Abstract
The gut microbiota plays a crucial role in various physiological functions, such as the production of microbial compounds and maintaining homeostatic equilibrium by complex host-microbial interactions. However, any shift in the constitution and diversity of the microbiota or abnormal interaction with the host can prompt the development of dysbiosis. This review thus illustrates that microbial metabolites, notably short-chain fatty acids, tryptophan metabolites, bile acids, and polyamines, exert significant regulatory effects on innate and adaptive immunological processes, immune response and intestinal barrier integrity through specific receptor activation, involving TLRs, NODs, GPCRs, nuclear receptors, and Wnt/β-catenin. It further explores the disruption of host signalling pathways, caused by dysbiosis, promoting the transcription of specific genes and activating pro-inflammatory pathways. Consequently, this suggests that microbiota acts beyond general health, eventually contributing to gastrointestinal, metabolic, and neurological disorders. Lastly, this review highlights therapeutic approaches required to restore balance and uphold physiological balance.
Introduction
Trillions of microbes in mammalian hosts contribute significantly to developing the immune system and regulating host metabolism, affecting numerous aspects of host physiology. The microbiome refers to microorganisms, their collective genetic elements, and their environmental interactions within a specific environment. 1 The human microbiome is a complex ecosystem of bacteria, viruses, fungi, and archaea that reside in and on the human body and play a pivotal role in maintaining health and regulating diseases. 1 Alterations to the normal microbiome called dysbiosis are associated with the onset and progression of several metabolic and inflammatory conditions, including obesity, insulin resistance, diabetes, inflammatory bowel disease, cancer, and infections. 1 Disruption of microbial balance can lead to widespread systemic effects, influencing distant organs and overall physiology. 2 Moreover, disparity of microbial communities in individuals is influenced by diet, lifestyle, genetics, and environmental exposures, which regulate disease susceptibility and therapeutic efficacy. 3 Hence, the microbiome has emerged as a leading edge for tailored healthcare and biomedical research. 3 Host cell signaling pathways, such as MAPK, NF-κB, PI3 K/Akt/mTOR, Wnt/β-catenin, and Notch, and GPCR-mediated cascades, regulate essential cellular functions, such as inflammation, immune responses, proliferation, and metabolism. 4 These pathways, predominantly controlled by endogenous signals, are increasingly acknowledged as targets for microbial-derived molecules such as short-chain fatty acids (SCFAs), lipopolysaccharides (LPS), and peptidoglycans. 5 This dynamic interaction underscores the role of the microbiome as an active modulator of host signaling, rather than a passive inhabitant. 6 Recent research underscores the impact of microbial metabolites and structural elements on host signaling networks, affecting metabolic, immune, and neurological outcomes. 7 Due to a lack of longitudinal studies and methodological challenges, an in-depth understanding of the mechanisms involved is limited. 8 Influencing host signaling pathways mediated by the microbiome provides potential therapeutic opportunities. Strategies such as fecal microbiota transplantation (FMT), probiotics, and microbiome engineering are being investigated to reestablish microbial equilibrium and target disease-specific signaling pathways. 9 Progress in multiomics and systems biology is aiding the creation of precision medicine approaches by pinpointing and altering microbial pathways that are functionally associated with disease conditions. 10
However, safety issues, individual microbiome variability, and ecosystem complexity must be addressed for successful clinical implementation. 7 This review explores the current understanding of how the microbiome modulates host cell signaling and its implications for therapeutic discovery. It defines the microbiome and its relevance to human health and provides an overview of host signaling systems and the influence of microbes. In addition to discussing potential treatments, current obstacles, and future directions in this quickly developing field, this review preferentially examines how dysbiosis interferes with host signaling pathways to trigger pro-inflammatory reactions that lead to neurological and metabolic disorders.5,6
Microbiome composition and functional diversity
A comparative overview of microbial communities across major human body sites, highlighting dominant taxa, metabolites, and associated host functions, is presented in Table 1
Comparative overview of microbiomes by body site – taxa, dominant metabolites, and associated host functions.

Microbial taxa across body sites
The human microbiome contains a variety of microbial communities, with different body sites carrying unique assortments of microbes that serve specific functions. For instance, the gut microbiota is primarily dominated by two phyla, Firmicutes and Bacteroidetes. Among these dominant phyla, several genera like Bacteroides, Faecalibacterium, Lactobacillus, and Bifidobacterium are noticeable, and their functions are to maintain smooth digestion, modulate the immune system, and maintain gut health and barrier. 11 The skin microbiota primarily comprises Actinobacteria such as Propionibacterium and Corynebacterium, and Firmicutes such as Staphylococcus. They ensure the skin is healthy and can protect against pathogens. 12 The oral microbiota is highly diverse, with the most common phyla being Firmicutes, Bacteroidetes, Proteobacteria, and Actinobacteria. The most predominant genera are Streptococcus, Neisseria, and Prevotella, which ensure good oral health and can fight diseases. 13 The acidic pH of the vagina is due to the most abundant bacterial species found in the vagina, which is Lactobacillus. The importance of the acidic pH is significant because it helps protect against infections and diseases, like bacterial vaginosis, which can occur when the acidic condition is hampered. 14
Functional roles: metabolism, immune regulation, and barrier integrity
Microbes play several roles in ensuring a good host physiology. They help metabolize dietary fibers into short-chain fatty acids (SCFA), such as acetate, propionate, and butyrate, which are used as energy sources for the host. 15 Microbes communicate with the host immune system, producing immune cells and modulating the inflammatory response. For example, SCFAs cause the differentiation of regulatory T cells, resulting in immune homeostasis. 16 Microbes also increase mucus and antimicrobial peptide production, and thus they strengthen epithelial barriers so that pathogens cannot move to a different location. 17
Tools for profiling microbiomes
Recently, sequencing and technological advancements have characterized microbes more precisely. For instance, by 16S rRNA Gene Sequencing, we can identify and classify bacteria. Despite being a cost-effective procedure, it only provides us with limited information about the various functions of a specific bacterium. 18 Metagenomics is a technique that sequences all microbial DNA and analyzes the genes and their potential functions. 19 Another analysis technique, metabolomics, analyzes all microbial metabolites, determines their functions, and analyzes their effects on the host. 20
Overview of key host cell signaling pathways
Microbial components and metabolites significantly influence host cellular functions by modulating diverse types of intracellular signaling pathways. The major host signaling pathways influenced by TLR/NOD, Wnt, GPCR, nuclear receptor, and Akt pathways are illustrated schematically in Figure 1

Toll-like receptor (TLR) and NOD-like receptor (NLR) signaling
The strong interplay between the gut microbiota and immune responses can maintain and regulate the internal environment of the gut. The cells of the gut can discriminate between pathogens via pattern recognition receptors (PRRs), which include families of toll-like receptors (TLRs), nucleotide-binding oligomerization domain (NOD) like receptors (NLRs), among others. TLRs are mainly located on the membrane of immune and epithelial cells and consist of eleven transmembrane proteins. 26 Whether the TLR family activates the immune system or induces tolerance depends on its ability to recognize pathogen-associated molecular patterns (PAMPs) produced by microbial invaders or microbe-associated molecular patterns (MAMPs) from commensal microbes. 26 Upon recognizing PAMPs, TLRs trigger the innate immune system by detecting pathogens from both Gram-negative and Gram-positive bacteria, activating several intracellular signalling cascades, leading to the production of cytokines, chemokines, and transcription factors, and dimerization of TLR2 with other TLR members to stimulate the NF-κB proinflammatory pathway to maintain gut homeostasis. 26 NODs, such as NOD1 and NOD2, are present in the cytoplasm of enteric cells and, like TLRs, can recognize PAMPs and MAMPs and also induce the innate immune system by the activation of NF-κB, MAPK, type I interferon, cytokines, and chemokines, as illustrated in Figure 1(a). 27
MAPK and Nf-κb pathways
Mitogen-activated protein kinases (MAPKs) are a highly conserved signalling pathway that regulates various cellular processes such as cell proliferation and differentiation. They also play a crucial role in angiogenesis and tumour metastasis. The path comprises three kinases: MAP kinase kinase kinase (MKKK), MAP kinase kinase (MKK), and MAPK, where MAPK is the most critical signalling for the survival and development of tumour cells. 28 MAPK in the cytoplasm can translocate to the nucleus and mediate the phosphorylation of both cytoplasmic and nuclear proteins, contributing to metastasis, resistance towards apoptosis, and abnormal cell growth. 29 Nuclear factor kappa-light-chain-enhancer of activated B cells (NF-κB) is a transcription factor family of five subunits forming distinct protein complexes, which regulates cellular processes by binding to promoter regions at DNA sequences. In an inactive state, NF-κB, found in the cytoplasm, is bound to three inhibitory factors (IκBα, IκBβ, and IκBε). 30 However, its activation depends on a series of phosphorylation events resulting in free NF-κB dimers. 30 This results in NF-κB dimers translocating from the cytoplasm to the nucleus, regulating gene transcription and inducing protein conformational changes, promoting cell proliferation, angiogenesis, suppressing apoptosis, and facilitating metastasis. 31
Wnt/β-catenin and notch pathways
Wnt is a family of 19 glycoproteins that modulates embryonic development, adult tissue homeostasis, and tissue regeneration. Activation of this signalling pathway induces the activation of both the canonical Wnt and non-canonical Wnt pathways of β-Catenin. 32 However, accumulation of β-Catenin in the cytoplasm can lead to the migration from cytoplasm to nucleus, binding to LEF-1/TCF4, promoting increased transcription of c-Myc and CyclinD-1. The induction of oncogenes thus facilitates the tumour progression as shown in Figure 1(b). 33 Another signalling network, the Notch pathway, regulates the development and homeostasis of multiple organs and modulates cell proliferation, differentiation, and apoptosis. 34 ligand binding causes NOTCH receptors to undergo three cleavages, releasing the Notch intracellular domain (NICD). This results in translocation into the nucleus to regulate transcriptional complexes containing the DNA-binding protein.34,35 However, aberrant activation can result in cancerous disease by disrupting homeostasis and immune responses. 35
GPCR and nuclear receptor pathways
G protein-coupled receptors (GPCR) are versatile and vital proteins of seven α-helical transmembrane proteins. They can facilitate various regulations and responses, including immune response, hormone regulation, and neurotransmission. Binding to ligands (microbial metabolites such as short-chain fatty acids) causes GPCRs to transduce extracellular signals and regulate intracellular second messengers by activating the downstream effectors of GPCR pathways, such as adenylate cyclase (AC), cyclic adenosine 3′,5′-monophosphate (cAMP), and protein kinase A (PKA), which regulate many physiological processes. Hence, this signalling pathway can influence the immune response as demonstrated in Figure 1(c). 36 The nuclear receptor gets activated when bound to hydrophobic ligands, including fatty acids, hormones, microbial-derived bile acids, and oxysterols. Examples of atomic receptors include proinflammatory-activated receptors (PPARs) and liver X receptors (LXRs), which, when stimulated, modulate cell differentiation, proliferation, and metabolism, contributing to cardiovascular diseases, cancer, and inflammation. The signalling cascade is represented in Figure 1(d).37–39
PI3K/Akt/mTOR pathway
The phosphatidylinositol-3 kinase (PI3 K), a signal transduction pathway, modulates cell survival, growth, and metabolism. 40 Its activation occurs when the activated receptor tyrosine kinase activates IRS, which stimulates PI3 K and catalyzes the conversion of PIP2 to PIP3. The events follow with the binding of PKB/Akt to PIP3, allowing PDK1 to phosphorylate, resulting in partial PKB/Akt activation. 41 This alteration activates mTORC1 and mTORC2. mTORC1 promotes protein synthesis and cell growth, while mTORC2 regulates cytoskeletal organization and cell survival. 40 Hence, any disruptions in this pathway can result in carcinogenesis. Figure 1(e) depicts the pathway. 40
Mechanisms of microbiome-mediated modulation of host signaling
Microbial communities influence host physiology by producing specific molecular signals that directly interacting with host receptors and signaling pathways. Table 2 summarizes major microbial-derived molecules and their targets within host signaling pathways. Figure 2 schematically illustrates the principal mechanisms through which microbial metabolites, molecular patterns, extracellular vesicles, and enzymes influence host signaling pathways by highlighting their roles in immune regulation, inflammation, and homeostasis.

Key microbial molecules and their known targets in host signaling pathways.

Microbial metabolites
The gut microbiota communicates with the host signalling pathways using microbial metabolites, like short-chain fatty acids (SCFAs), tryptophan metabolites, bile acids, and polyamines. 42
Short-chain fatty acids (SCFAs)
The gut microbiota metabolizes dietary fibers to metabolites such as short-chain fatty acids, acetate, butyrate, and propionate.
43
These metabolites can dictate the host's health through various mechanisms, including epigenetic regulation, immune modulation, and barrier integrity. Epigenetic regulation means altering the expression of genes without changing the DNA sequence. Metabolites such as butyrate and propionate are involved in the tryptophan metabolism of histone deacetylases. As a result, histone acetyltransferases can add acetyl groups to the histones to a greater extent, leading to an increased modulation of gene expression.
44
They also promote the differentiation of regulatory T cells.
44
SCFAs also regulate immune cell functions and inflammatory responses by activating G-protein coupled receptors like GPR41, GPR43, etc.
45
SCFAs can also increase the number of tight junction proteins and enhance mucin production. A study that aimed to determine the protective effects of SCFAs on intestinal barrier function used the immunoblotting technique to analyze the expression of ZO-1, a tight junction protein. Furthermore, it was revealed that after the treatment of the CACO 2 monolayer with SCFAs like acetate, propionate, and butyrate, the expression of ZO-1 increased.
46
Due to this, intestinal barrier function improves, leading to a well-balanced gut homeostasis
Tryptophan metabolites
The gut microbiota metabolizes tryptophan into compounds like indole derivatives, kynurenine, and serotonin. 48 These compounds influence host signaling in multiple ways. For instance, indole derivatives bind to aryl hydrocarbon receptor (AhR), a ligand-activated receptor, and then it moves to the DNA inside the nucleus and alters gene expression. This can lead to the modulation of immune responses and barrier function. 49 Not just this, tryptophan metabolism also triggers the synthesis and production of serotonin, a neurotransmitter, which influences gut motility and mood regulation. 50 The gut microbiota modifies the primary bile acids synthesized by the liver into secondary bile acids, which regulate host signaling in various ways. 51 Secondary bile acids trigger lipid metabolism, glucose homeostasis, and inflammatory responses by activating the farnesoid X receptor (FXR), a nuclear receptor, and G-protein coupled bile acid receptor (TGR5). 52 The antimicrobial properties of bile acids Gram-negative bacteria's outer membrane, which is LPS, directly influence host health. 53
Polyamines
Unlike other metabolites, polyamines such as putrescine, spermidine, and spermine are secreted by the host and the gut microbiota. They carry out multiple cellular functions. Polyamines play a role in maintaining intestinal mucosal integrity. They do this by triggering the proliferation and differentiation of epithelial cells. 54 An alteration in the levels of polyamines, which is associated with the onset of certain metabolic disorders like obesity and type 2 diabetes; hence, polyamines could be deemed as potential therapeutic targets of these metabolic disorders. 55 These metabolites explain the association of the gut microbiota with the host pathways, further demonstrating their potential as drug therapies.
Microbe-Associated molecular patterns (MAMPs)
The pattern recognition receptor (PRRs) recognize specific microbial structures called the Microbe-Associated Molecular Patterns (MAMPs), such as lipopolysaccharide (LPS), peptidoglycan, and flagellin, to generate an immune response and act as the host's first line of defense.56,57
Lipopolysaccharide (LPS)
A crucial component of the Gram-negative bacteria's outer membrane is LPS, composed of an oligosaccharide, lipid A, and an O-antigen. Lipid A activates the endotoxic activity of LPS. When released, LPS binds to the LPS binding receptor in the serum, which is then sent to CD14. CD14 presents this to the TLR4-MD2 complex on immune cells. This association triggers a downward signaling cascade, including NF-κB activation, resulting in the release of inflammatory cytokines like TNF-α and IL-6. 58
Peptidoglycans
Peptidoglycans comprise cross-linkages between peptides and glycans, and they are the structural components of the bacterial cell walls. Both intracellular and extracellular PRRs recognize these peptidoglycans. Peptidoglycan fragments are usually identified by the TLR2-TLR1 or TLR2-TLR6 complex on the cell surface, causing the production of pro-inflammatory cytokines. Intracellular receptors like NOD1 and NOD2 recognize specific peptidoglycan fragments, activating NF-κB and MAPK pathways, eventually producing cytokines and antimicrobial peptides. 59
Flagellin
The major bacterial flagella protein, Flagellin, is a MAMP recognized by extracellular and intracellular PRRs. TLR5, present on epithelial cells, recognizes extracellular flagellin, which triggers the activation of NF-κB; hence, various cytokines are produced. 60 Intracellular flagellin is detected by NAIP5/6 and the NLRC4 inflammasome complex, releasing mature interleukins. This initiates pyroptosis, a process by which inflammatory cells die, as demonstrated in Figure 2(b). 61
Microbial enzymes and effectors
Enzymes and effectors secreted from microbes provide ways to modulate host cell signalling pathways, which then lead to changes in the various physiological and pathological processes, such as immune responses, cell survival, and tissue homeostasis.
Proteases
Specific bacterial proteases, such as gingipains secreted from Porphyromonas gingivalis, can degrade host cytokines, including proinflammatory cytokines and immunoglobulins, which weakens the defense mechanisms of the host's immune system and also results in persistent inflammation. 62 Conversely, bacterial proteases like LasB from Pseudomonas aeruginosa and GelE from Enterococcus faecalis alter the host's inflammatory responses by activating protease-activated receptors on host cells. It also results in altered gut motility and permeability. 62
Glycosidases
Some microbes can get the nutrients necessary for survival by expressing glycosidases that break down the host glycan molecules. For instance, Bacteroides thetaiotaomicron produces glycosidases that break down host mucin glycans, compromising the mucosal barrier and resulting in immune responses. 63 The microbes express glycosidases based on the amount of glycans in the lipid bilayers, endowing microbes in different host environments. 63
Secreted effectors
Pathogenic bacteria use their inbuilt secretion systems to secrete effectors into the host cells, and these effectors then hijack the host's signaling pathways. Escherichia coli (EPEC) employs a type III secretion system to deliver effectors like NleE, NleC, and NleD into the host. These effectors manipulate and interfere with NF-κB and MAPK signaling pathways, which are central regulators of the inflammatory responses. As a result, the inflammatory responses are weakened, as mentioned in Figure 2(d). 64 Conversely, Legionella pneumophila secretes effectors to inhibit host protein synthesis, demonstrating that the NF-κB pathway is not activated. This results in prolonged NF-κB activation and modified cytokine production, which may lead to the pathogenesis of various diseases. 64
Extracellular vesicles and nanoparticles
Bacterial extracellular vesicles (BEV) are nanosized particles secreted by the bacteria. They comprise lipid bilayers enclosing various molecules like proteins, lipids, nucleic acids, and metabolites. They can manipulate host physiological processes remotely, one of the many pivotal roles in inter-kingdom communication. 65
Role in inter-kingdom communication
The gut microbiota can interact with the host in many ways, one of which is by BEVs. Various microbial components can be sent to host cells through BEVs, leading to the modulation of multiple signaling pathways and altered systemic physiology. For instance, a study revealed that Akkermansia muciniphila secretes BEVs that can trigger serotonin release in the colon and hippocampus, meaning that the gut and the brain can communicate through BEVs. 66 According to a study, BEVs can reach distal organs like the liver and adipose tissues, where they unload their cargo and trigger some specific signaling cascades, leading to insulin resistance and glucose intolerance. 66 BEVs can also alter host immune responses. They can interact with various receptors on host cells, like Toll-like receptors (TLRs) and nucleotide-binding oligomerization domain (NOD)-like receptors, which will eventually lead to either activation or suppression of immune signaling pathways. For instance, BEVs from Bacteroides fragilis interact with TLR2 and TLR4 and modulate their expression in epithelial cells, leading to the secretion of the anti-inflammatory cytokine IL-10 by CD4+ T cells. 67
Effects on immune and epithelial signaling
As mentioned before, BEVs play a role in influencing host immune response, but besides that, they can also influence epithelial cell signaling. They also achieve this action by activating or inhibiting signaling pathways. For example, BEVs from Escherichia coli Nissle 1917 upregulate tight junction proteins like zonula occludens-1 (ZO-1) and claudin-14, and downregulate claudin-2, which is a leaky protein, thus maintaining intestinal homeostasis
Host signaling responses to microbiome interactions
The human microbiome profoundly affects host physiology by regulating immune responses, cellular balance, barrier function, and neuroimmune signaling. Understanding these interactions is key to developing microbiome-based therapies. 68 Figure 3 illustrates key host responses to microbiome-derived signals, including immune modulation via PRRs, systemic effects of circulating metabolites on the brain, epigenetic regulation of gene expression, and epithelial barrier restoration through AHR signaling.
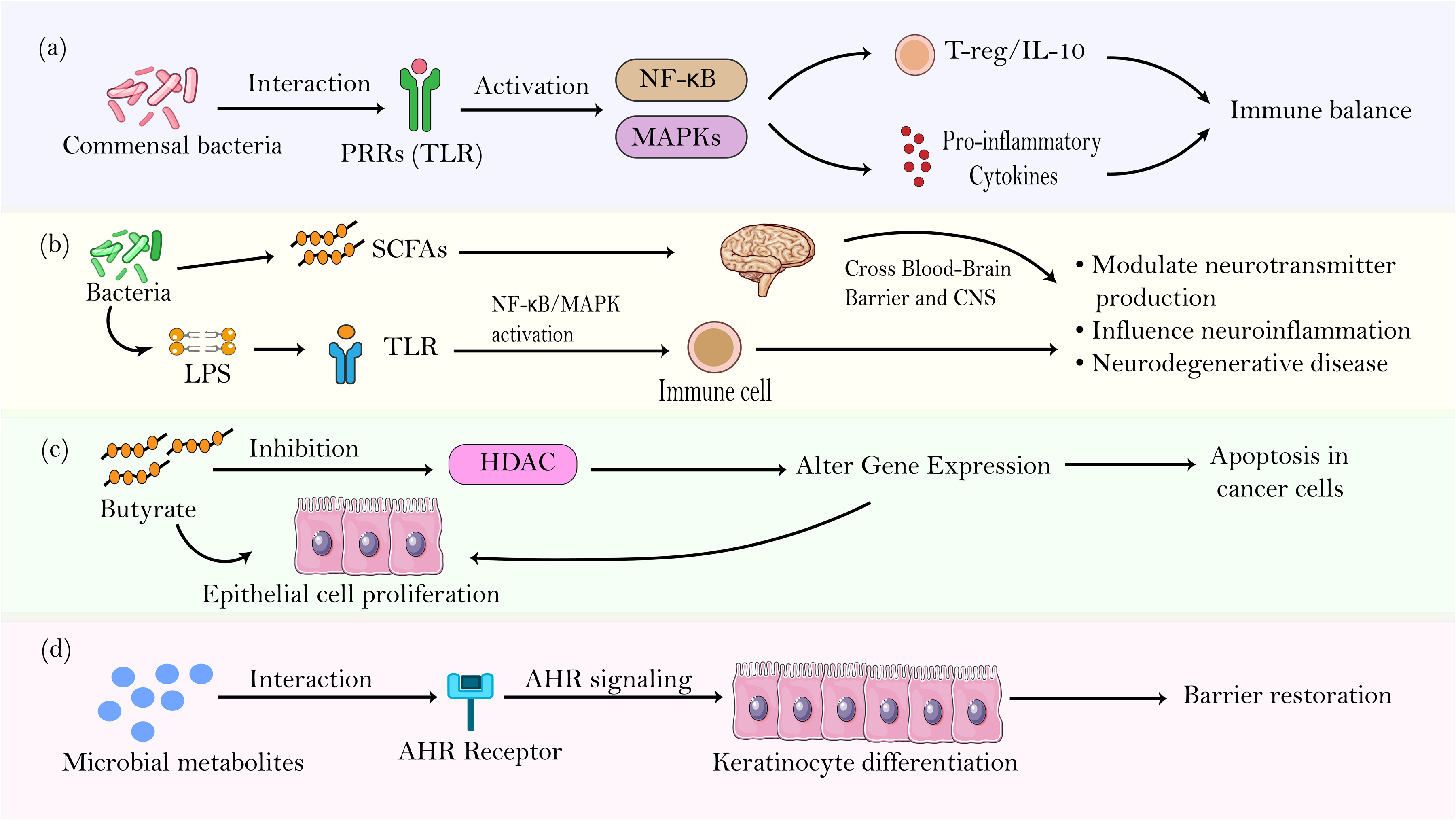
Inflammation and immune modulation
The microbiota is essential in shaping the host immune system. Commensal microbes engage pattern recognition receptors (PRRs) like TLRs and NOD-like receptors, triggering signaling pathways such as NF-κB and MAPKs. 26 Signaling pathways that release pro-inflammatory cytokines such as TNF-α, IL-1β, and IL-6 play an essential function in immune defense; however, an imbalance can cause chronic inflammation. In response, short-chain fatty acids (SCFAs) produced by microbial factors activate regulatory T cells (Tregs) and IL-10 synthesis to induce an anti-inflammatory response and maintain immune equilibrium, as depicted in Figure 3(a). 69
Cell proliferation, apoptosis, and differentiation
Microbial interactions affect host cell proliferation and apoptosis, which are vital for tissue maintenance and repair. For example, butyrate, a SCFA from dietary fiber fermentation, fuels colonocytes and modulates gene expression by inhibiting histone deacetylases (HDACs).
70
This modulation initiates a butyrate paradox process, which promotes healthy epithelial cell proliferation while inducing apoptosis in colorectal cancer cells.
71
Furthermore, signaling networks that influence cellular growth and differentiation, such as the TGF-β pathway, are activated, enhancing immune evasion and metastasis
Barrier function and epithelial repair
Epithelial barrier integrity, especially in the gut, is vital for blocking pathogens and maintaining immune tolerance. Commensal microbes support this by boosting tight junction proteins, mucins, and antimicrobial peptides. Disrupted microbiota can weaken the barrier, causing increased permeability, inflammation, and disease. 73 Microbial metabolites activate aryl hydrocarbon receptor (AHR), facilitating keratinocyte differentiation and enhancing barrier integrity, thus contributing to skin barrier repair as outlined in Figure 3(d). 74
Neuroimmune and gut-brain axis signaling
The gut-brain axis allows the interaction between the gut microbiota and the central nervous system, influencing neurodevelopment, behavioral processes, and the progression of neurological disorders. Microbial metabolites like SCFAs can cross the blood-brain barrier, influencing neuroinflammation and neurotransmitter activity. 75 SCFAs have been shown to influence serotonin production, affecting mood and gastrointestinal motility. 76 Additionally, microbial components such as lipopolysaccharides (LPS) can activate Toll-like receptors, inducing the release of pro-inflammatory cytokines, contributing to pathogenesis in neuroinflammatory disorders, including Alzheimer's and Parkinson's (Figure 3(b)). 77
Case studies: disease contexts
Inflammatory bowel disease (IBD)
Inflammatory bowel disease (IBD) encompasses essentially two main disease subtypes: Ulcerative colitis (UC) and Crohn's disease (CD) and is characterized by chronic inflammation of the gastrointestinal tract. 78 Emerging theoretical perspectives suggest that one potential contributor to IBD is the gut microbiome dysbiosis, often induced by diet, lifestyle, medications, or infections. This imbalance causes commensal bacteria to transition into pathobionts, compromising the mucosal barrier. Consequently, a heightened activation of the host's immune and metabolic responses occurs to restore balance between the gut and microbiome. 79 IBD is specified by a series of symptoms such as diarrhoea, abdominal pain, gastrointestinal bleeding, weight loss and fatigue, caused due to the narrowing of the gastrointestinal tract, inflammation and ulceration.
The quantitative or qualitative change in the composition of the gut microbiome is one of the most important factors regulating the intestinal immune system; it thus may influence the development and course of IBD. 80 A study examining the gut microbiome in IBD patients revealed the decreased representation of specific beneficial bacterium species, with an increase in detrimental bacteria, such as Bacteroides and Enterobacteriaceae species. 81 Another study analyzing the efficacy of fecal microbiota from UC patients and healthy groups observed heightened levels of cytokines and immune-related markers such as IL-6, TNF-α, IL-10, IL-12, CD-80, and CD-86 in UC patients, indicating a stronger inflammatory response compared to healthy groups. 82 Accumulating studies have concluded that IBD is associated with a generalized reduction in microbiota diversity and diminished microbiota composition, significantly increasing the host's immune responses."83,84 By understanding the functional properties of the intestinal microbiome, it is possible to restore health to a diseased microbiome through diet, prebiotics, probiotics, antibiotics, and/or FMT. Probiotics are live microorganisms that can provide health benefits by suppressing pathogen growth and T-cell proliferation, and include organisms such as Bifidobacteria, Lactobacilli, and Enterococci, among others. 85 One of the first probiotics that was administered to UC patients was Escherichia coli Nissle 1917, which exhibited beneficial effects. 86 Prebiotics are non-digestible dietary compounds such as lactulose, lactosucrose, oligofructose, and inulin, providing food for specific host microbiota, facilitating growth and enhancing gut function. 87 FMT influences multiple stages of the immune system, such as increasing regulatory T cells, while reducing TH17 cells and proinflammatory pathways.88,89 A study analysed the uses of rifaximin, metronidazole, clarithromycin, and ciprofloxacin antibiotics in Crohn's disease and observed improved patient clinical outcomes. 90
Cancer
Microbial metabolites produced during the metabolic process of microorganisms can influence host tumorigenesis and tumour progression by several mechanisms, including modulation of the immune system, cancer or immune-related signaling pathways, epigenetic modification of proteins, and DNA damage. For instance, a Methylobacterium causes a decrease in the production of TGF-β and CD8+ tissue-resident memory T-cells, leading to gastric cancer. 91 Candida albicans releases IL-22 by activating the aryl hydrocarbon receptor (AHR) and STAT3, resulting in cell metastasis and the formation of intratumoral blood vessels. 91 Moreover, tumor and microenvironment cells respond to signals from microbiota, regulating multiple host pathways related to carcinogenic processes. One such example is presented by Escherichia coli or Fusobacterium nucleatum, which induces DNA damage via the pks gene and genetic mutations in epithelial cells and modulation of the tumor microenvironment, leading to colorectal carcinogenesis. 92 There is a strong correlation between a patient's gut microbiota and their response to immunotherapy; however, whether this correlation is positive or negative depends on the abundance and range of organisms in the gut microbiota.
Treatment targeting synergistic inhibitory molecules that suppress the immune response, such as CTLA-4, PD-1, and PD-L1, is applied to reduce tumor progression. The presence of B. polymorpha, B. fragilis, and Burkholderia cepacia in one research has been associated favourably with the efficacy of CTLA-4-targeting therapy. 93
In another study, examining the effect of anti-PD-L1 treatment in melanoma patients showed an abundance of Faecalibacterium in responders. At the same time, an increase in Bacteroides thetaiotaomicron, Escherichia coli, and Anaerotruncus was observed in patients with poor clinical outcomes. 94 An analysis of the effect of anti-PD-1 efficacy in metastatic melanoma patients revealed a significantly improved response in patients with higher abundances of bacterial species such as Bifidobacterium longum, Collinsella aerofaciens, and Enterococcus faecium. 95 In contrast, patients with hepatocellular carcinoma responded poorly to anti-PD-1 treatment due to increased Proteobacteria and decreased Akkermansia muciniphila and Ruminococcaceae spp. 96 The studies strongly suggest that manipulating the profile and concentration of gut microbiota may modulate cancer immunotherapy.
Metabolic diseases
Several metabolic diseases, such as type 2 diabetes mellitus, non-alcoholic fatty liver disease, and atherosclerosis, have been implicated in dysbiosis. The gut microbiota triggers activation against bacterial components and metabolic production from dietary compounds to regulate metabolic health; thus, disruptions in the pathway cause low-grade inflammation and altered metabolic processes. 97
Impact on insulin signaling
Gut dysbiosis reduces the production of short-chain fatty acids (SCFA), with increased absorption of lipopolysaccharide (LPS) resulting in the activation of Toll-like receptor and LPS receptor CD14, leading to the induction of inflammatory cytokines, NF-κB, and MAPK pathways. With these activations, an impairment of the insulin signaling is observed with the increased serine phosphorylation of IRS by protein kinases such as c-Jun N-terminal kinase (JNK) and inhibitory κB kinase (IKK)-β, and decreased tyrosine phosphorylation of the insulin receptor, insulin receptor substrate (IRS), and Akt. Additionally, the plasma concentration of IL-6 is inversely proportional to insulin sensitivity. Other cytokines, IL-17 and IFN-γ, and tumor necrosis factor (TNF)-α can also affect insulin signalling. Thus, low-grade inflammation plays a key role in the mechanisms of insulin resistance.98,99
Impact on lipid metabolism
The gut microbiota regulates dietary lipid composition, digestion, and absorption, potentially altering intestinal lipoprotein formation. The lipid and lipoprotein metabolism can be affected by gut microbiota via bile acid receptors FXR and TGR5, short-chain fatty acids (SCFAs), branched-chain fatty acids (BCFAs), bile salt hydrolase (BSH), and lipopolysaccharides (LPS). Prevalent SCFAs, including acetate, propionate, and butyrate, are involved in cholesterol synthesis and lipid metabolism and have been observed in decreased amounts in patients affected by dysbiosis. Consequently, an increased proportion of lipopolysaccharide (LPS)-containing microbiota in the gut has been linked to increased glycemia and insulinemia by transporting lipids in lipoproteins. The findings suggest that influencing SCFA, LPS production, and cholesterol-metabolism-related gene expression can alter lipid metabolism.100,101 Bioactive compounds such as polyphenols, prebiotics, and FMT can reduce the effect on metabolic pathways. 102
Neurological disorders
Five separate studies established the communication process between the commensal microbiota, highlighting the onset of neurological disorders due to any disruptions in this pathway. 103
Microbiota-Gut-Brain axis
A strong interplay between gut microbiota and the brain via the microbiota-gut-brain axis(MGB) involves bidirectional communication, responsible for maintaining mood, cognition, and neurodevelopment by modulating immune, endocrine, neural, and metabolic pathways. 104 The gut microbiota produces crucial neurotransmitters such as GABA, dopamine, and glutamate, with approximately 90% of serotonin being synthesized in the gut. They also make short-chain fatty acids (SCFAs) like butyrate and propionate, which modulate the microglial activity in the brain, modify the blood-brain barrier, and impact neuroinflammation.104,105 Additionally, MGB affects the hypothalamic-pituitary-adrenal (HPA) axis, which maintains the stress response. 106 lastly, they can modulate host physiological processes through microbial constituents such as lipopolysaccharides or substrate metabolism, including bile acids. 106
Neuroinflammation and neurological disorders
Association between neurological disorders such as Alzheimer's disease, Parkinson's disease, and multiple sclerosis due to dysbiosis has been demonstrated in recent findings. 107 In Alzheimer's disease (AD), dysbiosis can accumulate amino acids like phenylalanine and isoleucine, inducing neuroinflammation by activating TH1 cells, cell differentiation, and microglial activation.108,109 In Parkinson's disease (PD), dysbiosis causes inflammation and oxidative stress, transmission of α-synuclein protein, which causes dopaminergic neuron death in the substantia nigra, leading to motor symptoms of PD. 110 Multiple sclerosis (MS) is characterized by the composition and range of gut bacteria, marked by elevated production of Methanobrevibacter, Akkermansia, and Bacteroidetes. At the same time, a reduced population of Bifidobacteria, Roseburia, Coprococcus, and Prevotella is observed in MS patients. Noticeably, short-chain fatty acids are produced by the present bacteria, contributing to chronic inflammation in MS.111,112 SCFA supplementation, probiotics, prebiotics, and FMT have been explored among the therapies. 113
Table 3 summarizes key microbiome–host signaling interactions across various disease contexts and highlights emerging therapeutic strategies aimed at modulating these pathways.
Summary of microbiome-host signaling interactions across diseases and potential therapeutic interventions.

Therapeutic opportunities and strategies
Probiotics, prebiotics, and postbiotics
Microbiome-targeted therapies like probiotics, prebiotics, and postbiotics modulate host signaling pathways to restore balance and treat diseases, with specific strains or metabolites targeting distinct pathways. Probiotics modulate host pathways like NF-κB and TLR, reducing inflammation and improving barrier function strains such as Lactobacillus rhamnosus and Bifidobacterium longum lower pro-inflammatory cytokines.114,115 Tailored microbial consortia enhance this effect, restoring microbial balance and immunity. Anaerobe consortia, for instance, promote regulatory T cells via SCFAs for gut anti-inflammatory responses.116,117 Prebiotics, non-digestible fibers, and oligosaccharides, selectively promote beneficial microbiota that produce metabolites modulating host pathways. 118 Insulin-type fructans increase SCFA production, activating GPR41/43 and inhibiting histone deacetylases, enhancing GLP-1 secretion and insulin signaling. 119 They also regulate TLR-mediated immune responses, fostering an anti-inflammatory state. Postbiotics modulate host signaling without live microbes, including microbial metabolites and secreted factors. SCFAs, bacteriocins, and exopolysaccharides regulate NF-κB, MAPK, and PI3 K/Akt pathways to reduce inflammation, strengthen epithelial barriers, and influence metabolism. 120 Butyrate, for instance, promotes regulatory T cell differentiation by inhibiting histone deacetylases, affecting gene expression for immune tolerance. Postbiotics offer a stable, well-defined alternative to probiotics, bypassing viability and colonization issues. 121 Multi-omics and systems biology will facilitate the accurate identification of microorganisms and metabolites that influence host signaling, hence aiding the creation of personalized, disease-specific microbiome therapies.
Microbiome-Derived metabolites as drug leads
Metabolites produced from the microbiome are increasingly acknowledged as possible therapeutic possibilities because they alter host signalling and maintain homeostasis. These substances offer potential treatments for metabolic, neurological, and inflammatory diseases and their synthetic derivatives. Notably, through GPR41/43 activation and HDAC inhibition, short-chain fatty acids like acetate, propionate, and butyrate control immune activity, metabolism, and epithelial integrity. 122 To improve stability, mimetics like tributyrin reduce inflammation and aid repair in models. 123 Synthetic HDAC inhibitors also show promise in modulating immune tolerance and metabolism. 124
Bile acids, converted by gut microbes, regulate metabolism and immunity via FXR and TGR5. 53 Obeticholic acid, an FXR agonist, treats biliary cholangitis and is under study for NASH. 125 TGR5-targeting analogs may improve metabolism by enhancing energy use and insulin sensitivity. 126 Microbiome metabolites like tryptophan catabolites modulate AhR and immune checkpoints, serving as drug leads. 127 Synthetic indole derivatives targeting AhR treat autoimmune diseases by modulating mucosal immunity. 128 Challenges in stability and bioavailability remain, but advances in chemistry and delivery will improve these therapies.
Engineered microbes and synthetic biology
Synthetic biology and microbial engineering enable precise delivery of bioactive molecules by programming microbes to sense host cues and produce therapeutics, allowing targeted modulation of host signaling for treating complex diseases. 129 Genetically modified microbes deliver cytokines, enzymes, or peptides to modulate immune and metabolic signaling. Lactococcus lactis secreting IL-10 reduced colitis inflammation by inhibiting NF-κB. 130 Synthetic biology enables microbes to sense biomarkers and release therapeutics conditionally, enhancing safety and restoring signaling balance. 131 Engineered microbes can deliver enzymes to metabolize pro-inflammatory compounds or correct metabolic issues. E. coli Nissle 1917 expressing bile salt hydrolase modulates FXR and TGR5 signaling, affecting metabolism and immunity. 132 Bacteria secreting antimicrobial peptides shape microbiota and host immunity, though challenges like stability, immunogenicity, and regulation remain. 133 Genome editing, biocontainment, and synthetic circuits enhance engineered microbes’ robustness and control, advancing clinical translation. 134
Fecal Microbiota transplantation
FMT restores microbial diversity, enhancing immunity, barrier integrity, and metabolism. 135 It is highly effective in recurrent C. difficile infection, with cure rates over 85%, surpassing antibiotics by restoring colonization resistance and reducing inflammation.136,137 Beyond rCDI, FMT may modulate host signaling in IBD, metabolic syndrome, and neuropsychiatric disorders. 138 In ulcerative colitis, it promotes regulatory T cells and reduces NF-κB inflammation. 139 However, efficacy varies due to donor differences, lack of protocols, and unclear mechanisms limiting optimization. 140 Safety concerns over pathogen transmission require strict donor screening and quality control. 141 The incomplete understanding of the long-term effects of FMT highlights the significance of long-term research. Efforts are being made to create synthetic microbiota and defined microbial consortia that can alter host signalling pathways to get beyond the current constraints. 142 Furthermore, it is anticipated that combining systems biology and multi-omics techniques would make it easier to identify critical microbial effectors, promoting the creation of individualised treatment plans.
Current challenges and future directions
Individual variability and personalized microbiome medicine
The microbiome's composition differs among individuals due to variations in genetic factors, diet, environment, and lifestyle, which complicates the creation of universal microbial biomarkers. 143
Causality versus correlation in microbiome studies
Distinguishing causation from correlation in microbiome-disease links is challenging. 143 Functional studies using germ-free animals and synthetic communities help infer causality, but applying these findings to humans is restricted by complexity and ethical considerations. 144
Need for integrated multi-omics and computational modeling
A precise understanding of microbiome-host interactions necessitates the integration of multi-omics data. 145 While computational tools can make predictions, there is still a need for standardized methods and large, validated cohorts. 146
Regulatory and ethical considerations
Microbiome therapies face regulatory challenges due to the complexity and variability of live microbial products. 147 Quality control, long-term safety, and risk mitigation, including horizontal gene transfer, must be prioritized in regulatory frameworks. Donor permission, safeguarding the privacy of microbiome data, and guaranteeing equitable access to customised treatments are all ethical considerations. 148 Individuals from different sectors, such as Physicians, Microbiologists, Bioinformaticians, and regulatory agencies, should affiliate to address these pressing concerns. Advances in single-cell technology, synthetic biology, and longitudinal studies may enhance treatment precision and understanding of causality. Realizing the full clinical potential of microbiome-based therapies requires integrating mechanistic understanding with robust ethical and regulatory frameworks.
Table 4 outlines the key obstacles in this field and summarizes proposed solutions aimed at advancing microbiome-focused therapeutic research
Challenges and proposed solutions in microbiome-based therapeutic research.

Conclusion
The delicate balance between the human microbiome and host cell signaling is essential in determining health and disease. The review highlights that microbial communities’ metabolites and structural components alter major host signaling systems such as NF-κB, MAPK, PI3 K/Akt, Wnt/β-catenin, and GPCR pathways. These interactions affect the immune responses, metabolism, neuroinflammation, epithelial integrity, and cancer progression. Notably, microbial products like short-chain fatty acids, bile acids, and tryptophan degradation products have therapeutic potential by acting on the inflammatory, metabolic, and neuroimmune routes. Treatments that promise to rectify dysregulated host signaling and reinstate microbial balance include probiotics, prebiotics, postbiotics, microbiome-derived metabolites, engineered microbes, and microbiota transplantation, including FMT. The indications of the successful use of these interventions include a wide range of diseases: inflammatory bowel disease and metabolic disorders, neurological disorders, and oncological disorders. However, these can be limited by the cross-individual microbiome biodiversity, incomplete mechanistic knowledge, safety issues, and a deficit of regulatory standardization. The ultimate outcomes of microbiome-based therapeutics should be to provide precision, patient-specific treatments and exploit a favorable microbiome-host interaction associated with minimal risks. The development of more of these personalized treatments is likely to be boosted by improving the state of multi-omics, synthetic biology, and computational modeling. Although these approaches have much potential, they must address shortcomings, including stability, delivery, and differentiating causality and correlation in microbiome research.
In summary, host signaling regulated by the microbiome is a paradigm shift in therapeutic discovery. The strengths are numerous, with specific modulation, low invasiveness, and overall systemic effects. However, the method has to deal with a complicated regulatory and biological environment. Future studies of microbial effectors, host responses, and delivery platforms will be instrumental in achieving the potential of microbiome-integrated medicine.
Footnotes
Acknowledgements
The author(s) would like to thank the Department of Biochemistry & Microbiology, School of Health & Life Sciences at North South University, Dhaka, Bangladesh, for providing the resources and support necessary for this work.
ORCID iDs
Author contributions
M.S. led the literature search, citation management, introduction writing/editing, figure/table preparation, final formatting, and review. C.F.T.Z.H. handled the abstract, signaling overview, disease case studies, and final review. T.B.H. wrote the host response, therapy, and future direction sections. R.T. contributed microbiome and mechanism sections and assisted with formatting. J.F.O. finalized the conclusion, figures, and legends. K.M.M.R., S.M.B.U.I., N.B., J.S.J., MAS.S., and A.S. reviewed and revised the draft. M.D.F. conceptualized the manuscript. All authors approved the final version.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
