Abstract
Urinary tract infections (UTI), associated with vesicoureteral reflux (VUR), can lead to chronic kidney disease. Genetic alterations in the innate immune defenses contribute to UTI risk. We investigated a novel gene, Dachsous Cadherin-Related 1 (DCHS1), in children with UTI. We determined absolute DNA copy number (CN) of DCHS1 in children with UTI. In this case-control study, we utilized multiple complementary methods to determine the genomic CN of DCHS1. Children with (n = 370) and without (n = 71) VUR from two well-phenotyped clinical trials of UTI were copy-typed and compared to 491 healthy controls with no known history of VUR or UTI. Less than 1% of controls had a single copy of DCHS1, while 31% of children with UTI and no VUR and 7% of children with UTI and VUR had a single copy of the DCHS1 gene. Using immunostaining, we localized expression postnatally to the bladder and renal epithelia. Mice were also challenged with two uropathogenic Escherichia coli strains, and Dchs1 mRNA was quantified. This study represents the first report of DCHS1 in association with pediatric UTI. We hypothesize that its role in innate immunity is critical to lower urinary tract defense. Further investigation is required to determine the role of DCHS1 in innate immunity.
Introduction
Urinary tract infection (UTI) is one of the most common childhood infections and is caused by uropathogenic Escherichia coli (UPEC) in 80% of cases. 1 Acute pyelonephritis has variable sequelae with or without renal scars, depending on the virulence of the organism and host defense mechanisms.2,3 Children with congenital anomalies of the kidney and urinary tract, such as vesicoureteral reflux (VUR), are at increased risk for UTI-associated sequelae such as chronic kidney disease and renal scarring. 4
The innate immune system plays a critical role in protecting the urinary tract from uropathogens.5,6 Innate immune responses are rapid and encompass a variety of components, including epithelial extracellular barriers and intracellular tight junctions, antimicrobial peptides, PRRs, inflammatory mediators, and phagocytes.6–8 PRRs are protein receptors on the surface of phagocytes, dendrites, and epithelial cells, which recognize microbial molecules and are capable of initialing immune responses called PAMPs. 9
A number of studies implicate a heritable component to UTI susceptibility.10,11 A potential mechanism for UTI heritability is genetic alterations in innate immune defenses. 12 For example, DNA copy number (CN) variations (CNVs) in the gene that encodes α-defensin 1 have been shown to be associated with variable susceptibility to UTIs and response to antibiotic prophylaxis. 13 CNVs occur when DNA sections between a few hundred and several million bp are deleted or duplicated and influence disease by a range of mechanisms, including gene dosage.14,15 We performed high-resolution genome-wide array comparative genomic hybridization (aCGH) comparing children with VUR and UTI to healthy controls. We identified a relative CNV in the gene Dachsous Cadherin-Related 1 (DCHS1) that segregated to patients with VUR. 16 A limitation of aCGH is that it only determines if DNA CN is relatively increased or decreased compared to a reference sample and does not define the absolute number of gene copies that are present.13,16 Thus, validation methods to determine true CNs are needed to rule out false-positives. 13
DCHS1 encodes for a calcium-dependent adhesion molecule, which has a role in kidney development and planar cell polarity.16,18 The gene has not been previously investigated in the context of UTI. The purposes of this study were to determine absolute DCHS1 CN in two pediatric clinical trial cohorts of UTI and to investigate its postnatal role in innate immune defenses.
Methods
Safety approval of human subjects
Approval of human subjects was obtained through the University of Tennessee Health Science Center Institutional Review Board Protocol 14-03325-XP. All clinical investigations were conducted according to the principles expressed in the Declaration of Helsinki.
Cohorts
We utilized genetic samples from three clinical cohorts of children and adults with and without a history of UTI. All genomic DNA samples were obtained from immortalized lymphoblastic cell lines. Each is described below.
Cohort of subjects with VUR and UTI
The Randomized Intervention for Children with Vesicoureteral Reflux (RIVUR) Clinical Trial (ClinicalTrial.gov Identifier NCT00405704) enrolled patients aged 2–71 mo with one to two UTI who were diagnosed with grade 1–4 VUR. 19 Patients were randomized to receive antibiotic prophylaxis or placebo and were followed for 2 yr for recurrent UTI. UTI was defined as a positive culture with a single organism of 50,000 CFU/ml on a catheter specimen or 100,000 CFU/ml on a clean catch specimen with evidence of pyuria on the urinalysis, as per guidelines by American Academy of Pediatrics (AAP) at the time of collection. 20 We utilized DNA obtained from 320 non-Hispanic Caucasian girls from the RIVUR study.
Cohort of subjects with UTI without evidence of VUR
The Careful Urinary Tract Infection Evaluation (CUTIE) Study observed 2–71 mo old pediatric patients who had one to two UTI without VUR for a 2 yr period. 21 The CUTIE Study was concomitant with the RIVUR study. We utilized 71 DNA samples from non-Hispanic Caucasian girls enrolled in the CUTIE Study.
Control cohort without VUR and UTI
We utilized 491 DNA samples of healthy, non-Hispanic Caucasian adult women with no known history of VUR and UTI.
Mice
In vivo murine studies were approved by Indiana University Institutional Animal Care and Use Committee (IACUC) under protocol # 11333. Wild-type C57BL/6J female mice (Jackson Laboratory, Bar Harbor, ME) were used for induction of pyelonephritis and cystitis in infection studies.
Absolute CN determination
Absolute CNs of DCHS1 were determined by digital PCR (dPCR) and were confirmed with Short Multiply Aggregated Sequence Homologies (SMASH) methods for seven reference individuals who were used on each plate as calibrators for relative quantitation for quantitative RT-PCR (qRT-PCR).
dPCR
To determine absolute DCHS1 CN, quantitative dPCR was performed on seven reference individuals at the Molecular Resource Center at the University of Tennessee Health Science Center on a Fluidigm BioMark System (San Francisco, CA). For complete methodology, please refer to the manufacturer’s protocol (Fluidigm PN 100-6896 B1). Briefly, we used dPCR 37 K integrated fluidic circuits (IFC). After filling IFC with standard control line fluid, we primed the IFC in Fluidigm MX. A pre-mix sample mixture, which included TaqMan gene expression master mix (Applied Biosystems, Foster City, CA), 20× GE Sample Loading Reagent, and TaqMan copy-typing primer/probe mixtures (Applied Biosystems), was used.
Specifically, RNASEP VIC-labeled probes were used as an internal reference for diploid genes and DCHS1-FAM labeled probes for our target gene (Applied Biosystems). We added 1.8 µl sample DNA (20 ng/µl) to 4.2 µl above mixture in each pre-mix well. After priming, IFC 37 K was loaded with 4.0 µl from each sample. Each sample was genotyped in triplicate. The data were analyzed with Fluidigm Digital PCR Analysis Software v4.1.2.
SMASH
We performed a next-generation sequencing-based method for CNV analysis by SMASH, as previously described. 22
qRT-PCR
Multiplex qRT-PCR was performed on all subjects and controls to determine the relative CN of the DCHS1 gene, as previously described.
13
Briefly, individuals with known CNs from dPCR and SMASH sequencing served as calibrators to determine CN for unknown samples using a standard curve for ΔΔCt. TaqMan copy-typing primer/probe mixtures (Applied Biosystems) were used. RNASEP-VIC-labeled probes were used as an internal reference for diploid gene, and DCHS1-FAM labeled probes for the target gene. Each sample was performed in triplicate. Experiments were performed on a Light Cycler 480 (Roche, Basel, Switzerland) Real-Time PCR System. Primers/probe sequences were as follows: DCHS1_F 5′-CGG TAA GGC AGG TTG GTG-3′; Probe FAM-
The comparative cycle threshold (Ct) difference method was used to determine the CN for the DCHS1 gene for each patient. 23 This was done using LCS480 v1.5.1.62 software and Microsoft Excel 2010. The method measured the ΔCt by subtracting the average Ct of DCHS1 as the target and the average Ct RNASEP as the reference and then calculating the ΔΔCt value by comparing the average ΔCt values of samples with the average ΔCt of the calibrator sample known to have two copies of the target sequence. 24 Known calibrators with two CNs of DCHS1 gene per diploid genome as determined by dPCR and SMASH were included on each plate to create a standard curve for CN in unknown. The amount of target normalized to endogenous reference and relative to calibrator is calculated by 2–ΔΔCt.
Immunofluorescence staining
Paraffin embedded human kidney and bladder sections (7 μm thick) were stained with polyclonal rabbit anti-DCHS1 Ab (cat. no. Ab203690; Abcam, Cambridge, MA) to locate DCSH1-positive cells in the kidney and bladder and V-ATPase E1 (cat. no. GW22284F; Sigma–Aldrich, St. Louis, MO) in conjunction with DCHS1 to locate intercalated cells positive for DCHS1 in the kidney. Human kidney samples with and without pyelonephritis were obtained from the Cooperative Human Tissue Network (Westerville, OH), as previously described. 25 DCHS1 primary Ab was added at a 1:50 dilution, and V-ATPase E1 was added at 1:300 dilution and incubated overnight at 4°C. To visualize, DCHS1 and anti-Rabbit Cy3 (1:600 final dilution) secondary Ab and to locate intercalated cells, anti-Chicken AF488 (1:600 final dilution) was used (Jackson ImmunoResearch, West Grove, PA) for 30 min at 20°C. Slides were mounted using DAPI containing a hard-set solution to visualize nuclear staining (Vector Laboratories, Burlingame, CA). Sections were visualized using a Keyence BZ9000 microscope (Keyence Corporation, Osaka, Japan).
Murine bacterial challenge experiments
To determine the mRNA in mouse models with UTI compared to control, E. coli pyelonephritis stain (UPEC; CFT073) and cystitis strain (UPEC; UTI89) were grown overnight on Luria-Bertani (LB) media at 37°C in a 20xg shaker and static incubator, respectively. OD600 measurements were recorded using the Eppendorf Biophotometer Spectrophotometer UV/Vis Reader (Eppendorf North America, Hauppauge, NY) to adjust the bacterial inoculum concentration to 1 × 108 CFU/ml per mouse. Bacterial cultures were made into two 1 ml aliquots and centrifuged at 5,500 g for 1 min. Supernatant was discarded without disturbing the pellet. Sterile PBS (1 ml) was then added to each tube. Tubes were vortexed several times in order to suspend the bacteria pellet in sterile PBS. The goal optical density of the bacterial solution was 0.073 OD600, which correlated to a concentration of 1 × 108 CFU/ml. Specifically, a mixture of 800 µl sterile PBS and 20 µl bacterial solution were combined and placed in a cuvette to be measured in the OD600 reader. From this baseline measurement, the bacterial/PBS suspension was added from the stock in 20 µl increments until an OD600 of 0.073 was reached. The mice were anesthetized with 2.5% isoflurane and then placed in the supine position. Bladders were palpated, and urine was expelled before transurethral inoculation was performed. Then, a sterile 1 cc syringe (Becton Dickinson, Franklin Lakes, NJ) with lubricated polyethylene-coated needle (Becton Dickinson, Franklin Lakes, NJ) was used to administer 50 µl of bacterial suspension transurethrally. The same volume amount of sterile PBS was inoculated transurethrally as a negative control. Mice were euthanized 6 h after inoculation of bacteria by CO2 inhalation for organ harvesting.
mRNA extraction and quantification
For quantifying the mRNA in mice, murine kidneys and bladders were placed into zirconium bead/PBS vials for homogenization in the Bullet Blender Storm 24 (Next Advance, Troy, NY) for 10 min at maximum speed. Following tissue homogenization, mRNA was extracted using an RNeasy Plus Mini Kit (Qiagen, Valencia, CA) and reverse transcribed using a High Capacity cDNA Reverse Transcription Kit (Applied Biosystems) as per the manufacturer’s protocol. qRT-PCR was performed using gene expression Dchs1 primers (Sigma–Aldrich) and Gapdh (Real Time Primers, LLC, Elkins Park, PA) as the housekeeping gene primer. The forward and reverse primer sequences for Dchs1 were 5′-
Statistical analyses
In genetics, the results were analyzed using SAS v9.4 (SAS Institute, Cary, NC). Categorical variables were compared using the chi-square test or Fisher’s exact test, depending upon the variable number in the box. The level of significance was defined as a P value of < 0.05. In mRNA comparisons, normalized Dchs1 mRNA expression data are presented as the mean ± SD. Statistical analysis by unpaired two-tailed t-test was used to compare the response between UPEC to saline control groups and plotted in GraphPad Prism 8 (GraphPad Software, La Jolla, CA). A P value of < 0.05 was considered statistically significant.
Results
Subject and control cohort characteristics
We determined DCHS1 gene CN per diploid genome using qRT-PCR in RIVUR (UTI and VUR; n = 320) subjects, CUTIE (UTI and no VUR; n = 71) subjects and controls (no UTI and no VUR; n = 491). All cohorts comprised non-Hispanic Caucasian females. The clinical characteristics of the groups are given in Table 1.
Clinical characteristics of subjects and controls.

Children with VUR and UTI had decreased CN of DCHS1 compared to controls
DCHS1 CN in children with RIVUR varied from 1 to 3, with a mean CN of 1.95 ± 0.30. DCHS1 CN for controls ranged between 1 and 4, with a mean CN of 2.49 ± 0.39. The distribution of CNs is shown in Figure 1. More RIVUR subjects (n = 23; 7%) had one copy of DCHS1 compared to controls (n = 3; 0.6%). In the RIVUR cohort, one copy individuals were 13 times more prevalent (odds ratio (OR) = 12.59; 95% confidence interval (CI) 3.75–42.32; P < 0.0001). Distribution of 1 versus ≥ 2 CNs in groups is given in Figure 2.
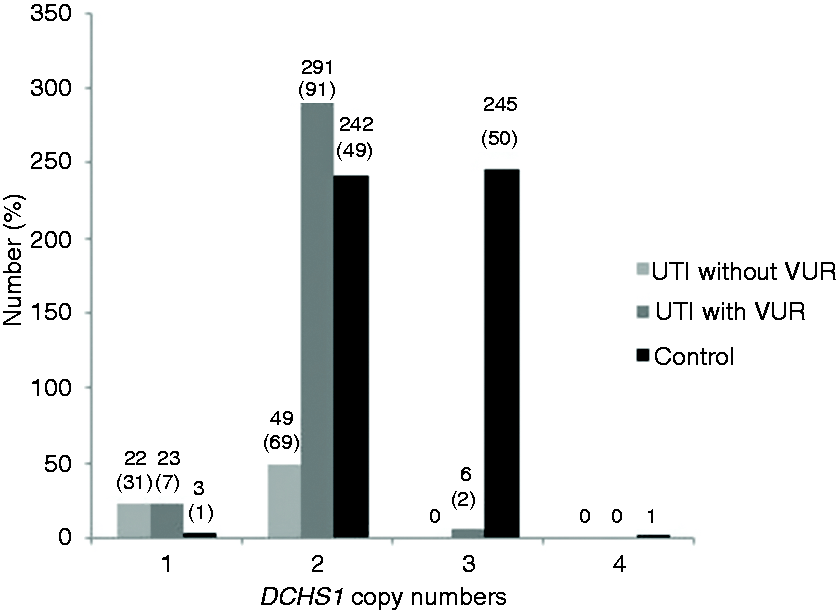
DCHS1 DNA copy number (CN) in subjects with UTI without vesicoureteral reflux (VUR), UTI with VUR, and controls. DCHS1 CN distribution and percentages in subjects with UTI without VUR (n = 71), UTI with VUR (n = 320), and controls (n = 491).
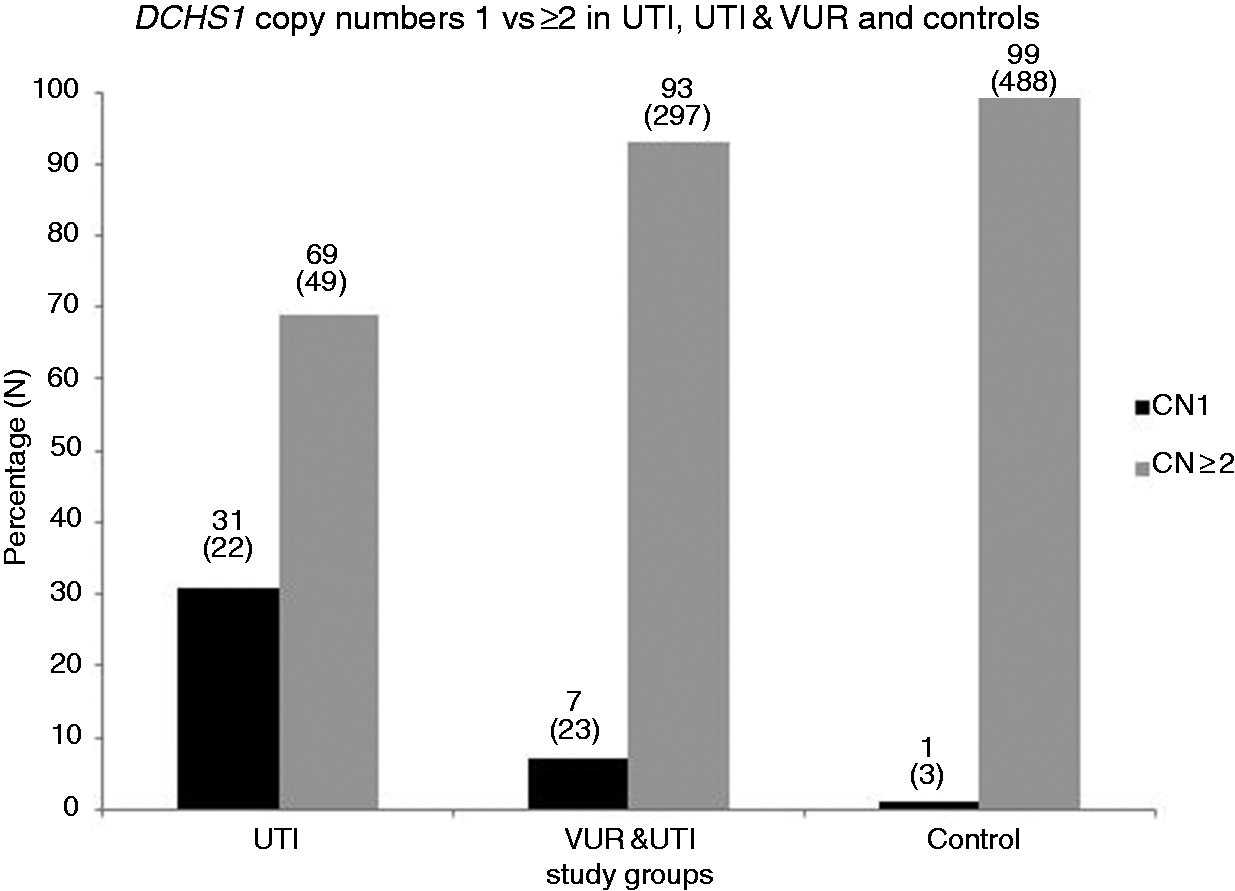
DCHS1 DNA CNs 1 vs. ≥ 2 in subjects with UTI without VUR, UTI with VUR, and controls. Percentage and distribution of 1 CN vs. ≥ 2 CN of DCHS1 in subjects with UTI without VUR (n = 71), UTI with VUR (n = 320), and controls (n = 491). CN = DNA CN of DCHS1.
Children with UTI and without VUR had a higher incidence of low DNA CN of DCHS1 compared to controls, as well as children with VUR
The DCHS1 CN in children with UTI and no VUR (CUTIE) varied from 1 to 2, with a mean CN of 1.69 ±0.47. A statistically significant number of children with UTI (n = 22; 31%) had one copy of DCHS1 compared to controls (n = 3; 0.6%). Individuals with one copy of DCHS1 compared to all other genotypes were 73 times more prevalent in the CUTIE cohort compared to controls (OR = 73.03; 95% CI 21.10–252.76; P < 0.0001). When comparing the two UTI cohorts, the CUTIE cohort (UTI and no VUR) had a statistically significant larger percentage of children with one copy of DCHS1 compared with RIVUR (UTI and VUR; 31% vs. 7%; OR = 5.8 95% CI 3.00–11.19; P < 0.0001).
DCHS1 protein is expressed in bladder epithelial and renal intercalated cells
DCHS1’s expression and location in postnatal kidney and bladder tissue has not been previously described. Because we discovered a significant genetic variation in DCHS1 in patients with UTIs, we aimed to localize DCHS1 protein expression using human tissue. In the normal human kidney and the kidney from a patient with pyelonephritis, we demonstrated DCHS1 was expressed by intercalated cells of renal-collecting duct epithelia. As many of the innate immune proteins are expressed in the kidney, DCHS1 localization in intercalated cells highlights its likely role in innate immunity. In the bladder, we demonstrated strong immunostaining in the bladder epithelia. All tissue immunostaining is demonstrated in Figure 3.
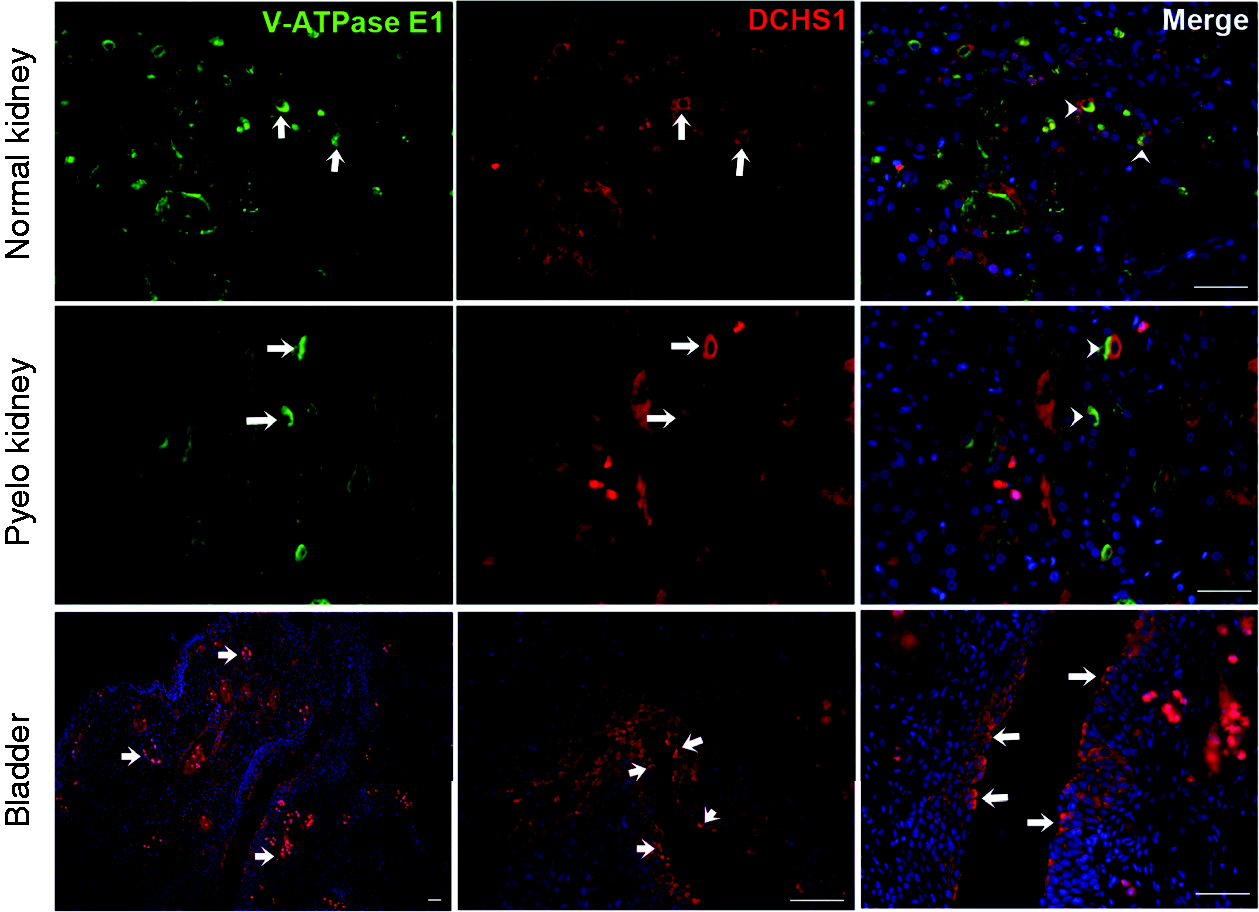
Immunolocalization of DCHS1 in human kidney (upper and middle panel) and bladder (lower panel). Upper-left panel: kidney section showing V-ATPase E1-positive intercalated cell in collecting duct (green, white arrow); middle panel: DCHS1-positive cells (red, white arrow); and right-panel: merged image (white arrowheads) showing DCHS1-positive intercalated cells. Lower panel: bladder staining for DCHS1 indicated by arrows; left panel: 10× magnification showing islands of DCHS1 positive infiltrating cells; middle and right panel: 40× magnification, showing DCHS1-positive bladder urothelium. Blue nucleus staining DAPI, scale bar 50 µm.
Dchs1 expression does not significantly increase in response to murine uropathogen challenge
Following a UPEC challenge in mice (n = 15 UTI89, n = 16 CFT073), we quantified the Dchs1 mRNA amount in both the bladder and kidney (Figure 4). The cystitis UPEC strain (UTI89) produced a larger increase in Dchs1 mRNA than the pyelonephritis UPEC strain did (CFT073). Compared to saline controls, neither kidney nor bladder Dchs1 mRNA transcripts were significantly increased. The bladder demonstrated the largest increase in UTI89-challenged mice but missed significance (P = 0.11).
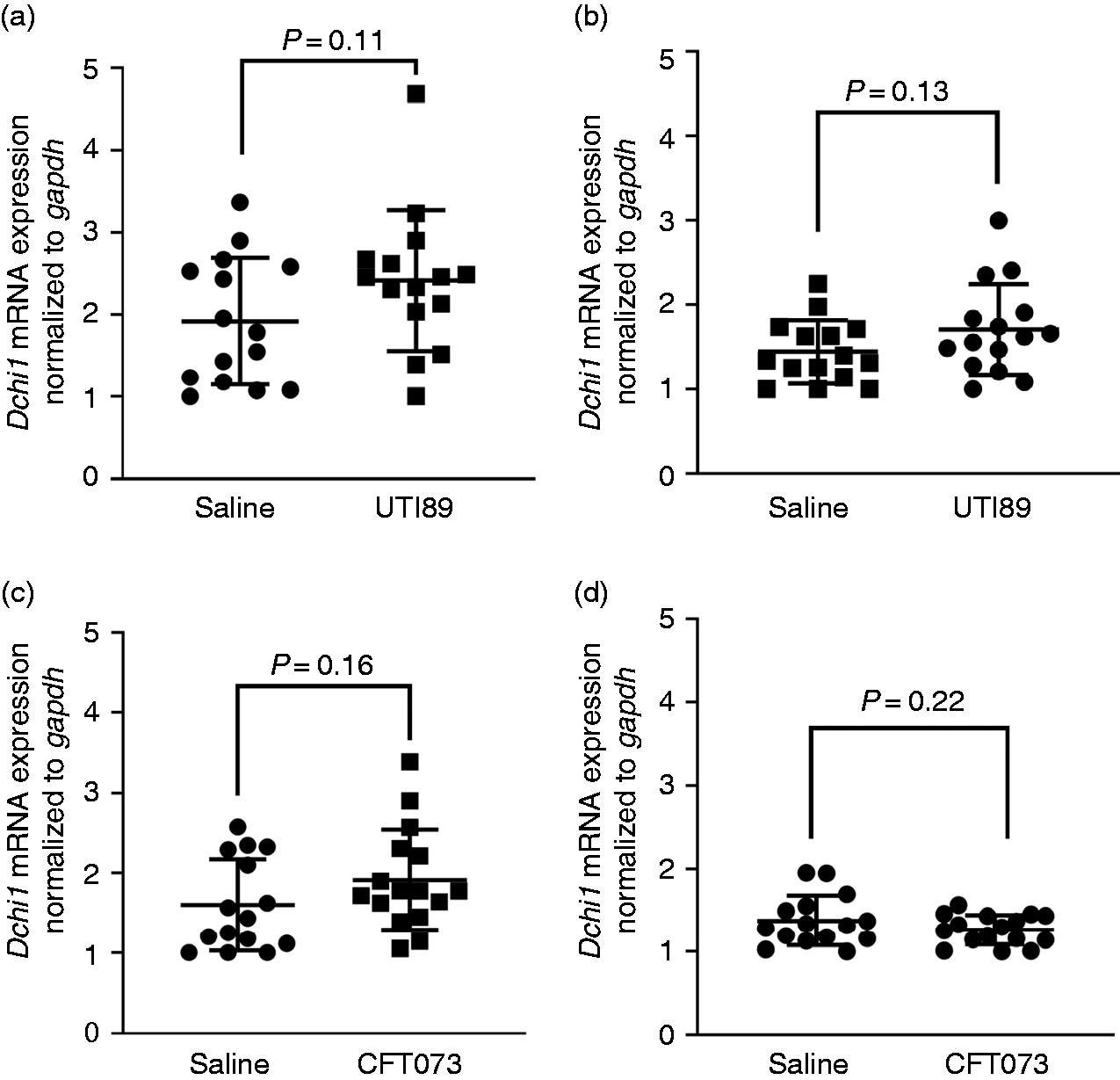
Dchs1 mRNA expression of murine bladder and kidneys in response to different uropathogenic E. coli strains. A and B graphs correspond to mRNA upregulation after 6 h on mouse bladder and kidneys infected with UTI89 compared to saline controls (n = 15 UTI89 and n = 15 saline). C and D graphs show mRNA transcripts of mouse bladder and kidneys upon inoculation with CFT073 compared to controls (n = 16 CFT073 and n = 15 saline), respectively. Long horizontal bars represent the mean, with upper and lower smaller horizontal bars representing ±1 standard deviation.
Discussion
This study used well-characterized pediatric cohorts from the previously published RIVUR and CUTIE studies.19,21 In a genome-wide study focused on a subset of UTI patients, our research group identified a novel association of DCHS1 relative gene CNV with UTI, which has not been reported previously. 16 In this report, we determined the absolute DNA CN of DCHS1 in two clinical cohorts of pediatric UTI to expand on our genome-wide screening study. We found that individuals with low DCHS1 CN (one copy) were 73 times more prevalent in the CUTIE (UTI without VUR) cohort compared to controls. In the RIVUR cohort (UTI with VUR), one copy individuals were 13 times more prevalent. Even in the absence of VUR, children with UTIs can develop renal scarring, thus showing that UTI-associated genes are important for children at risk. 21
DCHS1 encodes for atypical cadherin, which belongs to the cadherin superfamily and is a calcium-dependent cell–cell adhesion molecule. The encoded protein has 27 extracellular cadherin repeat domains, transmembrane domain, and a unique cytoplasmic region. 26 The Dchs1 gene was found to have role in kidney development in a murine transgenic model. 17 Dchs1 also has been shown to have a role in cell adhesion, growth, tissue patterning, and planar cell polarity. 18 To the best of our knowledge, the role of DCHS1 in the postnatal kidney has not been investigated.
This report serves as the first study to demonstrate the association of DCHS1 gene loss in patients with pediatric UTIs. Our immunostaining demonstrates a potential postnatal role of DCHS1 in the bladder and intercalated cells of the kidneys. Many key components of the innate immune system demonstrate a similar expression pattern. Thus, this finding supports the notion that it is critical in innate defenses.25,27 Because Dchs1 expression did not significantly increase following UPEC challenge, this gene is likely important in basal defenses and not inducible. Conversely, UPEC may subvert the epithelial response to invasion, although our experiments missed this possible role. Additional murine experiments are needed to dissect the pathways up- and downstream of DCHS1. Additionally, transgenic murine studies with bladder and renal collecting duct deletion will help to determine which tissue is key in innate defenses and if intercalated cell dysfunction exists with gene loss.
We can only hypothesize the precise mechanisms of UTI susceptibility with a single copy of DCHS1, and further studies are needed. Because DCHS1 is expressed on the apical surface of epithelia, DCHS1 may have a role in pattern recognition and/or inflammatory cascade signaling. Our research group and others have demonstrated the role of intercalated cells in the innate defense of the kidney and urinary tract. Because cell polarity is critical in epithelial cells such as intercalated cells of the collecting duct, we postulate that DCHS1 single-copy individuals may have subtle intercalated cell innate immune dysfunction leading to UTI susceptibility. 28
Because we observed the highest percentage of subjects in the UTI cohort without VUR (CUTIE Study cohort) with a single copy of DCHS1, we postulate that DCHS1 CNV is more relevant to infection and immunity than abnormal development of the kidney and urinary tract. Additionally, because genetic variations in DCHS1 were more prevalent in those with normal urinary tracts and an intact anti-reflux anatomy, the DCHS1 gene is likely more relevant to bladder as opposed to kidney innate immunity. The gene product may also prevent attachment in the bladder and/or urethra. Thus, in the context of VUR, bacteria may be able to ascend the urinary tract to the kidneys quickly, and the effect of DNA CNVs is blunted if the primary site of action postnatally is the bladder. If this speculation is true, the association of DNA CNVs in patients without VUR would be more consistent with the results of this study.
We do acknowledge some limitations in this study. First, our study was limited to Caucasian girls. So, results may not apply to other cohorts. Further investigations are needed on diverse cohorts to confirm these findings are consistent across study populations. Additionally, we utilized an adult cohort of women for controls. We utilized this cohort for two reasons. First, large control cohorts of children with no history of UTI are not publicly available. Second, an adult control cohort with no history of UTI should actually have a higher risk of UTI than a pediatric cohort, given the high incidence of UTI in adult women compared to children. The impact of this study may need to be tempered until we can perform additional studies to discover the role of this gene in the postnatal kidney and urinary tract.
Conclusion
This study represents the first report of DCHS1 in association with UTI. We speculate that the site of biological action may be in the lower urinary tract and “bypassed” to a certain extent in individuals with VUR. Thus, DCHS1 genetic variations associate with pediatric UTI risk, but the effect of these variations is masked by VUR. DCHS1 can be added to the growing body of disease-modifying genes such as DEFA1A3, in conjunction with patient characteristics such as VUR grade and bowel/bladder dysfunction to start creating UTI risk panels for patients at risk for UTI sequelae. 13 Further investigation is required to determine the role of DCHS1 and innate immune functions of the kidney and urinary tract.
Footnotes
Acknowledgements
We thank the Randomized Intervention for Children with Vesicoureteral Reflux Trial Investigators. We would like to thank the NINDS Human Genetics Resource Center and the Coriell Institute for providing healthy control DNA. D.S.H. initiated this project at Le Bonheur Children’s Hospital in Memphis, TN.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: A.L.S. and D.S.H. were funded by National Institutes of Health (NIH) Grants (1RC4DK090937-01, 1R01DK106286-01 and 1R01DK117934-01).
