Abstract
Background:
Notably, 5-aminosalicylates (5-ASA) are vital in treating inflammatory bowel diseases (IBD). The adverse events of 5-ASA rarely occur but they could be fatal.
Objectives:
We aimed to discover new genetic biomarkers predicting 5-ASA-induced adverse events in patients with IBD.
Design:
This was a retrospective observational study.
Methods:
We performed a genome-wide association study on patients with IBD in South Korea. We defined subset 1 as 39 all adverse events and 272 controls; subset 2 as 20 severe adverse events and 291 controls (mild adverse events and control); subset 3 as 20 severe adverse events and 272 controls; and subset 4 as 19 mild adverse events and 272 controls. Logistic regression analysis was performed and commonly found associated genes were determined as candidate single-nucleotide polymorphisms predicting 5-ASA adverse events.
Results:
Patients with Crohn’s disease (CD) were significantly negatively associated with the development of adverse events compared to patients with ulcerative colitis (UC) (5.3% versus 22.9%). However, sex and age at diagnosis were unassociated with the adverse events of 5-ASA. rs13898676 [odds ratio (OR), 20.33; 95% confidence interval (CI), 5.69–72.67; p = 3.57 × e−6], rs12681590 (OR, 7.35; 95% CI, 2.85–19.00; p = 3.78 × e−5), rs10967320 (OR, 4.51; 95% CI, 2.18–9.31; p = 4.72 × e−5), and rs78726924 (OR, 3.54; 95% CI, 1.69–7.40; p = 7.96 × e−5) were genetic biomarkers predicting 5-ASA-induced severe adverse events in patients with IBD.
Conclusion:
The adverse events of 5-ASA were more common in patients with UC than those with CD in our study. We found that novel rs13898676 nearby WSB2 was the most significant genetic locus contributing to 5-ASA’s adverse event risk.
Introduction
Inflammatory bowel diseases (IBD), including ulcerative colitis (UC), Crohn’s disease (CD), and intestinal Behçet’s disease (BD), are chronic refractory inflammatory disorders of the gastrointestinal tract. The crude prevalence of IBD is globally increasing. The treatment options for IBD have accordingly expanded.1,2 Nevertheless, 5-aminosalicylates (5-ASA) are still considered essential for the induction and maintenance of clinical remission in UC patients.3–5 The role of 5-ASA in treating UC is clear but not in CD. However, the 5-ASA is still widely used in routine clinical practice.6–8 In a recent network meta-analysis of various drugs used in treating CD, it was demonstrated that 5-ASA at a dose of 2.4 g or more was more effective than the placebo group; however, it was less effective than steroids or budesonide in patients with mild to moderate CD. 9 Regarding intestinal BD, 5-ASA is recommended for inducing and maintaining clinical remission. 10
In the era of many treatment options for patients with IBD, 5-ASA is a relatively safe and well-tolerable drug during long-term use. 11 In the recent Cochrane review, the incidence of overall pooled adverse events was not statistically different between 5-ASA and placebo.12,13 The most common adverse events of 5-ASA are mild, such as diarrhea, abdominal pain, nausea, headache, myalgia, and skin lesion, and occur in 5–10% of patients of IBD at 1–3 weeks after the start of 5-ASA14,15 However, rare, life-threatening severe adverse events, such as fever, liver toxicity, pancreatitis, interstitial pneumonitis, myocarditis, pericarditis, hemolytic anemia, and pancreatitis, can also occur. Ransford et al. 16 reported that three pancreatitis cases, three blood dyscrasias, and one hepatitis were issued in 2.8 million prescriptions for mesalamine in patients with IBD. Severe adverse events are rare in patients with IBD treated with 5-ASA; however, they can be fatal. The discovery of biomarkers could predict serious 5-ASA adverse events and allow appropriate patient selection without a fatal disease course.
The discovery of genetic biomarkers predicting thiopurine-induced leucopenia contributed to decreasing unpredictable treatment discontinuation and occasional morbidity. In Western countries, thiopurine S-methyltransferase (TPMT) gene polymorphism is prevalent, and its direct correlation with hematological toxicity of thiopurine has been widely demonstrated, which personalized thiopurine dosing based on TPMT genotyping as standard care for patients with IBD. 17 Despite the lower TPMT polymorphism, leucopenia is more frequent in East Asian patients with IBD. 18 Later, the nudix hydrolase 15 (NUDT15) gene polymorphisms were discovered to predict thiopurine-induced leucopenia, and this genotype-based treatment of thiopurines has been established in East Asian countries.19,20
Therefore, we aimed to discover new genetic variants predicting severe 5-ASA-induced adverse events in patients with IBD in South Korea using a genomic-wide association study (GWAS).
Materials and methods
Study population
We collected blood samples from patients diagnosed with UC, CD, or intestinal BD during routine outpatient visits at the IBD Center, Severance Hospital, between 2012 and 2022. We pooled 401 patients with IBD from two cohorts, 1 from 40 patients who experienced severe adverse events of 5-ASA (cohort 1) and the other from 361 patients treated with biologic agents (cohort 2). Notably, all patients in cohort 2 had been treated with 5-ASA. In all, 47 patients were excluded because of incomplete electronic medical records, changes in initial diagnosis, or only a single visit. In all, 18 patients who did not experience 5-ASA therapy were also excluded. GWAS analysis was performed on the final 311 samples, with 25 samples removed from the quality control (QC) step. The study is reported according to the Strengthening the reporting of observational studies in epidemiology (STROBE) guidelines. 21
Adverse events of 5-ASA
Clinical adverse events of 5-ASA were defined as cases judged by treating physicians as symptoms caused by the 5-ASA within 1 month of initiating therapy and when symptoms disappeared after discontinuation. 22 We included diarrhea, abdominal pain, nausea, headache, myalgia, skin lesions, fever, liver toxicity, pancreatitis, interstitial pneumonitis, and pancreatitis. We also classified mild adverse events as diarrhea, abdominal pain, nausea, headache, myalgia, skin lesions, and severe adverse events such as fever, liver toxicity, interstitial pneumonitis, myocarditis, pericarditis, hemolytic anemia, and pancreatitis. 15
DNA extraction and genotyping
Genomic DNA was extracted from whole blood samples using Qiagen’s DNA Blood Maxi Kit (Santa Clarita, CA, USA). Quant-iT™ PicoGreen® dsDNA Reagent and Kits (Invitrogen) and Nanodrop (ND-1000 Spectrophotometer, Thermo Scientific, Wilmington, USA) were chosen to measure the quality and quantity of the DNA. A total of 336 samples were genotyped using Infinium™ Global Screening Array-24 beadchip (v3.0) (Illumina, San Diego, CA, USA), which contains 654,027 single-nucleotide polymorphism (SNP) markers, according to Illumina’s protocols. Based on the initial genotype data, we further removed samples and markers using PLINK (v 2.0) software (Complete Genomics, USA). 23 Before genotype calling, samples with low call rate and low 10% GenCall (GC) score that outliers from the main population were excluded. After genotype calling, samples with a call rate <95% and samples with an outlying heterozygosity rate >3 standard deviations (SDs) were excluded. Hardy–Weinberg equilibrium further removed the SNPs (p < 1 × e−4) and minor allele frequency <0.01. A quantile–quantile (Q–Q) plot analysis was conducted to verify whether the distribution of the inflation-corrected p values deviated from the expected distribution under the null hypothesis of no genetic association. After QC control, 364,441 SNPs and 311 samples with IBD were considered available for further analysis.
Genome-wide association analyses
We performed GWAS of the four subsets separately (Figure 1). Subset 1 had 39 adverse event cases (Casemild + Casesevere) and 272 controls (Control); subset 2 had 20 severe adverse event cases (Casesevere) and 291 controls (Casemild + Control); subset 3 had 20 severe adverse event cases (Casesevere) and 272 controls (Control); and subset 4 had 19 mild adverse event cases (Casemild) and 272 controls (Control). We performed a logistic regression analysis of the discovery GWASs with three subsets using gender and diagnosis to identify susceptibility SNPs for adverse events of 5-ASA. Candidate variants were defined with p < 1 × e−3. To investigate the possible functions of the recently discovered SNPs, we conducted functional annotations using 3DSNP (database linking noncoding SNP) (https://www.omic.tech/3dsnpv2/). 24
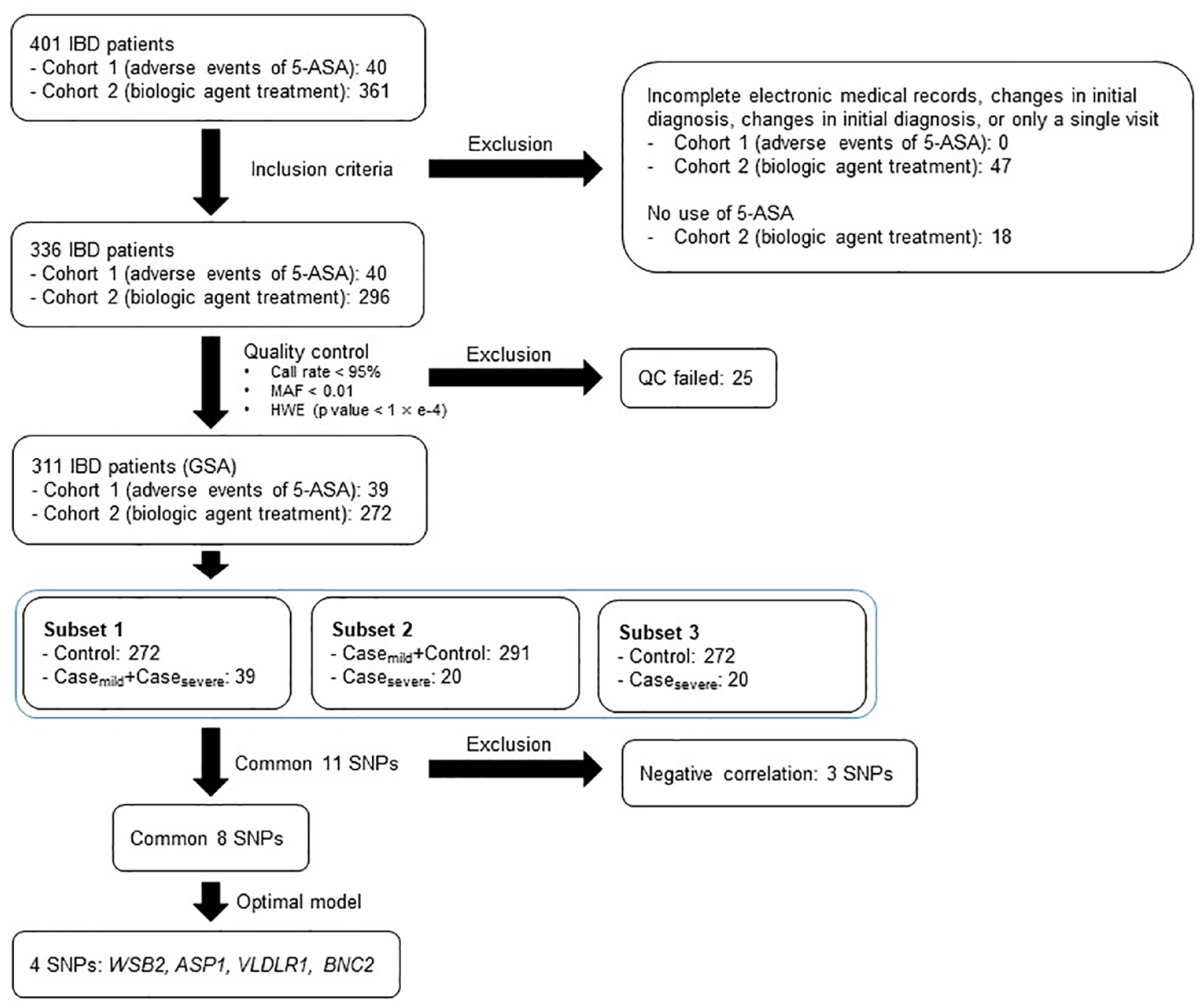
Steps of genome-wide association study. Before genotype calling, samples with low call rate and low 10% GC score that outliers from the main population were excluded. After genotype calling, samples with a call rate <95% and samples with an outlying heterozygosity rate >3 SDs were excluded. HWE further removed SNPs (p < 1 × e−4) and MAF < 0.01. We performed GWAS of the three subsets separately. Subset 1 has 39 adverse event cases (Casemild + Casesevere) and 272 controls (Control); subset 2 has 20 severe adverse event cases (Casesevere) and 291 controls (Casemild + Control); and subset 3 has 20 severe adverse event cases (Casesevere) and 272 controls (Control).
Statistical analysis
Means and SDs, or medians and ranges, were calculated for all continuous variables, as appropriate. Categorical variables were expressed as percentages and statistical analyses were performed to compare the groups of variables. Logistic regression analysis was performed, adjusting for various confounders with PLINK (v 2.0) software to identify the susceptibility SNPs for adverse events of 5-ASA. Statistical significance was set at a p value of <1 × e−3. The Bonferroni correction was applied to adjust for multiple comparisons. We used Locus Zoom to generate the regional plot, and Manhattan plots were generated using R software (3.4.0) (R Development Core Team). 25
Results
Patient characteristics
The mean patient age was 28.5 years, and 207 (66.5%) were male. The mean body mass index was 22.3 kg/m; 17 patients (5.5%) had a smoking history, and 61 patients (19.6%) had abdominal surgery history. There were 122 UC patients (39.2%), 188 CD patients (60.5%), and 1 intestinal BD patient (0.3%). At diagnosis, UC patients presented with the following disease extent: proctitis (8.2%), left-sided colitis (23.8%), and pancolitis (68.0%). CD patients presented with the following disease location: ileal (23.9%), colonic (6.4%), and ileocolonic (69.7%) and disease behavior: inflammatory (36.2%), stricturing (27.1%), and penetrating (36.7%). A total of 121 patients (64.4%) had perianal lesions (Table 1).
Baseline characteristics of patients.

Variables are expressed as mean ± SD or n (%).
CD, Crohn’s disease; UC, ulcerative colitis.
Adverse events of 5-ASA
A total of 49 adverse events of 5-ASA were observed in 39 patients. Of 39 patients with 5-ASA adverse events, 2 patients prescribed sulfasalazine, and 37 patients prescribed mesalamine. The most common types of adverse events were skin lesions and fever. The mild adverse events included diarrhea, abdominal pain, nausea, headache, myalgia, and skin lesions, and severe adverse events included fever, liver toxicity, pancreatitis, interstitial pneumonitis, myocarditis, pericarditis, hemolytic anemia, and pancreatitis (Table 2). CD patients were significantly less likely to develop adverse events compared to UC patients [5.3% versus 22.9%, odds ratio (OR), 0.186; 95% confidence interval (CI), 0.086–0.298; p value < 0.001]. Sex and age at diagnosis were not associated with adverse events of 5-ASA.
Adverse events of 5-aminosalicylate.

Variables are expressed as numbers of events.
New genetic biomarkers predicting adverse events of 5-ASA
A Q–Q plot analysis between genetic data and adverse events of 5-ASA with subset 1, subset 2, or subset 3 was performed to verify whether the distribution of the inflation-corrected p values deviated from the expected distribution under the null hypothesis of no genetic association (Supplemental Figure 1).
Manhattan plots were generated for subset 1, subset 2, and subset 3 (Figure 2 and Supplemental Figure 2) to offer a comprehensive overview of the disease-associated regions. We identified 172 SNPs suggesting statistical significance (p < 1 × e−3) in subset 1, 173 SNPs suggesting statistical significance in subset 2, and 169 SNPs suggesting statistical significance in subset 3. Nevertheless, due to the limited sample size, none of the SNPs retained their significance following the most conservative Bonferroni correction. We categorized adverse events into mild and severe adverse events to assess the association of SNPs based on the severity of the phenotype. Since severe phenotype is expected to have a more significant association between phenotype and genotype compared to mild phenotype, we selected 11 common SNPs, all present in subsets 1, 2, and 3, to identify possible candidate SNPs (Supplemental Table 1). Of these, three SNPs were negatively associated with adverse events of 5-ASA, and only eight were positively associated with adverse events of 5-ASA. Of the eight SNPs, only four candidate SNPs were located nearby the known genes (Table 3). rs13898676 (subset 1, p = 3.57 × e−6; subset 2, p = 8.88 × e−5; subset 3, p = 9.84 × e−5) was on chromosome 12 and located upstream of WD (also known as the Trp-Asp) repeat, and suppressors of cytokine signaling (SOCS) box containing 2 (WSB2), rs12681590 (subset 1, p = 3.78 × e−5; subset 2, p = 1.98 × e−5; subset 3, p = 1.31 × e−5) was located in the intron of ArfGAP with SH3 domain, ankyrin repeat, and PH domain 1 (ASAP1) on chromosome 8, rs10967320 (subset 1, p = 4.72 × e−5; subset 2, p = 1.40 × e−5; subset 3, p = 8.87 × e−5) was located in the intron of very low-density lipoprotein receptor (VLDLR) on chromosome 9, and rs78726924 (subset 1, p = 7.96 × e−5; subset 2, p = 1.38 × e−5; subset 3, p = 1.21 × e−5) was located in the intron of basonuclin 2 (BNC2) on chromosome 9 (Figures 2 and 3, and Table 3). Among these SNPs, rs13898676, which has not yet been published, exhibited the lowest p value and the largest OR value (subset 1: OR, 20.33; 95% CI, 5.69–72.67; p = 3.57 × e−6; subset 2: OR, 10.52; 95% CI, 2.63–42.14; p = 8.88 × e−4; subset 3: OR, 23.34; 95% CI, 4.78–113.89; p = 3.57 × e−5) independently in the subset analysis (Table 3 and Supplemental Table 1). The four SNPs were predicted to have biological functions using 3DSNP, and the 3DSNP scores were summarized in Table 3. We described the top candidate SNPs from each analysis (subsets 1, 2, and 3) in Supplemental Tables 2–4, and additionally analyzed the mild adverse events compared to the controls as subset 4 (Supplemental Table 5).

A genome-wide Manhattan plot of adverse events with 5-aminosalicylate in inflammatory bowel disease patients. Subset 1: N = 39 (Casemild + Casesevere), N = 272 (Control).
Novel genome-wide significant loci for adverse events of 5-aminosalicylate in inflammatory bowel disease patients.

Subset 1: N = 39 (Casemild + Casesevere), N = 272 (Control).
The total score of functionality for this SNP in 3DSNP.
CHR, chromosome number; CI, confidence interval; OR, odds ratio; SNP, single-nucleotide polymorphism.

Regional plots of the area around genome-wide significant and candidate SNPs. (a) rs13898676 is on chromosome 12 and located upstream of WSB2. (b) rs12681590 is in the intron of ASAP1 on chromosome 8. (c) rs10967320 is in the intron of VLDLR on chromosome 9. (d) rs78726924 is in the intron of BNC2 on chromosome 9.
Discussion
A few studies have attempted GWAS analyses to discover genes associated with 5-ASA-induced adverse events in patients with IBD. We performed the first genetic analyses predictive of mild and severe adverse events of 5-ASA. The analysis was divided into mild and severe adverse events of 5-ASA because severe adverse events are clinically more important and presumed to be more related to genetic factors. We discovered new genetic biomarkers predicting 5-ASA-induced adverse events in patients with IBD in GWAS (rs12681590, rs10967320, rs78726924, and rs13898676 near ASAP1, VLDLR, BNC2, and WSB2, respectively).
The exact mechanisms of action of 5-ASA remain unknown. However, it is believed that mesalazine reduces the synthesis of prostaglandins and leukotrienes, thereby influencing the inflammatory responses associated with the cyclooxygenase and lipooxygenase pathways.26,27 Also, 5-ASA seems to exert its effects specifically on the mucosa of the intestine. Consequently, 5-ASA-associated adverse events have not been fully classified; however, it has been reported as an allergic reaction. 15 Drug-induced lymphocyte stimulation test (DLST) could be performed to definite diagnosis of drug allergy of 5-ASA, and successful desensitization therapy of 5-ASA was reported in the previous study. 28 However, the DLST and desensitization therapy of 5-ASA are performed after the symptoms have already appeared and are not preventive practices. The early recognition and prediction of severe adverse events of 5-ASA are very important in regular clinical practice but there were few genetic studies regarding 5-ASA-induced severe adverse events. There was a GWAS study to figure out genetic biomarkers associated with 5-ASA-induced diarrhea and fever in Japanese patients with IBD, and rs144384547 in the upstream of RGS17 was a suggested novel biomarker. 22 Another study also suggested rs3135349 and rs3135356 as genetic biomarkers associated with 5-ASA-induced nephrotoxicity in UK patients with IBD. 29 However, these SNPs were not replicated in our cohort, presumably due to different ethnic groups with different definitions of adverse events and the small sample size of the individual study. To discover genetic biomarkers more specific to the hypersensitivity reaction, we included mild and severe adverse events of 5-ASA and found four candidate SNPs. Also, the adverse events of 5-ASA were more common in UC patients in our study. The reason is unclear, but if it is true that it occurs more in UC, there is a possibility that the candidate gene for causing side effects of the drugs and the gene for disease development are related.
WSB2 is one of the SOCS box-containing proteins, which contributes to the decreased cytokine signaling cascades, such as the JAK (Janus kinases)-STAT (signal transducer and activator of transcription protein) pathway. 30 In the previous study, WSB2 was related to the opacification subgroup of chronic rhinosinusitis and is known as a negative regulator of the interleukin-21 receptor, which was also associated with atopic dermatitis susceptibility. 31 Also, it was reported to be one of the downregulated genes by Saccharomyces boulardii in the 2,4,6-trinitrobenzene sulfonic acid-induced colitis. 32 Therefore, WSB2 can be speculated to contribute to a hypersensitivity reaction in the IBD treated with 5-ASA. rs138986796, a novel SNP with the highest functionality score (3DSNP score: 138.2, Table 3), is a regulatory region variant located in the promoter region of the WSB2. This promoter region contains 38 transcription factor binding sites, including RNA polymerase II subunit A (3DSNP promoter score: 100.00).
rs12681590, an intron variant of ASAP1, is located in the enhancer region of ASAP1 (3DSNP enhancer score: 26.85) and was found to be a potential candidate in the GWAS analysis. ASAP1 is associated with encoding a regulator of dendritic cell migration, and it was susceptible to tuberculosis in the previous study. Also, ASAP1 is one of the hub proteins in IBD and colorectal cancer and the only gene actively engaged in other diseases such as tuberculosis, pulmonary, and bladder neoplasm diseases. 33 In this regard, the ASAP1 gene was revealed as a key aspect of the gene–disease association. Therefore, it could be suggested as a novel genetic biomarker predicting 5-ASA-induced adverse events related to various organs in patients with IBD.
VLDLR, the closest gene to rs10967320, another gene associated with 5-ASA-induced adverse events, is a low-density lipoprotein receptor family member. In the previous study, VLDLR attenuates mite-induced airway inflammation through dendritic cell-mediated adaptive immunity. 34 The phenotype of IgE increase, eosinophilic increase, lymphocyte inflammation, and mucus cell metaplasia in VLDLR−/− mice suggested a possible hypersensitivity role of this gene. 34 Also, one study demonstrated the anti-inflammatory role of VLDLR in the dextran sodium sulfate-induced mouse colitis model, which suggested a possible association in patients with IBD. 35
rs78726924 in linked to enhancer region of BNC2 gene (3DSNP enhancer score: 0.02). BNC2, which encodes Basonuclin 2 protein, was studied in the previous GWAS study to find genetic biomarkers for lamotrigine-induced skin rash, comparing 1080 controls with 33 lamotrigine-induced skin rash cases in Korean epilepsy patients. 36 Also, using a machine learning prediction model, BNC2 is 1 of the new 32 potential signature genes predicting IBD from healthy cases. 37 In summary, rs78726924 could be a novel biomarker-associated ASA-induced adverse event in patients with IBD.
We discovered rs12681590, rs10967320, rs78726924, and rs13898676 as new genetic biomarkers predicting 5-ASA-induced adverse events in patients with IBD using GWAS analysis that compared mild adverse events group and/or controls with severe adverse events of 5-ASA. Severe phenotype is expected to show a more significant association between phenotype and genotype compared to mild phenotype. Nevertheless, mild adverse events could have a different genetic mechanism compared to severe adverse events. Genetic analysis between the group with mild side effects and the control group showed different results from the analysis based on severe side effects, suggesting that the mechanisms involved in the development of mild and severe side effects may be different.
There were a few limitations in our study. First, the sample size was small in our study. Nevertheless, our study is the first to determine a genetic predictor of 5-ASA adverse events, including very rare lethal events. Second, the phenotypes of adverse events were heterogeneous. However, severe adverse events of 5-ASA are rare, and they could be specific events closely associated with specific genetic variants. Third, the discovered genes were not validated in this study because of the limited sample size. Further prospective studies should be warranted to confirm whether our novel genetic biomarkers indeed have the alleged biological significance.
In conclusion, our study identified novel genetic biomarkers (rs12681590, rs10967320, rs78726924, and rs13898676) to predict adverse events of 5-ASA in patients with IBD. Further replication studies are warranted to validate the novel genetic biomarkers further.
Supplemental Material
sj-doc-1-tag-10.1177_17562848241227029 – Supplemental material for New genetic biomarkers predicting 5-aminosalicylate-induced adverse events in patients with inflammatory bowel diseases
Supplemental material, sj-doc-1-tag-10.1177_17562848241227029 for New genetic biomarkers predicting 5-aminosalicylate-induced adverse events in patients with inflammatory bowel diseases by Jihye Park, I. Seul Park, Ji Hyung Kim, Jung Hyun Ji, Soo Jung Park, Jae Jun Park, Tae Il Kim, Seung Won Kim and Jae Hee Cheon in Therapeutic Advances in Gastroenterology
Supplemental Material
sj-docx-4-tag-10.1177_17562848241227029 – Supplemental material for New genetic biomarkers predicting 5-aminosalicylate-induced adverse events in patients with inflammatory bowel diseases
Supplemental material, sj-docx-4-tag-10.1177_17562848241227029 for New genetic biomarkers predicting 5-aminosalicylate-induced adverse events in patients with inflammatory bowel diseases by Jihye Park, I. Seul Park, Ji Hyung Kim, Jung Hyun Ji, Soo Jung Park, Jae Jun Park, Tae Il Kim, Seung Won Kim and Jae Hee Cheon in Therapeutic Advances in Gastroenterology
Supplemental Material
sj-jpg-2-tag-10.1177_17562848241227029 – Supplemental material for New genetic biomarkers predicting 5-aminosalicylate-induced adverse events in patients with inflammatory bowel diseases
Supplemental material, sj-jpg-2-tag-10.1177_17562848241227029 for New genetic biomarkers predicting 5-aminosalicylate-induced adverse events in patients with inflammatory bowel diseases by Jihye Park, I. Seul Park, Ji Hyung Kim, Jung Hyun Ji, Soo Jung Park, Jae Jun Park, Tae Il Kim, Seung Won Kim and Jae Hee Cheon in Therapeutic Advances in Gastroenterology
Supplemental Material
sj-jpg-3-tag-10.1177_17562848241227029 – Supplemental material for New genetic biomarkers predicting 5-aminosalicylate-induced adverse events in patients with inflammatory bowel diseases
Supplemental material, sj-jpg-3-tag-10.1177_17562848241227029 for New genetic biomarkers predicting 5-aminosalicylate-induced adverse events in patients with inflammatory bowel diseases by Jihye Park, I. Seul Park, Ji Hyung Kim, Jung Hyun Ji, Soo Jung Park, Jae Jun Park, Tae Il Kim, Seung Won Kim and Jae Hee Cheon in Therapeutic Advances in Gastroenterology
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
