Abstract
Periodontitis is induced by periodontal dysbiosis characterized by the predominance of anaerobic species. TLRs constitute the classical pathway for cell activation by infection. Interestingly, the Toll/IL-1 receptor homology domain adapters initiate signaling events, leading to the activation of the expression of the genes involved in the host immune response. The aim of this study was to evaluate the effects of Porphyromonas gingivalis on the expression and protein-protein interactions among five TIR adapters (MAL, MyD88, TRIF, TRAM and SARM) in gingival epithelial cells and endothelial cells. It was observed that P. gingivalis is able to modulate the signaling cascades activated through its recognition by TLR4/2 in gingival epithelial cells and endothelial cells. Indeed, MAL-MyD88 protein-protein interactions associated with TLR4 was the main pathway activated by P. gingivalis infection. When transient siRNA inhibition was performed, cell viability, inflammation, and cell death induced by infection decreased and such deleterious effects were almost absent when MAL or TRAM were targeted. This study emphasizes the role of such TIR adapter proteins in P. gingivalis elicited inflammation and the precise evaluation of TIR adapter protein interactions may pave the way for future therapeutics in both periodontitis and systemic disease with a P. gingivalis involvement, such as atherothrombosis.
Introduction
Periodontitis is an infectious inflammatory disease associated with periodontal tissue destruction and is considered the main cause of tooth loss. Such disease is characterized by increased periodontal pocket depth (PPD), gingival bleeding, and clinical attachment loss. 1 Periodontitis is a highly prevalent disease affecting more than 40% of the US population. 2 Nevertheless, its severe form is found in around 11% of the worldwide population with a peak incidence around 38 yr of age. 3 Aside from its impact on periodontium and masticatory functions associated with a decreased oral health-related quality of life, 4 an association with several systemic diseases, such as cardiovascular diseases or diabetes, has been observed. 5 , 6 Periodontitis is induced by dysbiotic periodontal microbiome characterized by the predominance of anaerobic species, including Porphyromonas gingivalis. The establishment of a dysbiosis is associated with an imbalance between the bacterial insult and the host immune response, leading to periodontal breakdown. 7 Indeed, the role of keystone pathogens such as P. gingivalis has been highlighted, with such bacterium demonstrating a large panel of virulence factors, including LPS or protein-modifying enzymes such as peptidylarginine deiminase. 8
In the oral cavity, gingival epithelial cells represent the largest area of contact and, therefore, constitute the first line of host defense; they are constantly challenged by dysbiotic flora. P. gingivalis has been shown to be able to invade GEC, to replicate intracellularly, and to modulate the cell cycle and programmed cell death through modulation of P53 and JAK kinase/STAT and APAF-1 apoptosome-related pathways.9–12 All these mechanisms are part of the bacterial strategy, aiming to escape the host immune response, thus contributing to the establishment of chronic inflammation and periodontal tissue destruction. 7 , 13 Nevertheless, P. gingivalis is also able to spread in other cell types, such as fibroblasts and endothelial cells (EC), 10 , 14 and, at distance, can be detected in several organs such as the liver or gut 15 and blood vessels. 16 , 17 Indeed, its implication in the development of systemic diseases, such as cardiovascular diseases and, particularly, atherothrombosis, is reported as several studies demonstrated that P. gingivalis is able to amplify and sustain inflammation, contributing to the development of atheromatous plaque in vitro and in vivo. 14 , 15 ,18–20
TLRs play a key role in the activation of the innate immune response against invading pathogens. 17 , 21 TLRs activate a molecular cascade that induces the secretion of inflammatory cytokines and adhesion molecules, such as IL-1β, IL-6, TNF-α, NF-κB, and other molecules involved in cell death. 22 , 23 The signal transmitted by TLRs must therefore be tightly controlled, and it is clearly established that, in the case of TLRs over-activation, infectious and inflammatory diseases may develop.24–26 P. gingivalis is able to activate or modulate signaling pathways initiated by TLRs in different cell types, especially those regulated by TLR2 and TLR4 recognition. 12 , 21 , 27 TLR signaling involves a family of five adapter proteins, which activate downstream protein kinases and ultimately lead to the activation of transcription factors such as NF-κB and members of the IFN regulatory factor (IRF). 28 A key signaling domain, unique to the TLR system, is the Toll/IL-1 receptor (TIR) domain, located on the cytosolic side of each TLR as well as in adapters28–30 such as MyD88, MyD88-adaptor-like (MAL, also known as TIRAP), an adapter protein containing an IFN-inducing TIR domain (TRIF, also known as TICAM1), a molecule TRIF-related adapter (TRAM, also known as TICAM2), and a protein called sterile α- and armadillo-motif-containing protein (SARM). 29
The identification of the activated pathways may pave the way for their targeting by inhibitors as new potential pharmacological management of periodontitis and give a better insight in the understanding of P. gingivalis-associated systemic effect. Therefore, the aim of this study was to evaluate the involvement of TIR domains in the TLR2/TLR4 signaling pathways in GEC and EC infected with P. gingivalis.
Materials and methods
Ethics statement, study population, and collection of gingival tissue samples
For the analysis of ex-vivo biopsied samples, a total of 20 patients agreed to participate in this study (French Ministry of Research, Bioethic department authorization DC-2014-2220). All patients gave written and informed consent prior to enrollment. Complete medical and dental histories were considered from all the subjects. None of the patients had systemic diseases, were smokers, or had taken any medication such as antibiotics or antimicrobial mouthwashes in the last 3 months. Clinical periodontal examination included measurement of PPD and bleeding on probing (BOP) at six sites around each tooth with a manual periodontal probe. The healthy group (H) consisted of nine patients (five males and four females, mean age 37.8 +/- 17.3 yr) and the chronic periodontitis group (CP) of 11 patients (four males and seven females, mean age 62.4 +/- 7.3 yr) based on the Armitage classification. 31 In the periodontitis group, all patients were diagnosed with severe chronic generalized periodontitis, and gingival biopsies were collected during periodontal surgeries performed at sites with PPD > 5mm + BOP (open access flap). Samples from the control group (healthy tissues) were collected from healthy patients during extraction of third molars mainly for orthodontic or prosthodontic reasons. In control patients, harvested tissues correspond to sites with PPD < 4mm without BOP. Mean PPD in the periodontitis group was 6.8 mm (+/-0.8) at sites of harvesting. All samples were placed in a sterile tube and stored at -80°C until RNA extraction.
P. gingivalis culture
P. gingivalis strain 33277 (ATCC, Manassas, VA, USA) was cultured under strict anaerobic conditions at 37°C in brain-heart infusion medium (Sigma, Saint-Quentin Fallavier, France), supplemented with hemin (5 mg/ml) and menadione (1 mg/ml). On the day of infection, bacteria were collected and counted as described previously. 6
Cell culture
Human oral epithelial cells (GEC) used in this study were derived from the TERT-2 OKF-6 cell line (BWH Cell Culture and Microscopy Core, Boston, MA, USA) and were cultured in KSFM culture medium (serum-free medium for keratinocytes, Gibco, Aachen, Germany). Human umbilical vein EC (C-12200, PromoCell, Heidelberg, Germany) and IncuCyte® CytoLight Green HUVEC (4453, Essen Bioscience, Royston, Hertfordshire, United Kingdom) were cultured in EGM2 medium (Promocell) supplemented with 10% FBS. To reduce the risk of contamination, 100 units/ml of penicillin and 100 μg/ml of streptomycin were added to the KSFM culture medium and no antibiotics were added to the medium for EC. Both were grown at 37°C in a humidified atmosphere with 5% CO2 as described previously. 11 , 18
Infection with P. gingivalis and stimulation with its LPS
On the day of infection, bacteria were collected and counted as described. 6 Twenty-four h before the experiment, 2 × 105 cells were seeded in a 24-well plate. On the day of the experiment, GEC and EC were washed twice with PBS and infected for 24 h with P. gingivalis at a multiplicity of infection (MOI) of 100. After 2 h of infection, medium was removed, and cells were washed three times with PBS to remove non-adherent and external bacteria. Then, metronidazole (200 μg/ml) was added for 1 h to kill the external bacteria and, after washing, fresh bacteria-free medium was added in each well.32–34 To assess the effect of LPS, GEC and EC were stimulated with ultrapure P. gingivalis LPS (Pg-LPS) (InvivoGen, San Diego, CA, USA) at a concentration of 1 μg/ml for 24 h.
RNA isolation, reverse transcription and quantitative real-time PCR (qPCR)
Total RNA was extracted using Tri reagent (Sigma-Aldrich) according to the manufacturer’s instructions. The total RNA concentration was quantified using NanoDrop 1000 (Thermo Fisher Scientific, Illkirch-Graffenstaden, France). Reverse transcription was performed using iScript Reverse Transcription Supermix (Bio-Rad, Miltry-Mory, France) according to the manufacturer’s instructions. To quantify mRNA expression, qPCR was performed on the cDNA samples. PCR amplification and analysis were performed with CFX ConnectTM Real-Time PCR Detection System (Bio-Rad, Miltry-Mory, France). Amplification reactions have been performed using iTAq Universal SYBR Green Supermix (Bio-Rad). β-actin was used as an endogenous RNA control (housekeeping gene) in all samples. Primers related to β-actin, TNF-α, IL-6, TLR2, TLR4, P. gingivalis, and to the five TIRs (MyD88, MAL, TRAM, TRIF, SARM) were synthesized (Thermo Fisher) (Table 1). Expression levels were calculated after normalization to the housekeeping gene expression.
Sequence of primers used for real-time quantitative RT-qPCR.

Pull-down assay
Interactions between MAL and TRAM on cell lysates after infection were evaluated following previously described protocols 35 , 36 and using the kit containing the EZ-Link Sulfo-NHS-LC-Desthiobiotin enzyme according to manufacturer’s instructions (Thermo Fisher). Briefly, removal of excess desthiobiotin reagent from samples was performed using Zeba spin columns (Thermo Fisher). After coupling the desthiobiotinylated bait protein to the streptavidin resin, cell lysates were incubated for at least 1 h to finally elute the desthiobiotinylated bait protein and captured proteins. Obtained samples were then analyzed by SDS-PAGE and stained with Coomassie Blue G-250 (Thermo Fisher).
Co-immunoprecipitation
To identify interactions between the five TIR domain containing proteins and TLR2 or TLR4, co-immunoprecipitation using DynabeadsTM Protein G (Thermo Fisher) was performed. Abs against TLR2 (SC-10741, anti-rabbit, Santa Cruz Biotechnology), TLR4 (SC-10739, anti-rabbit, Santa Cruz Biotechnology), MyD88 (PA1-9072, anti-goat, Thermo Fisher), MAL (PA1-12815, anti-rabbit, Thermo Fisher), TRIF (PA5-20030, anti-rabbit, Thermo Fisher), TRAM (PA5-23115, anti-rabbit, Thermo Fisher), and SARM (PA5-20059, anti-rabbit, Thermo Fisher) were incubated with Dynabeads Protein G. The bead-bound Abs were then used for immunoprecipitation in the presence of cell lysates. Bound material was analyzed using a Western blot.
Western blotting
In order to detect the protein levels of TLR2, TLR4, MyD88, MAL, TRIF, TRAM, SARM, and SOCS-1, SDS-PAGE followed by immunoblotting was performed as described. 14 Briefly, cells were lysed for 5 min on ice in 200 μl of ice-cold RIPA buffer (65 mM Tris–HCl, pH 7.4, 150 mM NaCl, and 0.5% sodium deoxycholate) supplemented with phosphatase inhibitor cocktails I and II and a protease inhibitor cocktail (Sigma, Darmstadt, Germany). Lysates were centrifuged at 10,000 g at 4°C for 10 min and supernatants were collected. Both lysates or co-immunoprecipitation samples were quantified using the Bradford protein assay (Bio-Rad, Hercules, CA, USA). To perform SDS-PAGE and immunoblotting, 15 to 25 μg of proteins were used for each condition. Abs against TLR2, TLR4, MyD88, MAL, TRIF, TRAM, SARM, and SOCS-1 (38-5200, Anti-rabbit, Thermo Fisher) were used. Secondary alkaline phosphatase conjugated Abs (anti-goat [1:3000]; anti-rabbit [1:5000]) were purchased from Bethyl Laboratories (Montgomery, Texas, USA) and Thermo Fisher. All Abs were used at the dilutions recommended by the manufacturer.
ELISA
In order to detect secreted TNF-α, a sandwich ELISA was performed. Briefly, the first step involved attachment of a capture Ab to the microplate (goat anti-Human TNF-α, PeproTech, Rocky Hill, NJ, USA) (4 μg/mL). Then, samples containing unknown amounts of TNF-α were added and bound to the capture Ab. After washing, a detection Ab (biotinylated goat anti-Human TNF-α, PeproTech) (0.5 μg/ml) was added to the wells. Plates were washed extensively and a HRP conjugate (streptavidin) was added to the protein-Abs sandwich. Finally, a substrate solution was added and converted by the enzyme (HRP) for detection using a spectrophotometer at 450 nm. The intensity of this signal was compared with a standard curve (Recombinant Human TNF-α, PeproTech) (from 0 to 1000 pg).
Small interfering RNA transfection
One d prior to the transfection, 5 × 106 of GEC or EC was seeded in a six-well plate. Transfection with 5 nM of Silencer™ Select Pre-Designed small interfering RNA (siRNA) targeted against MAL (ID: s195607, Ref: 4392420, Thermo Fisher), TRAM (ID: s51477, Ref: 4392420, Thermo Fisher), or with a negative control was performed using Lipofectamine 3000 and the Silencer™ siRNA Transfection II Kit (Thermo Fisher) according to the manufacturer’s protocol. Transfected cells were then infected with P. gingivalis according to the experimental design.
Metabolic activity assay
An alamarBlue test was used to evaluate the metabolic activity of GEC and EC. Briefly, 200 μl of incubation media was transferred separately to a 96-well plate and absorbance was measured at OD570 and OD595 nm by a spectrophotometer (Multiskan, Thermo Fisher) in order to determine the percentage of alamarBlue reduction as indicated by the manufacturer.
Immunofluorescence
After infection, cells were fixed by immersion in 4% paraformaldehyde in PBS. Sections were incubated with primary Ab against TLR2 (1:250), TLR4 (1:250), MyD88 (1:500), MAL (1:500), TRIF (1:500), TRAM (1:500), SARM (1:600), and SOCS-1 (1:600), respectively, at 4°C for 24 h. After incubation, sections were incubated with the secondary Ab for 1 h at room temperature (1:250 to 1:500 of Alexa Fluor 594 or Alexa Fluor 488 (Invitrogen, Thermo Fisher, Illkirch, France)). Nuclei were stained with DAPI 200 nM (Euromedex, Souffelweyersheim, France) and β-actin was labeled with phalloïdin Alexa Fluor 546 (Thermo Fisher) in GEC. Finally, slides were mounted (Dako, Trappes, France) and observed with a fluorescence microscope (Leica DM4000B).
Statistical analysis
Statistical analysis was performed using a pairwise ANOVA test, a post-hoc Tukey’s test, and Spearman’s rank order correlation analysis. The statistical significance level was considered as P < 0.05. Data was analyzed using PRISM 9.1 (GraphPad, La Jolla, CA, USA). All experiments have been performed at least three times with at least triplicate samples (biological and technical replicates).
Results
TNF-α and IL-6 expression is increased in diseased tissues and their secretion is increased by P. gingivalis infection in GEC and EC
TNF-α and IL-6 are key markers for periodontal inflammation. As expected, the mRNA expression of TNF-α and IL-6 was significantly increased, as well as P. gingivalis detection in clinical samples from periodontitis patients in comparison with healthy ones (fold change 2.8, 3.7 and 7.2, respectively; P < 0.05) (Figure 1A).
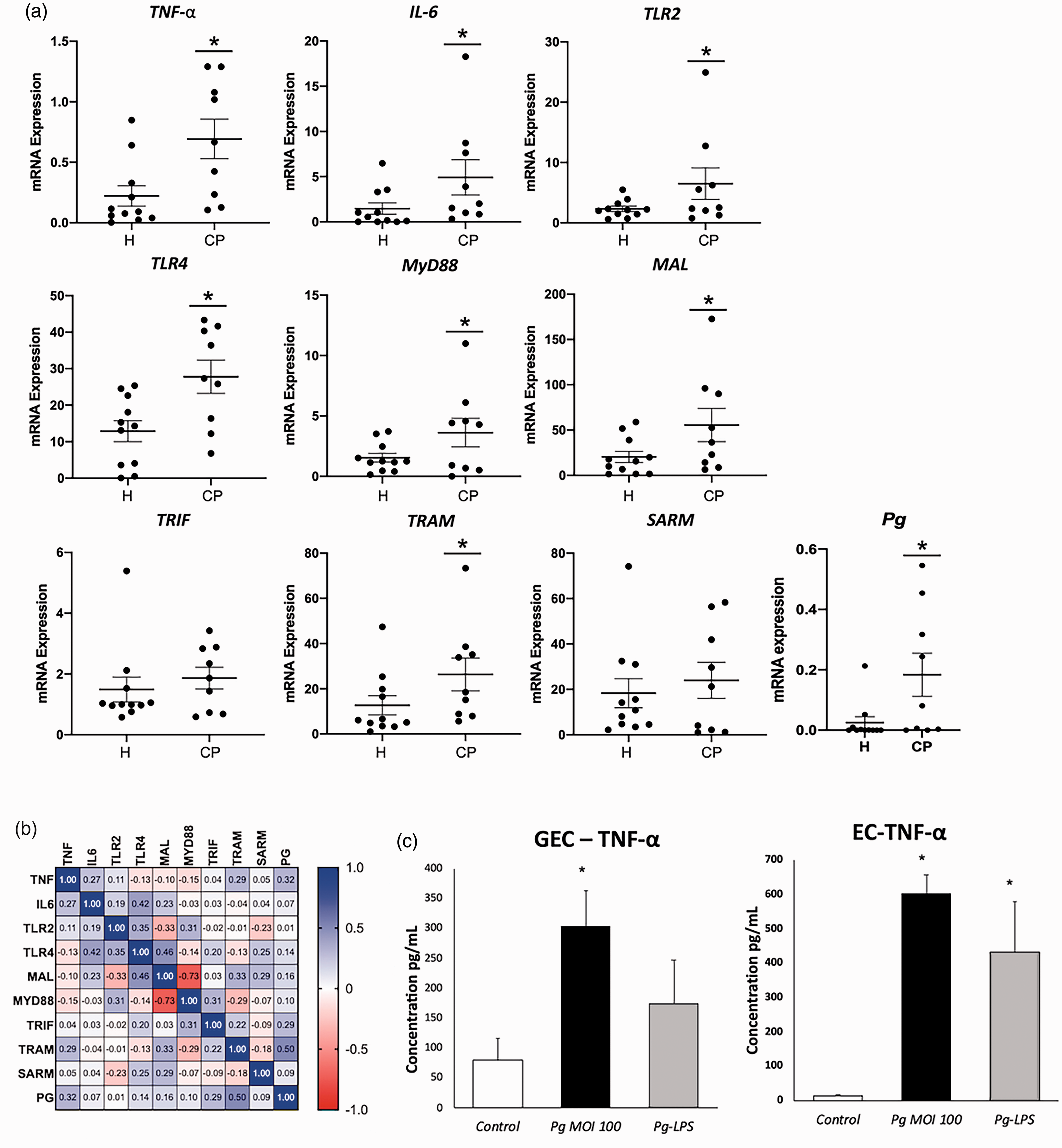
P. gingivalis infection induces an inflammatory response. (a) mRNA expression of inflammation related genes: TNF-α, Il-6, TLR2, TL4, MyD88, MAL, TRIF, TRAM, and SARM in gingival tissues harvested from chronic periodontitis (CP) and healthy patients (H). The presence of bacteria (Pg) was also quantified by TaqMan Real-Time PCR in gingival tissues. (b) Spearman’s rank correlation between the expression of all evaluated targets. (c) TNF-α secretion in supernatant of GEC and EC, in response to P. gingivalis (Pg) (MOI:100) and 1 μg/ml Pg-LPS for 24 h was measured by ELISA. All data were expressed as the mean mRNA expression normalized with the β-actin expression ± SD from three independent experiments *: P < 0.05 between CP and H samples and between cells infected or stimulated against control (unstimulated cells).
Interestingly, their increased expression was correlated with the presence of P. gingivalis (TNF-α: r = 0.32; IL-6: r = 0.07) (Figure 1B). The increased expression of TNF-α was also confirmed in response to P. gingivalis infection and stimulation with Pg-LPS in both GEC and EC, confirming their pro-inflammatory properties (fold change 3.8 and 40, respectively; P < 0.05) (Figure 1C). TLR2 and TLR4 have been identified as the principal signaling receptors for bacterial components involved in periodontal inflammation. In inflamed gingival tissues, a significant increase of TLR2 and TLR4 mRNA expression was observed (fold change 2.8 and 2.2, respectively; P < 0.05) concomitantly to the increased cytokine gene expression (Figure 1A). To evaluate the potential implication of TIR domain containing proteins in periodontal inflammation, their mRNA expression was evaluated in healthy and diseased tissues (Figure 1A) and a significant increase of MyD88, MAL, and TRAM was measured in tissues harvested from periodontitis patients (fold change 2.4, 2.8, and 2.3, respectively; P < 0.05) (Figure 1A).
Increased expression of TIR domain containing proteins in P. gingivalis-infected cells
In GEC and EC, infection with P. gingivalis up-regulated TLR2 and TLR4 mRNA expression (Figures 2 and 3).
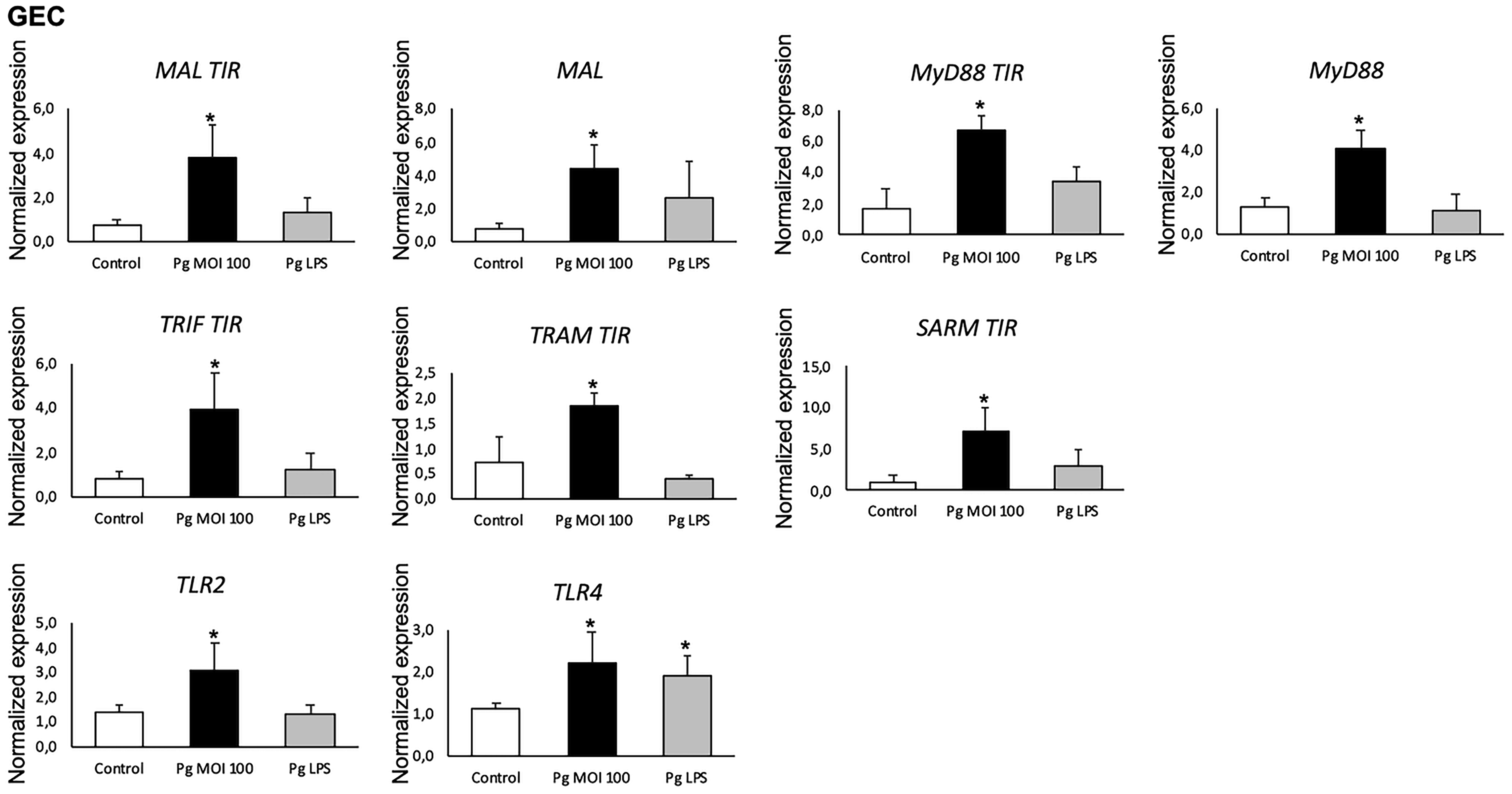
P. gingivalis significantly increased mRNA expression of TIR domain containing proteins in GEC. The mRNA expression of MAL and MyD88 with and without their TIR terminal domain, TLR2, TLR4 ,and other TIR domain containing proteins: TRIF, TRAM, and SARM in GEC infected with P. gingivalis (Pg) (MOI:100) and stimulated with Pg-LPS (1 µg/ml) for 24 h was measured by qRT-PCR. All data was expressed as the mean mRNA expression normalized with the β-actin expression ± SD from three independent experiments *: P < 0.05 between cells infected or stimulated against control (unstimulated cells).
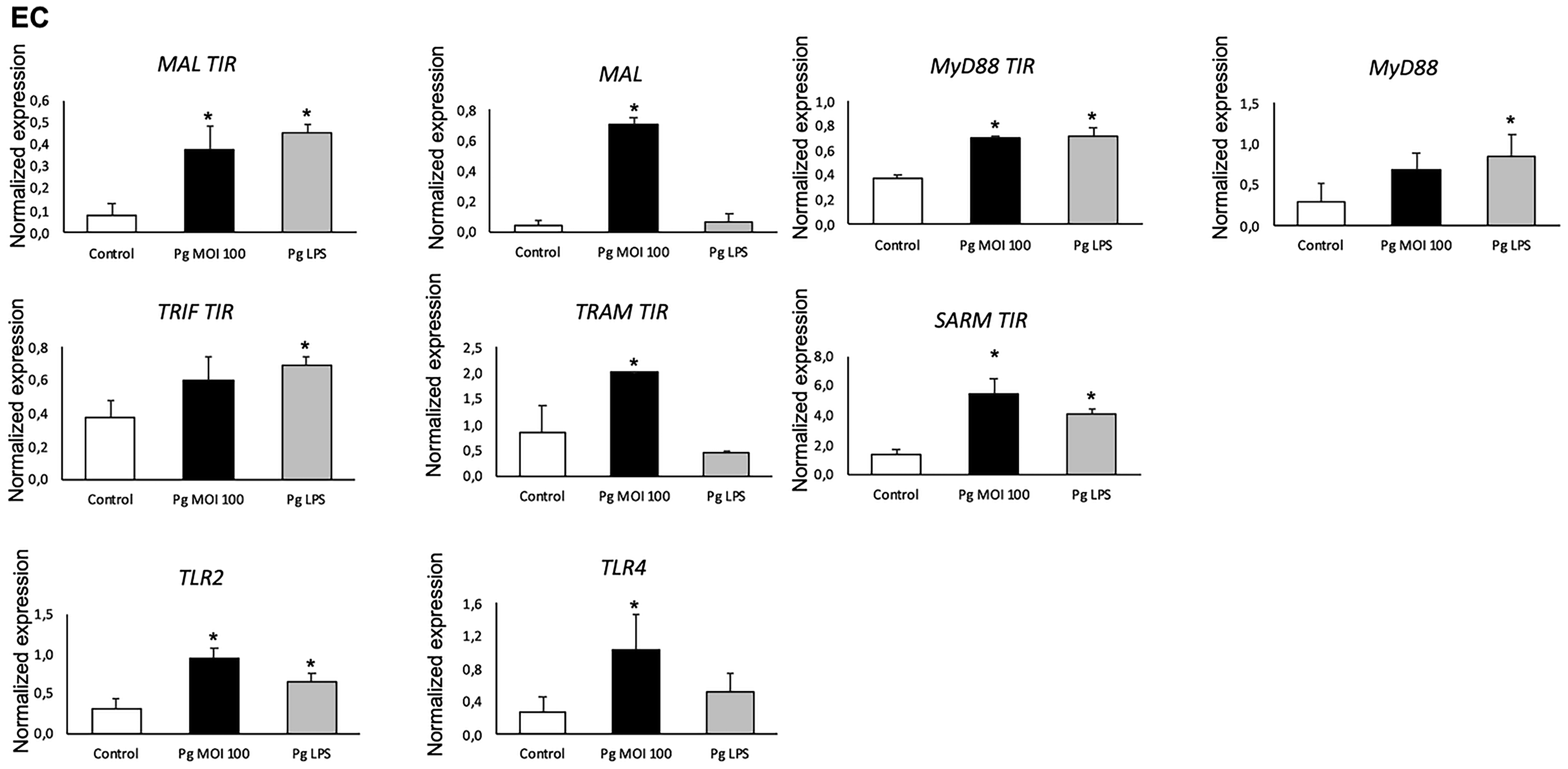
P. gingivalis significantly increased mRNA expression of TIR domain containing proteins in EC. The mRNA expression of MAL and MyD88 with and without their TIR terminal domain, TLR2, TLR4, and other TIR domain containing proteins: TRIF, TRAM, and SARM in GEC infected with P. gingivalis (Pg) (MOI:100) and stimulated with Pg-LPS, 1µg/mL) for 24 h was measured by qRT-PCR. All data was expressed as the mean mRNA expression normalized with the β-actin expression ± SD from three independent experiments *: P < 0.05 between cells infected or stimulated against control (unstimulated cells).
However, at the protein level, while both TLR2 and TLR4 were increased in GEC, only TLR4 was significantly up-regulated in EC (Figure 4).
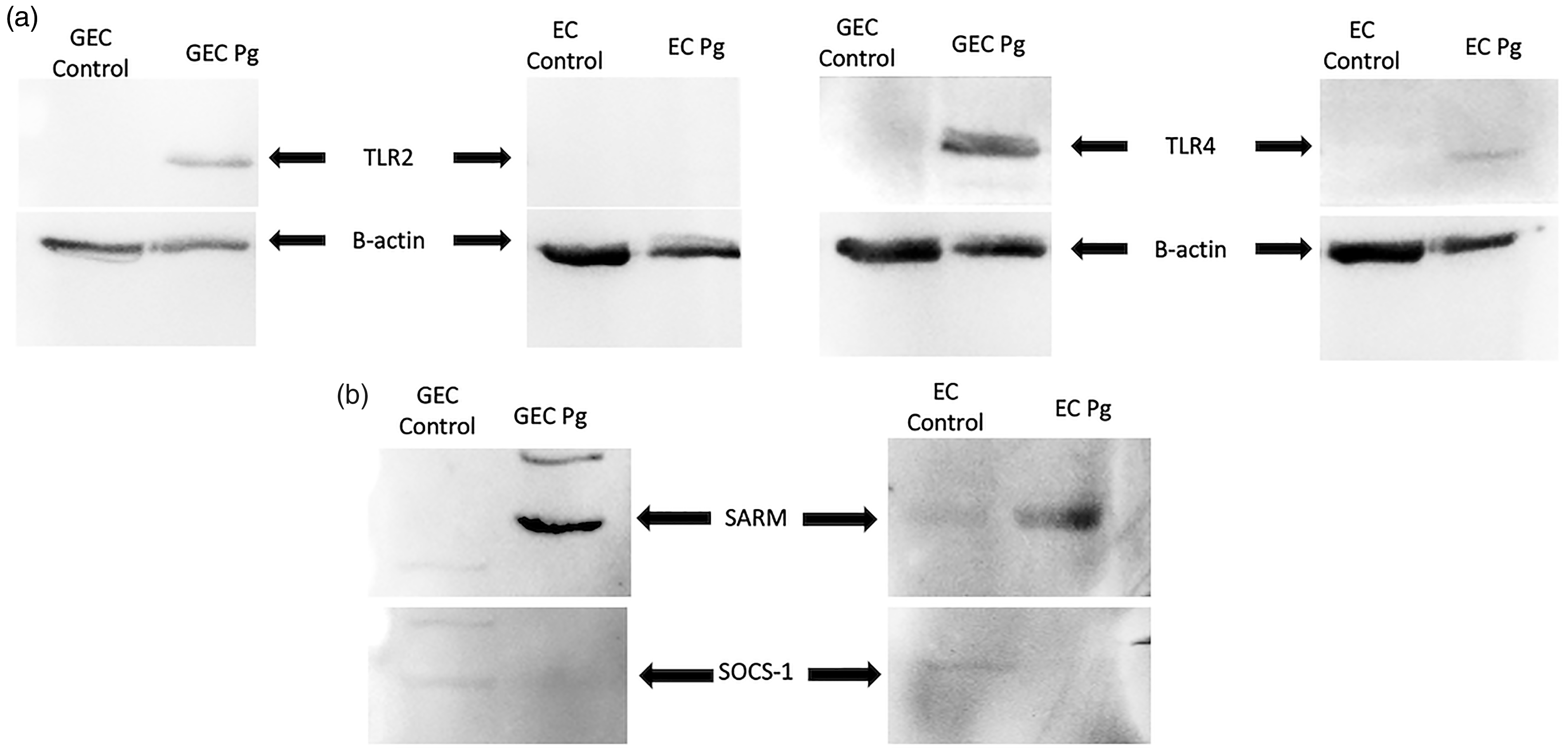
P. gingivalis induced the activation of TLR2 and TLR4, activation of the SARM adapter, and the inhibition of SOCS-1 protein expression. (A) Protein expression of TLR2, TLR4 in response to P. gingivalis (Pg) (MOI:100) for 24 h was evaluated by Western blot. (B) Protein expression of SARM and SOCS-1 in response to P. gingivalis (MOI:100) for 24 h in GEC and EC was evaluated by Western blot. All bands were compared and normalized with β-actin expression.
Additionally, an increased mRNA expression of the five adapters was also observed in GEC and EC after infection by P. gingivalis. Interestingly, MAL and TRAM showed a high increase in both cell types (Figures 2 and 3). The expression of the MAL and MyD88 adapters using primers designed for the adapter with and without its TIR terminal domain was evaluated. It showed that the TIR terminal domain is not mandatory for the expression of these two molecules at the mRNA level in response to P. gingivalis infection (Figure 2). Interestingly, Pg-LPS induced a specific profile of TIR domain containing protein expression in GEC and EC (Figures 2 and 3).
Similar patterns of TIR domain containing protein expression and TLR2/TLR4 after P. gingivalis infection
To evaluate the activation of TLR-associated pathways (Figure 2, 3, 4A) and the correlated involvement of TIR domain containing proteins, their expression was evaluated by immunofluorescence. Regarding the protein expression and the localization of molecules involved in TLR-associated pathways, after infection, a similar pattern of protein expression of adapters with TLRs was observed in both cell types (Figures 4A, 5-8).
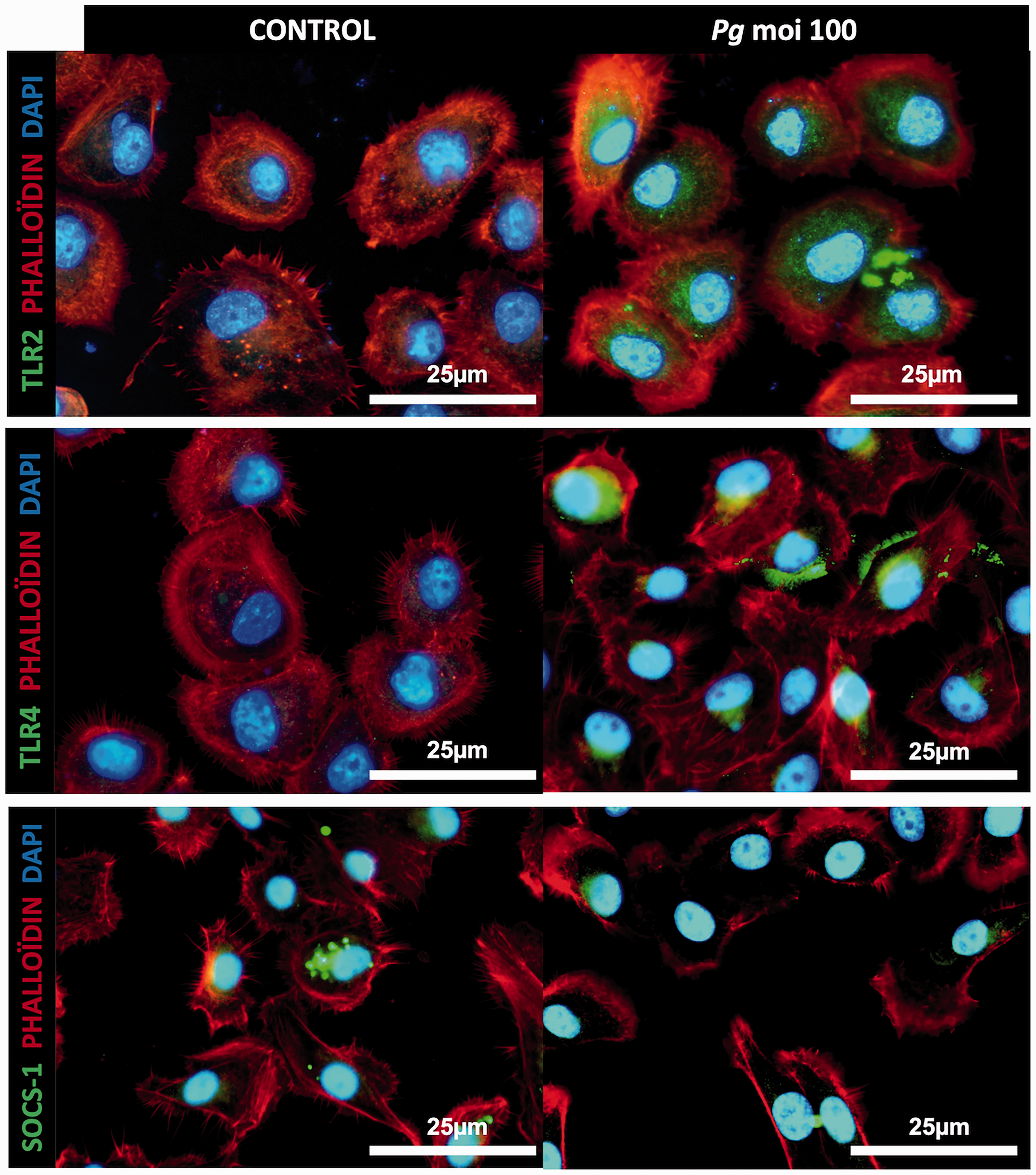
P. gingivalis infection induces an increased expression of TLR2 and TLR4 and decreased SOCS-1 expression in GEC. Immunofluorescence staining of uninfected and infected GEC with P. gingivalis (Pg) (MOI:100) for 24 h. In green: TLR2, TLR4, and SOCS-1; in red: phalloidin; and in blue: DAPI (nuclear staining). All images were acquired at 20× magnification.
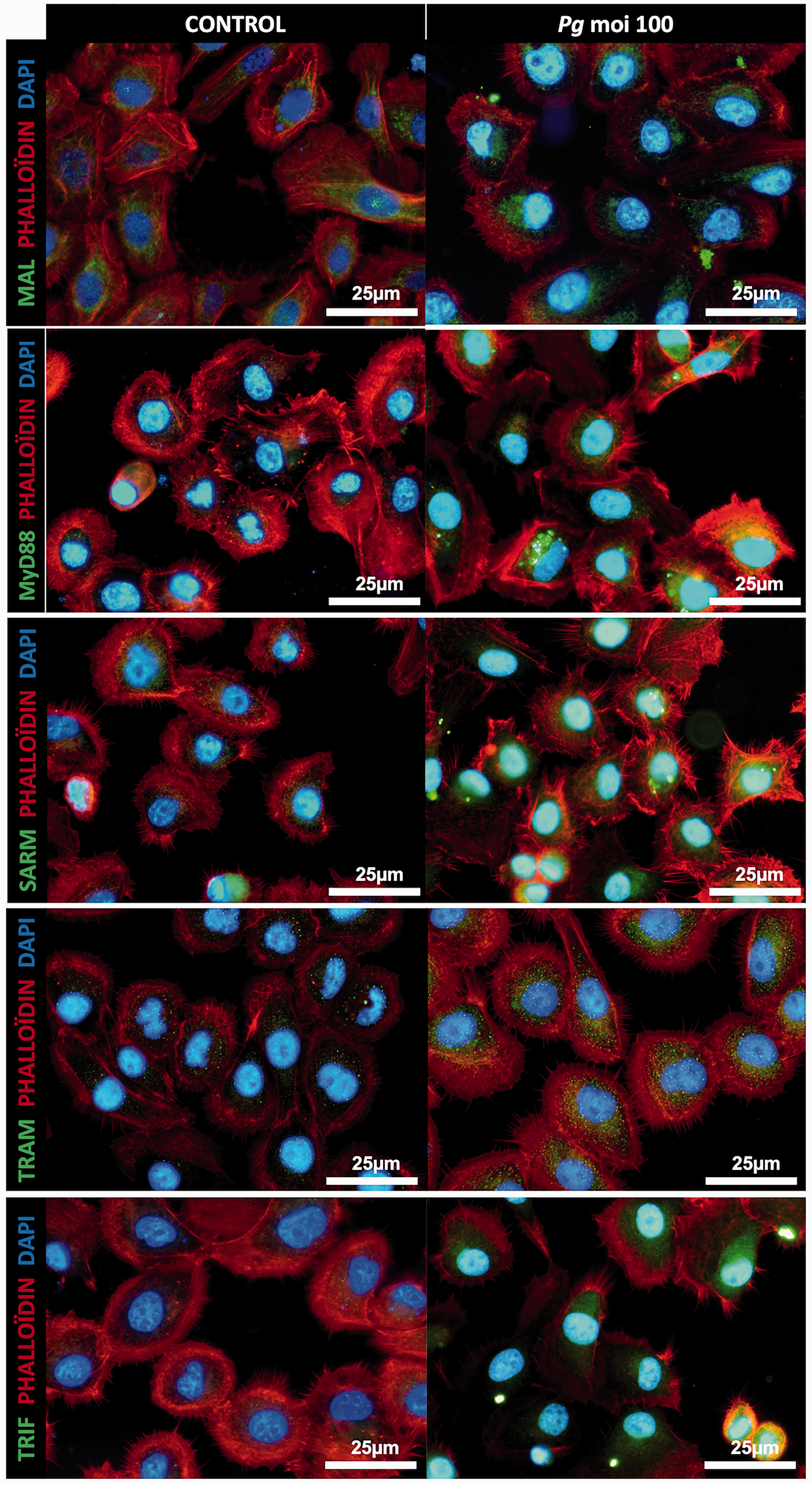
P. gingivalis infection induces an increased expression of the TIR domain containing proteins in GEC. Immunofluorescence staining of uninfected and infected GEC with P. gingivalis (Pg) (MOI:100) for 24 h. In green: MAL, MyD88, SARM, TRAM, and TRIF; in red: phalloidin; and in blue: DAPI (nuclear staining). All images were acquired at 20× magnification.
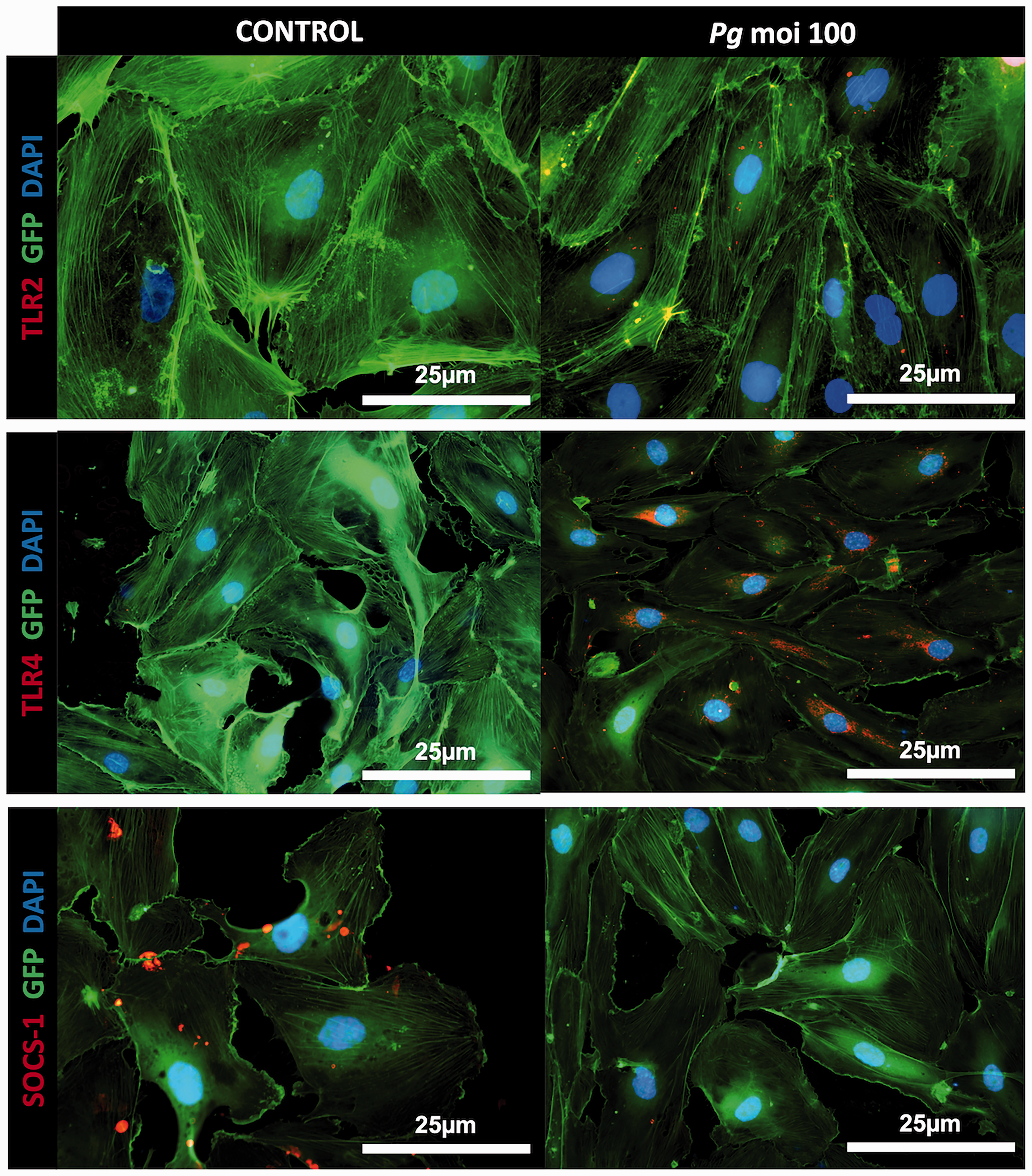
P. gingivalis infection induces an increased expression of TLR4 and decreased SOCS-1 expression in EC. Immunofluorescence staining of uninfected and infected EC with P. gingivalis (Pg) (MOI:100) for 24 h. In red: TLR2, TLR4, and SOCS-1; and in blue: DAPI (nuclear staining). GFP-EC were used in this experience. All images were acquired at 20× magnification.
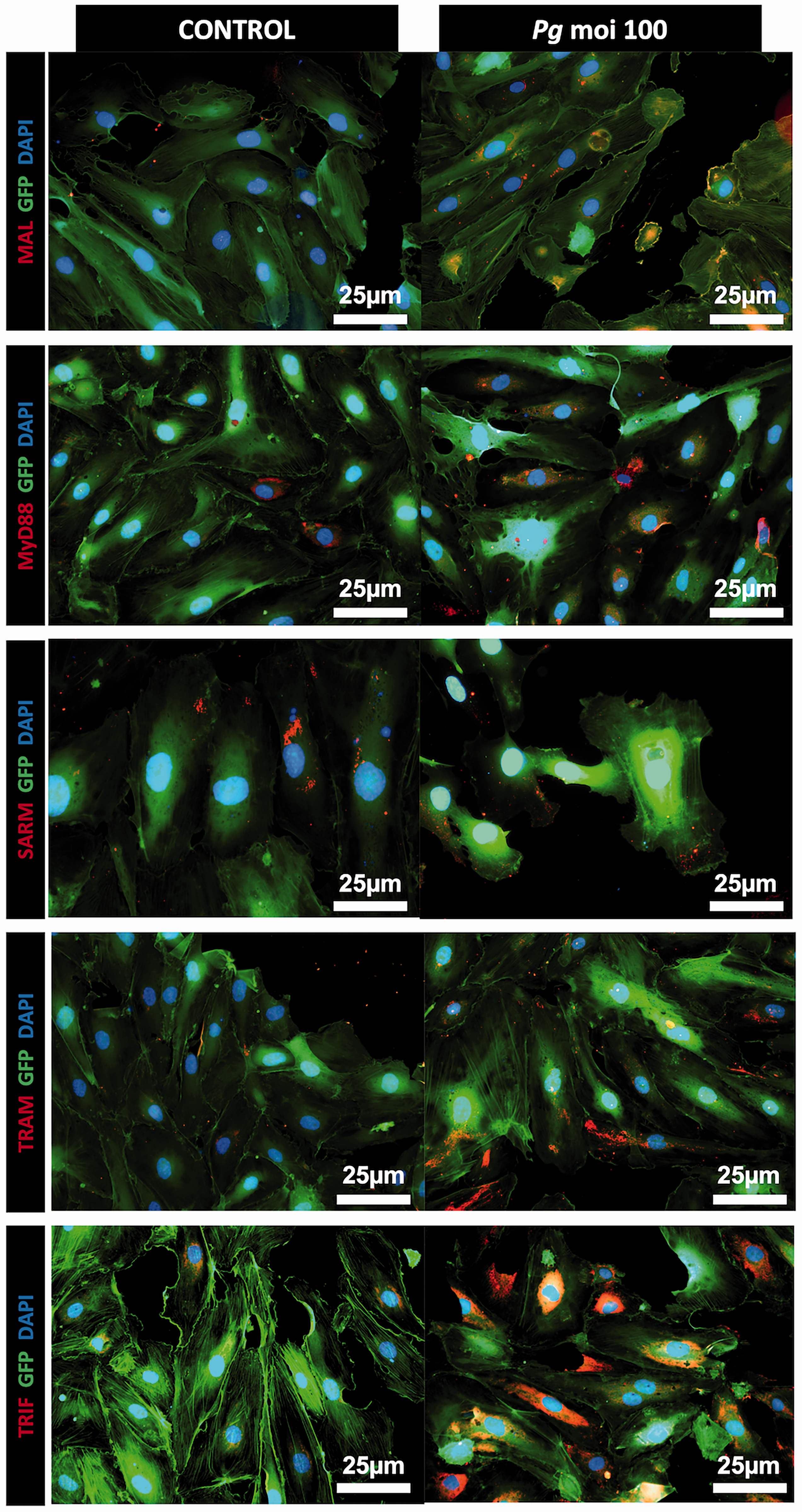
P. gingivalis infection induces an increased expression of the TIR domain containing proteins in EC. Immunofluorescence staining of uninfected and infected EC with P. gingivalis (Pg) (MOI:100) for 24 h. In red: MAL, MyD88, SARM, TRAM, and TRIF; and in blue: DAPI (nuclear staining). GFP-EC were used in this experience. All images were acquired at 20× magnification.
Moreover, this activation of TIR domain containing proteins was associated with a concomitant activation of the SARM adapter and the inhibition of SOCS-1 protein expression after P. gingivalis infection (Figure 4B).
MAL/MyD88 and TRAM/TRIF interactions are observed when TLR4 is activated by P. gingivalis infection
Following this significant increase of TLR2 and TLR4 expression in GEC and EC, both receptors were isolated by immunoprecipitation, and protein-protein interactions with the five TIR domain adapters were analyzed by co-immunoprecipitation (Figure 9) and pull-down assays (Figure 10) in GEC and EC infected with P. gingivalis (Table 2).
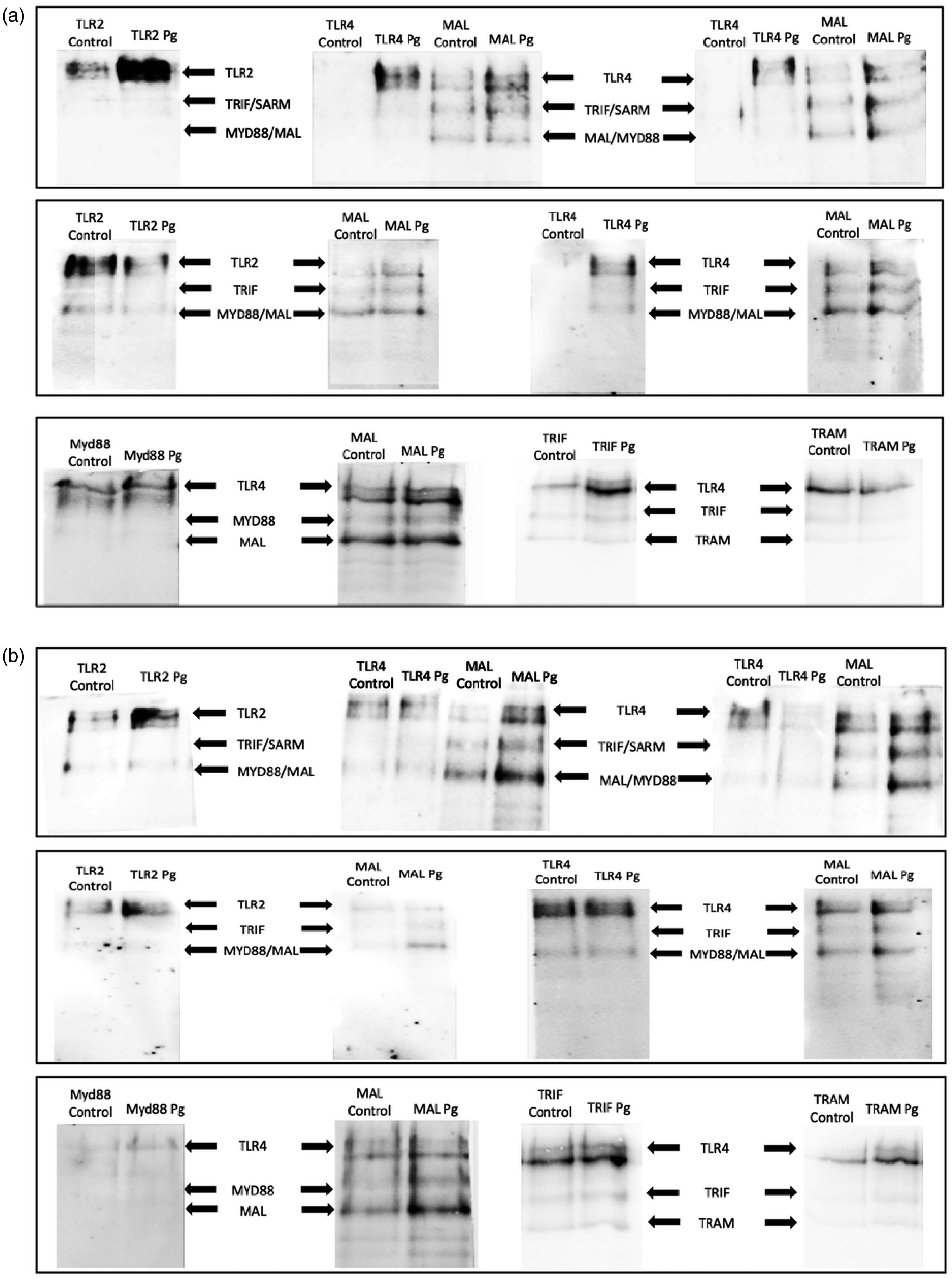
Co-immunoprecipitated proteins using Dynabeads Protein G, identifying interactions between the five TIR domains containing proteins (MyD88, MAL, TRIF, TRAM, SARM) and TLR2 and TLR4 in GEC (A) and EC (B), by SDS-PAGE and blotting of the protein complexes. In each column, the primary precipitated protein by the Ab capture is indicated in the upper part. Then, in each band, the presence and expression of the different proteins that are interacting at the time of precipitation are indicated with the arrows.
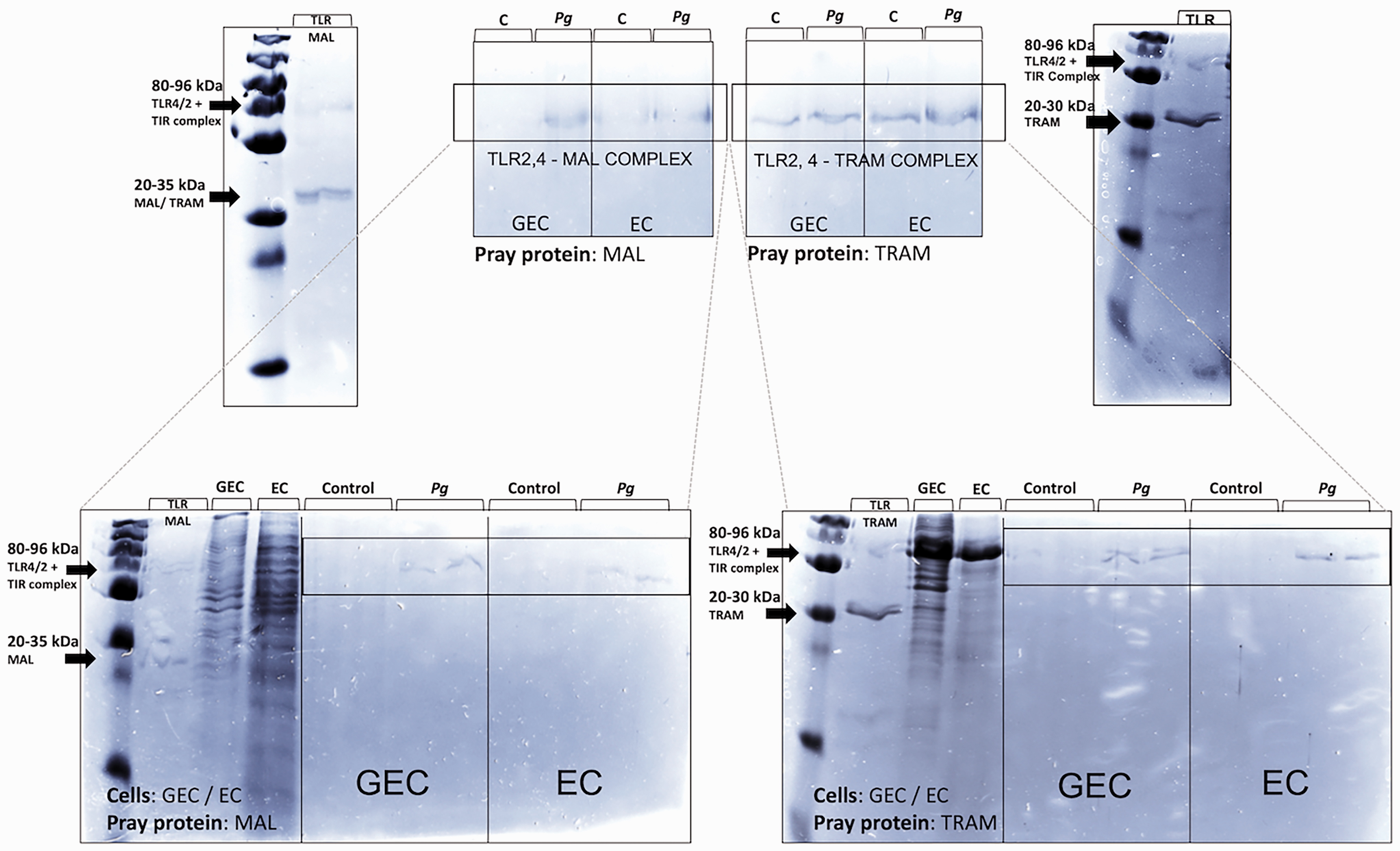
Analysis of pull-down assay of GEC and EC lysates untreated and infected with P. gingivalis (Pg) (MOI:100), considering the coupling of the desthiobiotinylated bait proteins MAL and TRAM to the streptavidin resin. All the final samples were analyzed by SDS-PAGE and stained by Coomassie Blue G-250 after pulling down proteins.
Summary of Western blot, pull-down assay and co-immunoprecipitation in GEC (A) and EC (B) lysates of untreated and infected with P. gingivalis (Pg) (MOI:100) cells. +++ when the protein was highly increased, ++ when proteins were moderately increased, + when proteins were slightly increased or +/- not significantly increased. ND: not determined.

TLR2 or TLR4 activation was mainly associated with the increase of MAL and MyD88 adapter interactions and protein expression (evaluated through their presence on the co-immunoprecipitated protein complex) and very slightly with TRIF and TRAM adapters in both cell types (Figures 9-10 and Table 2).
Pull-down assays demonstrated that, following P. gingivalis infection, an increase of TLR-MAL or TLR-TRAM complexes is detected in both cell types, emphasizing a putative critical role for these two adapters in the development of the immune response (Figure 10).
MAL and TRAM silencing decreased P. gingivalis elicited inflammation
To evaluate the impact of MAL and TRAM on the inflammatory process induced by P.gingivalis, GEC and EC were transfected with MAL and TRAM-targeted siRNA (Figure 11).
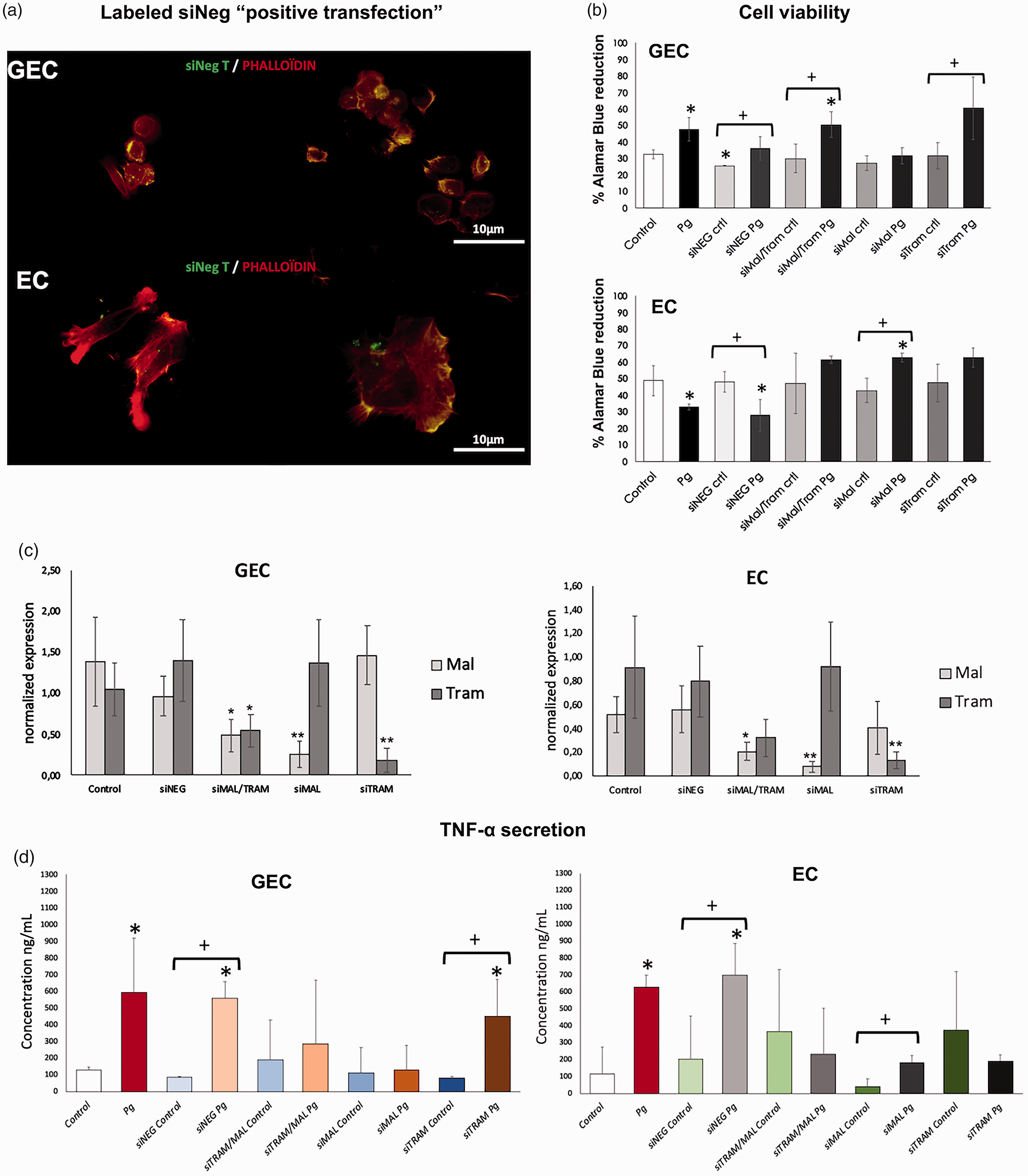
Decrease of P. gingivalis inflammatory effects in GEC and EC through MAL and TRAM-targeted inhibition. (A) Immunofluorescence staining of siRNA transfected GEC and EC for 8 h. In green: siRNA fluorescence negative control; and in red: phalloïdin. All images were acquired at 40× magnification. (B) Metabolic activity of GEC and EC transfected with siRNA against MAL, TRAM, or both and then infected with P. gingivalis (Pg) at MOI of 100 for 24 h. (C) mRNA expression of TRAM and MAL after siRNA silencing in GEC and EC. All data was expressed as the mean mRNA expression normalized with the β-actin expression. (D) TNF-α secretion in supernatant of GEC and EC, in response to P. gingivalis (Pg) (MOI:100) for 24 h was measured by ELISA after transfection with siRNA against MAL, TRAM, or both. *: P < 0.05 for differences versus control. **: P < 0.01 for differences versus control. +: P < 0.05 between selected conditions.
As observed previously, P. gingivalis infection modulated cell metabolic activity in a cell-dependent manner. Indeed, it induced an increase in GEC metabolic activity while a decrease of EC metabolic activity was measured (Figure 11B). Interestingly, such results were not observed in siRNA treated cells (especially EC) emphasizing a role for MAL and TRAM in the effect induced by P. gingivalis (Figure 11B). Nevertheless, TNF-α secretion was also significantly decreased in transfected cells infected with P. gingivalis (Figure 11C).
Discussion
In this study, the pro-inflammatory impact of P. gingivalis was confirmed in both epithelial and EC. Such an effect was associated with the modulation of the expression of TIR domains containing proteins, especially MAL and TRAM. Indeed, when MAL and TRAM silencing was performed, P. gingivalis-induced inflammation was significantly decreased.
The deleterious impact of P. gingivalis on cellular homeostasis has been demonstrated in several tissues and cells emphasizing its pathogenicity in periodontal destructive processes, but also, at distance, in vascular cells. This bacterium possesses a wide array of virulence factors and develops different strategies, allowing its survival and spreading through the induction of a sustained inflammation. Indeed, a role for TIR domain-containing adapters, especially MAL and TRAM, was highlighted in the inflammatory response triggered by P. gingivalis. P. gingivalis is highly prevalent in patients with severe periodontitis. 37 , 38 Its multiple virulence factors activate inflammation and immune system responses, resulting in the subsequent destruction of the periodontium. 10 , 39 Indeed, its recognition by TLRs, TLR2 and TLR4, induces the modulation of several biological processes, such as apoptosis and the cell cycle, that are correlated with the exacerbated secretion of pro-inflammatory mediators such as TNF-α, interleukins, and also proteolytic enzymes such as matrix metallo-proteinases, allowing it to escape the immune system and to induce the formation of a niche. 9 , 11 , 38 , 40 , 41 Such effects have been observed in periodontal tissues but also in the endothelium, where P. gingivalis is frequently detected 12 , 14 , 18 , 39 , 40 and elicits pro-senescent and pro-oxidative effects that may be part of the development of atherothrombosis. 14 , 19 , 42 In this study, analysis of the response of epithelial cells and EC to bacterial insult has been performed as both cell types are considered the first line of defense against pathogens, at the gingival level for epithelial cells and at the vascular level for EC. 43 , 44 Here, we confirmed the pro-inflammatory effect of P. gingivalis in both cell types and the involvement of TLRs in early bacterial recognition, 45 and suggest a role for TIR domain-containing adapters in the activation of the host response to such periodontal pathogens.
Due to their involvement in downstream signaling, TIR domains containing proteins play an important role in the innate immune response of mammals and other types of animals and appear to be a target for several pathogens to subvert it. Indeed, this subversion or hijacking of the host response was observed for bacteria such as Brucella melitensis, 47 Escherichia coli, 48 and periodontal pathogens such as P. gingivalis. 43 In human gingival samples, it was observed an increased expression of TIR domain-containing adapters in periodontitis affected gingival tissues in comparison with healthy tissues. Such an observation emphasizes their instrumental role in inflammation and immune responses to pathogenic biofilms. 50 Here, we observed that P. gingivalis was able to differentially modulate the expression of TIR domain-containing adapters, mainly associated with the TLR4 signaling pathway in GEC and EC. Such modulation of their expression was closely related to the secretion of pro-inflammatory cytokines such as TNF-α, especially for MAL and TRAM. The role of MAL in P. gingivalis elicited TNF-α, but IL-8 secretion was also observed in neutrophils where transfection with MAL siRNA significantly reduced their secretion. 51 The role of MAL in the induction of inflammatory responses is well described and several pathways have been suggested, one of them involving the recruitment of TRAF6 to the plasma membrane allowing the transactivation of TLR2 and TLR4. 52 Congruently, the transfection with TRAM siRNA significantly reduced the secretion of TNF-α induced by P. gingivalis infection, especially in EC. TRAM has been demonstrated as being over-expressed in cases of bacterial insult such as LPS stimulation, notably in macrophages. 53 Interestingly, the potential targeting of such pathways was demonstrated with Annexin A2, which protects the host from excessive inflammatory damage through the activation of TRAM-dependent endosomal signaling, thus promoting the release of anti-inflammatory cytokines such as IL-10. 54
Aside from the increase in their expression, a differential localization pattern of TIR was observed between GEC and EC. Indeed, SARM and SOCS-1 are predominantly nuclear in GEC but not in EC. As demonstrated for MyD88, subcellular localization plays a key role in the signaling function as well as its tridimensional conformation. 55 Interestingly, regarding SARM, earlier studies showed that it associates with the outer membrane of the mitochondria, however, its expression in the nucleus and elsewhere in the cell was also observed. 56 In the study of Paneerselvam et al., focusing on the effect of SARM in apoptosis, it was observed that the mutation of one amino acid in the SARM targeting sequence (Arg 14 ) reduced its apoptotic potential by 50%, demonstrating that its mitochondrial localization is important but not the sole mechanism involved as some other residues or other factors could be involved. Therefore, a specific investigation should be performed in the future to determine the importance of the localization of adapters in the host response modulation and if such a process is cell-dependent.
In this study, we focused on P. gingivalis infection, considering its high proportion in severe forms of the disease and its role as a keystone pathogen. 7 , 24 However, complex biofilms are found within periodontal pockets as well as in atherothrombotic-related endothelium,57–59 sustaining the need to assess the impact of multispecies infection on TIR domain activation and signaling. Nevertheless, as immune cells are key effectors in the periodontal host response and periodontal destruction, TIR domain activation should also be investigated in such cells (neutrophils, macrophages, …) and results need to be confirmed in vivo in experimental periodontitis models to overcome limitations associated with in vitro models. As we observed a difference in terms of gene expression in human tissues, it should be mentioned that those observed patterns of expression should be analyzed in a clinical trial, considering all potential influencing factors (age, gender, local risk factors, stage, and grade of periodontitis).
Finally, due to the involvement of TIR domain-containing adapters, especially MAL and TRAM, in the adverse cellular response to P. gingivalis their targeting could be envisaged as a potential new therapeutic tool for periodontitis management. The development of specific drugs should, therefore, be developed.
Supplemental Material
sj-pdf-1-ini-10.1177_17534259211013087 - Supplemental material for Implication of Toll/IL-1 receptor domain containing adapters in Porphyromonas gingivalis-induced inflammation
Supplemental material, sj-pdf-1-ini-10.1177_17534259211013087 for Implication of Toll/IL-1 receptor domain containing adapters in Porphyromonas gingivalis-induced inflammation by Isaac M Bugueno, Nadia Benkirane-Jessel, Olivier Huck in Innate Immunity
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Agence Nationale pour la Recherche (ANR ENDOPAROMP ANR- 17-CE17-0024-01).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
