Abstract
Background:
Non-small cell lung cancer (NSCLC) remains the most common cause of cancer-related mortality worldwide. The introduction of targeted therapies against oncogenic drivers, particularly EGFR and KRAS mutations, has significantly improved patient outcomes. However, next-generation sequencing (NGS), the current gold standard for molecular profiling, is not always accessible in routine clinical practice, emphasizing the need for noninvasive surrogate biomarkers. Radiomics has emerged as a promising imaging-based approach that extracts a large number of quantitative features from standard modalities such as [18F]FDG PET/CT. By capturing tumor heterogeneity and biological characteristics, radiomics can provide clinically relevant insights and holds potential for identifying predictive biomarkers. Recent studies suggest that CT radiomic features related to heterogeneity, texture, and shape may predict EGFR and KRAS mutation status, while the integration of metabolic parameters from [18F]FDG PET radiomics may further enhance predictive performance and offer a more comprehensive characterization of tumor biology.
Objectives:
To assess the putative role of [18F]FDG PET/CT radiomic features for the prediction of mutated NSCLC.
Study design:
This retrospective observational study included patients with histologically confirmed NSCLC, molecularly profiled by NGS and who underwent baseline [18F]FDG PET/CT scans at Fondazione IRCCS San Gerardo dei Tintori, Monza. Tumor segmentation and radiomic feature extraction were performed on PET images using Pyradiomics, generating 766 quantitative features. Feature selection for EGFR and KRAS mutation association was conducted via repeated random subsampling and LASSO logistic regression.
Data source and methods:
Patients’ histological, clinical and PET/CT imaging data were obtained from the electronic clinical database and picture archiving and communication system of IRCCS Fondazione San Gerardo dei Tintori di Monza between January 2023 and December 2024. Data from 105 patients with biopsy-proven NSCLC and available NGS and [18F]FDG PET/CT scans were analyzed to identify radiomic features from PET images associated with specific mutations. Two different PET/CT scanners were used (Discovery IQ and Discovery MI, GE Healthcare), and radiomic features were extracted using IBSI-compliant algorithms, generating 766 features per tumor. Features correlated with mutations were selected using the Discovery MI dataset (55 patients) and subsequently evaluated on the independent Discovery IQ dataset (50 patients).
Results:
No radiomic features were identified as associated with EGFR mutation in the Discovery MI dataset. Among the features correlated with KRAS mutation in the Discovery MI dataset, FBS_glcm_MCC—a measure of image texture complexity—was confirmed to be associated with KRAS mutation in the independent Discovery IQ dataset, with an AUC of 0.68, p = 0.04, and an odds ratio of 0.65.
Conclusion:
The potential of [18F]FDG PET radiomics as a surrogate for genetic profiling in NSCLC is promising; however, these preliminary findings require further validation in larger cohorts.
Plain language summary
Lung cancer is the leading cause of cancer-related deaths worldwide, and non-small cell lung cancer (NSCLC) is the most common type. Modern treatments can target specific genetic mutations, such as EGFR and KRAS, which can improve patient outcomes. Detecting these mutations typically requires a tissue biopsy analyzed with next-generation sequencing (NGS). However, biopsies are not always possible, especially when tumor samples are small, and the tests are costly and not available everywhere. In this study, we explored a noninvasive approach called radiomics, which uses computer algorithms to analyze PET/CT images and extract detailed information not visible to the human eye. These image features may reflect tumor characteristics and even hint at underlying genetic mutations. We analyzed 105 NSCLC patients who had both PET/CT scans and NGS results. More than 700 features were extracted from the images to evaluate whether any could predict EGFR or KRAS mutations. No features were linked to EGFR mutations, but one texture-related feature, FBS_glcm_MCC, consistently correlated with KRAS mutations. Tumors with KRAS mutations showed more complex patterns of glucose uptake on PET scans, suggesting that radiomics may capture subtle biological differences related to these mutations. Although the predictive accuracy is not yet sufficient to replace genetic testing, these findings suggest that PET/CT scans, which are already used in routine care, could provide additional information about tumor genetics. If confirmed in larger studies, radiomics could help guide treatment decisions, support noninvasive mutation detection, and reduce the need for repeated biopsies, especially when tissue is limited or hard to obtain.
Introduction
Lung cancer accounts for approximately one-quarter of cancer-related deaths worldwide.1,2 Non-small cell lung cancer (NSCLC) is the most frequent subtype, representing around 85% of cases and predominantly adenocarcinomas. 3 Despite the poor prognosis associated with conventional chemotherapy,4–6 the recent introduction and approval of targeted therapies against specific driver mutations have rapidly and dramatically changed the management and survival of these patients. 7 These advances have necessitated coupling routine cytological and histological evaluations of suspected lung cancer samples with more advanced genetic analyses, 8 with some international oncology societies already recommending a “minimal” panel of actionable biomarkers to be tested at diagnosis. 9 Detecting these genetic alterations is not only critical for selecting patients who may benefit from targeted therapies but also for excluding those who could be exposed to potentially detrimental novel immunotherapies. 10
However, several limitations continue to impede the broad and uniform application of this approach in NSCLC assessment. 11 These include the inability to obtain adequate biopsy samples (e.g., scant material or frail patients), 12 the wide spectrum of available testing techniques—from time-consuming and lower-performance targeted methods such as PCR or Sanger sequencing to more expensive and less accessible high-throughput approaches like next-generation sequencing (NGS), 13 and the requirement for specialized infrastructure and trained personnel, which are not always available at all centers. While some of these challenges have found potential solutions (e.g., liquid biopsy as an alternative or complementary approach when tissue is inaccessible), 14 this scenario has led to a heterogeneous and decentralized local testing landscape, which is counterproductive in the era of precision medicine. In this context, surrogate biomarkers capable of identifying patients harboring driver mutations, particularly the most frequent ones such as EGFR and KRAS, could help reduce costs and facilitate the management of NSCLC patients.
Radiomics is a high-throughput imaging analysis technique that extracts a large number of quantitative features from standard-of-care medical images, such as [18F]FDG PET/CT. This methodology employs computational approaches to analyze patterns within these images, revealing subtle features associated with tumor characteristics, tissue heterogeneity, and disease progression.15–17 By correlating these image-derived features with clinical outcomes or patient subgroups, radiomics has the potential to identify predictive biomarkers that can inform personalized treatment strategies. 18 Previous studies have demonstrated the ability of radiomics to identify NSCLC patients with high expression of biomarkers such as PD-L1, 19 further underscoring its clinical relevance.
Recent evidence suggests that radiomics applied to CT images may serve as a useful tool for predicting EGFR and KRAS mutations in NSCLC. 20 While CT radiomic features related to tumor heterogeneity, texture, and shape have been proposed as putative predictors, this approach remains exploratory. [18F]FDG PET radiomics may further improve mutation prediction by incorporating metabolic information, enhancing accuracy, and providing a more comprehensive assessment of tumor biology.
The aim of this work is to evaluate whether [18F]FDG PET radiomics can serve as a noninvasive approach for mutation detection in NSCLC, thereby complementing existing molecular diagnostics by offering additional imaging-based insights into the tumor’s phenotypic characteristics. Integrating molecular and radiomic data may provide a more complete understanding of tumor biology, improve the prediction of treatment responses, and support more personalized and effective patient management.
Materials and methods
Patients’ selection
The study was conducted in accordance with the STROBE (Strengthening the Reporting of Observational Studies in Epidemiology) guideline. 21
We retrospectively reviewed a consecutive cohort of patients with NSCLC diagnosed at the Department of Molecular Pathology, Fondazione IRCCS San Gerardo dei Tintori, University of Milano-Bicocca, Monza, between January 2023 and December 2024. Inclusion criteria were as follows: histologically confirmed NSCLC, availability of baseline [18F]FDG PET/CT scans suitable for radiomic analysis, and molecular profiling data obtained through NGS of lung lesions. PET/CT scans performed within 1 month of the cyto/histological diagnosis of NSCLC were included. Exclusion criteria were: insufficient imaging quality or unavailable PET/CT scans, absence of molecular profiling data, tumors not suitable for accurate segmentation, prior treatments or interventions that could influence baseline imaging features, PET/CT scans performed more than 1 month after cyto/histological diagnosis, NGS analysis not derived from lung lesions, or molecular testing performed at an external institution.
None of the enrolled patients underwent repeat biopsy or had a subsequent surgical specimen available for assessing diagnostic concordance between specimen types; however, the laboratory’s distinction between NSCLC and small cell lung cancer was consistent with previously reported accuracy rates (89%–97%).22,23
From the starting cohort of 140 selected patients, 35/140 patients were excluded because target lesions could not be accurately delineated or segmented on PET images (e.g., pulmonary lesions inseparable from adjacent lymphadenopathy or scarce FDG uptake).
The final study cohort, therefore, consisted of 105 patients. Demographic and clinical data were collected and analyzed, including age and sex at diagnosis, biopsy site, histological subtype, 24 and results of genetic testing (Table 1). All patients were diagnosed with NSCLC and were eligible for NGS testing according to the most recent guidelines. 25
Characteristics of the patients population enrolled in the study.

Written informed consent for participation in the study and for the publication of data was obtained from all subjects prior to study initiation, in accordance with the approval of the local Ethics Committee.
Genetic analysis
Mutational analysis was conducted using Thermo Fisher’s IonTorrent PGM technology with NGS Oncomine Focus Assay panel (Thermo Fisher Scientific, Waltham, MA, USA), which is a targeted, multi-biomarker assay (52 genes) that enables detection of hotspots, SNVs, indels, CNVs, and gene fusions from DNA and RNA in a single workflow, from tumor tissue obtained from formalin-fixed paraffin-embedded (FFPE) tissue samples or cytological samples. The extraction of nucleic acids was performed using Promega’s DNA FFPE and RNA FFPE kits, through the automated Maxwell CSC extractor (Promega Corporation, Madison, WI, USA). Prior to the NGS run, the libraries were verified by electrophoresis quantification with the High Sensitivity D1000 Screen Tape assay on Agilent’s 2200 TapeStation (Agilent Technologies, La Jolla, CA, USA). Subsequently, the files generated by the NGS analysis were applied to the bioinformatics analysis. Data inspection was carried out automatically by using the Ion Reporter Torrent Suite version 5.14 with a dedicated analysis workflow optimized for somatic annotation of solid tumors. For the identification of the somatic variants, the read depth of at least 1000× was respected. In a subset of cases, we used orthogonal methods to confirm the presence of the detected genetic alteration found with NGS, mainly with IHC for ALK evaluation (4%) and real-time PCR for MET exon 14 skipping assessment (7%). Moreover, a small proportion of cases (4%) underwent NGS reanalysis mainly due to pre-analytical and analytical reasons, as previously described. 26
PET/CT image acquisition and reconstruction
[18F]FDG PET/CT scans were acquired according to European Association of Nuclear Medicine Procedural Guidelines 27 at baseline on two PET/CT scanners (50 scans on Discovery IQ (DIQ) and 55 scans on Discovery MI (DMI), GE Healthcare, Milwaukee, WI, USA). Patients were injected with 3,7 MBq/Kg of [18F]FDG, fasting for at least 6 h and with a blood glucose level <150 mg/dL immediately before tracer administration; scans started approximately 60 min after the radiotracer injection. PET images were all reconstructed on a common 2.73 × 2.73 × 3.27 mm3 voxel grid with the same reconstruction algorithm (VUE point high-definition system, VPHDS).
Tumor segmentation and radiomic feature extraction
Target lesions were segmented on PET images using a commercial segmentation tool based on an iterative thresholding method implemented in PET-VCAR (Volume Computed Assisted Reading, GE Healthcare, Milwaukee, WI, USA) and already utilized for lung cancer segmentation.28,29 Radiomics features were computed with the IBSI-compliant Pyradiomics tool. 30 Images were resampled with Bspline interpolation on a 0.8 × 0.8 × 0.8 mm3 voxel grid for morphologic feature computation and on a 3 × 3 × 3 mm3 voxel grid for texture feature computation. 766 radiomics features were computed:
-14 morphologic features (including MTV)
-18 first-order features (including SUVmax, SUVmean)
-59 higher order features computed on the original image with fixed bin number (FBN) quantization (64 bin)
-59 higher order features computed on the original image with fixed bin size (FBS) quantization (bin width 0.3125 SUV)
-77 features computed on the HHH coif1 wavelet sub-band with FBN quantization (16 bin)
-77 features computed on the LLL coif1 wavelet sub-band with FBN quantization (16 bin)
-77 features computed on LoG filtered images (sigma = 1 mm) FBN quantization (64 bin)
-77 features computed on LoG filtered images (sigma = 2 mm) FBN quantization (64 bin)
-77 features computed on LoG filtered images (sigma = 3 mm) FBN quantization (64 bin)
-77 features computed on LoG filtered images (sigma = 1 mm) FBS quantization (bin width 0.3125 SUV)
-77 features computed on LoG filtered images (sigma = 2 mm) FBS quantization (bin width 0.3125 SUV)
-77 features computed on LoG filtered images (sigma = 3 mm) FBS quantization (bin width 0.3125 SUV)
Different quantization methods, specifically FBN and FBS, were explored in combination with filtering techniques such as wavelet transforms and Laplacian of Gaussian (LoG) filtering with varying sigma values, as each approach captures distinct aspects of tumor heterogeneity.
Given the limited sample size, particularly the small number of patients with mutations, radiomic feature harmonization between the two scanners was not feasible. Therefore, we selected features that correlated with mutations in the Discovery MI dataset and subsequently evaluated whether these features also correlated with mutations in the independent Discovery IQ dataset, without defining any optimal threshold. The RQS 31 associated with the analysis flowchart is 9 (25%).
Radiomics feature selection
Radiomic features associated with EGFR or KRAS mutations were selected from the Discovery MI dataset (Figure 1). Patients were randomly divided 100 times into five equally sized subgroups, each maintaining a similar outcome distribution. Within each of the resulting 500 subgroups, the association between radiomic features and outcome was assessed using the non-parametric Wilcoxon–Mann–Whitney test. Features with a Spearman correlation coefficient greater than 0.5 were discarded, and LASSO feature selection was then applied (logistic regression model with L1 penalty, with the regularization parameter C optimized through 5-fold cross-validation over a grid of values ranging from 0.01 to 100). In each iteration, we recorded which features were selected (non-zero coefficients). Final feature selection was based on stability; only features that appeared in >35% of the 500 LASSO models were retained. On selected features, Rank-Biserial Correlation was computed to assess the effect size. It’s worth noticing that although Wilcoxon–Mann–Whitney tests were computed across individual features during the resampling procedure, feature selection was ultimately based on LASSO regularization and stability across 500 subsets, not on statistical significance. As such, multiple testing correction was not applied.
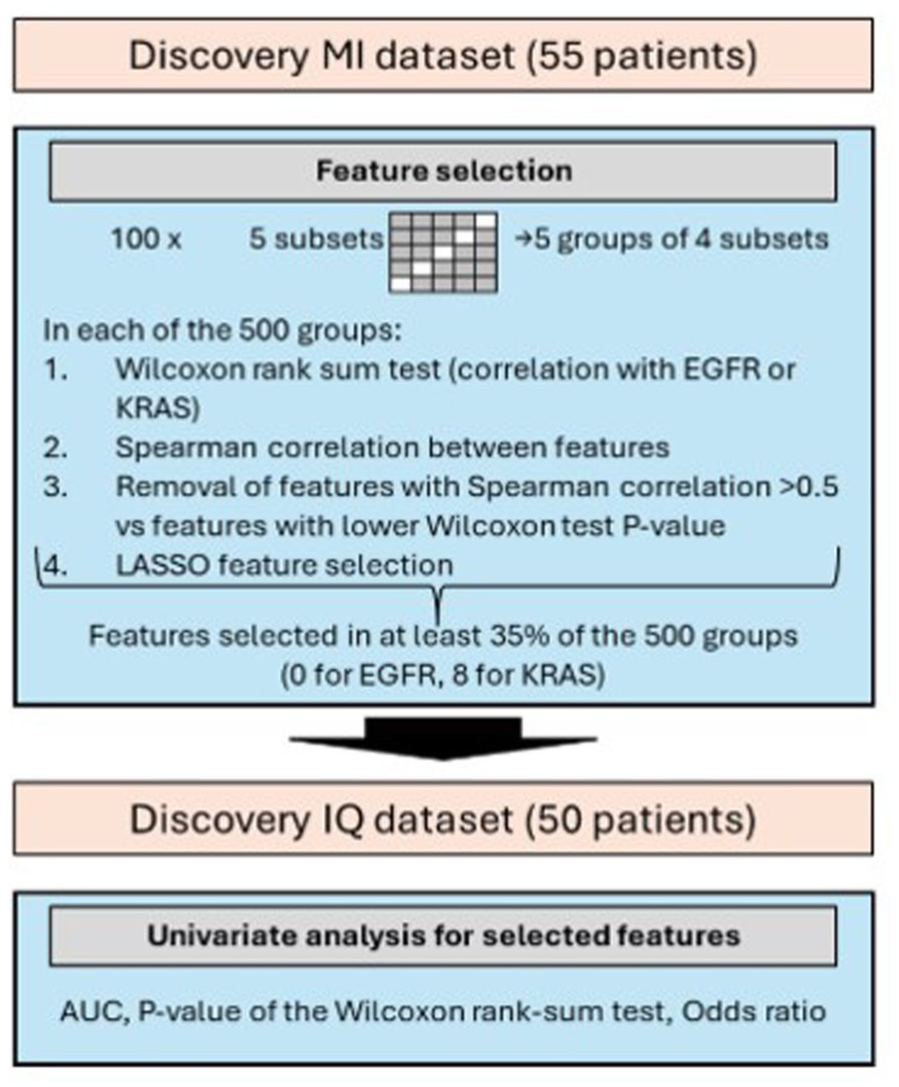
Analysis flowchart.
Selected radiomic feature assessment (test) on an independent dataset
Radiomic features selected from the Discovery MI PET dataset were evaluated on the external Discovery IQ dataset (Figure 1) in terms of Area Under the Curve (AUC), Wilcoxon rank-sum test p-value, and odds ratio.
Results
Patients
Table 1 summarizes the main characteristics of the study cohort. The diagnosis was established in 38/105 cases (36%) based on histological biopsy, in 51/105 (49%) on surgical specimens, and in the remaining 16/105 (15%) on cytology. A final diagnosis of adenocarcinoma (ADC) was available in 91 cases (87%), whereas the remaining 14 cases (13%) included various histologies: poorly differentiated carcinomas (n = 8), non-small cell carcinoma not otherwise specified (n = 5), and adenosquamous carcinoma (n = 1). Results of mutation analysis are reported in Table 2. PET scans were acquired using the Discovery MI scanner in 55/105 cases (52%) and the Discovery IQ scanner in 50/105 cases (48%). Overall, NGS analysis revealed KRAS mutations in 44/105 cases (46%) and EGFR mutations in 14/105 cases (13%) (Table 2). In 21/105 cases (20%), other mutations were identified, including MET, TP53, ALK, BRAF, CDKN2A, PIK3CA, and ERBB2 (see Table 2 for further details).
Mutation and histology distributions on the two analyzed datasets, Discovery MI group (55 patients) and Discovery IQ group (50 patients).

1 case of EGFR and four cases of KRAS had a TP53 co-mutation.
The cases were distributed as follows: seven with MET mutation (including one with a TP53 co-mutation), four with ALK mutation, three with BRAF mutation, three with CDKN2A mutation, two with PIK3CA mutation (including one with a TP53 co-mutation), and two with ERBB2 mutation (including one with a TP53 co-mutation).
Prediction of EGFR and KRAS mutation status with radiomics
Whereas no radiomic features were found to correlate with the presence of mutations overall, or specifically with EGFR mutations, more promising results emerged in KRAS-positive versus KRAS-negative cases. In this context, the radiomic features selected on the Discovery MI dataset and their evaluation on the Discovery IQ dataset are summarized in Table 3. The radiomic feature that demonstrated consistent correlation with KRAS mutation across both the Discovery MI and Discovery IQ datasets was
Radiomics features selected as correlated with KRAS mutation on the Discovery MI scanner are reported (with selection percentage on the 500 trials and Rank-Biserial Correlation), together with their assessment on the Discovery IQ dataset in terms of p-value of the Wilcoxon rank-sum test, AUC and Odds Ratio with lower and upper 95% CI.

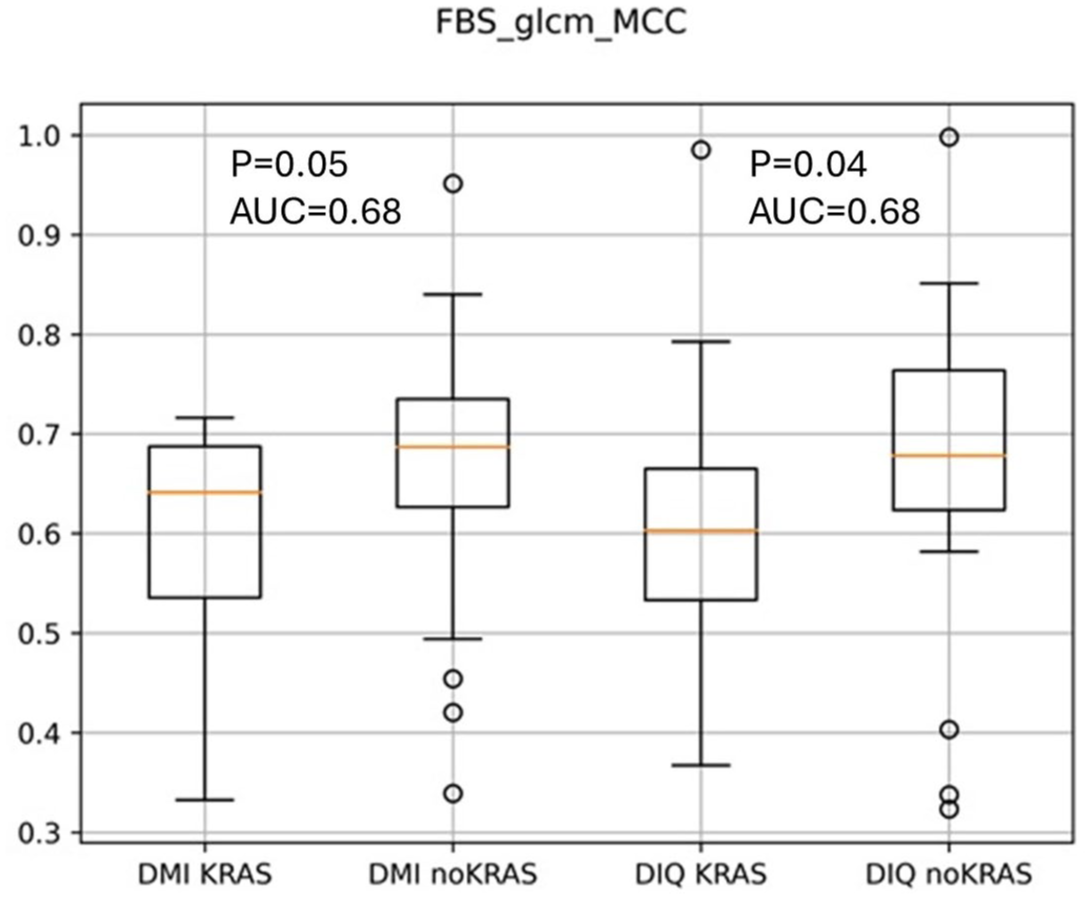
Boxplot representing the distribution of the radiomic feature FBS_glcm_MCC on KRAS-mutated and non-KRAS-mutated lung cancer PET images on the two PET/CT scanners. KRAS-mutated tumors appear to have a lower FBS_glcm_MCC value, that is, a more complex texture of the FDG uptake inside the tumor.
Discussion
The detection of genetic mutations is increasingly becoming the gold standard in the routine assessment of NSCLC, both at diagnosis and during disease monitoring. While targeted approaches (e.g., PCR or Sanger sequencing) perform well in detecting the most frequent genetic aberrations, more comprehensive methods such as NGS provide a broader and more accurate characterization of the complex mutational landscape of cancer, although their use remains limited by cost and availability. Several previous studies have already investigated the potential of radiomics applied to [18F]FDG PET/CT for the detection of the most common NSCLC mutations, namely EGFR and KRAS. Le et al. 32 demonstrated that PET/CT radiomics can effectively distinguish EGFR- and KRAS-mutated tumors, with textural and intensity-based features playing a pivotal role. Similarly, Shiri et al. 33 developed a PET/CT radiomic sequencer that successfully predicted EGFR and KRAS mutations in NSCLC patients. These studies underscore the potential of radiomics to advance precision medicine by enabling noninvasive tumor characterization and treatment stratification. In our NSCLC cohort, we investigated whether radiomic features could correlate with EGFR or KRAS mutation status.
KRAS mutation has evolved from being considered merely a negative predictor for EGFR-targeted therapy in colorectal cancer 34 to the recent approval of anti-KRAS drugs in NSCLC 35 . KRAS mutations are associated with poor prognosis and resistance to targeted therapies; thus, their early detection may support treatment selection and risk stratification. In the population analyzed in this study, KRAS mutations were present in 50% of patients scanned on the Discovery MI and in 32% of those scanned on the Discovery IQ. Eight radiomic features were identified as correlated with KRAS mutation in the Discovery MI dataset; among these, FBS_glcm_MCC retained a significant correlation with KRAS mutation in the independent Discovery IQ test set (AUC 0.68, p = 0.04, odds ratio 0.65). Tumors harboring KRAS mutations exhibited lower FBS_glcm_MCC values, consistent with a more complex intratumoral FDG uptake texture.
Although the biological interpretation of textural features is inherently indirect, it has been suggested that they may reflect underlying tumor heterogeneity at the microscopic level, including variations in cellular density, stromal architecture, and metabolic gradients.36,37 KRAS is among the most frequently mutated genes, driving alterations in metabolic pathways such as enhanced nutrient uptake, increased glycolysis, elevated glutaminolysis, and augmented synthesis of fatty acids and nucleotides. However, the mechanisms underlying KRAS-driven metabolic reprogramming remain incompletely understood. 38 These metabolic alterations may result in heterogeneous FDG uptake patterns, which could potentially be captured through radiomic analysis. Although this association remains hypothetical, it is consistent with prior evidence suggesting that radiomic heterogeneity features may serve as surrogates for underlying molecular and microenvironmental complexity. 39 Our findings are consistent with those reported by Zhang et al., who, in a study conducted exclusively on lung adenocarcinomas, developed a PET-based KRAS mutation prediction model incorporating three radiomic features and smoking status. 40 The model achieved an AUC of 0.731 (95% CI: 0.619–0.843) in the training set and 0.750 (95% CI: 0.248–1.000) in the validation set. The selected features included wHLH_fo_IR, wHLH_glrlm_SRHGLE, and wHLH_glszm_SAHGLE, all of which characterize tumor heterogeneity. Specifically, IR (Interquartile Range) quantifies the spread of voxel intensities within the lesion, reflecting intensity variability; SRHGLE (Short Run High Gray Level Emphasis) captures short, high-intensity voxel runs, representing fine, bright intratumoral structures and potentially indicating localized areas of high FDG uptake; and SAHGLE (Small Area High Gray Level Emphasis) quantifies small clusters of high-intensity voxels, serving as a marker of localized metabolic hotspots. Collectively, these features capture spatial heterogeneity and intensity patterns in FDG uptake, which may reflect underlying tumor biology.
Despite the lower AUC obtained in our analysis compared with Zhang et al., 40 and although we did not evaluate the same specific features, our findings corroborate their results by confirming a relationship between radiomic indicators of tumor heterogeneity and KRAS mutation.
Although
A recent meta-analysis evaluated the predictive value of [18F]FDG PET/CT radiomics for EGFR mutations in NSCLC, including 17 studies with a total of 3763 patients. 44 The study concluded that although PET/CT radiomics shows promise for predicting EGFR status, substantial variability in predictive performance exists across studies, underscoring the need for standardized protocols to improve reproducibility and clinical utility.
Several limitations of our study must be acknowledged. First, its retrospective design and relatively small sample size may limit the generalizability of the results. The major limitation is the low number of available cases, which increases the risk of overfitting. To mitigate this, we applied repeated cross-validation, correlation filtering, and stability-based LASSO selection, and we further assessed the selected features on an independent dataset. In addition, we intentionally restricted the analysis to primary lung cancer cases (excluding lymph nodes) with available NGS data, discarding those assessed with more targeted techniques (e.g., PCR or Sanger sequencing), to obtain a more robust and comprehensive representation of the true genetic landscape of these tumors. Moreover, several cases were excluded from the initial cohort due to segmentation issues and low metabolic volumes, which may represent potential sources of variability within the study population. We acknowledge that this reduction in sample size may have affected the statistical power of the study. Radiomic feature reproducibility (e.g., through ICC analysis) was not assessed; however, in this retrospective setting, we did not have access to test–retest or multiple-segmentation datasets required for such evaluation. The inclusion of two different PET/CT scanners may also have introduced variability in feature extraction and reproducibility. In addition, mutation prevalence differed between the two scanner cohorts. Although AUC and odds ratios are relatively robust to such shifts, differences in class prevalence could still impact generalizability, underscoring the need for further validation in larger, well-matched populations.
Finally, this study focused exclusively on static PET features. Future work may benefit from integrating dynamic PET parameters, which provide additional insights into tracer kinetics and tumor physiology over time. Moreover, combining information from multiple imaging modalities—such as CT, MRI, and PET—through fusion radiomics could capture complementary structural, functional, and metabolic characteristics. Such multimodality approaches may substantially improve the robustness and predictive accuracy of radiomic models, particularly in complex or heterogeneous tumors. Ultimately, prospective, multicenter trials will be required to validate these findings and support their clinical translation.
Conclusion
Our study contributes to the growing body of research exploring [18F]FDG PET radiomic features as noninvasive biomarkers for predicting KRAS mutation status in NSCLC patients. If validated in larger cohorts, radiomics-driven tumor profiling could play a crucial role in personalized oncology, guiding treatment selection, informing prognosis, and monitoring tumor evolution. The results presented here should be considered preliminary, as they were derived from a relatively small patient cohort; further investigation in larger series is warranted.
