Abstract
Interest in artificial intelligence (AI) applications for ulcerative colitis (UC) has grown tremendously in recent years. In the past 5 years, there have been over 80 studies focused on machine learning (ML) tools to address a wide range of clinical problems in UC, including diagnosis, prognosis, identification of new UC biomarkers, monitoring of disease activity, and prediction of complications. AI classifiers such as random forest, support vector machines, neural networks, and logistic regression models have been used to model UC clinical outcomes using molecular (transcriptomic) and clinical (electronic health record and laboratory) datasets with relatively high performance (accuracy, sensitivity, and specificity). Application of ML algorithms such as computer vision, guided image filtering, and convolutional neural networks have also been utilized to analyze large and high-dimensional imaging datasets such as endoscopic, histologic, and radiological images for UC diagnosis and prediction of complications (post-surgical complications, colorectal cancer). Incorporation of these ML tools to guide and optimize UC clinical practice is promising but will require large, high-quality validation studies that overcome the risk of bias as well as consider cost-effectiveness compared to standard of care.
Plain language summary
Ulcerative colitis (UC) is a chronic inflammatory disorder of the colon. The clinical care of patients with UC and research efforts to better understand the disease has inevitably produced a significant quantity of diverse and complex datasets ranging from electronic health records, laboratory values, images (endoscopy, radiology, histology) to gene expression. The size and complexity of datasets derived from UC poses a significant challenge to accurately and effectively predict clinically meaningful endpoints in order to ultimately improve UC outcomes. Artificial intelligence through the application of machine learning tools has the potential to improve the analysis of large, complex, high-dimensional datasets and reveal novel, deeper insights compared to traditional analytical tools. Here, we provide an updated and comprehensive summary of AI applications in UC.
Introduction
Ulcerative colitis (UC) is a chronic inflammatory disorder of the gut without a medical cure that affects nearly 1 million Americans. 1 Inflammatory bowel disease (IBD) is characterized by intestinal dysbiosis and immune dysregulation. Environmental factors, particularly diet, are thought to play a key role in disease pathogenesis, particularly via impact on the gut microbiome.2,3
Current therapeutics control IBD via broad immunosuppression but do not address the underlying intestinal dysbiosis. Further, despite our best therapies, most patients do not achieve long-term remission highlighting the need for improved disease monitoring and personalized therapeutic interventions. 4 There is a growing interest in utilizing deep, multi-omics phenotyping in IBD including whole exome sequencing, transcriptomics, proteomics, and metagenomics of the microbiota. Additionally, there is a rapid expansion in the growth of clinical images from endoscopy and pathology samples. The resultant rapid expansion of data has led to interest in the application of artificial intelligence (AI) to IBD.
AI is a multidisciplinary field that seeks to apply computer software to mimic human intelligence. Machine learning (ML) is a subset of AI that uses statistical methods to recognize patterns from datasets and can be done through supervised and unsupervised methods. Supervised learning relies on labeled input to give accurate classification or prediction of the outcome of interest. Examples of supervised learning include regression, K-nearest neighbor, and random forest (RF). Unsupervised learning, in contrast, does not require labeled input and is used to reduce dimensionality and allow for clustering.5,6 Deep learning (DL) utilizes artificial neural networks (ANN), which mimic brain logic structures, to perform complex learning tasks by utilizing layers of representation and subsequent transformation to highlight aspects of the input which improves task performance. Examples of DL include virtual assistants and image recognition. 6
Over the last decade, there has been increased application of AI in IBD. 7 In particular, computer vision in endoscopy in UC has been a key area of growth.8–11 The purpose of this review is to provide an updated and comprehensive evaluation of recent advances in AI in UC, with a particular focus on the prediction and diagnosis of new UC, prediction of response to therapy, disease monitoring, and identification of disease complications. We also review challenges to the translation of these novel technologies into the clinic and discuss future directions.
Literature search strategy
We performed a literature review using PubMed (MEDLINE) from inception until July 30, 2023, of all studies applying AI in UC. Our search strategy consisted of the following combinations: (((((((ulcerative colitis [Title]))) AND (artificial intelligence [Title])) OR (computer-assisted [Title])) OR (computer-aided [Title])) OR (machine learning [Title])) OR (deep learning [Title]). We included studies that used AI in the (1) prediction and diagnosis of UC, (2) prediction of response to therapy in UC, (3) monitoring disease activity in patients with UC, and (4) prediction of complications of UC. We excluded reviews, studies with non-human subjects (animal models), or studies that did not provide objective measures of the efficacy of AI applications (e.g. area under the curve (AUC), sensitivity, specificity, etc.).
Results
Our search strategy yielded 97 studies that applied AI to UC, of which 61 studies met our inclusion criteria. In total, 54 (88.5%) of studies were published in the last 5 years. Eighteen studies focused on the prediction and diagnosis of new UC, 11 studies predicted response to therapy, 15 evaluated disease monitoring, and 14 focused on prediction of UC complications. The AI methods utilized include: linear regression (LR), lasso regression, gradient boosted machine (GBM), principal component analysis (PCA), RF, linear discriminant analysis (LDA), support vector machines (SVM), segment anything model (SAM), ANN.
Prediction and diagnosis of new UC
Identification of biologic pathways in UC
What is already known?
While previous research has identified some common genetic, environmental, and microbial risk factors for UC, the associations are neither strong enough nor consistent enough to be clinically useful.12–16 The use of AI has enormous potential for assessing risk and identifying biologic pathways enriched in UC compared to the general population.
What do current studies show?
Table 1 summarizes studies that applied AI to the diagnosis of new UC. Four studies utilized omics data, including genetic/genomic (n = 3) and transcriptomic (n = 1) data sets. While this is a growing area of research interest, only a few studies have specifically focused on prediction of UC from a healthy population.
Prediction and diagnosis of new UC.

ANN, artificial neural network; AUC, area under the curve; CI, confidence interval; CNN, convolutional neural networks; CRP, C-reactive protein; DEGs, differentially expressed genes; DL, deep learning; GBM, gradient boosted machine; GIF, guided image filtering; IBD, inflammatory bowel disease; LASSO, least absolute shrinkage and selection operator; MCC, Matthews correlation coefficient; PCA, principal component analysis; QWK, quadratic weighted kappa; RGB, red, green, blue; RF, random forest; RNN, recurrent neural network; SAM, segment anything model; SVM-RFE, support vector machines recursive feature elimination; UC, ulcerative colitis; UCEIS, UC endoscopic index of severity.
In a cross-sectional study of colon biopsy samples from 298 active UC and 76 healthy control patients by Tang et al., a combination of three ML algorithms—including least absolute shrinkage and selection operator (LASSO), SVM recursive feature elimination (SVM-RFE), and RF—identified seven differentially expressed cell death-related genes (average AUC 0.859) to build a prediction model of UC diagnosis. The resulting nomogram had good predictive performance with an AUC of 0.982 in the validation set. 17 In a separate cross-sectional study of 387 UC and 139 healthy patients by Zhang et al., 2 useful genes (OLFM4 and C4BPB) were identified using a combination of 6 ML methods including SVM, LASSO, RF, GBM, PCA, and ANN. OLFM4 and C4BPB were found to be of diagnostic values as determined by an average AUC of 0.865 based on their performance in training, test, and independent validation sets. Notably, both genes were significantly correlated with M1 macrophages, M2 macrophages, activated mast cells, resting mast cells, monocytes, and activated natural killer cells (p < 0.05). 18 Another cross-sectional analysis of 259 UC and 41 healthy patients by Li et al. utilized RF to identify differentially expressed genes (DEGs) with highest contribution to UC occurrence from sets of mucosal transcriptomic profiles from rectal biopsies and used an artificial neural net to calculate DEG weights to UC. The algorithms demonstrated excellent prediction performance of AUC 0.9506, which also agreed with that of an independent data set. 19 In a separate cross-sectional study of 178 patients (143 UC, 35 healthy control), Han et al. constructed a disease classifier from 41 genes using SVM-RFE for diagnosis of UC. The model demonstrated high accuracy of 96.5% and performed excellently in training and validation sets with an AUC of 0.999 and average AUC of 0.832, respectively. 20
What could AI add in the future?
The prevalence of UC is increasing, and despite this, our understanding of the pathophysiology of UC is still limited. Bench scientists have increasingly applied a systems biology approach to study disease pathogenesis, and there has been a resultant explosion in the volume of scientific data. While the current studies have applied traditional ML methods to these datasets, there is an opportunity to apply DL methods to these data to generate novel insights.
AI in diagnosis of UC
Traditionally, evaluation and diagnosis of UC involve comparing clinical symptoms to relevant laboratory data, radiographic imaging, and endoscopic reports via index colonoscopy. 29 In recent years, many studies have begun exploring the potential of AI methodologies to enhance prediction and accuracy of UC diagnosis, improve treatment outcomes through early diagnosis, and discovery of novel pathways associated with UC pathogenesis.
Of the 14 total studies, 3 used AI to assist in the analysis of diagnostic labs and radiographic data, 6 involved the use of AI to aid in the discovery of novel biomarkers for diagnosis, and 5 studies utilized AI or computer vision in index colonoscopy.
AI analysis of laboratory, pathology, and radiographic data
What is already known?
Current clinical practice for diagnosis of UC relies on a combination of laboratory testing and radiographic data in combination with endoscopic and histological evaluation. Laboratory testing includes assessment of serum inflammatory markers such as leukocyte count and differential, platelet count, and C-reactive protein (CRP) as well as stool tests such as fecal calprotectin or lactoferrin levels which are stronger indications of activation of immune pathways in the gut. 30 Radiographic techniques using magnetic resonance imaging, computed tomography, and abdominal ultrasound can be used to rule out small bowel involvement and distinguish UC from other gastrointestinal pathologies. 31 Despite the best application of current technology, approximately 5%–10% of patients with IBD are initially diagnosed with indeterminate colitis.
What do current studies show?
The overarching goal of research in this area is to use AI techniques to create an objective model for evaluation of clinical labs and radiographic data to improve accuracy and precision of UC diagnosis. Of the studies that employed AI for the analysis of labs and radiographic data, data modalities included electronic health records (n = 2 studies) and imaging datasets (n = 1 study).
In a prospective cohort study of 702 medical records belonging to 372 IBD patients (180 UC, 192 CD) conducted by Kraszewski et al., an RF algorithm was used to create an UC diagnostic prediction model based on routine blood, urine, and fecal tests compared to diagnosis based on CRP alone. While the RF ensemble achieved a mean average precision of 91% for UC, the comparison to CRP alone does not represent typical clinical practice; no comparison was made to physician diagnosis. 21 In a separate cross-sectional study of 74 pediatric colonic IBD (56 UC and 18 colonic-CD) patients, Dhaliwal et al. used an RF classifier that accurately distinguished UC from colonic-CD in 97% of patients in the training set, and 100% of the patients in the validation set of patients when given a combination of baseline clinical, endoscopic, radiologic, and histologic data. 22
Jiang et al. demonstrated diagnostic ability of a guided image filtering (GIF) algorithm in a cross-sectional study of 60 patients with suspected IBD and 60 non-IBD patients undergoing radiologic examination via CT scan. The improved GIF algorithm accurately diagnosed 98.3% of UC cases. Despite the smaller sample size, the performance characteristics are promising and show the capability of AI to enhance diagnostic accuracy when applied to CT images. 8
What could AI add in the future?
The three studies on the application of AI to the diagnosis of UC have significant limitations. One study did not compare their model to physician diagnosis; all studies focused on differentiating patients who clearly had either UC or CD from each other, which is not a problem a practicing gastroenterologist typically faces. Rather, given the expanding repertoire of IBD medications, some of which are specific to UC, applying AI to accurately classify UC or CD among patients initially diagnosed with indeterminate colitis would be more clinically relevant.
AI-assisted discovery of novel biomarkers for diagnosis
What is already known?
Leading biomarkers for UC include serum CRP and fecal calprotectin.32,33 While both of these biomarkers indicate inflammatory states at a systemic and gastrointestinal level, respectively, elevated levels of these markers are not sufficient to diagnose UC without additional more invasive testing through endoscopic and histological evaluation. 33 There is ongoing and increasing interest in the identification of novel biomarkers that are specific and sensitive enough to have strong clinical relevance to UC diagnosis. The lack of specific diagnostic signatures for UC has been noted as a potential barrier to early detection. 34 Of the six studies which focused on using AI to assist in the discovery of novel biomarkers for UC diagnosis, data modalities included genetic/genomics (n = 2 studies), transcriptomics (n = 3 studies), and proteomics (n = 1 study).
What do current studies show?
In a cross-sectional study of blood samples from 20 UC and 20 healthy patients, Duttagupta et al. applied a SAM to identify 31 differentially expressed platelet-derived miRNAs from whole genome maps of circulating miRNAs from PBMC, micro-vesicles, and platelets. They then used SVM to evaluate biomarker performance using non-probabilistic binary linear classification, which revealed predictive scores with 92.8% accuracy and specificity and sensitivity of 96.2% and 89.5%, respectively. Candidate biomarkers independently validated by qPCR assays run on pooled patient and control samples and demonstrated 88% success. 25
Lu et al. created a logistic regression model based on five genes (REG3A, REG1A, DEFA6, REG1B, and DEFA5) determined to be strongly associated with UC occurrence based on analysis of a microarray of colonic biopsies from 106 UC and 21 healthy patients. The logistic model demonstrated strong performance at predicting UC with average AUC of 0.850, and AUC of 0.721 when evaluated in an independent set of 137 unseen samples. 23 Khorasani et al. took a similar approach, using a SVM classifier to distinguish between healthy controls and patients with UC by gene expression, but the model had poor precision for identifying inactive UC. 24 In a separate cross-sectional study involving microarray expression data of 193 UC and 42 healthy control patients, Wang et al. identified 64 upregulated and 38 downregulated genes then used LASSO regression and SVM-RFE to identify 5 diagnostic genes with strong ability to distinguish UC cases from normal samples. They found UC samples had significantly higher expression levels of DUOX2 and DMBT1 (AUC 0.985 and 0.896, respectively) and lower expression of CYP2B7P, PITX2, and DEFB1 (AUC 0.966, 0.968, and 0.966, respectively) compared to samples from healthy patients. These genes were also found to be associated with infiltration of regulatory T cells, CD8 T cells, activated and resting memory CD4 T cells, activated natural killer cells, neutrophils, activated and resting mast cells, activated and resting dendritic cells, and M0, M1, and M2 macrophages. 25 Using RF to identify 54 feature genes from expression profiles of 55 UC and 35 healthy patients, Chen et al. constructed a LASSO regression model to screen for diagnostic markers of UC. The model performed well in the training set but when validated in an external data set, model performance was not found to be clinically useful (AUC = 0.650). 27
What could AI add in the future?
Current studies have sought to identify novel biomarkers for the diagnosis of UC. Patients with UC typically do not experience significant diagnostic delay, and there is no meaningful clinical action a gastroenterologist could take even if a high-risk patient was identified prior to developing overt symptoms. Therefore, the current approach has limited clinical utility and is more likely to have an impact by aiding our understanding of disease pathogenesis.
Computer vision in index colonoscopy
What is already known?
Endoscopic and histological evaluation via index colonoscopy is the gold standard for confirming UC diagnosis, and it is frequently analyzed in collaboration with clinical symptoms, laboratory, and radiological findings. 35 However, endoscopic scoring is inherently subjective despite attempts to create consistent scoring systems, leading to observed high rates of inter- and intra-observer variability and general lack of widespread use among endoscopists.10,35 AI techniques such as ML and computer vision are promising in the creation of an objective approach to analyzing endoscopic and histological data for early and accurate diagnosis of UC at index colonoscopy. 10 The studies included in this section have different applications. Some studies focused on automated scoring of disease severity and others focused on distinguishing UC and CD.
For included studies exploring computer vision and ML for evaluation of endoscopic data, data modalities included imaging and endoscopic datasets (n = 4 studies), combined endoscopic and histological datasets (n = 1 study), electronic health data (n = 1 study), metagenomics (n = 1 study), and metabolomics (n = 1 study).
What do current studies show?
Using a CNN to clean and extract abnormality features, Gottlieb et al. trained a recurrent neural network to predict UC severity in a prospective cohort study using 795 full-length endoscopy videos (19.5 million image frames) from 249 patients enrolled in a phase II trial of mirikizumab. The model’s predictions agreed strongly with endoscopic scoring by centralized human readers demonstrated by quadratic weighted kappa score of 0.844 (95% confidence interval (CI): 0.787–0.901) for endoscopic Mayo score and 0.855 (95% CI: 0.80–0.91) for the UC endoscopic index of severity (UCEIS). Notably, the performance metrics met or exceeded those previously published for endoscopic Mayo score and the UCEIS scores. 9
In one cross-sectional study, Sutton et al. used DL CNNs to discriminate between UC and non-UC pathologies with high accuracy when compared against review by consensus labeled data from a single gastroenterologist and three medical trainees. The initial diagnostic classification model based on 2643 pathological endoscopy images was only able to make predictions of majority class with 72.02% accuracy, compared to the final diagnostic model for grading endoscopic severity of UC, which had prediction accuracy of 87.50%. The final model was based on 851 images from diagnostic colonoscopies with endoscopic Mayo scores of 0–3 and had stronger overall performance with AUC of 0.90. 10 In a separate cross-sectional study, Chierici et al. applied a prototype DL framework based on ResNet architectures merged by ensemble learning to 14,226 three-channel RGB (red, green, blue) endoscopic images of 11,404 IBD (4388 UC, 5949 CD, and 1067 other IBD) and 2822 healthy patients to identify disease patterns and distinguish endoscopic images of UC (Matthews Correlation Coefficient = 0.931) from healthy patients. 11
In a cross-sectional study of endoscopic and histological data from 287 pediatric patients (178 CD, 80 UC, 29 IBD unclassified) at time of diagnosis, Mossotto et al. identified four new subgroups of disease based on colonic disease using unsupervised PCA and multidimensional scaling. They then applied supervised linear SVM with RFE fivefold cross-validation to construct a model to discriminate UC from CD with 82.7% accuracy (AUC 0.87) based on a combination of histological and endoscopic data when compared against physicians. While the model performed well overall, this still falls below the requirement needed for clinical application. Notably, this combined model outperformed models that relied on either endoscopic or histological data alone in terms of accuracy (71% and 76.9%, respectively) and AUC (0.78 and 0.82, respectively). However, even the optimized model was able to identify Crohn’s disease more precisely versus UC. 28
What could AI add in the future?
The is considerable inter- and intra-observer variability in endoscopic scoring in UC, and rates of agreement with agreement for the endoscopic Mayo score and UCEIS reported to be as low as 0.58. Computer vision offers a promising avenue for recording disease severity, allowing for standardized scoring between different providers and institutions. It may also serve as an alternative to centralized reading for IBD clinical trials, potentially allowing for significant cost savings.
Predicting response to medical therapy
What is already known?
Rational selection of therapy in UC is an area of great promise and interest. Investigators have applied ML techniques to omics and clinical data in order to develop models that can accurately predict response to therapy a priori with varying degrees of success. Prediction of response to medical therapy in UC has focused on prediction of response to anti-tumor necrosis factor (TNF) therapy (n = 7), though some studies exist for thiopurines (n = 1) and anti-integrin therapy (n = 3). Overall, these efforts have been limited by small datasets and study quality is variable. A key limitation of all the studies discussed is that they do not definitively show that the models predict response to a specific drug; for example, a model predicting response to anti-integrin may not in fact identify features that specifically are associated with response to anti-integrin therapy, and rather as associated with response to any form of IBD therapy. The studies are summarized in Table 2.
Predicting response to medical therapy.
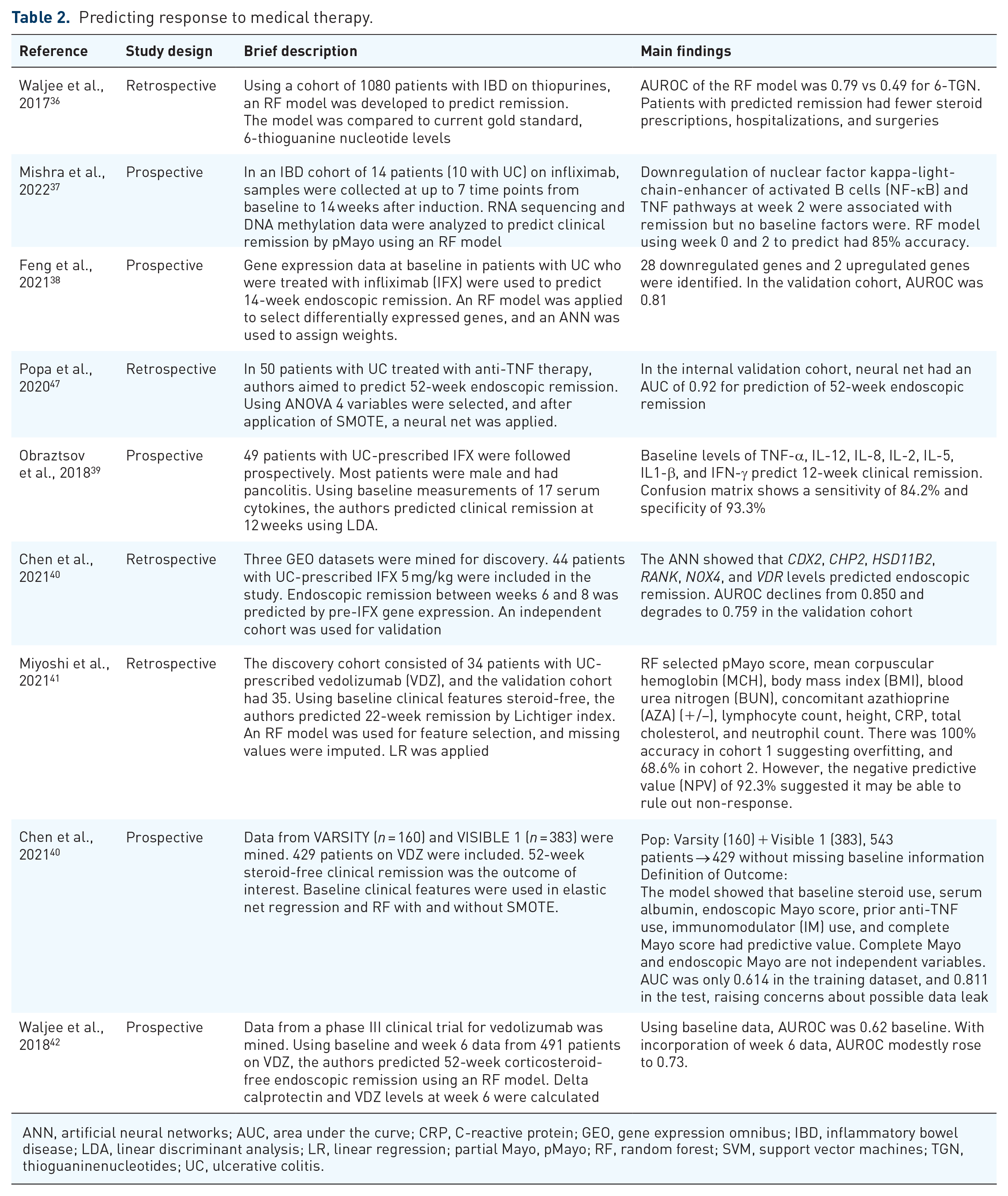
ANN, artificial neural networks; AUC, area under the curve; CRP, C-reactive protein; GEO, gene expression omnibus; IBD, inflammatory bowel disease; LDA, linear discriminant analysis; LR, linear regression; partial Mayo, pMayo; RF, random forest; SVM, support vector machines; TGN, thioguaninenucleotides; UC, ulcerative colitis.
What do current studies show?
Thiopurines
Thiopurines are antimetabolite drugs that function as immunomodulators. 6-Mercaptopurine and its prodrug azathioprine, are enzymatically metabolized to 6-thioguaninenucleotides (6-TGN) which reduces gut inflammation. Thiopurines have a narrow therapeutic window, and traditionally therapeutic drug monitoring (TDM) has been utilized. 43 However, two small, albeit underpowered, randomized controlled trials failed to show a clinical difference between TDM-guided and weight-based dosing regimens.44,45 Up to half of patients on thiopurines discontinue treatment within 2 years of initiation due to either adverse event or failure of therapy. 43 To address these shortcomings a ML algorithm that could predict clinical remission was developed. A RF model was trained using approximately 1000 patients, and the authors showed that the model was superior to 6-TGN in predicting remission (AUROC 0.79 vs 0.49); patients with algorithm-predicted remission had lower rates of steroid prescription, hospitalization, and surgery. 36
Anti-tumor necrosis alpha therapy
Most studies that applied ML techniques to UC have focused on omics data (n = 5), rather than clinical data. These studies are promising and provide key insights into the underlying mechanisms of disease but are far from being clinically applicable. Only two studies utilized clinical data that is readily available to the practicing clinician. The studies are summarized in Table 2.
In a study by Mishra et al., the authors aimed to predict clinical remission by partial Mayo scores at week 14 using whole blood samples to obtain RNA sequencing and DNA methylation data. Data was obtained from a discovery cohort of 14 patients; all but 1 patient was prescribed infliximab. They applied an RF model using data obtained at baseline and 2 weeks after induction. Downregulation of NF-κB and TLF signaling at week 2 predicted response to therapy (accuracy 85%), but no baseline findings accurately predicted response. 37 Feng et al. used colonic mucosal gene expression, from gene expression omnibus (GEO) datasets, at baseline to predict endoscopic remission at week 14. They utilized RF for feature selection and then applied an ANN to assign weights to the DEGs. They tested in a separate cohort with an AUC of 0.81. The datasets were small and may have contributed to large uncertainty in the test set. 38 Obraztsov et al. evaluated 49 patients with UC treated with IFX; most patients were male and had pancolitis. They used a pre-specified 17 cytokine panel to predict clinical remission at 12 weeks by using baseline data. Applying LDA, they showed that TNF-α, IL-12, IL-8, IL-2, IL-5, IL1-β, and IFN-γ levels predicted remission. The confusion matrix showed a sensitivity of 84.2% and a specificity of 93.3%. 39 Finally, Chen et al. mined three GEO datasets for discovery, utilizing only patients receiving 5 mg/kg of infliximab. Baseline mucosal gene expression prior to IFX infusion was used to predict 8-week endoscopic remission. Given the small datasets, synthetic bootstrapping was used and then an ANN was applied. A model incorporating CDX2, CHP2, HSD11B2, RANK, NOX4, and VDR levels was shown to have an AUC of 0.850, but AUC declined to 0.759 in an independent cohort. 40
Two studies aimed to predict response to anti-TNF therapy with clinical variables. Xiojun et al. used a heterogeneous group of 420 patients with UC on a variety of therapies including aminosalicylates, thiopurines, and biologics. They used demographics, laboratory measurements, and medicines to predict endoscopic remission, but the time points for the input data were not clearly defined. They used inferential analysis for feature selection, and then applied multiple models including LR, RF, and SVM to the data; to address under-sampling of patients in remission and those with mild disease SMOTE was utilized. The final model had an AUC of 0.80. 46 A second study using clinical data by Popa et al. used baseline data from a cohort of 50 patients with UC to predict endoscopic remission at 52 weeks. Feature selection was done with ANOVA and the final model incorporated four variables: neutrophil count, platelet distribution width, CRP, and alpha-1-globulins. SMOTE and cross-validation were performed to reduce overfitting and imbalance data with a small dataset as much as possible. The model had an AUC of 0.92 in a validation dataset of five patients from the same center. 47
Overall, there are multiple promising models for assessing response to anti-TNF therapy, but overall any clinical application is currently limited either by use of data that is not widely available or derivation from small samples that have unclear external validity.
Anti-integrin therapy
Studies predicting response to anti-integrin therapy are less numerous than those predicting response to anti-TNF therapy. A study by Miyoshi et al. utilized baseline demographic, IBD, laboratory, and prescription data to predict 22-week remission by Lichtiger index. Data was trained on 34 patients at a single hospital and tested on 35 patients at a different institution. Missing data was imputed, carrying a risk of bias given the small sample size and retrospective data. RF was used for feature selection, and ultimately eight features were included in the LR model (MCH, BMI, BUN, Concomitant AZA use, lymphocyte count, height, CRP, total cholesterol, and neutrophil count). The model only had 68.6% accuracy in the validation cohort suggesting overfitting but had an NPV of 92.3%. It may be valuable to rule out non-response. 41 Chen et al. combined data from VARSITY and VISIBLE 1, resulting in a dataset of 429 patients. Fifty-two-week steroid-free remission by Mayo score was the outcome of interest, and baseline clinical features were used as the predictors. Elastic net regression was compared with RF, with and without SMOTE on a 75:25 split dataset. Baseline steroid use, albumin, endoscopic Mayo score, prior anti-TNF use, IM use, and complete Mayo score were included; notably complete Mayo is not an independent predictor from endoscopic Mayo. AUC was 0.614 in the training set, and 0.811 in the test set raising the possibility of data leak given the unexpected increase in performance. 48 Finally, Waljee et al. used baseline and week 6 clinical data from a phase III clinical trial to predict 52-week corticosteroid-free endoscopic remission. An RF model was used with a 70:30 split of data. AUC using only baseline data was 0.62, and AUC with addition of week 6 data was 0.73. 42 Overall, no model had adequate predictive characteristics to inform clinical decision making at this time.
What could AI add in the future?
Rational selection of IBD therapy is both an area of research interest and significant clinical need. Current models suffer from significant limitations as noted above. AI, likely in conjunction with fundamental advances in basic science, has the potential to bring the era of precision medicine to patients with IBD by providing true class-specific predictions on the likelihood of response to medications.
Monitoring disease activity in patients with established UC
Endoscopy, histologic assessment, and laboratory testing play important roles in the surveillance of UC. There have been numerous AI and ML applications aimed toward evaluating endoscopic lesions, predicting histological indices to grade severity of UC activity, and identifying biomarkers of active disease. These studies are summarized in Table 3.
Monitoring disease activity in patients with established ulcerative colitis.

AI, artificial intelligence; ASUC, acute severe ulcerative colitis; AUC, area under the curve; CAD, computer-assisted diagnosis; CDS, cumulative disease score; CNN, convolutional neural networks; GEO, gene expression omnibus; IBD, inflammatory bowel disease; ICC, intraclass correlation coefficient; MES, Mayo endoscopic score; ML, machine learning; PHRI, PICaSSO Histologic Remission Index; RD, red density; UC, ulcerative colitis; UCEIS, UC endoscopic index of severity; VCE, virtual chromoendoscopy; WLE, white-light endoscopy.
AI applications in endoscopic monitoring
What is already known?
Endoscopy is a cornerstone of assessing UC disease activity, and several endoscopic scores have been developed to define disease activity. The most commonly used and extensively validated endoscopic scores include the Mayo endoscopic score (MES), the UCEIS, and the UC colonoscopic index of severity. 72 However, these endoscopic scores are limited by their qualitative nature, subjectivity, and corresponding interobserver variability. Further, these scores typically report the maximum severity observed and fail to capture the heterogeneity of disease and total disease burden. Thus, AI applications in UC endoscopy have focused on identifying signs of inflammation on endoscopy as well as standardizing the interpretability of endoscopic findings in UC disease surveillance.
What do current studies show?
An initial study by Ozawa et al. constructed a computer-assisted diagnosis (CAD) system using a CNN that accurately identified endoscopic disease remission from still images. 49 A subsequent study by Stidham et al. 50 employed a CNN constructed as a DL model to differentiate endoscopic remission (MES 0 or 1) from moderate-to-severe disease (MES 2 or 3) from still images (AUC 0.966). Both studies are significantly limited due to their applications to still images, which does not represent typical clinical practice in which decisions are made based off of the entire colonoscopy. However, neural networks have since been applied to analysis of full-length endoscopic video data and have been demonstrated to reliably predict endoscopic disease severity with reasonably high rates of agreement with expert reviewers.9,51–53 In one prospective study, AI-assisted colonoscopy was able to stratify patients with UC in clinical remission into higher and lower risk groups for clinical relapse of UC, evidencing the prognostic potential of AI-assisted endoscopy to predict clinical outcomes and accordingly influence disease management. 54 A recent publication by Stidham et al. showed that computer vision could be used to calculate a cumulative disease score (CDS) by assigning MES to all frames of adequate quality for a given colonoscopy that were mapped to an estimated location; CDS was defined as the sum of MES-squared values. The authors showed that CDS was more sensitive than MES for detecting change; CDS required 50% fewer participants to demonstrate a difference in the endoscopic outcome, a finding with clear cost implications for clinical trials. 69
What could AI add in the future?
Computer vision may augment the capabilities of general gastroenterologists, allowing them to perform at a similar level as IBD specialists. Future studies will be needed to evaluate if AI assessment will obviate the need for virtual or dye-based chromoendoscopy. Use of tools such as the CDS may be able to increase power and decrease cost for clinical trials in IBD.
AI applications in histology assessment
What is already known?
In addition to assessing endoscopic outcomes in UC, AI has also been applied to histological evaluation. Histological signs of inflammation, even in the absence of endoscopic inflammation, have been associated with adverse clinical outcomes in UC, making histologic remission an important adjunct goal of UC treatment.73–75 To this end, there have been widespread efforts to apply AI and ML techniques to predict histologic activity in patients with UC.
What do current studies show?
In an initial study, Takenaka et al. 55 constructed a deep neural network using colonoscopy images and biopsy results from a cohort of patients with UC, which was then able to predict endoscopic remission with 90.1% accuracy as well as histologic remission with 92.9% accuracy in the validation cohort. In a follow-up study using a prospective cohort of 875 patients, mucosal healing predicted by the deep neural network algorithm based on endoscopic and histologic remission was shown to be correlated with significantly lower risk of hospitalization, colectomy, steroid use, and clinical relapse. 76 In another study, Maeda et al. 56 developed a CAD system to predict histologic inflammation using endocytoscopy with a sensitivity of 74%, specificity of 97%, and accuracy of 91% when compared to pathologist interpretation of corresponding biopsies. Najdawi et al. developed a CNN which was compared against the Nancy index. The CNN showed strong correlation with pathologist-determined Nancy index (r = 0.89) and was highly accurate at determining histologic remission (accuracy 97%). Notably, the CNN was only designed to assess disease activity and was unable to identify other clinically important features including signs of infection or dysplasia. 70 Iacucci et al. also developed a CNN which was compared against Nancy index, PICaSSO Histologic Remission Index, and Robarts. The CNN when compared against these three indices had sensitivities ranging from 89% to 94% and specificities ranging from 76% to 85%. This algorithm was also unable to identify other clinically important features including infection and dysplasia. 71
Other studies have incorporated additional endoscopy features, such as red density lighting and virtual chromoendoscopy into ML algorithms to predict measures of endoscopic and histologic inflammation.57,58 Models to assess histologic inflammation have also been developed from direct analysis of biopsies themselves, using image processing techniques to detect eosinophils, neutrophils, and other histologic features.59–61 These models have consistently demonstrated a high degree of agreement with scoring by independent experts.
What could AI add in the future?
There are a variety of scoring systems for pathology in IBD, but these scoring methods are not standardized across institutions and there are issues with interobserver variability which may be addressed with AI. Further, with continued improvement, these tools may expand access to specialist care by enabling general pathologists to evaluate IBD specimens at a similar level to GI pathologists at referral centers.
AI applications in novel biomarker discovery
What is already known?
ML techniques have also been applied toward multi-omics data sets including genetic, transcriptional, and microbiome data to identify novel biomarkers of UC disease activity. Most of these datasets have a very high number of predictors (omics output) derived from a small cohort of patients; this problem, called “big-p, little-n,” causes significant issues for prediction, and require specialized data preparation and proper algorithms to properly handle in the input.
What do the current studies show?
In a study by Morilla et al., 62 microarray data was utilized to build a deep neural network-based classifier consisting of nine miRNAs and five clinical factors that accurately discriminated patients with acute severe UC as responders versus non-responders to steroids (accuracy 93%, AUC = 0.91), infliximab (accuracy 84%, AUC = 0.82), and cyclosporine (accuracy 80%, AUC = 0.79). In another study by He et al., 63 ML algorithms were used to identify differentially expressed mRNAs to serve as diagnostic biomarkers for UC, which were then further validated in cell lines and mouse models of colitis. Whole blood transcriptomic data has been used to develop a qPCR-based classifier that stratified patients into high and low-risk groups associated with earlier need for treatment escalation (hazard ratio 3.12) and more escalations over time in UC patients. 64 Notably, endoscopic severity at baseline did not predict need for treatment escalation in this cohort, highlighting the ability of ML algorithms to impact treatment decisions in ways that would be undetectable by conventional methods of disease surveillance. Transcriptomic data has also been leveraged to characterize different subtypes of UC patients, which were associated with various relevant clinical features including Mayo scores, calprotectin levels, and histological severity scores. 65 Other studies have used ML techniques to investigate blood-based, genome-wide association studies, and microbiome biomarkers related to UC diagnosis, phenotypes, and disease severity.66–68 Overall, the application of ML techniques to biomarker discovery in UC has revealed promising biomarkers related to diagnosis, disease subtyping, and prognostication. Further testing is required to determine the clinical translatability of these biomarkers.
What could AI add in the future?
Even though calprotectin is a reliable biomarker, between 5% and 10% of patients have discordant results when compared with colonoscopy. Application of AI for the identification of biomarkers for more reliable non-invasive clinical monitoring would be extremely clinically valuable. Further, while any individual biomarker is unlikely to rival the diagnostic accuracy of colonoscopy, an AI tool that can combine multiple biomarkers may be able to provide similar accuracy.
Predicting complications of UC
The literature regarding applications of AI to complications of UC is limited. There were four broad areas that the literature focused on: predicting the need for colectomy (n = 2), predicting postoperative complications (n = 2), predicting colorectal cancer (CRC) (n = 2), and prediction of COVID-19 outcomes (n = 1). These studies are summarized in Table 4.
Predicting complications of ulcerative colitis.
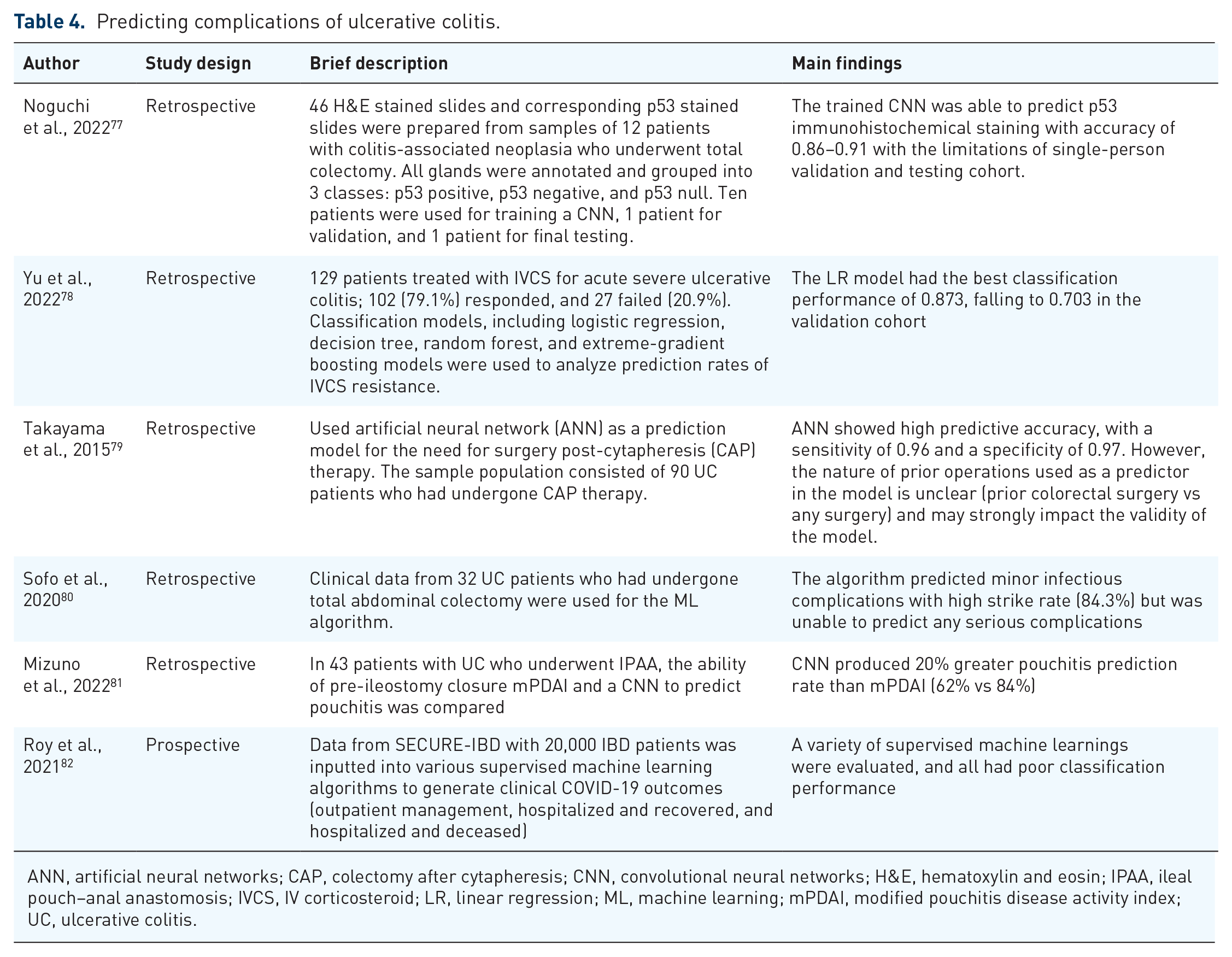
ANN, artificial neural networks; CAP, colectomy after cytapheresis; CNN, convolutional neural networks; H&E, hematoxylin and eosin; IPAA, ileal pouch–anal anastomosis; IVCS, IV corticosteroid; LR, linear regression; ML, machine learning; mPDAI, modified pouchitis disease activity index; UC, ulcerative colitis.
Predicting the need for colectomy
What is already known?
Colectomy is used to treat medically refractory acute severe UC. We know that approximately 10%–15% of patients with UC will undergo colectomy during their lifetime. While traditional epidemiologic methods have found risk factors associated with colectomy, these models are unable to predict risk for a given patient.
What do current studies show?
Two studies aimed to construct and validate models that could predict post-treatment complications that require follow-up treatment. In one study by Yu et al., traditional LR models and ML models were compared as predictive models for IV corticosteroid (IVCS) resistance in patients with acute severe ulcerative colitis (ASUC). UCEIS and CRP level at day 3 of IVCS therapy were independent predictors of IVCS response. No ML method was able to outperform traditional LR (AUC of 0.703 in the validation cohort). The study was limited by small sample size from a single patient population and the relatively poor classification performance of the algorithms. 78 In a second study by Takayama et al., an ANN was utilized to predict the need for colectomy after cytapheresis (CAP) therapy based on 13 input factors using a training data set (n = 54) and validation data set (n = 36). The prediction model identified four key factors: history of prior admissions, prior operations, use of immunomodulators, and response to CAP therapy. The model had a sensitivity of 0.96 and specificity of 0.97. The nature of prior operations used as a predictor in the model are unclear (prior colorectal surgery vs any surgery) and may strongly impact the validity of the model. 79
What could AI add in the future?
Predicting which patients with ASUC will require colectomy is an area of clinical need. Patients who fail IVCSs are often given rescue therapy, typically infliximab or cyclosporine. By identifying patients who are not likely to respond to medical therapy, algorithms may help clinicians avoid unnecessary immunosuppression prior to surgery. A significant barrier to AI for this application is the relative rarity of ASUC, and the lack of large databases for training models for this end use.
Prediction of postoperative complications
What is already known?
Patients with acute severe UC or treatment-refractory UC often undergo surgery. These patients have a high risk of post-surgical complications, and there is a clear clinical benefit of being able to predict surgical outcomes. In particular, pouchitis is a vexing complication that can lead to persistence of symptoms and impaired quality of life after colectomy. Two studies applied AI methodology to this clinical problem.
What do the current studies show?
In one study by Mizuno et al., the researchers aimed to determine whether a CNN could accurately predict pouchitis development after ileal pouch–anal anastomosis (IPAA) in UC patients. Modified pouchitis disease activity index (mPDAI) before ileostomy closure was compared with a CNN model based on the endoscopic imaging of a retrospective cohort of 43 patients with 5-fold cross-validation. The predictive rates for pouchitis of mPDAI prior to ileostomy closure and the CNN model were estimated by ROC. mPDAI had an accuracy of 62% and the CNN had an accuracy of 84%. Limitations include a small number of images and variation in image scoring due to use of different endoscopists, which could be overcome in the future with a multicenter design and standardization of imaging methods. Nevertheless, the findings suggest that CNN models may predict pouchitis, and allow for early intervention. 81 In a second study, Sofo et al. looked at a cohort of high-risk UC patients who had undergone a total colectomy and aimed to predict various types of postoperative complications using data available before surgery. This simulated a prospective study and was able to predict minor infectious complications accurately, but major infection and non-infectious complications were not predicted as accurately, greatly limiting the clinical utility of this study. 80
What could AI add in the future?
Pouchitis is a common complication after IPAA. While many patients respond to a single course of antibiotics, a subset develops chronic pouchitis, a devastating complication that affects quality of life after a theoretically curative colectomy. Being able to predict complications like pouchitis may help surgical planning.
Colorectal cancer surveillance
What is already known?
Although patients with long-standing UC have a higher risk of developing CRC and there is significant literature regarding CAD in general, there is limited data on the application of AI to CRC risk in UC patients. 83
What do the current studies show?
One study by Uttam et al. aimed to aid early detection by applying three-dimensional nanoscale nuclear architecture mapping to detect advanced dysplasia or neoplasia in normal-appearing rectal biopsies of patients with both UC and Crohn’s disease prior to detection by conventional history. They applied SVM as a binary classifier and the final model had an AUROC of 0.870. 84 Noguchi et al. 77 used a CNN to predict p53 immunohistochemical staining from hematoxylin and eosin stained slides without dedicated p53 stains. The trained CNN was able to predict p53 immunohistochemical staining with accuracy of 86%–91%. Although the results are promising, the study did not validate the CNN in an external dataset, and the sample size was small with only 12 patients, with strong risk of overfitting.
What could AI add in the future?
Further studies should incorporate external validation and larger sample sizes in order to develop strong predictive models for colitis-associated dysplasia, and biomarkers aside from p53 should be investigated. 77 Surveillance in patients with dense pseudopolyposis is technically challenging and represents an area in which computer vision may prove to be useful.
COVID-19 outcomes
UC is often treated with immunosuppressants, which may lead to higher risk of infection. The outcome of COVID-19 in UC patients is of significant interest and there have been numerous studies which have applied traditional epidemiologic methods. However, there is a paucity of studies which have applied AI methods to this patient population. A single study by Roy et al. 82 addresses this issue by using the SECURE-IBD database. They applied a variety of and supervised learning methods, but the best performing model only had an accuracy of only 70%.
Conclusion
AI shows great promise in UC, and there has been burgeoning interest in the field. ML and DL techniques have been applied to a wide range of meaningful clinical problems in UC, including the identification of new UC, personalized therapy, monitoring of disease activity, and prediction of complications. Despite the considerable promise of AI in UC, there are also key limitations; many studies have small sample sizes and biases that risk overfitting. There have been limited validation of studies in truly independent external datasets. On the whole, rarely have the developed models had adequate performance characteristics to justify potential clinical deployment. Given the current status of the field of AI in UC, future research should include: (1) robust, large scale external validation of models to overcome the many limitations and bias that come with using small internal training datasets, (2) studies that predict clinically meaningful outcomes that are in line with current standard of care, such as endoscopic remission rather than clinical remission, (3) studies that evaluate the cost-effectiveness of model-guided therapy compared to the current standard of care, (4) head-to-head studies of models which predict the same outcomes to guide clinical implementation, and (5) randomized controlled trials of AI models to determine if they meaningfully impact clinical outcomes.
