Abstract
Vaping is an increasing health threat in the US and worldwide. The damaging impact of vaping on the human distal lung has been highlighted by the recent epidemic of electronic cigarette or vaping use-associated lung injury (EVALI). The pathogenesis of EVALI remains incompletely understood, due to a paucity of models that recapitulate the structural and functional complexity of the human distal lung and the still poorly defined culprit exposures to vaping products and respiratory viral infections. Our aim was to establish the feasibility of using single cell RNA-sequencing (scRNA-seq) technology in human precision-cut lung slices (PCLS) as a more physiologically relevant model to better understand how vaping regulates the antiviral and pro-inflammatory response to influenza A virus infection. Normal healthy donor PCLS were treated with vaping extract and influenza A viruses for scRNA-seq analysis. Vaping extract augmented host antiviral and pro-inflammatory responses in structural cells such as lung epithelial cells and fibroblasts, as well as in immune cells such as macrophages and monocytes. Our findings suggest that human distal lung slice model is useful to study the heterogeneous responses of immune and structural cells under EVALI conditions, such as vaping and respiratory viral infection.
Introduction
The use of electronic cigarette (e-cig) product or vaping in the general population especially in the youth has steadily risen in the past decade. 1 Research using the human airway epithelial cell culture and animal models suggests the detrimental health effects of vaping including more severe respiratory virus infection such as influenza A virus (IAV).2–4 The distal lung is the major site of lung injury, inflammation and tissue remodeling or destruction in chronic obstructive pulmonary disease (COPD) associated with cigarette smoke exposures. We recently studied the impact of vaping on human distal airway epithelium, and found that vaping increased human small airway epithelial pro-inflammatory responses to IAV infection. 5 However, the current research models have limitations in mimicking the heterogeneous and simultaneous responses of various immune and structural types of cells residing in the human lung. Recently, human precision-cut lung slices (PCLS) have been used to study the health effect of various environmental factors such as tobacco smoke and respiratory viruses.6,7 PCLS contain all the types of lung cells in situ and keeps the cell-cell interactions intact. The 3-dimensional nature of PCLS culture coupled with its architectural integrity and high fidelity to mimic human distal lung responses to various environmental hazardous factors makes it a unique and highly physiologically relevant model to study the effects of vaping. 8 The goal of this report was to determine the effect of vaping on human lung transcriptomic responses to IAV infection at the single cell level. To achieve this goal, we applied the single cell RNA-sequencing (scRNA-seq) technology to profile transcriptomic changes of genes involved in host responses to vaping and IAV in human PCLS.
Materials and methods
Precision-cut lung slice (PCLS) processing
The upper lobe of the right lung from a healthy, non-smoking donor (male, 51 years old) was obtained from the Colorado Donor Alliance. The Institutional Review Board (IRB) at National Jewish Health, Denver, Colorado approved our work as nonhuman subject research. The lung lobe was inflated with 1.5% low-melting agarose (42°C), cooled on ice, cut into tissue cores with a diameter of about 1.5 cm. and then sliced into consecutive 450 µm thickness sections using a Compresstome® VF-300 vibratome (Precisionary Instruments, Boston, MA, USA). The slices were transferred to 24-well plates filled with Dulbecco's Modified Eagle Medium (DMEM) (0.5 ml/well) supplemented with penicillin, streptomycin and amphotericin.
Culture of human PCLS
JUUL Labs (Washington D.C.) Virginia tobacco flavor at 3% nicotine strength was used for this study. JUUL e-vapor was generated using a JUULbattery and JUULpod attached to tubing on a MasterFlex L/S Economy Variable Speed Drive.9,10 Vaping extract (VE) was prepared by bubbling e-cig vapor for 15 min into 25 ml tissue culture medium (DMEM plus antibiotics and antifungal agents), resulting in 100% VE. After passing through a 0.22 µm filter, 100% VE was diluted to 10% VE with DMEM for tissue culture. The final nicotine concentration in 10% VE was 17.5 µg/ml, a concentration within the range of airway nicotine concentrations in human vapers where serum nicotine concentrations were determined to be about 1000 times lower than those in the airway epithelial lining fluid.11–13 Pandemic A/California/04/2009 (CA04) IAV (3 × 105 pfu/slice) and 10% VE were added to PLCS in 24-well plates with DMEM media. PCLS were removed after 72 h of IAV infection. For single cell isolation, two pieces of lung slices from each treatment condition were transferred into a 1.5 ml tube filled with 500 µl of digestion solution containing collagenase (2 mg/ml), DNase I (200 µg/ml), RNase inhibitor (1 µl/ml), and DMEM with penicillin and streptomycin. After incubation at 37°C for one hour, lung slices were processed by gently pushing it through a tissue metal sieve over a dish. The flow-through cells were centrifuged at 4°C for 5 min. The resulting cell pellets were resuspended in 1 ml of Trypsin/EDTA and incubate at 37°C for 5 min and then passed through 70 µm pore size cell strainers to remove mucus and cell aggregates. After centrifugation at 4°C for 5 min, cells were resuspended in 50 µl of PBS with an RNase inhibitor (40 U/ml). An aliquot (5 µl) of cells was used for staining with trypan blue to count cells and assess viability and single cell distribution.
Single cell RNA-Sequencing (scRNA-Seq)
Single cells from PCLS were processed for scRNA-seq using the 10x Genomics Chromium Controller. 14 Cells were captured using the 10X Chromium solution, reversed transcribed, bar-coded for each cell, and processed to build cDNA libraries using the 10x Genomics Chromium Next GEM 3′ Single Cell Reagent kits v3.1 for sequencing on an Illumina sequencer with approximately 300 million read pairs per sample.
Single cell data analysis and cell type annotations
The sequencing reads were processed using 10x Genomics Cell Ranger analytical pipeline (v6.1.2) and human GRCh38 reference. Dataset aggregation was performed using the cellranger aggr function and then normalized for the total number of confidently mapped reads across libraries to enable comparisons between cell types under different conditions.
The resulting gene expression count matrix was used as the input for Seurat 4.0 and Azimuth (R programs, R version 4.2.1) for reference-based cell type annotations. 15 First, we further cleaned the data, excluding cells with >10% of unique molecular identifiers (UMIs) associated with mitochondrial genes, with <750 UMI counts, or with <250 or >10,000 expressed genes. We then used the SCTransform function16,17 in Seurat to pre-process the filtered dataset, followed by finding anchors between our input dataset and the Human Lung Cell Atlas v2 reference 18 and label transfer of cell type annotations. The Human Lung Cell Atlas v2 reference was built by integrating data from 107 individuals from 14 datasets. The cells in the integrated atlas were re-annotated based on originally published labels and inputs from 6 experts, resulting in consensus on 58 cell identities in healthy human lungs, along with robust marker genes. The reference has multiple classification levels of cell annotations. The default level 3 was used for annotations for all cell types except for T cells, for which we adopted the finer level 4 classification that further separates CD4 and CD8 positive T cells. The markers used are illustrated in Table 1. To ensure high-quality results, we kept only the cells with high-confidence cell-type annotations (prediction score > 0.75 when mapped to the level 3 reference) supported by multiple consistent anchors during reference mapping. The resulting 17,175 high-quality cells from all 4 treatment conditions were used in subsequent differential expression and pathway analysis.
Gene markers for human lung cells.

Uniform manifold approximation and projection (UMAP) visualization
We used the cellranger reanalyze function to generate the feature-barcode matrix and the cloupe file for the 17,175 high-quality cells from the original aggregate sample generated by the cellranger aggr function described above. UMAP was created by sample integration with Harmony, 19 an algorithm (R program) that projects cells into a shared embedding in which cells were grouped by cell types rather than dataset-specific conditions. Specifically, we processed the feature-barcode matrix of the 17,175 cells with Seurat 4.0 (NormalizeData, FindVariableFeatures using the top 2000 variable genes, ScaleData, Run PCA with npcs = 20) and conducted Harmony integration. We ran UMAP (reduction = ”harmony”, dims = 1:20), FindNeighbors (reduction = ”harmony”, dims = 1:20), and FindClusters (resolution = 0.5) for clustering.
Differential expression analysis
The differential expression analysis was done for the 17,175 high-quality cells using the “Locally Distinguishing” significant feature comparison and the calculator tool of the 10x Genomics Loupe Browser (v6.4.1). We compared the differential gene expression of the following cell types (AT1, AT2, airway basal epithelial cells, vascular endothelial cells, fibroblasts, mast cells, macrophages/monocytes, CD8 T cells and CD4 T cells) between VE + IAV and IAV samples and between VE and control (-) samples. We did not do differential expression analysis for ciliated epithelial cells because there was a significant reduction in the number of ciliated epithelial cells in the VE + IAV sample. To test for differences in mean expression between groups of cells, Loupe Browser uses the exact negative binomial test of the sSeq method. 20 When the counts become large, Loupe Browser switches to the fast asymptotic negative binomial test used in edgeR. 21 For each comparison, we obtained the log2 fold change and adjusted p-value (adjusted using the Benjamini Hochberg correction for multiple tests) for all 32,603 genes.
Pathway enrichment analysis
The differentially expressed genes between conditions for each cell type that met the criteria, adjusted p-value <0.05, were used as input for KEGG enrichment analysis using the enrichKEGG function in the clusterProfiler R program. 22 There were too few differentially expressed genes in mast cells between conditions, so mast cells were not included in the pathway analysis. We analyzed up-regulated and down-regulated gene lists separately for each of the 8 cell types. In the pathway summary plots for all comparisons, the pathways with the lowest 20 adjusted p-values that met the criteria, adjusted p-values <0.05, for each cell type were added to the summary plots.
Results
Cell viability and counts in human PCLS
PCLS obtained from a normal human donor lung were incubated with VE and/or IAV for 3 days, and then processed for single cell isolation. Cytospin slides made from human PCLS showed that single cells were successfully obtained (Figure 1). As shown in Figure 2, more than 105 cells per condition were obtained. After 72 h of viral infection, viability of isolated cells was above 90% and was similar across all the 4 conditions.

Single cells isolated from a human precision-cut lung slice (hPCLS) were centrifuged onto a cytospin slide stained with Diff-Quick solution. Cells shown in the figure included ciliated epithelial cells, macrophages and mononuclear cells (monocytes).

Cell count (
Cell composition in human PCLS by scRNA-Seq analysis
The identity of cells was determined based on the scRNA-seq cell markers (Table 1) as described in the Methods section. 18 Using this tool, we found 16 identifiable cell populations under 4 culture conditions, shown in the UMAP plots (Figure 3). We observed that the cell types annotated by Azimuth matched the clusters from Harmony integration very well. Using the total number of cells that met the criteria for scRNA-seq data analysis under each experimental condition (control [–], n = 4502 cells; VE, n = 4192 cells; IAV, n = 3714 cells and VE + IAV, n = 4767 cells), we noted that vaping extract (VE) exposure and IAV infection increased the % of alveolar type 1 (AT1) cells, while decreasing the % of AT2 cells, airway basal and ciliated epithelial cells, as well as macrophages/monocytes compared to other three conditions (Figure 4). The % of T cells and structural cells (vascular endothelial cells, fibroblasts and smooth muscle cells) was similar among the 4 treatment conditions.

UMAP showing distribution of 14 different types of cells in human precision-cut lung slice (hPCLS) under 4 treatment conditions.
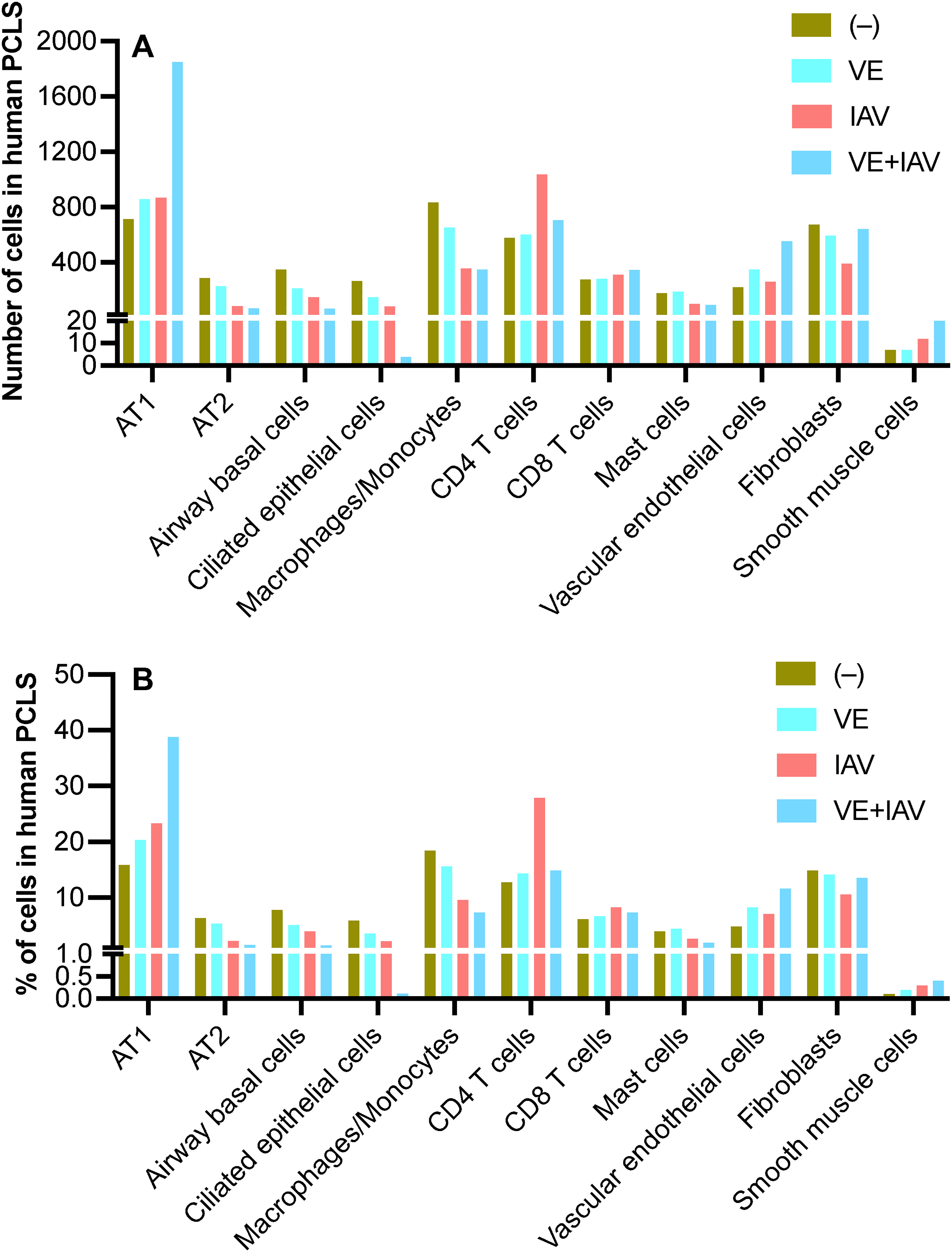
Cell counts analyzed by scRNA-Seq in human precision-cut lung slice (PCLS). (
scRNA-Seq data focusing on the effect of VE on IAV-mediated antiviral and pro-inflammatory responses
All the genes that were significantly (adjusted p < 0.05) up-regulated or down-regulated by VE in the presence or absence of IAV infection in the following 8 different cell types were provided in

Volcano plots showing up- and down-regulated genes by vaping extract (VE) and influenza A virus (IAV) as compared to IAV alone in alveolar type 1 (AT1) epithelial cells (

KEGG pathway analysis showing up-regulated genes in human precision-cut lung slice (PCLS) treated for 72 h with vaping extract (VE) and influenza A virus (IAV) as compared to IAV alone.

KEGG pathway analysis showing down-regulated genes in human precision-cut lung slice (PCLS) treated for 72 h with vaping extract (VE) and influenza A virus (IAV) as compared to IAV alone.
There were 945 genes significantly up-regulated, and 1458 genes down-regulated by VE in IAV-infected cells (Figure 5(b)). Multiple antiviral and pro-inflammatory molecules were increased by VE in IAV-infected basal cells. These include IFNL1, 2 and 3, IFI6, 27 and 35, CXCL1, 6, 8, 9 and 17, IL-6 and GDF15. Notably, S100A4, a pro-inflammatory mediator, 31 was also significantly up-regulated by VE in IAV-infected cells. Several pathways including oxidative stress were affected by VE and IAV. Some of the down-regulated genes are related to RNA degradation and autophagy. VE alone increased the expression of chemokines CCL2 and CXCL2, but decreased the anti-inflammatory mediator TGF-β2, and antimicrobial substance SLPI.
There were 48 genes significantly up-regulated, and 277 genes down-regulated by VE in alveolar macrophages and lung monocytes isolated from IAV-infected PCLS. The up-regulated genes are related to viral infection, chemokine and Toll-like receptor signaling, including IFI6, CCL4, CCL4L1 and CCL4L2, CXCL9 and TNF. VE alone had a minimal effect on the inflammatory gene expression.
Discussion
This report represents the first study combining human PCLS and scRNA-seq approaches to unravel the direct effects of vaping on transcriptomic changes of different types of cells in the lung infected with respiratory viruses. Our results suggest the feasibility of this innovative research approach in the human lung exposed to vaping products and viral infection. Importantly, we were able to successfully infect the PCLS with IAV as the viruses robustly increased antiviral (e.g., IFNL, IFI6, ISG15) and pro-inflammatory (e.g., CXCL10) genes in epithelial cells and immune cells, which is consistent with the known effects of IAV infection in cultured human lung epithelial cells and in mouse models.32,33 Importantly, cells isolated from the lung treated with IAV alone or both IAV and VE for 3 days had viability of over 90%, suggesting that PCLS is a viable approach to study the vaping health effect in human lungs during respiratory viral infection. The current study did not extend the viral infection to more than 3 days. However, given the ability of PCLS to survive in culture media at least for 15 days, 34 future studies can be performed to evaluate the long-term effect of vaping on viral infection. With our model system, it might also be possible to investigate the vaping effect on transcriptomic changes of infection with other respiratory viruses such as SARS-CoV-2 and rhinovirus that are commonly seen in patients with various lung diseases.
There are several interesting findings from this study. Airway epithelial cells appear to be more vulnerable to IAV infection with or without e-cig exposure than alveolar type 1 epithelial cells as the % of airway epithelial cells particularly ciliated cells was reduced by more than 2-fold. While we did not determine the underlying mechanisms of significant reduction of airway epithelial cells in PCLS treated with IAV alone and particularly with VE + IAV, a previous airway epithelial cell culture 35 and our own observation in cultured human airway epithelial cells at air-liquid interface suggest that IAV infection may result in the loss of airway epithelial cells especially ciliated cells. We found that in PCLS treated with both VE and IAV, there was an increased % of AT1 cells. It is possible that AT1 cells become overrepresented as a result of death of airway epithelial cells and AT2 cells, which are highly susceptible to IAV infection/replication.36–38 In our recent publication, 39 we also found that e-cig exposure significantly increased the release of lactate dehydrogenase (LDH) in IAV-infected human PCLS. Vaping similarly amplified IFN responsive genes in airway epithelial cells, alveolar epithelial cells, and macrophages/monocytes. Nonetheless, AT1 cells appeared to have a more robust antiviral pro-inflammatory response than AT2 cells after 72 h of VE and IAV treatment. Due to small number of ciliated cells, goblet cells and club cells in PCLS treated with both VE and IAV, we were not able to determine if they responded differently from airway epithelial basal cells to VE and IAV. Interestingly, although fibroblasts are considered as a typical type of structural cells, they responded to both VE and IAV in a similar manner to epithelial cells and macrophages/monocytes. Moreover, fibroblasts increased the expression of genes contributing to smooth muscle contraction and fibrosis following VE and IAV, suggesting the potential of VE exposure to induce distal lung tissue remodeling and airway obstruction during IAV infection. There was a minimal impact of VE (versus control) on gene differential expression and enriched pathways in almost all cell types. This suggests a minimal effect of VE alone on lung immune response. Our data along with previous studies using other experimental models suggest that vaping is less cytotoxic and has a weaker pro-inflammatory response than tobacco smoke in human lung cells.40,41 We found that e-cig exposure in PCLS did not increase the release of LDH at 24 h and 48 h time points, but moderately increase the LDH release at 72 h (p = 0.08). 39 Thus, the 72 h chosen in the current study should not have missed the peak effect of VE treatment alone. One of the reasons for the minimal effect of VE alone on the pro-inflammatory effect may be related to less toxic chemicals contained in VE as compared to tobacco (regular) cigarettes.42–44
There are several limitations to our study. First, we only included one human donor in this pilot study, requiring validation in future experiments. Second, although we focused on the effects of vaping and viral infection on the expression of genes representing the IFN or antiviral and pro-inflammatory responses, two most robust pathways regulated by viral infection, our pathway analysis also demonstrated the involvement of other pathways such as ribosomal signaling, oxidative stress, cell adhesion pathways. The potential damaging effect of vaping on lung structure and function needs to be further investigated. Third, human lungs contain more than 40 different types of cells. 45 While our study focused on some of the most abundant types of cells in the lung, due to limited number of minor types of cells such as ionocytes, the transcriptomic responses of those minor types of cells to vaping and viral infection were not revealed in this study. Fourth, Unlike the fresh non-cultured lung tissue, some short-lived cells such as neutrophils may not survive during the culture of PCLS and thus the transcriptomic profiling of these cells was not available. Fifth, our study did not measure the viral load in PCLS or cultured supernatants. Nonetheless, our recent publication 39 demonstrated that e-cig exposure significantly increased IAV levels in the tissue as well as in the supernatants. Finally, the transcriptomic changes detected by scRNA-seq analysis has not been confirmed by in situ hybridization or spatial transcriptomic analysis or by immunostaining of protein expression.
In summary, our pilot study may have provided a new physiologically relevant ex vivo human lung tissue culture model to study the effect of vaping on human health. It also provides a novel understanding at a single cell level of the effect of e-cig exposure in modulating lung immune responses to viral infection in a complex multicellular lung environment. Evaluation of responses of different types of lung immune and structural cells in this model helps us further understand how cell-cell interactions may play a role in the pathogenesis of vaping-related lung diseases. This report may enable the research community to further determine the role of vaping and other environmental risk factors such as tobacco smoke and air pollutants in modulating human distal lung response to viral infections at the single cell level.
Supplemental Material
sj-xlsx-1-ini-10.1177_17534259231181029 - Supplemental material for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health
Supplemental material, sj-xlsx-1-ini-10.1177_17534259231181029 for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health by Taylor Crue, Grace Yihua Lee, Joyce Yao-chun Peng, Niccolette Schaunaman, Hina Agraval, Brian J. Day, Kris Genelyn Dimasuay, Diana Cervantes, Hamid Nouri, Taylor Nichols, Paige Hartsoe, Mari Numata, Irina Petrache and Hong Wei Chu in Innate Immunity
Supplemental Material
sj-xlsx-2-ini-10.1177_17534259231181029 - Supplemental material for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health
Supplemental material, sj-xlsx-2-ini-10.1177_17534259231181029 for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health by Taylor Crue, Grace Yihua Lee, Joyce Yao-chun Peng, Niccolette Schaunaman, Hina Agraval, Brian J. Day, Kris Genelyn Dimasuay, Diana Cervantes, Hamid Nouri, Taylor Nichols, Paige Hartsoe, Mari Numata, Irina Petrache and Hong Wei Chu in Innate Immunity
Supplemental Material
sj-xlsx-3-ini-10.1177_17534259231181029 - Supplemental material for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health
Supplemental material, sj-xlsx-3-ini-10.1177_17534259231181029 for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health by Taylor Crue, Grace Yihua Lee, Joyce Yao-chun Peng, Niccolette Schaunaman, Hina Agraval, Brian J. Day, Kris Genelyn Dimasuay, Diana Cervantes, Hamid Nouri, Taylor Nichols, Paige Hartsoe, Mari Numata, Irina Petrache and Hong Wei Chu in Innate Immunity
Supplemental Material
sj-xlsx-4-ini-10.1177_17534259231181029 - Supplemental material for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health
Supplemental material, sj-xlsx-4-ini-10.1177_17534259231181029 for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health by Taylor Crue, Grace Yihua Lee, Joyce Yao-chun Peng, Niccolette Schaunaman, Hina Agraval, Brian J. Day, Kris Genelyn Dimasuay, Diana Cervantes, Hamid Nouri, Taylor Nichols, Paige Hartsoe, Mari Numata, Irina Petrache and Hong Wei Chu in Innate Immunity
Supplemental Material
sj-xlsx-5-ini-10.1177_17534259231181029 - Supplemental material for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health
Supplemental material, sj-xlsx-5-ini-10.1177_17534259231181029 for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health by Taylor Crue, Grace Yihua Lee, Joyce Yao-chun Peng, Niccolette Schaunaman, Hina Agraval, Brian J. Day, Kris Genelyn Dimasuay, Diana Cervantes, Hamid Nouri, Taylor Nichols, Paige Hartsoe, Mari Numata, Irina Petrache and Hong Wei Chu in Innate Immunity
Supplemental Material
sj-xlsx-6-ini-10.1177_17534259231181029 - Supplemental material for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health
Supplemental material, sj-xlsx-6-ini-10.1177_17534259231181029 for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health by Taylor Crue, Grace Yihua Lee, Joyce Yao-chun Peng, Niccolette Schaunaman, Hina Agraval, Brian J. Day, Kris Genelyn Dimasuay, Diana Cervantes, Hamid Nouri, Taylor Nichols, Paige Hartsoe, Mari Numata, Irina Petrache and Hong Wei Chu in Innate Immunity
Supplemental Material
sj-xlsx-7-ini-10.1177_17534259231181029 - Supplemental material for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health
Supplemental material, sj-xlsx-7-ini-10.1177_17534259231181029 for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health by Taylor Crue, Grace Yihua Lee, Joyce Yao-chun Peng, Niccolette Schaunaman, Hina Agraval, Brian J. Day, Kris Genelyn Dimasuay, Diana Cervantes, Hamid Nouri, Taylor Nichols, Paige Hartsoe, Mari Numata, Irina Petrache and Hong Wei Chu in Innate Immunity
Supplemental Material
sj-xlsx-8-ini-10.1177_17534259231181029 - Supplemental material for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health
Supplemental material, sj-xlsx-8-ini-10.1177_17534259231181029 for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health by Taylor Crue, Grace Yihua Lee, Joyce Yao-chun Peng, Niccolette Schaunaman, Hina Agraval, Brian J. Day, Kris Genelyn Dimasuay, Diana Cervantes, Hamid Nouri, Taylor Nichols, Paige Hartsoe, Mari Numata, Irina Petrache and Hong Wei Chu in Innate Immunity
Supplemental Material
sj-xlsx-9-ini-10.1177_17534259231181029 - Supplemental material for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health
Supplemental material, sj-xlsx-9-ini-10.1177_17534259231181029 for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health by Taylor Crue, Grace Yihua Lee, Joyce Yao-chun Peng, Niccolette Schaunaman, Hina Agraval, Brian J. Day, Kris Genelyn Dimasuay, Diana Cervantes, Hamid Nouri, Taylor Nichols, Paige Hartsoe, Mari Numata, Irina Petrache and Hong Wei Chu in Innate Immunity
Supplemental Material
sj-xlsx-10-ini-10.1177_17534259231181029 - Supplemental material for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health
Supplemental material, sj-xlsx-10-ini-10.1177_17534259231181029 for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health by Taylor Crue, Grace Yihua Lee, Joyce Yao-chun Peng, Niccolette Schaunaman, Hina Agraval, Brian J. Day, Kris Genelyn Dimasuay, Diana Cervantes, Hamid Nouri, Taylor Nichols, Paige Hartsoe, Mari Numata, Irina Petrache and Hong Wei Chu in Innate Immunity
Supplemental Material
sj-xlsx-11-ini-10.1177_17534259231181029 - Supplemental material for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health
Supplemental material, sj-xlsx-11-ini-10.1177_17534259231181029 for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health by Taylor Crue, Grace Yihua Lee, Joyce Yao-chun Peng, Niccolette Schaunaman, Hina Agraval, Brian J. Day, Kris Genelyn Dimasuay, Diana Cervantes, Hamid Nouri, Taylor Nichols, Paige Hartsoe, Mari Numata, Irina Petrache and Hong Wei Chu in Innate Immunity
Supplemental Material
sj-xlsx-12-ini-10.1177_17534259231181029 - Supplemental material for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health
Supplemental material, sj-xlsx-12-ini-10.1177_17534259231181029 for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health by Taylor Crue, Grace Yihua Lee, Joyce Yao-chun Peng, Niccolette Schaunaman, Hina Agraval, Brian J. Day, Kris Genelyn Dimasuay, Diana Cervantes, Hamid Nouri, Taylor Nichols, Paige Hartsoe, Mari Numata, Irina Petrache and Hong Wei Chu in Innate Immunity
Supplemental Material
sj-xlsx-13-ini-10.1177_17534259231181029 - Supplemental material for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health
Supplemental material, sj-xlsx-13-ini-10.1177_17534259231181029 for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health by Taylor Crue, Grace Yihua Lee, Joyce Yao-chun Peng, Niccolette Schaunaman, Hina Agraval, Brian J. Day, Kris Genelyn Dimasuay, Diana Cervantes, Hamid Nouri, Taylor Nichols, Paige Hartsoe, Mari Numata, Irina Petrache and Hong Wei Chu in Innate Immunity
Supplemental Material
sj-xlsx-14-ini-10.1177_17534259231181029 - Supplemental material for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health
Supplemental material, sj-xlsx-14-ini-10.1177_17534259231181029 for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health by Taylor Crue, Grace Yihua Lee, Joyce Yao-chun Peng, Niccolette Schaunaman, Hina Agraval, Brian J. Day, Kris Genelyn Dimasuay, Diana Cervantes, Hamid Nouri, Taylor Nichols, Paige Hartsoe, Mari Numata, Irina Petrache and Hong Wei Chu in Innate Immunity
Supplemental Material
sj-xlsx-15-ini-10.1177_17534259231181029 - Supplemental material for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health
Supplemental material, sj-xlsx-15-ini-10.1177_17534259231181029 for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health by Taylor Crue, Grace Yihua Lee, Joyce Yao-chun Peng, Niccolette Schaunaman, Hina Agraval, Brian J. Day, Kris Genelyn Dimasuay, Diana Cervantes, Hamid Nouri, Taylor Nichols, Paige Hartsoe, Mari Numata, Irina Petrache and Hong Wei Chu in Innate Immunity
Supplemental Material
sj-xlsx-16-ini-10.1177_17534259231181029 - Supplemental material for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health
Supplemental material, sj-xlsx-16-ini-10.1177_17534259231181029 for Single cell RNA-sequencing of human precision-cut lung slices: A novel approach to study the effect of vaping and viral infection on lung health by Taylor Crue, Grace Yihua Lee, Joyce Yao-chun Peng, Niccolette Schaunaman, Hina Agraval, Brian J. Day, Kris Genelyn Dimasuay, Diana Cervantes, Hamid Nouri, Taylor Nichols, Paige Hartsoe, Mari Numata, Irina Petrache and Hong Wei Chu in Innate Immunity
Footnotes
Acknowledgements
The authors thank Nicole Pavelka, Shaan Gellatly, Elysia Min, Kelly Schweitzer, Patrick Hume, William Janssen, Kara Mould and Christina Lisk for their technical assistance in precision-cut lung slice methodology, donor lung processing and nicotine measurement.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by the following grants from the National Institutes of Health, (grant number R01AI150082, R01AI152504, R01AI161296, R01HL144396, U19AI125357).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
