Abstract
A greater demand for food animal production without antibiotics has created the common practice of feeding food animals dietary immunomodulatory feed additives (IFA) throughout their life cycle. However, little is known about the impact of IFA on cytokine and chemokine signaling in non-stressed, non-pathogen-challenged food animals during the early feeding period. We evaluated the expression of 82 genes related to cytokine and chemokine signaling in the whole blood of growing Angus heifers to determine the effect of IFA supplementation on cytokine and chemokine signaling during the first 28 d of feeding. One gene (CCL1) was significantly up-regulated and 14 genes (17%) were significantly down-regulated by IFA feeding during the entire early feeding period including 5 of 21 (24%) evaluated chemokine and IL receptors (CCR1, CCR2, IL1R1, IL10RA, IL10RB). These data when taken together suggest providing an IFA in the diet of growing beef cattle during the early feeding period may suppress the inflammatory response through cytokine–cytokine receptor signaling.
Introduction
Food animals can encounter expected and unexpected stressful events such as heat stress through multiple phases of their productive life cycles. Stress can lead to immunosuppression, which may correspond to greater incidences of disease during a pathogen challenge. 1 Cattle management can assist in limiting the effects of stressful events; feeding an immunomodulatory feed additive (IFA) designed to support animal health has become a practical, efficient means of improving the immune response during expected or unexpected stressful events. 2 One IFA product available for use in the livestock industry, OmniGen-AF® (OG; Phibro Animal Health Corporation, Teaneck, NJ, USA), provides evidence of a supportive effect on innate immunity activity in sheep, predominantly an increase in protein abundance of l-selectin (CD62L; cluster of differentiation 62 ligand) in neutrophils 3 and IL-8 receptor (IL8R) gene expression in whole blood 4 after 4 wk of feeding. Given that the induction of the genes coding for these two receptors represents a limited aspect of the innate immune response, it is important to note that these proteins are markers for neutrophil functions, and thus provide a physiological mechanism for the cell-mediated cascade leading to pathogen clearance also reported with OG supplementation.5–8 Specifically reported in dairy cattle, an increase in expression of CD62L and IL8R in leukocytes coincides with improved neutrophil-mediated killing capacity against pathogenic bacteria associated with mastitis.7,9,10
Another aspect of the process of neutrophils or other immune cells responding to a pathogen challenge is the production of cytokines and chemokines. IL-1β in neutrophils 3 and IL8R in whole blood 4 are known to be up-regulated by OG after 28 d of feeding (late immunomodulatory feeding period). This investigation will focus on the immunomodulatory impact of OG supplementation on the gene expression of 82 cytokines and chemokines and their receptors (see supplemental appendix for list of genes, their abbreviations, and their full names) in whole blood during the first 4 wk of feeding (early immunomodulatory feeding period). In comparison, our previous studies focused on the immunomodulatory OG effect after more than 28 d of feeding.4,11
Most studies with IFA utilize animals experiencing specific pathogen challenges under controlled stress conditions such as heat stress; these animal models are useful to determine immunomodulatory effects of specific pathogens but do not represent most beef cattle during their life cycle phases. Thus, the current study will focus on food animals without an induced stressor or specific pathogen challenge. The focus of this study is to (1) expand the scope of immune function gene markers in beef cattle, especially genes coding for cytokines/chemokines and their receptors that are regulated by feeding OG in a non-stressor condition; and (2) discover changes in immune function gene markers induced in the early immunomodulatory OG feeding period.
The central hypothesis of this project is that supplementation with OG will induce changes in gene expression of cytokines/chemokines or their receptors in whole blood before 28 d of supplementation. The objective of this study was to monitor changes in expression of genes coding for cytokines and cytokine receptors in circulating blood cells in growing, purebred Angus heifers fed OG sampled on d 3, 5, 10, 14, 21 and 28 of supplementation. A relatively large number of immune function gene markers was evaluated with the use of gene expression profiling array-based technology for greater insight into the immunomodulatory effect of OG.
Materials and methods
Animal care
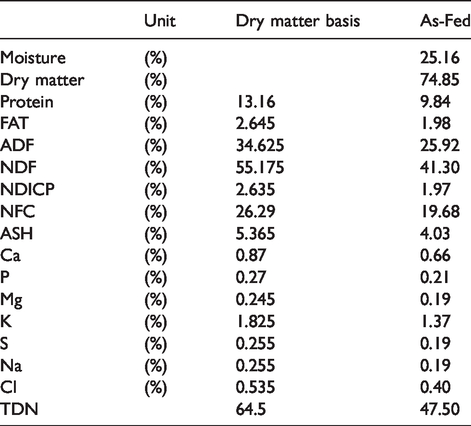
All animals were cared for in accordance with OmniGen Research Animal Handling Guidelines. These guidelines follow the Guide for the Care and Use of Agricultural Animals in Research and Teaching. 12 Eight purebred Angus heifers (mean: 326 kg; standard deviation: 4 kg; 10.5 mo old; range: 9–11 mo) were housed in a freestall barn (Corvallis, OR) with access to a Calan Broadbent Feeding System (American Calan, Northwood, NH) and fed a forage-based diet comprising grass hay and alfalfa hay. A nutrient composition table is shown in Table 1. Feed was mixed in a custom mix wagon and offered to cattle twice daily (9:00 and 15:00). Cattle were allowed a 7-d acclimatization period and then were randomly divided into two treatment groups: control animals (no additive) and the proprietary IFA OG (56 g/hd/d top-dressed; OmniGen-AF® Phibro Animal Health Corporation, Teaneck, NJ), which comprised a blend of ingredients that include silicon dioxide, aluminosilicate, dehydrated Saccharomyces cerevisiae, fermentation product of the fungus Trichoderma longibrachiatum, B complex vitamins, choline, and vitamin K precursors. 13
Nutrient composition of experimental diet.
Blood collection, RNA purification and reverse transcription
Prior to morning feeding, blood samples (n = 56) were collected on the first day of the trial (before supplementation began) and on d 3, 5, 10, 14, 21 and 28 of supplementation. Approximately 3 ml of whole blood was collected via jugular venipuncture into Tempus Blood RNA Tubes (Cat no 4342792, Life Technologies, Carlsbad, CA, USA). Immediately after collection, tubes were shaken vigorously for 15 s and stored at –20°C until RNA isolation (less than 2 mo). RNA was isolated using the Tempus Spin RNA Isolation kit (Cat no 4380204, Life Technologies); upon completion, RNA samples were stored at –80°C until reverse transcription.
RNA was evaluated for purity and concentration using a Thermo Scientific Multiskan Go microplate spectrophotometer (Cat no. 51119300; Thermo Fisher Scientific, Carlsbad, CA, USA) and a µDrop Plate (cat no N12391, Thermo Fisher Scientific). Only RNA with an absorbance ratio (260 nm: 280 nm) above 2.0 was used (n = 54). One μg total RNA was completely used as a template for cDNA synthesis in a RT2 First Strand Kit following the manufacturer’s protocol (cat no 330401; QIAGEN, Valencia, CA, USA). Upon completion, cDNA was stored at –20°C until use.
RTqPCR
We combined 102 µl cDNA with 1350 µl RT 2 SYBR Green qPCR Mastermix (cat no 330503; QIAGEN) and 1248 µl molecular-grade water (Cat no 338132; QIAGEN) and then thoroughly mixed the solution. Next, 25 µl of the mixed solution was pipetted into each well of a RT2 Profiler PCR array- Cow Inflammatory Cytokines and Receptors plate (PABT-011Z; QIAGEN; full gene listing found in supplemental appendix). Well plates (one plate/sample; 56 plates total) were placed in a Bio-Rad C1000 series thermocycler (cat no 184-1100, Bio-Rad, Richmond, CA, USA) and read using a CFX96 Real-Time PCR Detection System (cat no Bio-Rad 184-5097). Thermocycling protocols were as follows: 10 min at 95°C, 40 cycles with 15 s at 95°C and 1 min at 60°C. A melting curve was completed directly after thermocycling (95°C to 65°C to 95°C in 0.5°C increments).
qPCR data analysis
Data were analyzed using LinReg software 14 to account for efficiency of amplification and normalized by a normalization factor calculated by geometrical mean of three internal control genes (HPRT1, TBP and YWHAZ), which had M values of 1.2, 1.3 and 1.3, respectively. Internal control genes were selected by testing with geNorm. 15 Briefly, all internal reference genes available in the plate (ACTB, GAPDH, HPRT1, TBP, YWHAZ) were analyzed and the mean expression stability value (M value) was < 1.5. The determination of the optimal number of control genes for normalization indicated that the use of three internal control genes offered the largest stability among the possible combinations (i.e. V value = 0.326).
RTqPCR data were natural log-transformed prior statistical analysis. Presence of outliers was evaluated using the Studentized residuals using PROC REG of SAS (v9.3, SAS, Inc., Cary, NC, USA); when Studentized residuals t > 2 samples were removed from the final data set. The final data set (82 genes: 19 chemokines, 13 chemokine receptors, 16 IL, eight IL receptors, one IL receptor agonist, 24 other cytokines, one cytokine decoy receptor; two genes IL-17 F (IL17F) and chemokine ligand 11 (CCL11) were excluded from the analysis because the majority of samples were below the detection limit) was subjected to ANOVA analysis with treatment, time, and treatment × time as main effect and animal as random using JMP Genomics (SAS institute, NC, USA). Significance was deemed with a false discovery rate (FDR)-adjusted P-value < 0.10. All statistical tests were two-sided.
To identify specific inflammation-associated pathways altered by OG supplementation, we performed pathway analysis of our microarray using Dynamic Impact Approach (DIA) 16 and Database for Annotation, Visualization and Integrated Discovery (DAVID). 17 For the pathway analysis, we only considered inflammation-associated pathways with at the least two differentially expressed genes (FDR-adjusted P-value < 0.10) and two genes measured by the microarray. To identify significantly enriched inflammation-associated pathways, we used Expression Analysis Systemic Explorer (EASE) score of < 0.10 as cut-off for DAVID and a FDR-adjusted P-value < 0.10 as cut-off for DIA.
Results
Feeding OG altered expression of 15 out of 82 measured genes (18%) during the entire early feeding phase; of those, one (CCL1) was up-regulated (FDR-adjusted P = 0.03; Figure 1) and 14 were down-regulated (FDR-adjusted P < 0.10; Figure 2). In addition, three genes (VEGFA, NAMPT and CXCR1) were decreased by OG supplementation starting d 14 (Figure 3). Transcripts repressed by OG supplementation included three of 19 chemokines (C5, CCL26, CXCL2), two of 13 chemokine receptors (CCR1, CCR2), three of 16 IL (IL1B, IL3, IL9), three of eight IL receptors (IL1R1, IL10RA, IL10RB), two of 24 other cytokines (CSF1, BMP2), and the only receptor antagonist (IL1RN) (Figure 2). Of the 21 receptors evaluated, six (29%) were influenced by OG feeding. Significant time × treatment interactions (FDR-adjusted P < 0.10) were detected for five of 82 genes: TNFRSF11B, VEGFA, CX3CR1, NAMPT, and CXCR1 (Figure 3).
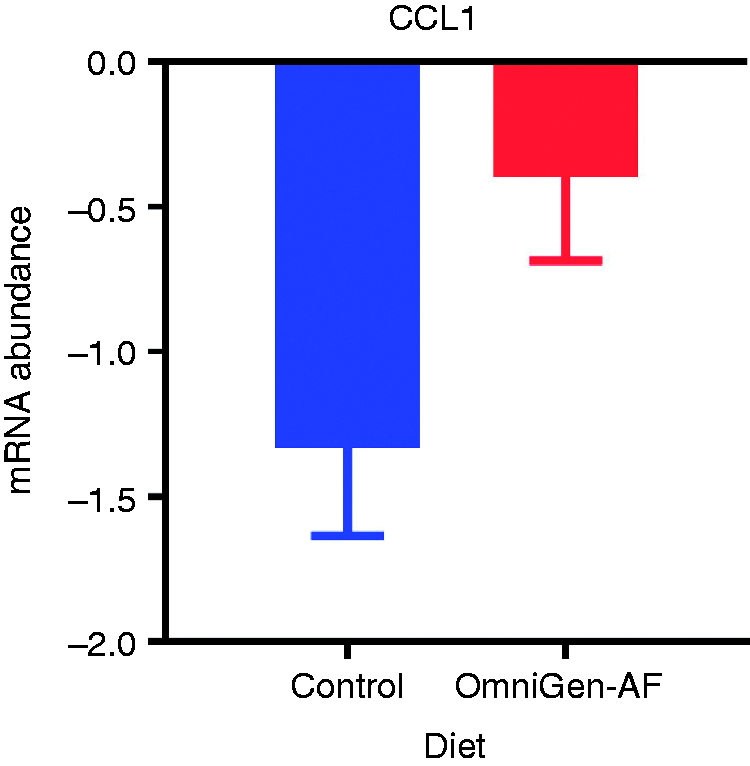
Down-regulation of CCL1 expression is attenuated in growing beef cattle supplemented with the immunomodulatory feed additive OmniGen-AF® during the first 28 d of supplementation.
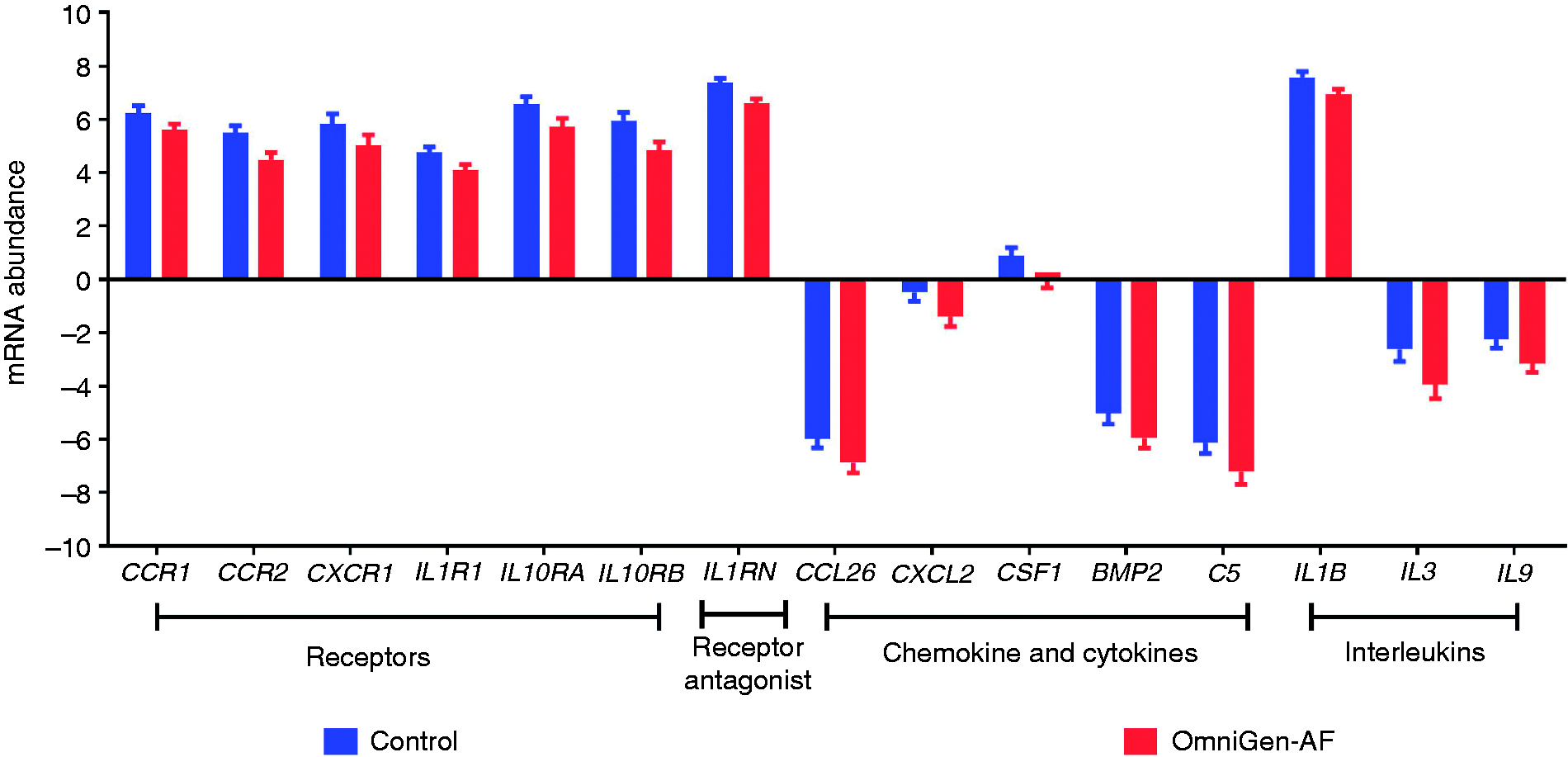
Genes suppressed by feeding the immunomodulatory feed additive OmniGen-AF® during first 28 d of supplementation.
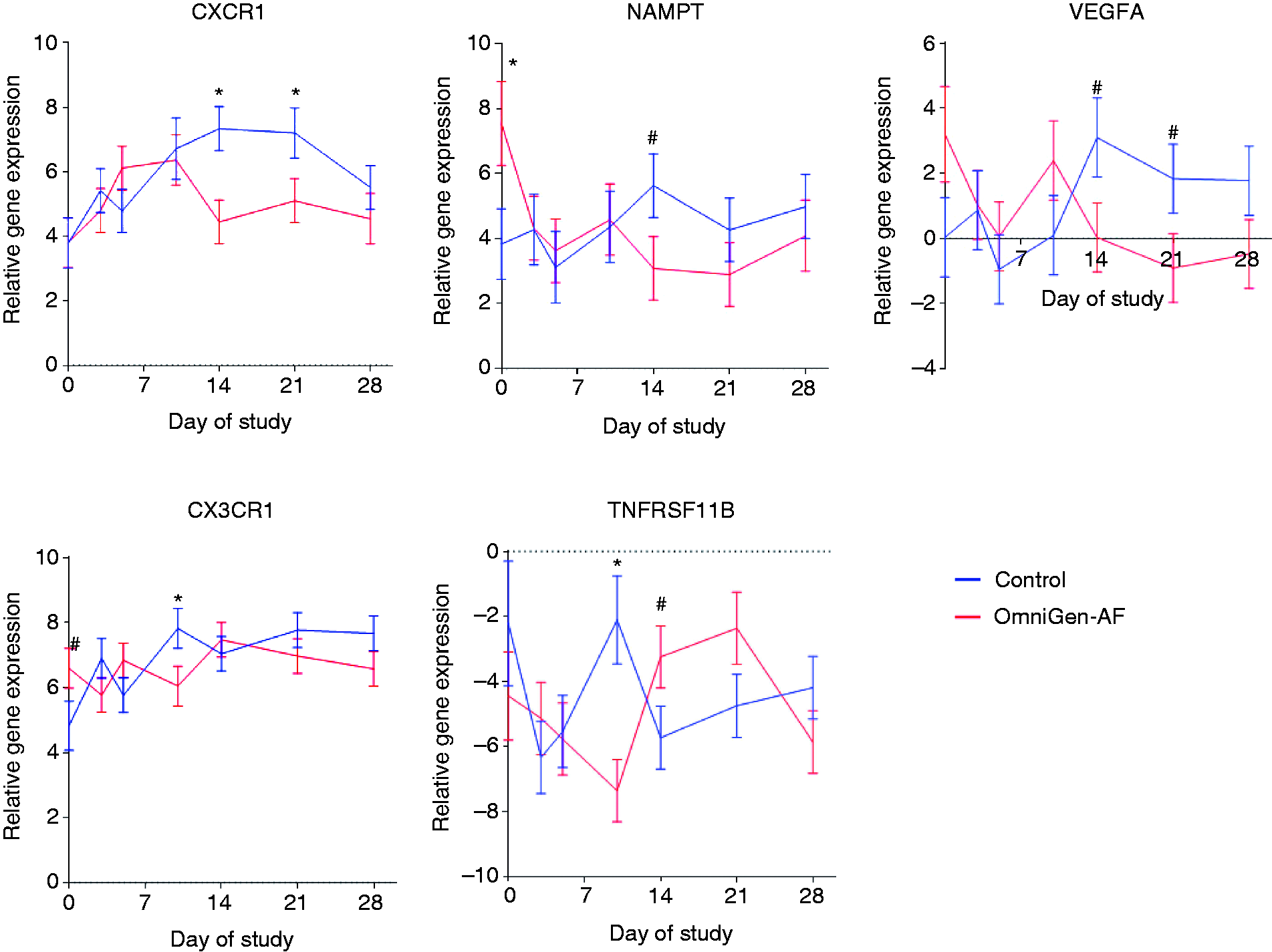
Effect of feeding the immunomodulatory feed additive OmniGen-AF® and the interaction of time on expression of immune function-related genes in whole blood during the first 28 d of supplementation. *Indicates comparison between Control and OmniGen-AF® is different (P < 0.05) while # indicates comparison has tendency to be different (0.05 < P < 0.10).
Using DIA, we detected an overall inhibition by OG of several KEGG pathways during the early immunomodulatory feeding period encompassing cell signaling and immune system development/response. Specific inflammation-associated pathways suppressed by OG supplementation (over 14% of genes within pathway significantly decreased) were: cytokine–cytokine receptor interaction (nine of 64 evaluated genes), chemokine signaling pathway (five of 34 evaluated genes), JAK–STAT signaling pathways (four of 19 evaluated genes), TNF-α signaling (three of 11 evaluated genes), hematopoietic cell lineage (four of 13 evaluated genes), osteoclast differentiation (three of eight evaluated genes), and MAPK signaling (two of five evaluated genes). DAVID identified the significantly suppressed inflammation-associated KEGG pathways: “chemokine signaling” and “hematopoietic cell lineage” (results not shown).
Discussion
Feeding livestock IFA has increased as a management tool for prevention of diseases and subsequent production losses. 18 Despite increased use of IFA throughout the lifecycle of food animals, to our knowledge, little is known about the immunomodulatory effects of these additives on non-stress, non-pathogen-challenged animals. This study focuses on the effect of OG supplementation on gene expression of cytokines/chemokines and their receptors during the feeding period under “normal” management conditions.
Effect of feeding OG on cytokine/chemokine receptor gene expression
The effect of OG on the expression of genes coding for cytokine/chemokine receptors is prominent. A total of 24% of all gene receptors (five out of 21 analyzed receptor genes) evaluated on the array were down-regulated by OG. A closer look at the function of these receptors indicates that many have promiscuous binding patterns (Table 2). For example, ILR1, CCR2, IL10RB and CCR1 can bind three, seven, eight and 12 unique cytokines as a part of a heterodimer receptor, respectively.19–22 The ligand receptor interaction of most cytokines results in an irreversible binding and leads to intracellular changes resulting in a biologic response, which varies by cytokines and includes gene expression, changes in cell cycles, and release of mediators. 23 The down-regulation of numerous cytokine receptors in this study, which have the capability of initiating multiple, differentiated effects on the cellular signaling of inflammation could be crucial to regulate the immune system and prevent widespread, uncontrolled inflammation. A pro-inflammatory metabotype has been identified as a contributing factor to several infectious and metabolic diseases including mastitis, retained placenta, metritis, displaced abomasum and ketosis. 24 It is possible that dietary OG may decrease disease incidence by limiting uncontrolled inflammation.
Cytokine receptors regulated by feeding the immunomodulatory feed additive OmniGen-AF® (or the interaction of OmniGen-AF® and time) can be promiscuous and bind to multiple cytokines.
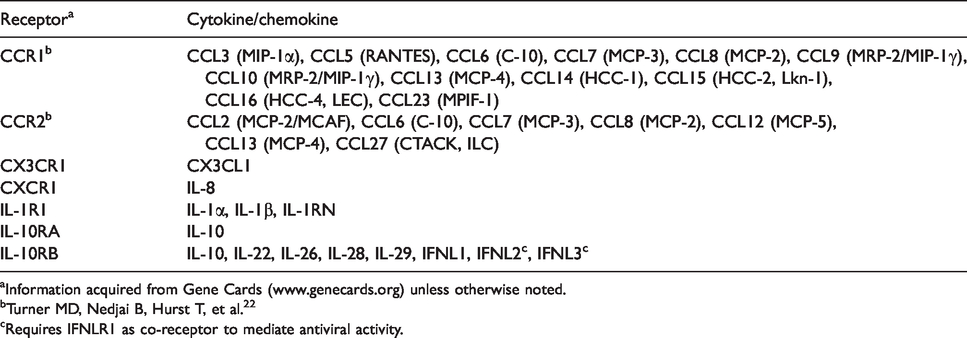

aInformation acquired from Gene Cards (www.genecards.org) unless otherwise noted.
bTurner MD, Nedjai B, Hurst T, et al. 22
cRequires IFNLR1 as co-receptor to mediate antiviral activity.
Previous studies focused on the effect of OG supplementation after the first 28 d of OG supplementation in dairy cattle and reported that CXCR1, a gene coding for IL-8 receptor type 1 (IL8R), is up-regulated in leukocytes or whole blood in dairy cattle fed OG after 2–4 mo of supplementation.4,7 In comparison, this study focused on the effect of OG in growing beef heifers during the first 28 d of supplementation. Thus, the down-regulation of CXCR1 reported in this study might be due to a difference in cattle type (dairy vs. beef) or length of OG supplementation.
Effect of OG supplementation on gene expression of cytokines/chemokines and their ligands
Less pronounced was the effect of OG supplementation on cytokine/chemokine vs. cytokine/chemokine receptor expression, as eight out of 58 (14%) genes were altered by OG supplementation. Six genes (BMP2, C5, CSF1, IL1B, IL3 and IL9) were repressed throughout OG supplementation. In addition, NAMPT and VEGFA were repressed by OG supplementation starting d 14. BMP2, 25 C5, 26 CSF1, 27 IL1B, 28 IL3, 29 NAMPT 30 and VEGFA 31 are considered pro-inflammatory cytokines/chemokines, whereas IL-9 has been shown to resolve chronic inflammation. 32 The results could indicate an overall down-regulation of inflammation through OG supplementation. We previously showed that OG-supplemented ewes had increased IL-1β protein expression in circulating immune cells after a dexamethasone and moldy feed challenge and the feeding period. 3 Possible explanations for the differences in results could include species differences, protein vs. mRNA expression, presence vs. absence of immune challenge, and sampling time (length of the feeding period).
CCL1 was the only gene up-regulated by OG supplementation during the 28 d experimental period (Figure 1). Secreted by activated T cells, CCL1 binds to CCR8 33 and acts as a chemokine for monocytes, NK cells and dendritic cells. 34 The up-regulation of CCL1 could suggest an increased ability to attract monocytes into tissue for macrophage differentiation and ultimately improved clearing of cellular debris and wounds. However, it is important to note CCL1 is not the only cytokine with monocyte chemotactic properties and that it is the only monocyte chemotactic agent up-regulated in this study. To determine if OG supplementation increases monocyte migration, a functional cell assay needs to be conducted to assess diapedesis rates of OG-supplemented cattle compared with their control counterparts.
Two genes, CCL26 and CXCL2, suppressed by OG supplementation are neither functionally connected nor their respective chemokine receptor altered by feeding OG. CCL26 binds to CCR3 and has been proposed as indicator of the end of a local inflammatory response in beef cattle. 35 CXCL2 binds to CXCR2 36 and has been shown to be up-regulated in bovine immune cells in response to bacterial infections.37,38 Thus, the suppression of these genes supports the concept of a general down-regulation of the chemokine signaling pathway through OG supplementation; indicating a general regulation of the innate immune system.
Pathway analysis
OG is a proprietary, natural bioactive feed additive composed of multiple ingredients with potential immunomodulatory effect; thus, no pathway-specific response can be expected. Pathway analysis showed that feeding cattle OG repressed the gene expression of multiple and diverse pathways such as cytokine–cytokine receptor interaction, chemokine signaling pathway, JAK–STAT signaling pathways, TNF-α signaling, hematopoietic cell lineage, osteoclast differentiation and MAPK signaling.
Feeding cattle OG repressed the expression of multiple genes associated with the inflammatory response. The detection of changes in cytokine–cytokine receptor interaction, chemokine signaling pathway and TNF-α signaling may be expected due to the nature of the targeted array used in this study; an array designed to evaluate differences in cytokines and cytokine receptors associated with the inflammatory response. However, the significant enrichment of these pathways in DAVID is indicative of these pathways being biologically significant, because the overrepresented approach used by DAVID would be indifferent to the type of array used if the proper background is used. 17 The interaction of cytokines and chemokines is biologically relevant for studies of the immune system. The broad scope of the cell pathways identified here requires future research regarding the nature of cytokine–cytokine receptors on specific immune cells.
The OG altered the highly conserved MAPK and JAK–STAT signaling pathways. These signaling pathways are highly conserved across species and integral in numerous biological functions including playing a broad role in the innate immune response.33,39 This was surprising given the low amount of the dietary feed additive (i.e. 0.56% of diet) and part of the supplement likely metabolized by ruminal microbial fermentation. Along with the general, non-specific responses in signaling, we also saw a regulation in the hematopoietic cell lineage and osteoclast differentiation pathways. Both of these pathways are utilized in a holistic system to respond to general biological insults, predominantly through the innate immune system. Previous work conducted with OG showed a diverse in vivo response, including a decrease of a variety of infectious and metabolic adverse health conditions. 40 The role of OG supporting innate immune pathways has also been identified in model animals; 13 however, the current study is our first to measure the effect of OG supplementation of signaling pathways involved in the innate immune response. It is possible that the regulation of general, highly conserved pathways regulated in this study during the feeding period could explain the general improvements in animal health of OG-supplemented cattle (i.e. fewer disease events including fewer cases of mastitis, metritis, ketosis, late-term abortion, and cow deaths, lower milk somatic cell count). 40
Other considerations
Natural bioactive feed additives can be subdivided based on the chemical structure of their components (e.g. bioactive lipids, glycoproteins, oligosaccharides, minerals, vitamins, etc.) 41 or their potential functional properties to decrease adverse health events (e.g. induce innate and/or adaptive immune response, attenuate uncontrolled inflammation or oxidative stress, microbiome modifiers, immunometabolism modifiers). 42 These functional response markers, alone or in combination, could be used to discover molecular targets and validate the efficacy of natural bioactive feed additives.
The array utilized here for gene expression profiling limits the data generated to aspects of immune cellular communications via cytokine and cytokine receptors, an area we wanted to focus on to detect novel molecular response markers of feeding OG to growing beef cattle. This targeted approach allows for a close investigation of cellular communication in circulating immune cells; however, the absence of transcription factors or second messenger expression limits the clarity of data interpretation and usefulness of biological pathways involved in the immune cell response regulated by OG supplementation.
Whole blood was the tissue type used for this experiment. Although whole blood provides a useful foundation for identifying novel response markers and examining immune cell communication, the lack of hematological data and differential cell counts limits the biological interpretation. Biological implications of OG supplementation on cellular communication involved in the inflammatory response remain speculative, because differences seen in receptors or cytokines may be due to underlying differences in cellular populations and may not translate into attenuated uncontrolled inflammation as functional endpoint. Nevertheless, the results of this study help in understanding how natural bioactive feed additives may support animal productivity and health, and suggest OG as a natural bioactive with potential anti-inflammatory properties in beef cattle via altering cellular communication in circulating immune cells.
Conclusion
Our results indicate a non-specific repression of genes involved in multiple pathways associated with inflammation (cytokine–cytokine receptor interaction, chemokine signaling pathway, JAK–STAT signaling pathways, TNF-α signaling, hematopoietic cell lineage, osteoclast differentiation and MAPK signaling) by feeding OG during the growth phase in replacement beef heifers. This is novel, as, with the exception of CXCR1 and IL1B, the majority of genes discussed have not been previously associated with feeding OG. Specifically, the high prevalence of repression of decoy receptors and cytokine receptors with promiscuous binding patterns suggests providing OG in the diet of growing beef cattle may repress the inflammatory response via cytokine receptor signaling during the feeding period. If translated to protein function, those changes could have a diverse impact on the immune system and may explain the previously observed protective effect of dietary OG to a variety of infectious adverse health events.
Supplemental Material
INI887232 Supplemetal Material - Supplemental material for A natural bioactive feed additive alters expression of genes involved in inflammation in whole blood of healthy Angus heifers
Supplemental material, INI887232 Supplemetal Material for A natural bioactive feed additive alters expression of genes involved in inflammation in whole blood of healthy Angus heifers by Shelby A Armstrong, Derek J McLean, Massimo Bionaz and Gerd Bobe in Innate Immunity
Footnotes
Declaration of conflicting interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship and/or publication of this article: This study involved a student researcher who is an employee of Phibro Animal Health Corporation and includes data collected by Phibro Animal Health Corporation. Phibro Animal Health Corporation develops feed additives, supplements and diagnostics to support animal health.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by Phibro Animal Health Corporation.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
