Abstract
Background
Killer immunoglobulin-like receptors (KIRs) are key molecules used by natural killer (NK) cells to interact with target cells. These receptors exhibit extensive genotypic polymorphism which has been associated with varying outcomes in immune responses against diseases. This study aimed to investigate the relationships between KIR genotypes and haplotypes with acute lymphoblastic leukemia (ALL) in Saudi patients.
Methods
A total of 259 Saudi subjects including 145 cases of acute lymphoblastic leukemia (ALL) and 114 healthy controls living in Riyadh were genotyped for 16 KIR genes and the two HLA-C1 and -C2 allotypes using PCR-SSP genotyping method.
Results
A significant high frequency of the two inhibitory KIR genes; 2DL1 (OR = 2.4; p < 0.0001) and 3DL1(OR = 10.87; p = 0.0068) in ALL compared to healthy group was observed. In contrast, the activating 2DS4 gene was significantly higher in healthy controls (OR = 0.15, p < 0.0001) compared to ALL patients. Haplotype analysis shows that BX haplogroup was strongly associated with the occurrence of ALL (OR = 4.39; p < 0.0001). Further combinatory analysis of KIR genes with their HLA-C1 and -C2 ligands demonstrated strong statistically protective effect of the 2DS1-C2 combination from ALL (OR = 0.06; p = 0.0003).
Conclusion
This study presents strong evidence supporting the connection between certain KIR genotypes, haplotypes, and KIR-HLA combinations with acute ALL in the Saudi population. The heightened occurrence of inhibitory KIR genes (2DL1 and 3DL1) and the BX haplotype in ALL patients indicates a possible involvement of these genetic variability with the dysfunctional of NK cells in the context of ALL disease.
Introduction
According to the International Agency for Research on Cancer, the top five cancers leading to mortality in 2020 were colorectal, breast, thyroid, non-Hodgkin lymphoma and leukemia. 1 In Saudi Arabia, leukemia was ranked as the 5th type of cancer cases among both genders with an overall prevalence of 7.6% in males and 4.4% in females. 2 According to the same source, leukemia appears as the 1st cancer disease among the most common types of cancers in children. In the Middle East, the Gulf Cooperation Council (GCC) reports that leukemia was classified 4th among the most common tumors in the GCC countries. Among the fourth common types of leukemia, acute lymphoblastic leukemia (ALL) is the most prevalent cancer in children worldwide. 3 ALL is characterized by a rapid proliferation of poorly differentiated lymphoid progenitor cells in the bone marrow. The incidence of ALL follows a bimodal allocation, the first peak occurring in childhood (between 2 and 5 years of age) and a second steady increase at about age 50 years. 4 In Saudi Arabia, the total incidence of childhood had increased from 1.58/100,000 in 2001 to 2.35/100,000 in 2014. 5
Although the cause in most cases of childhood leukemia is not yet known, a lot of factors including obesity, high birth weight, viral infections, prenatal and postnatal exposure to irradiation, toxin, tobacco and viral infection, sedentary life style and other environmental factors in a context of genetic background has been reported to predispose to this disease.6–8
Natural killer cells (NK cells) are innate immune cells of the lymphoid lineage that mediate antitumor and antiviral responses.9–11 NK cells play a pivotal role in inhibiting early tumor formation and promoting anti-tumor immune responses through cytotoxicity and cytokine production.9,10 The killing activity of NK cells is regulated by cytokines and a number of expressed transmembrane receptors from various families. The killer cell immunoglobulin-like receptor (KIR) family is one of the most polymorphic receptors of the NK cells that mediate cytotoxicity by interacting with opposing signals to trigger or inhibit cytotoxic activity.12,13 Individuals with impaired NK cell activities exhibit a notable incidence of malignancy`. 14 Their effector effects are regulated by a balance of signals from inhibitory and activating receptors, such as KIRs, which recognize self or modified HLA class I ligands on target cells. 15 The KIR locus is composed of 16 genes and pseudogenes spanning ∼10–16 kb in chromosome 19q13.4 within the leukocyte receptor complex. 16 The nomenclature of KIR genes is based on the number of extracellular Ig-like domains, which can be either 2 (2D) or 3 (3D), and on the length of the cytoplasmic tail, which can be long (L) or short (S). This nomenclature also reveals the function of the receptor that can be inhibitory or activating, respectively.
Thus, the inhibitory genes were named KIR2DL1, KIR2DL2, KIR2DL3, KIR2DL5, KIR3DL1, KIR3DL2, and KIR3DL3. While the activating genes were identified as KIR2DS1, KIR2DS2, KIR2DS3, KIR2DS4, KIR2DS5, KIR2DS5 and KIR3DS1. An exception for the KIR2DL4 which was reported to act as inhibitory or activating gene. Two pseudogenes were also characterized; KIR2DP2 and KIR3DP1.13,17 KIR3DL3, KIR3DP1, KIR2DL4, and KIR3DL2 are always present in the genome and are so-called, framework genes. 18
Based on the gene content, two types of KIR haplotypes have been identified, named A and B haplotypes. There is no single criterion that separates all A and B haplotypes, however, one or more of the following genes characterize group B haplotypes: KIR2DL5, KIR2DS1, KIR2DS2, KIR2DS3, KIR2DS5, and KIR3DS1. On the other hand, group A haplotypes are defined by the lack of all these genes.19,20 Consequently, the number of the B haplotypes is quite greater than the A haplotypes.
KIRs allows NK cells to identify target cells mainly through HLA class I, including HLA-C and HLA-B. 21 HLA-C molecules class I ligand was classified into two allotypes depending on the amino acid at position 80 in the α1 domain. 22 Thus, HLA-C1 allotype is carrying asparagine and the -C2 allotype is carrying lysine at this position. The recognition of HLA-C1 is mediated by the inhibitory KIR2DL2 and KIR2DL3 and the activating KIR2DS2 receptor, while the inhibitory KIR2DL1 and the activating KIR2DS1 receptors bind to the HLA-C2 group.18,23,24 Malignant or infected cells use a variety of strategies to evade the host's immune response, including downregulation of MHC-I molecule expression. 25
The number and type of KIR genes are variable between haplotypes, which diversify humans in their KIR gene content. KIR genotyping has been increasingly used for epidemiological studies that showed links between some KIR genes and the risk of developing certain human diseases. Numerous studies have confirmed the relationships between KIR genes polymorphisms and HLA–KIR gene combinations with diseases such as cancer, autoimmune disease, viral infections and so on.17,26–28 The aim of this study is to assess the relationships between KIR genotype and haplotypes with ALL in a Saudi population. Unveiling KIR and HLA class I gene polymorphisms that regulate the NK cell repertoire in patients with ALL could help in optimizing hematopoietic stem-cell donor selection and eventually NK-cell-based immunotherapies.
Materials and methods
Study subjects and blood collection
This study included 145 pediatric patients (Males = 87 and Females = 58) diagnosed with ALL between April 2017 until March 2020. The majority of patients were recruited from the pediatric hematology-oncology unit of Department of Pediatrics, King Khaled Hospital University (KKHU), Riyadh, Saudi Arabia. The health control (HC) group consisted of 114 ethnically matched, unrelated healthy volunteers (Males = 77 and Females 38) without any history for cancer or immunological diseases. The average age of ALL patients was 19.20 ± 14.8 years, whereas the average age of HC was 22.45 ± 20.18 years. Three ml of peripheral blood samples were collected from both patients and controls in EDTA-coated vials and stored at −80°C until analysis.
Ethics statement
This study was approved by the ethics committee of King Khaled Hospital University (KKHU) in Riyadh, Saudi Arabia (IRB approval number: E-20-5346). Written informed consent was obtained from all participants or their legal guardians prior to inclusion in the study. The study was conducted according to the ethical guidelines of King Saud University and the principles of the Declaration of Helsinki:
Genomic DNA extraction and KIR genotyping
Genomic DNA was extracted from peripheral blood samples using a genomic DNA purification kit (Applied Biosystems, Thermo Fisher Scientific, USA) according to the manufacturer's instructions. DNA concentration and purity were measured using a NanoDrop 8000 spectrophotometer (Thermo Scientific, Wilmington, DE, USA). Genotyping of 16 KIR genes and HLA-C1/C2 allotypes was performed using duplex sequence-specific primer polymerase chain reaction (SSP-PCR) according to previously described protocols.29,30 Negative control and positive control with known genotypes were included in each PCR reaction. KIR genotypes of randomly selected samples were confirmed using a commercial KIR genotyping kit (Miltenyi Biotec, Inc, Germany). PCR was performed on a Bio-Rad T100 thermocycler. Amplicons were resolved via electrophoresis on 2% agarose gels stained with ethidium bromide and visualized using a BioDocAnalyze (BDA) Gel Analysis system (Biometra GmbH, Germany).
HLA-C ligand genotyping
All patients and controls individuals were genotyped for the for HLA-C1 and HLA-C2 allotypes suing the same primers and protocol as reported before by Tajik et al. 30 A negative PCR control and positive control samples for each HLA-C allotype were used in each amplification.
Statistical analysis
KIR genotypes were identified based on the Allele Frequency Net database (AFND; https://www.allelefrequencies.net/kir6002a.asp accessed in February 2024). 31 Frequencies of KIR genes, KIR AA and BX genotypes, and HLA-C ligands were calculated for the patient and control groups. Statistical analyses were performed using SigmaPlot v11 statistical tools. Associations were analyzed using X2 and Fisher's exact tests with p-values adjusted for multiple comparisons via Holm-Bonferroni correction. Odds ratios and 95% confidence intervals were computed using a logistic regression model to evaluate associations between KIR genotypes, ligand combinations and ALL risk.
Results
Distribution of KIR genotypes in ALL and healthy groups
Sixteen KIR genes comprising 5 activating, 8 inhibitory genes and 2 pseudogenes were genotyped in ALL patients and HC (Table 1). Frequency of individual KIR genes were assessed in both groups. The framework KIR genes (3DL2, 3DL3, 2DL4 and 3DP1) were present in all individuals (Table 1).
Distribution of the KIR genes in ALL patients and healthy controls.

KIR: killer immunoglobulin-like receptor; HC: healthy controls; ALL: acute lymphocyte leukemia; N: number samples; X2: Pearson's chi-squared test; OR: odds ratio; CI: confidence interval; NA not applicable.
**Significant associations after Bonferroni correction are shown in bold (*significance was lost after correction).
When comparing ALL patients with controls, significant differences were found in the frequency of two inhibitory KIR genes (2DL1 and 3DL1) and one activating KIR gene (2DS4). In ALL cases, a high increase in the occurrence of the inhibitory KIR genes 2DL1 (OR = 2.4, p < 0.001) and 3DL1 (OR = 10.87, p = 0.0068) was observed. Conversely, the activating KIR-2DS4 gene was highly increased in HC compared to ALL cases (OR = 0.15, p < 0.001), suggesting a protective effect against ALL. The frequencies of KIR 2DS3, 2DS5 and 3DS1 genes were slightly higher in ALL patients compared to HC but the differences were not significant.
Comparative analysis based on the number of activating and inhibitory KIR genes
Comparison of the frequencies of total activating (aKIR) and inhibitory (iKIR) KIR genes between ALL patients and controls showed that HC had a slightly higher frequency of one aKIR compared to patients (OR = 0.52; 95% CI = 0.30–0.92; p = 0.03), but this association was not significant after Bonferroni correction. Conversely, patients with two aKIRs were significantly more frequent than controls (OR = 2.32; 95% CI = 1.25–4.30; p = 0.008). However, there were no statistically significant differences in aKIR frequencies ranging from 3–6 aKIRs (Table 2). Analysis of iKIR frequency showed that individuals with six iKIRs were more frequent in HC compared to ALL patients (OR = 0.48; 95% CI = 0.27–0.84; p = 0.01), but this association was also not significant after Bonferroni correction (Table 2).
Comparative distribution of activating KIR genes (aKIR) and inhibitory KIR genes (iKIR), between ALL patients and healthy control.
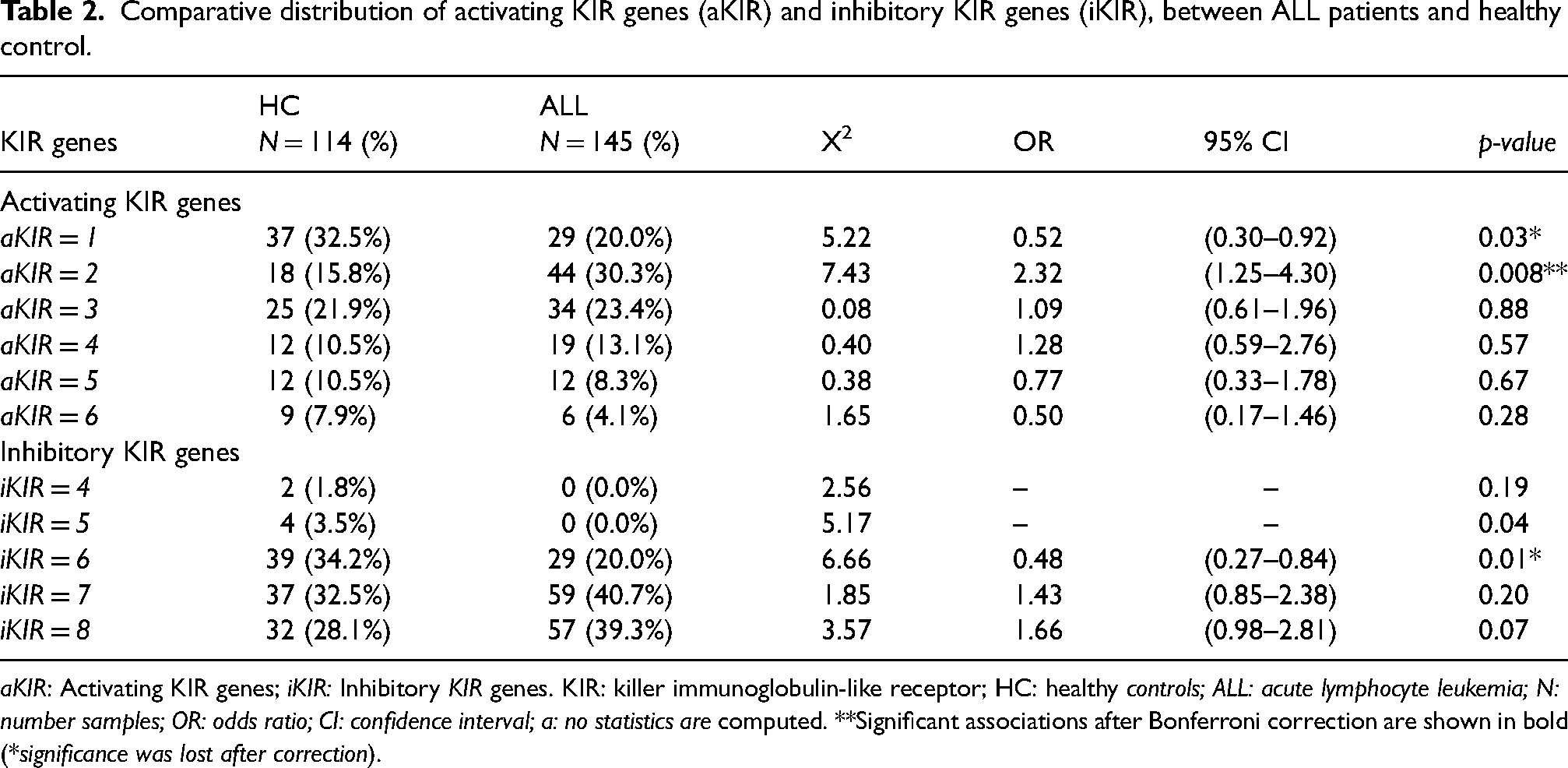
aKIR: Activating KIR genes; iKIR: Inhibitory KIR genes. KIR: killer immunoglobulin-like receptor; HC: healthy controls; ALL: acute lymphocyte leukemia; N: number samples; OR: odds ratio; CI: confidence interval; a: no statistics are computed. **Significant associations after Bonferroni correction are shown in bold (*significance was lost after correction).
Distribution of HLA-C genotypes in ALL patients compared to HC
Comparative analysis of HLA-C1 and HLA-C2 distributions is shown in Table 3. The five genetic models (codominant, dominant, recessive, overdominant, and log-additive) tested for association between HLA-C1/C2 polymorphism and ALL risk showed no significant association with disease for all tested models (Table 3).
Genotypic association of HLA-C C1/C2 alleles polymorphisms with ALL following, dominant, recessive, overdominant, and log-additive genetic models.

HC: healthy control; ALL: acute lymphoblastic leukemia; OR: odds ratio; CI: confidence interval; AIC: Akaike information criterion; BIC: Bayesian information criterion.
Combinatory analysis of the distribution of KIR genes and their HLA-C1 and C2 ligands frequency in ALL patients and HC
The combined correlation of the KIR genes and HLA-C ligands with ALL is reported in Table 4. The 2DL3, 2DS2, 2DS1, and 2DL2 KIR genes were found to be associated with various HLA-C ligands. It is worthy to notice that the 2DL3 and 2DS2 in the absence of their C1 ligand showed a risk for ALL (2DL3: OR = 1.84; 95% CI = 1.05–3.20; p = 0.04) and (2DS2: OR = 2.02; 95% CI = 1.03–3.93; p = 0.04). On the other hand the 2DS1 KIR gene showed a high protective effect from ALL in the absence of its C2 ligands, (OR = 0.06; 95% CI = 0.01–0.46; p = 0.0003).
Combinatory analysis of KIR genes in the presence and absence of their C1/C2 ligands between ALL patients and healthy controls.

KIR: killer immunoglobulin-like receptor; HLA: human leukocyte antigen; C+: allele C present; C−: allele C absent; HC: healthy controls; ALL: acute lymphoblastic leukemia; N: number samples; X2: Pearson's chi-squared test; OR: odds ratio; CI: confidence interval; NS: not significant. *Significant associations after Bonferroni correction are shown in bold (*p ≤ 0.05, **p ≤ 0.005 and ***p ≤ 0.0005).
In the presence of their C1 ligand, the combination of KIR 2DL2+/DS2-/HLA-C1 + showed a risk effect for ALL (OR = 3.34; 95% CI = 1.39–8.01, p = 0.005). However, KIR 2DL2-/2DS2-/HLA-C1 + combination showed a protective effect (OR = 0.32; CI = 0.15–0.864; p = 0.002).
Comparative analysis of KIR haplotypes in ALL patients and HC
Based on the gene content, a total of 50 different genotypes were observed, including 6 AA and 44 Bx group genotypes (Figure 1). In healthy group there are 31 genotypes versus 32 genotypes in ALL individuals. Only 14 genotypes are common between the two groups. The most frequent genotype was the AA ID1 in control group (21.04%) and was the second most frequent genotype in ALL group (6.2%). Thus, a significant difference in AA ID1 genotype was found between the two groups (p = 0.00056). The Bx ID110 was the most frequent genotype in ALL patients and was missing in HC (p = 0.00009) (Figure 1).

Main KIR gene content diversity among patients with ALL and healthy controls. Gene contents are shown by presence (shaded boxes) or absence (white boxes) of 16 KIR genes. For each genotype ID, the haplotype associated groups (AA and Bx), and the genotype subset are shown. Genotypes were obtained from Allele Frequency Net Database (https://www.allelefrequencies.net/kir6002a.asp, accessed in February 2024).
The frequencies of KIR haplotypes among ALL and HC are reported in Table 5. Group A haplotype is less represented in ALL (32.1%) than in HC (43.9%). The frequency of Group A haplotype is lower in the ALL group (32.1%) compared to the HC group (43.9%). In contrast, the occurrence of group B haplotype was notably greater in ALL patients (67.9%) compared to HCs (56.1%). The analysis of KIR genotypes revealed a substantial drop in the presence of the AA genotype in patients with ALL (6.9%) compared to HC (24.56%). Nevertheless, there were no notable disparities in the frequencies of AB and BB genotypes between ALL patients and HC.
Comparison of haplotypes genotypes, and linkage groups between ALL and control groups.
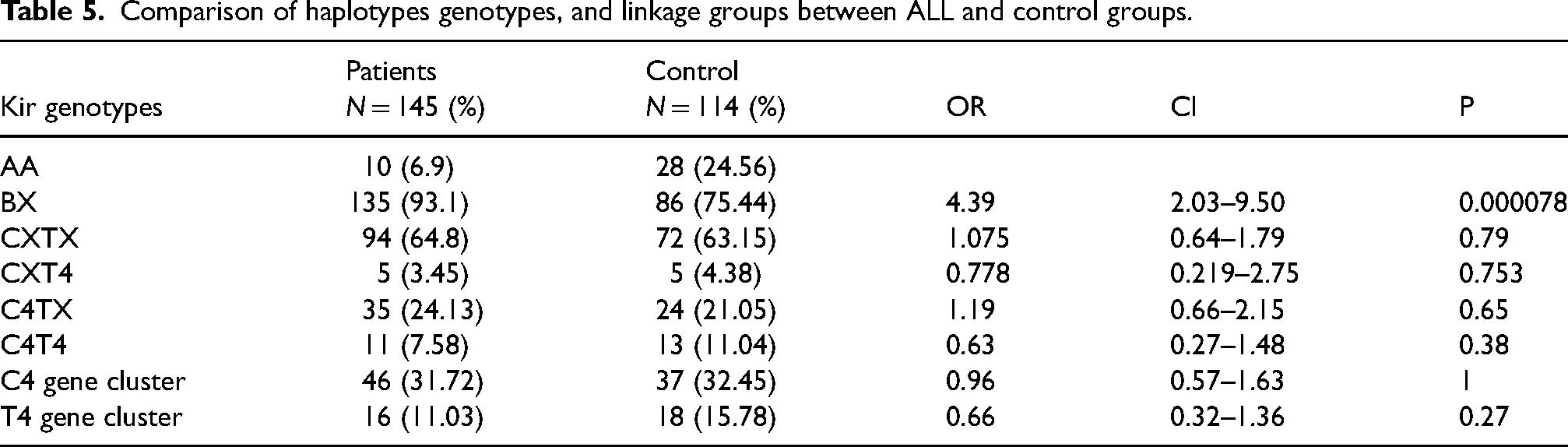
Furthermore, the four examined Bx subsets (C4T4, C4Tx, CxT4, CxTx) (x can be either an A or B haplotype) were compared among the different groups. No significant difference was observed between the healthy control group and ALL cases. The C4T4 variable was slightly higher among HC (11.04%) compared to ALL group (7.58%) but the difference did not reach significance.
Discussion
In the present study we have examined the genetic distribution of 16 KIR genes and their HLA-C ligands among Saudi patients with acute lymphoblastic leukemia (ALL) and a control group. Significant higher occurrence of inhibitory 2DL1 and 3DL1 KIR genes in ALL patients was observed, indicating that these genes might contribute to an elevated risk for ALL by suppressing the immune system's ability to detect and eliminate leukemic cells. Conversely, a notably lower occurrence of the activating 2DS4 KIR gene in ALL patients suggests a potential protective effect. In addition, the Bx haplotype was found to be strongly associated with occurrence of ALL. Some studies have been conducted on the correlation between KIR genes and hematological malignancy in some populations.32–38 Despite this limited number of published studies, conflicting results have been noted. Recently, Halimi et al. reported in Iranian population an increased frequency of 3DS1 KIR gene in patients with ALL and lower frequency of the inhibitory combinations 2DL1
Additionally, our findings showed no statistically significant differences in the HLA-C1 and C2 genotypes between patients with ALL and HC. However, combinatory analysis of the distributions of the KIR genes with their C1/C2 ligands showed that 2DL2+/2DS2-/-C1 + and 2DL2/2DS2-HLA-C1 + and 2DL2-/2DS2-/C1 + and C2 are associated with the occurrence of ALL. Inversely, the combination 2DS1 + C2 was found to protect from ALL. The mechanisms underlying HLA-C1 allotype recognition by 2DL2 and 2DL3 are different. 40 Similarly, HLA-C1 allotypes vary significantly in their ability to suppress primary NK cell activation. These functional variations are attributed in part to 2DS2 KIR molecule, suggesting that the 2DL2 and 2DL3 KIR molecule binding geometries work in tandem with other parameters to differentiate HLA-C1 functional recognition. 40 Hirayasu, Ohashi 41 found a significant association between 2DL3-HLA-C1 and cerebral malaria, and implications for KIR and HLA co-evolution. Knapp, Warshow 42 reported consistent beneficial effects of 2DL3 and HLA-C1 following exposure to the hepatitis C virus. Zuo, Yu 43 found that the interaction between HLA-C1 and 2DL2/L3 increases the incidence of acute graft-versus-host disease (aGVHD) after hematopoietic stem cell transplantation. This was caused by the promotion of 2DL2/L3 single-positive/NKG2C-positive NK cell reconstitution. 43 2DS1 receptor, through its recognition of HLA-C2 ligand can contribute to both the activation and tolerance of NK cells. 44 When This activating gene interacts with high levels of HLA-C2 in people with the HLA-C2/C2 genotype, the NK cells become less reactive. 45 Thus, this could explain the protective effect of this gene when its cognate receptor is missing. Considering the haplotype profiles, our results showed that the frequency of the KIR AA genotype was significantly higher in healthy controls than in patients with ALL. This result is in agreement with that reported by Verheyden et al. on Belgian Caucasians with chronic myeloid leukemia. 46 However, these results contrasted with that reported by de Smith et al., which found a significant association between ALL cases and AA haplotype in non-Hispanic white subjects. 33
When comparing the A and B haplotypes, the B haplotype was more common in ALL cases. Individuals with Bx genotypes are more prone to developing ALL than those with AA genotypes. These results are agreeing with those conducted by Misra et al. on ALL among North Indians. 34
In conclusion, this study provides compelling evidence of the significant role that specific KIR genes play in the outcome of ALL within the Saudi population highlighting a strong association between, the B haplotype and increased risk for ALL. To further elucidate the intricate relationship between KIR genotype diversity and ALL, future studies should focus on larger, more comprehensive cohort analyses that explore the NK receptor expression profiles and their impact on disease progression and treatment responses.
Footnotes
Acknowledgments
The authors extend their appreciation to the Researchers Supporting Project number (RSP 2025R75), King Saudi University, Riyadh, Saudi Arabia
Data availability
All data relevant to the study are included in the article
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Informed consent statement
Informed consent was obtained from all subjects involved in the study.
Institutional review board statement
the study was conducted according to the guidelines of the Declaration of Helsinki, and approved by the Institutional Review Board of the Ethic Committee of King Saud University under the number, 20-0795.
