Abstract
Background
Genome-wide association studies have implicated dozens of genes with migraine susceptibility, but it remains unclear in which nervous system cell types these genes are expressed.
Methods
Using single-cell RNA sequencing data from the central and peripheral nervous system, including the trigeminal ganglion, the expression of putative migraine-associated genes was compared across neuronal, glial and neurovascular cell types within these tissues.
Results
Fifty-four putative migraine-associated genes were expressed in the central nervous system, peripheral nervous system or neurovascular cell types analyzed. Six genes (11.1%) were selectively enriched in central nervous system cell types, three (5.5%) in neurovascular cell types, and two (3.7%) in peripheral nervous system cell types. The remaining genes were expressed in multiple cell types.
Conclusions
Single-cell RNA sequencing of the brain and peripheral nervous system localizes each migraine-associated gene to its respective nervous system tissue and the cell types in which it is expressed. While the majority of migraine-associated genes are broadly expressed, we identified several cell-type-specific migraine-associated genes in the central nervous system, peripheral nervous system, and neurovasculature.
Introduction
Migraine is a complex polygenic disease modified by environmental factors. The pathophysiology of migraine involves sensitization of trigeminovascular afferents via cortical hyperexcitability (e.g. cortical spreading depression) and activation of brainstem and thalamic nuclei that transmit trigeminal signals to sensory cortex (1,2). Susceptibility to common migraine could in principle result from genetic variation in central and/or peripheral nervous system components of these pathways.
Large-scale genome-wide association studies have identified 38 susceptibility loci associated with migraine (3), but how genomic variants contribute to migraine pathophysiology is not clear. Analysis of gene expression data from homogenized tissue observed significant enrichment of migraine-associated genes in vascular and smooth muscle tissues (3), but recent analysis of gene expression data from single-cell RNA sequencing of human cortex observed expression of migraine-associated genes in brain cell types as well (4). Specifically, significant enrichment of several migraine-associated genes was observed within neurons, glia, astrocytes, and vascular cells. While this study focused only on the cortex and was limited by the small number of cells analyzed, the rapid expansion of single-cell RNA sequencing datasets has allowed for direct comparisons of gene expression in diverse cell types across multiple nervous system structures. Our present study examines the enrichment of migraine-associated gene expression in both the central (CNS) and peripheral nervous system (PNS) as well as the distinct cell types within structures that have been implicated in migraine pathophysiology.
Methods
Protein coding genes associated with migraine were identified by Gormley et al. (3) Genes that contain the migraine-associated variant within their coding region or are <20 kb from the coding region were included (see supplementary Table 8 in Gormley et al. for the full list of genes). Genes with an established role in migraine pathophysiology or rare migraine subtypes were also analyzed. This included genes associated with the ligand or receptor of CGRP (
Gene expression analysis of CNS, PNS, and neurovascular cell types
The expression of each migraine-associated gene was obtained from single-cell RNA sequencing data of approximately 500,000 cells across multiple brain regions, dorsal root ganglia, sympathetic neurons and enteric neurons (5). Our analysis focused on the following cell types thought to be involved in migraine pathophysiology: central nervous system (excitatory and inhibitory neurons, astrocytes, oligodendrocytes, microglia); peripheral nervous system (peptidergic nociceptors, non-peptidergic nociceptors, large diameter neurofilament-positive mechanoreceptors, C low-threshold mechanoreceptors (cLTMR), sympathetic noradrenergic cells, sympathetic cholinergic cells, Schwann cells); and neurovascular cell types (leptomeningeal, smooth muscle, endothelial cells). Gene expression data from parasympathetic neurons were not available.
Of the 107 genes within 20 kb of the statistically significant migraine-associate genomic loci, 88 genes were annotated in the single-cell RNA sequencing dataset (5). To determine whether a gene was expressed in the cell types analyzed, we chose a minimum expression cutoff of ≥0.2 normalized mRNA counts. Because single-cell gene expression values are non-normally distributed and notoriously zero-inflated, the expression cutoff was chosen to include genes that may be of biological importance. The expression cutoff for each single-cell RNA-sequencing dataset required a gene to be expressed in at least one cell type at >2/3 the median of a distribution of gene expression values. This distribution of gene expression values consisted of the normalized mean expression value for each gene in the cell type in which it is most highly expressed. This resulted in inclusion of 54 out of 88 migraine-associated genes that were above the expression cutoff in the dataset from Zeisel et al.
Gene expression analysis of trigeminal ganglion cell types
The single-cell RNA sequencing dataset from Nguyen et al. includes about 7000 cells from mouse trigeminal ganglion (TG), of which 3500 are neuronal (6). Of the 107 genes identified from Gormley et al., 83 genes were annotated in the TG single-cell RNA sequencing data. There was good overlap between the availability of genes between the two datasets (81 genes were found in both datasets; five genes found in the Zeisel et al. dataset were not available in the Nguyen et al. dataset; and three genes available in the Nguyen et al. dataset were not identified in the Zeisel et al. dataset). The same criterion for exclusion of lowly expressed genes was used for the Nguyen et al. dataset. Given the distribution of gene expression values in this dataset, a value of 0.11 normalized mRNA counts was calculated as the minimum expression threshold, such that a gene had to have at least one cell type in which its expression was ≥0.11 normalized mRNA counts. This resulted in the inclusion of 54 out of 83 migraine-associated genes annotated in Nguyen et al.
Statistical analysis
Enrichment
Results
Comparison of CNS, PNS, and neurovascular cell types
To explore the nervous system cell types in which putative migraine-associated genes are expressed, we analyzed gene expression from a recent single-cell RNA-sequencing resource generated from multiple cell types of the mouse CNS and PNS. We focused on CNS cell types likely to be involved in cortical spreading depression and PNS cell types from the DRG involved in peripheral pain sensitization (this dataset did not sequence cells from trigeminal ganglion). We found that the approximately half of migraine-associated genes (28/54, 51.9%) have significant enrichment in cell types within both the CNS and PNS (Figures 1 and 2 and Supplemental Figure 1). Eleven (20.4%) genes are enriched in cell types in the CNS, PNS and neurovascular cell types. Expression of the remaining 17.7% of genes displays tissue-selective enrichment within the CNS ( (a) Heatmap of migraine-associated gene expression (mRNA counts, row normalized) across CNS, PNS (dorsal root ganglion) and neurovascular cells (Zeisel et al.). (b) Heatmap of migraine-associated gene expression (mRNA counts, row normalized) across trigeminal ganglion cells (Nguyen et al.). Genes expressed above threshold in either the Zeisel or Nguyen datasets are included. Genes in which all values in a row were <0.01 mRNA counts are in white (MRVI1, HCK, NOTCH3). Symp: sympathetic; norad: noradrenergic; chol: cholinergic; PEP: peptidergic nociceptor; NF: neurofilament positive nociceptor; NP: non-peptidergic nocicpetor; cLTMR: C low threshold mechanoreceptors. Venn diagram of migraine-associated genes that are significantly enriched in CNS, PNS and neurovascular cell types.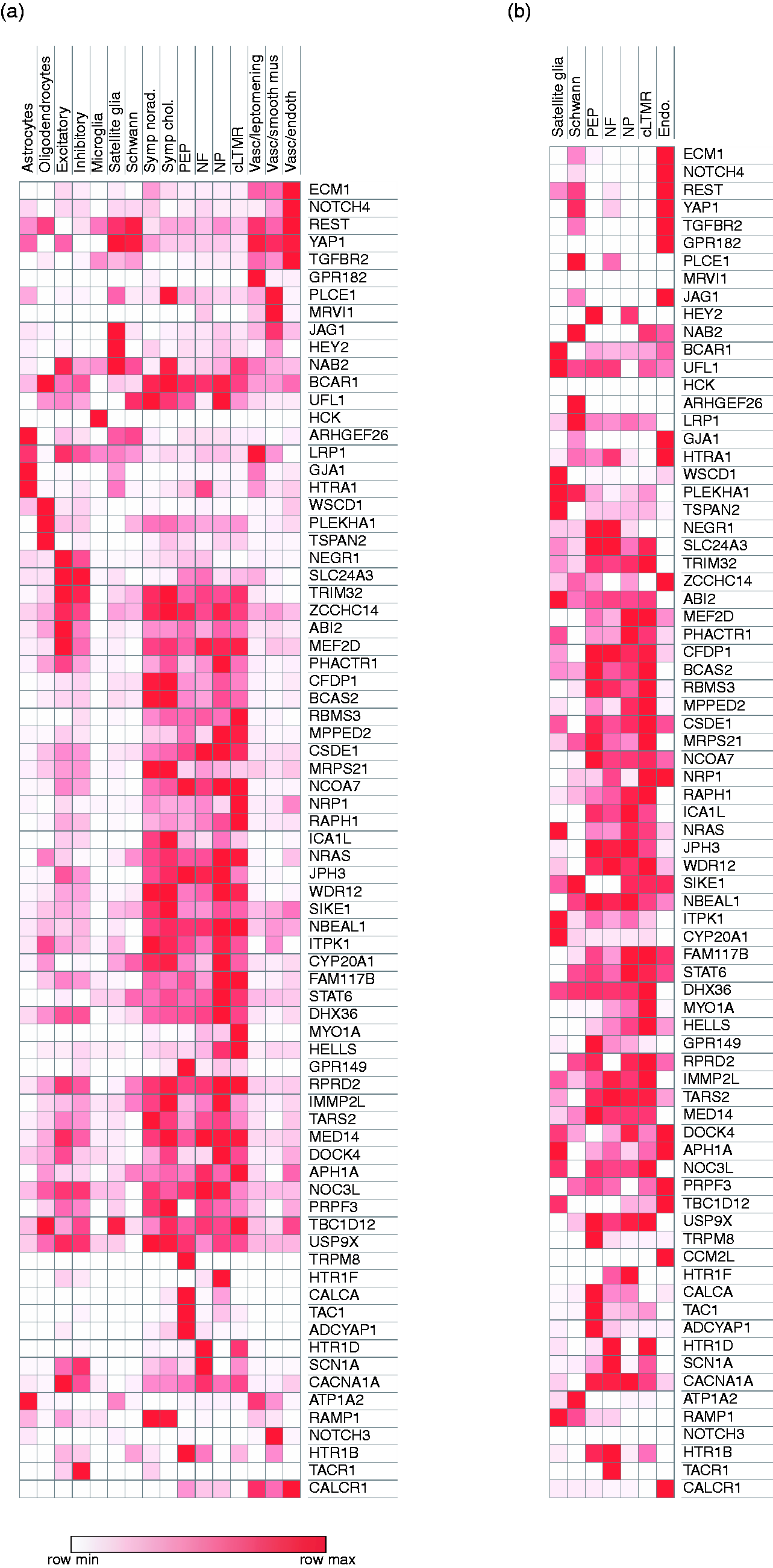

Most migraine-associated genes that are significantly enriched in the PNS are expressed within multiple PNS neuronal subtypes (68.5% of genes are expressed in non-peptidergic nociceptors, 59.3% in peptidergic nociceptors, 51.8% in large diameter neurofilament-positive mechanoreceptors, 59.3% in cLTMRs, 61.1% in sympathetic cholinergic cells, and 61.1% in sympathetic noradrenergic cells) (Figure 1(a)). Three genes are enriched in a single PNS cell type (expressed >2-fold as compared to the next most common cell type);
There is limited overlap between the CNS cell types in which migraine-associated genes are expressed, as 55.5% of genes are enriched in neurons only, 31.5% are enriched in non-neuronal CNS cells only, and 13.0% of genes are enriched in both neuronal and non-neuronal cells. Of the genes significantly enriched in CNS cells,
The migraine-associated genes enriched in neurovascular cell types are observed in smooth muscle (22.2%), leptomeningeal (18.5%), and endothelial (18.5%) cell types. While 33.3% of the migraine-associated genes enriched in neurovascular cells are observed across all three subtypes of neurovascular cells,
In addition to the putative migraine-associated genes identified in Gormley et al., we also explored the expression of genes known to be involved in migraine pathophysiology or familial causes of migraine.
Comparison of trigeminal ganglion cell types
Migraine pathophysiology typically involves the activation of the trigeminal system (8), although high cervical DRGs may also contribute (9,10). While the neuronal subtypes in the DRG are similar to those of TG, there are also some notable gene expression differences such as TRPM8, which is more highly expressed in TG than DRG. We therefore analyzed the gene expression patterns of migraine-associated genes within a recently published mouse trigeminal ganglion single-cell RNA-sequencing dataset (6). As expected, there is great similarity with regards to the cell types in which genes are expressed in the DRG and TG, although the degree of enrichment is different in some cases (Figure 1(a) and 1(b), Supplemental Figure 2). For example,
Conclusion
Putative migraine-associated genes are expressed in CNS, PNS and neurovascular cell types. This is consistent with the observation from familial hemiplegic migraine and CADASIL that dysfunction of different nervous system cell types can lead to migraine. In contrast to these diseases, which are caused by highly penetrant single-gene mutations, common migraine susceptibility is thought to be driven by a summation of common genetic variants that each exert a small effect on migraine susceptibility (11). Some common migraine-associated genes are enriched within specific cell types (17.7%), raising the possibility that individuals harboring variants in these cell-type-specific genes may have dysfunction in the respective cell type that disproportionately contributes to migraine susceptibility. That said, the majority of migraine-associated genes (82.7%) are expressed broadly, so there are likely functional changes occurring in multiple cell types that contribute to migraine susceptibility in most patients. Quantifying the extent of polygenic risk within each CNS or PNS cell type may provide important clues to an individual’s migraine pathophysiology and help stratify migraine patients through routine genotyping.
While our current analyses take advantage of advances in single-cell RNA sequencing and incorporate many more cells (∼500,000 cells) compared to the previous study (<3000), the localization of migraine-associated variants to specific nervous system cell types should be interpreted within the context of its limitations. First, migraine-associated genomic variants have been linked to putative migraine-associated genes largely by the distance between the variant and gene, so additional epigenetic studies are needed to validate these associations. Second, the single-cell RNA sequencing data used in this study was generated from mice, so the cell-type-specific gene expression patterns described here should be revisited once analogous human data become available. Although there are certainly differences in gene expression between species, recent single-cell RNA-sequencing studies in neurons (12) and pancreas (13) have found that the cell types in which a given gene is expressed is largely conserved. Indeed, new multi-dimensional gene expression algorithms are able to identify and annotate conserved cell types across species (14–16), and recent genome-wide association studies have now used single-cell RNA sequencing data from mice to estimate the likely brain cell types in which they are expressed in humans (17–19).
Our study leverages advances in single-cell RNA sequencing to characterize the cell types in which migraine-associated genes are expressed within the CNS and PNS using the largest available datasets to date. The correct cell type localization of known migraine-associated genes (e.g.
Key findings
Putative migraine-associated genes are expressed in cell types of both the central and peripheral nervous system. Approximately 17.7% of migraine-associated genes are expressed selectively within a specific CNS or PNS cell type. Localizing the cell types in which migraine susceptibility genes are expressed will guide future mechanistic studies into their function.
Supplemental Material
CEP877834 Supplemetal Material1 - Supplemental material for Migraine-associated gene expression in cell types of the central and peripheral nervous system
Supplemental material, CEP877834 Supplemetal Material1 for Migraine-associated gene expression in cell types of the central and peripheral nervous system by Angeliki Vgontzas and William Renthal in Cephalalgia
Supplemental Material
CEP877834 Supplemetal Material2 - Supplemental material for Migraine-associated gene expression in cell types of the central and peripheral nervous system
Supplemental material, CEP877834 Supplemetal Material2 for Migraine-associated gene expression in cell types of the central and peripheral nervous system by Angeliki Vgontzas and William Renthal in Cephalalgia
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
