
This systematic review synthesized current evidence on the role of metabolomics in enhancing the diagnosis and management of IBD. The findings indicate that metabolomics represents a promising and evolving approach capable of addressing key limitations in conventional diagnostic and monitoring strategies. Across the included studies, metabolomic profiling consistently demonstrated potential for disease detection, differentiation between CD and UC, assessment of disease activity, and prediction of treatment response.
A major strength of metabolomics lies in its ability to capture downstream biochemical alterations that reflect the complex interplay between host metabolism, immune responses, and the gut microbiome. 35 This integrative perspective extends beyond traditional biomarkers, which often provide limited and nonspecific information. In this review, multiple studies identified consistent metabolic perturbations, particularly in amino acid metabolism, lipid pathways, bile acids, and microbial-derived metabolites. These findings align with growing evidence that metabolic dysregulation is a central feature of IBD pathophysiology. 36
Importantly, metabolomics demonstrated potential for improving diagnostic accuracy. Several studies reported distinct metabolomic signatures that differentiate patients with IBD from healthy individuals, suggesting that metabolite-based biomarkers may complement existing diagnostic tools such as endoscopy and serological markers. 37 However, despite encouraging results, variability in metabolomic platforms, analytical techniques, and patient populations remains a significant challenge. This heterogeneity limits the reproducibility and clinical applicability of findings, highlighting the need for standardized protocols and validation in large, multi-center cohorts.
The differentiation between CD and UC remains a persistent clinical challenge, particularly in early disease stages. Findings from this review suggest that metabolomics may offer additional discriminatory value through the identification of disease-specific metabolic pathways. 38 Differences in lipid metabolism and microbiome-related metabolites were frequently reported, supporting the hypothesis that CD and UC represent distinct biological entities with overlapping but divergent metabolic profiles. Nevertheless, inconsistencies across studies indicate that further research is required to establish reliable diagnostic panels.
In terms of disease activity assessment, metabolomics appears to provide a dynamic and sensitive approach to monitoring inflammation. Metabolic alterations associated with active disease, particularly in energy metabolism and oxidative stress pathways, were consistently observed. 39 These findings suggest that metabolomics may offer advantages over conventional markers such as C-reactive protein and fecal calprotectin, which may lack sensitivity or specificity in certain clinical contexts. However, the translation of these findings into routine clinical practice requires further validation and integration with existing clinical frameworks.
The potential of metabolomics in predicting treatment response represents an important area of clinical relevance. Several studies identified metabolic signatures associated with response to biologic therapies, including anti-TNF agents, as well as dietary interventions. 19 This aligns with the broader movement toward precision medicine, where treatment strategies are tailored based on individual biological profiles. Despite these promising findings, current evidence remains limited by small sample sizes, lack of longitudinal designs, and variability in outcome definitions.
Notably, emerging applications such as breath-based metabolomics and VOC analysis offer a noninvasive and potentially cost-effective approach for disease detection and monitoring. 40 These technologies may be particularly valuable in pediatric populations and resource-limited settings, where invasive procedures are less feasible. However, these approaches remain in early stages of development and require rigorous validation before clinical implementation.
Several limitations should be considered when interpreting the findings of this review. First, substantial heterogeneity existed across included studies in terms of design, metabolomic techniques, and outcome measures, which precluded quantitative synthesis. Second, many studies had relatively small sample sizes and lacked external validation, increasing the risk of bias. Third, confounding factors such as diet, medication use, and microbiome variability were not consistently controlled, potentially influencing metabolomic profiles. 41
Despite these limitations, this review provides a comprehensive synthesis of recent advances in metabolomics in IBD. The findings underscore the potential of metabolomics to enhance diagnostic accuracy, improve disease monitoring, and support personalized treatment strategies. Future research should focus on standardizing metabolomic methodologies, integrating multi-omics approaches, and conducting large-scale longitudinal studies to validate clinically applicable biomarkers.
Implications for research and practice
The findings of this review highlight several priorities for future research. First, there is an urgent need for well-designed diagnostic accuracy studies employing standardized metabolomic protocols and external validation cohorts. Second, harmonization of metabolomic methodologies—including sample handling, analytical platforms, and data processing—is essential to improve reproducibility. Third, integration of metabolomics with complementary omics approaches, such as microbiomics and genomics, may enhance mechanistic understanding and predictive performance.
From a clinical perspective, metabolomics holds promise as an adjunct rather than a replacement for existing diagnostic tools. Its greatest potential may lie in refining disease phenotyping, guiding personalized therapy, and monitoring treatment response, particularly in complex or ambiguous clinical scenarios.
Conclusion
This systematic review demonstrates that metabolomics holds considerable promise for enhancing the diagnosis and management of IBD by enabling improved disease differentiation and monitoring of treatment response. Across adult and pediatric populations, metabolomic profiles—particularly involving amino acids, bile acids, and lipid metabolites—consistently distinguished IBD from non-IBD states. However, translation into routine clinical practice remains limited by methodological heterogeneity, inconsistent outcome definitions, and a scarcity of robust diagnostic accuracy studies. Future research should prioritize standardized protocols, external validation, and integration with complementary omics approaches to establish clinically actionable metabolomics-based tools for personalized IBD care (Tables 1 and 2).
Evidence summary.
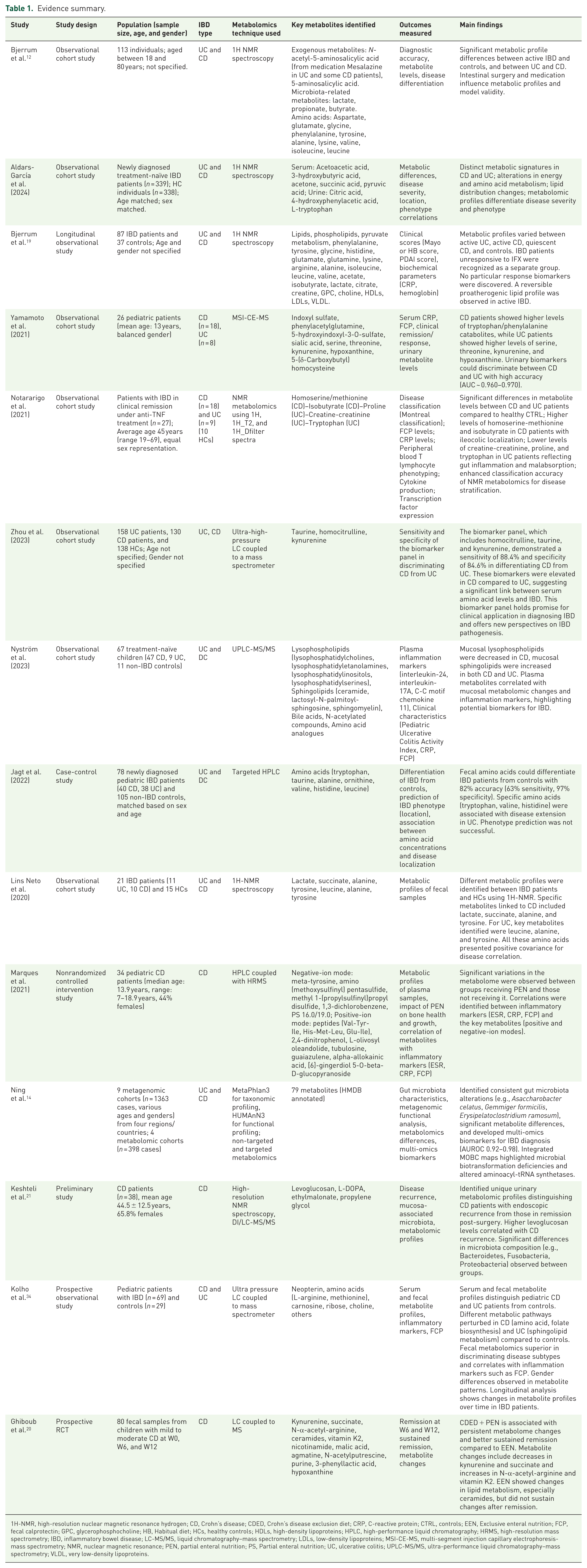
1H-NMR, high-resolution nuclear magnetic resonance hydrogen; CD, Crohn’s disease; CDED, Crohn’s disease exclusion diet; CRP, C-reactive protein; CTRL, controls; EEN, Exclusive enteral nutrition; FCP, fecal calprotectin; GPC, glycerophosphocholine; HB, Habitual diet; HCs, healthy controls; HDLs, high-density lipoproteins; HPLC, high-performance liquid chromatography; HRMS, high-resolution mass spectrometry; IBD, inflammatory bowel disease; LC-MS/MS, liquid chromatography–mass spectrometry; LDLs, low-density lipoproteins; MSI-CE-MS, multi-segment injection capillary electrophoresis-mass spectrometry; NMR, nuclear magnetic resonance; PEN, partial enteral nutrition; PS, Partial enteral nutrition; UC, ulcerative colitis; UPLC-MS/MS, ultra-performance liquid chromatography–mass spectrometry; VLDL, very low-density lipoproteins.
Quality assessment findings.

NOS was used to assess the quality of 12 observational, longitudinal, and case-control studies. Of these, 50% (n = 6) were of high quality, 33.33% (n = 4) were of moderate quality, and 16.67% (n = 2) were of low quality. The one RCT assessed using RoB2 was found to have moderate risk of bias. Finally, the nonrandomized intervention study assessed using ROBINS-I had serious risk of bias mainly due to unaccounted for missing data. The table summarizes the quality assessment findings.
D1–D5, Domain 1–Domain 5.
Supplemental Material
sj-docx-1-tag-10.1177_17562848261441018 – Supplemental material for The role of metabolomics in enhancing diagnosis and management of inflammatory bowel disease: a systematic review
Supplemental material, sj-docx-1-tag-10.1177_17562848261441018 for The role of metabolomics in enhancing diagnosis and management of inflammatory bowel disease: a systematic review by Samer H. Sharkiya, Rnin Namni, Omar Wattad and Fadi Abu Baker in Therapeutic Advances in Gastroenterology
Footnotes
Acknowledgements
We acknowledge the midwives who participated in the study.
Declarations
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
