Abstract
Innate recognition of Brucella spp. is a key step in the activation of inflammation. CD14 binds PAMPs and is involved in LPS-induced pro-inflammatory cytokine release. Previously we showed that knock down of CD14 in RAW264.7 macrophages disrupted Brucella–host interactions. However, its effect on the macrophage microRNA (miRNA) expression profile, especially after stimulation by Brucella infection, is still unclear. To identify miRNAs involved in the macrophage response to Brucella infection, we performed miRNA expression profiling of CD14 knock-down RAW264.7 (224.3) macrophages infected with Brucella melitensis, and demonstrated, for the first time, that CD14 knock down significantly up-regulated the expression of mmu-miR-199a-3p and mmu-miR-183-5p in these conditions. These miRNAs have a well-characterized association with the target genes involved in immune response, inflammatory response, innate immune response, apoptosis processes, anti-apoptosis, cytokine production and cytokine-mediated signaling pathways. Among the 104 inflammation-related candidate target genes of mmu-miR-199a-3p and mmu-miR-183-5p in the 224.3+ B. melitensis group cells, the expression of the Cbl-b, a potential target of mmu-miR-199a-3p, was confirmed to be down-regulated using qRT-PCR and Western blot analysis. Our findings suggest that CD14 functions in the Brucella–host interaction may be through altered miRNA expression, and regulation of Cbl-b proteins.
Keywords
Introduction
Brucellosis is an important zoonosis throughout the world. Brucella spp., the pathogen responsible for brucellosis, survive and replicate predominantly in macrophages. LPS is one of the PAMPs and the key virulence determinant of Brucella spp. 1 CD14 is present as a membrane-bound receptor in myeloid cell lines or in a soluble form in serum. When CD14 binds LPS, CD14–LPS complex binds to TLR4 and triggers the release of pro-inflammatory cytokines.2–5 Our previous study suggested that lentivirus-mediated RNAi targeting mCD14 inhibits TNF-α secretion, iNOS expression, and NO production in RAW264.7 cells stimulated by B. melitensis in vitro. 6
MicroRNAs (miRNAs), 18–25 nucleotide non-coding RNAs, post-transcriptionally regulate gene expression. miRNAs play important roles in Brucella infections.7–9 It has been reported that a high-throughput sequencing approach resulted in identification of 57 miRNAs that were differentially expressed between mock- and Brucella-infected RAW264.7 cells. 10 In macrophages, Brucella abortus infection leads to down-regulation of miR-125b-5p. miR-125b-5p targets A20, an inhibitor of NF-kB activation. In the presence of miR-125b-5p, B. abortus survival is attenuated. 11 A miRNA array-based initial screen followed by quantitative reverse transcription PCR (qRT-PCR) validation was used to identify the differentially expressed miRNAs in PBMCs of patients with acute or chronic brucellosis and healthy controls. The results indicated that the expression of miR-1238-3p was increased while expression of miR-494, miR-6069 and miR-139-3p were decreased. Thus, these miRNAs have the potential to be markers for chronic cases. 12 Using the same strategy, the differentially expressed miRNAs were identified in CD8+ T cells of patients with acute or chronic brucellosis and healthy controls. miR-126-5p and miR-4753-3p were decreased, and are therefore thought to have the potential to be regulators of CD8+ T-cell-related marker genes for chronic brucellosis infections. 13 However, in the presence of Brucella melitensis infection, the effects of mCD14 on the miRNA expression profile is unclear.
In this study, to identify miRNAs affected by Brucella spp. infection in the absence CD14, we analyzed the miRNA expression profiles of mCD14 knockdown RAW264.7 cells challenged with B. melitensis M5-90 using a miRNA array. We observed up-regulation of mmu-miR-199a-3p and mmu-miR-183-5p, and confirmed this by qRT-PCR. Interestingly, among the predicted target genes of mmu-miR-199a-3p and mmu-miR-183-5p, the significantly enriched gene ontology (GO) terms were apoptosis process, immune response, inflammatory response, innate immune response, anti-apoptosis, cytokine production and cytokine-mediated signaling pathway. Among the 104 candidate target genes, the expression of the Cbl-b gene in mCD14 knock-down RAW264.7 cells challenged with B. melitensis M5-90 was validated to be down-regulated.
Materials and methods
Cell culture and Brucella melitensis infection
The RAW264.7 cell line, used as a control, was obtained from the Cell Bank of the Chinese Academy of Science (Shanghai, China), and routinely grown, as previously described. 6 The 224.3 cell line, which is a mCD14 knock-down RAW264.7 cells line, and NC, which is a RAW264.7 stable cell line carrying shRNA-NC, were established and identified in our previous study. 6 The same number of 224.3, RAW264.7 and NC cells (5 × 105 cells per well) were infected with B. melitensis M5-90 for 6 h, as previously described. 6
miRNA array profiling and analysis
RNA preparation and quantitation were performed as previously described. 14 The small RNAs were extracted from the 224.3, RAW264.7 and NC cells, after infection with B. melitensis M5-90, and miRNA microarray analyses were undertaken by LC Sciences as previously described. 14
qRT –PCR for miRNAs
To validate the differential expression, as previously described, 14 small RNAs were extracted from the cells and qRT-PCR was performed to determine miRNA expression levels.
Target gene prediction and GO
To predict targets of significantly differentially expressed miRNAs, TargetScan, PicTar, miRDB and microRNA.org were used. GO (http://www.geneontology.org/) assignments were conducted as previously described. 14
qRT-PCR for mRNAs
To measure mRNA levels, high-quality total RNA was isolated from the 224.3, RAW264.7 and NC cells, after infection with B. melitensis M5-90, and subjected to qRT-PCR as previously described. 6
Western blot analysis
The total proteins were harvested from the 224.3, RAW264.7, and NC cells, after infection with B. melitensis M5-90, and Western blot analysis was performed as described in a previous study. 6 Primary Abs included Cbl-b (G-1) mouse monoclonal IgG1 (1:500 dilution; Santa Cruz Biotechnology, Santa Cruz, CA, USA) and GAPDH rabbit polyclonal IgG (1:1000 dilution; Cell Signaling Technology, Danvers, MA, USA). Secondary Abs used were HRP-labeled goat anti-mouse IgG (1:4000 dilution; Zymed Laboratories, San Francisco, CA, USA) and HRP-labeled goat anti-rabbit IgG (1:3000 dilution; Santa Cruz Biotechnology).
Data analysis
The Student’s t-test and one-way ANOVA were performed for data analysis as previously described. 6 A P-value of < 0.05 was considered significantly different and a P-value of < 0.01 was considered very significant.
Results
Brucella melitensis-induced differential miRNA levels in mCD14 knockdown RAW264.7 cells
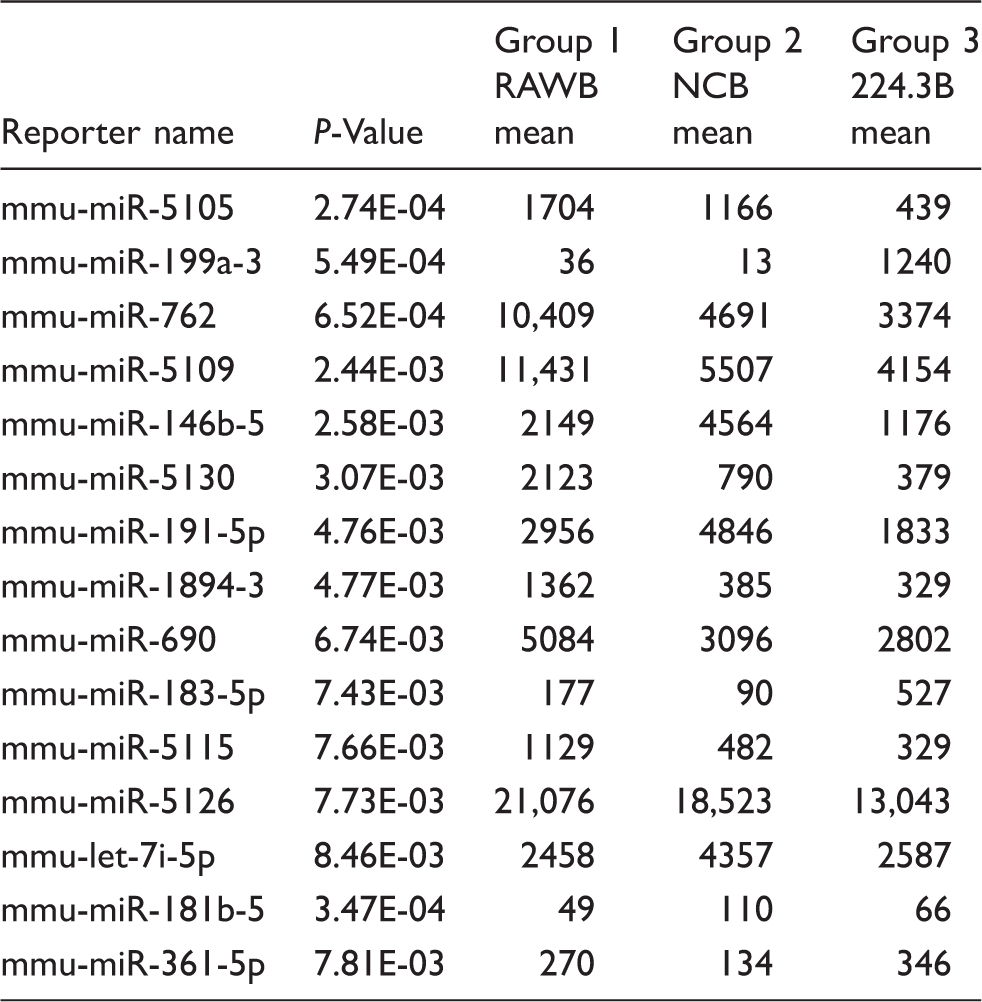
RNA samples obtained from 224.3, RAW264.7 and NC stimulated by B. melitensis infection were compared using microarray analysis. Figure 1(A) shows the relative expression of the 15 differentially expressed genes, namely mmu-miR-5105, mmu-miR-199a-3p, mmu-miR-762, mmu-miR-5109, mmu-miR-146b-5p, mmu-miR-5130, mmu-miR-191-5p, mmu-miR-1894-3p, mmu-miR-690, mmu-miR-183-5p, mmu-miR-5115, mmu-miR-5126, mmu-let-7i-5p, mmu-miR-181b-5p and mmu-miR-361-5p. Among them, three miRNAs (mmu-miR-199a-3p, mmu-miR-183-5p and mmu-miR-361-5p) were up-regulated and 10 miRNAs (mmu-miR-5105, mmu-miR-762, mmu-miR-5109, mmu-miR-146b-5p, mmu-miR-5130, mmu-miR-191-5p, mmu-miR-1894-3p, mmu-miR-690, mmu-miR-5115 and mmu-miR-5126) were down-regulated in the 224.3+ B. melitensis group compared with the control RAW264.7+ B. melitensis group and NC + B. melitensis group. Because of the unreasonable data obtained from the array, two miRNAs (mmu-let-7i-5p, mmu-miR-181b-5p) were considered noise signals (Table 1).
Up-regulation of mmu-miR-199a-3p and mmu-miR-183-5p expression. (a) Heat-map representation of data from two-way cluster analysis of differentially expressed miRNAs (P < 0.05) measured by an array-based screen of 224.3, RAW264.7 and NC cells, which are stimulated by B. melitensis infection. Green indicates suppression and red indicates induction compared with in control cells. (b) Validation of mmu-miR-199a-3p and mmu-miR-183-5p up-regulation by qRT-PCR. The significantly differentially expressed miRNAs by microarray analysis.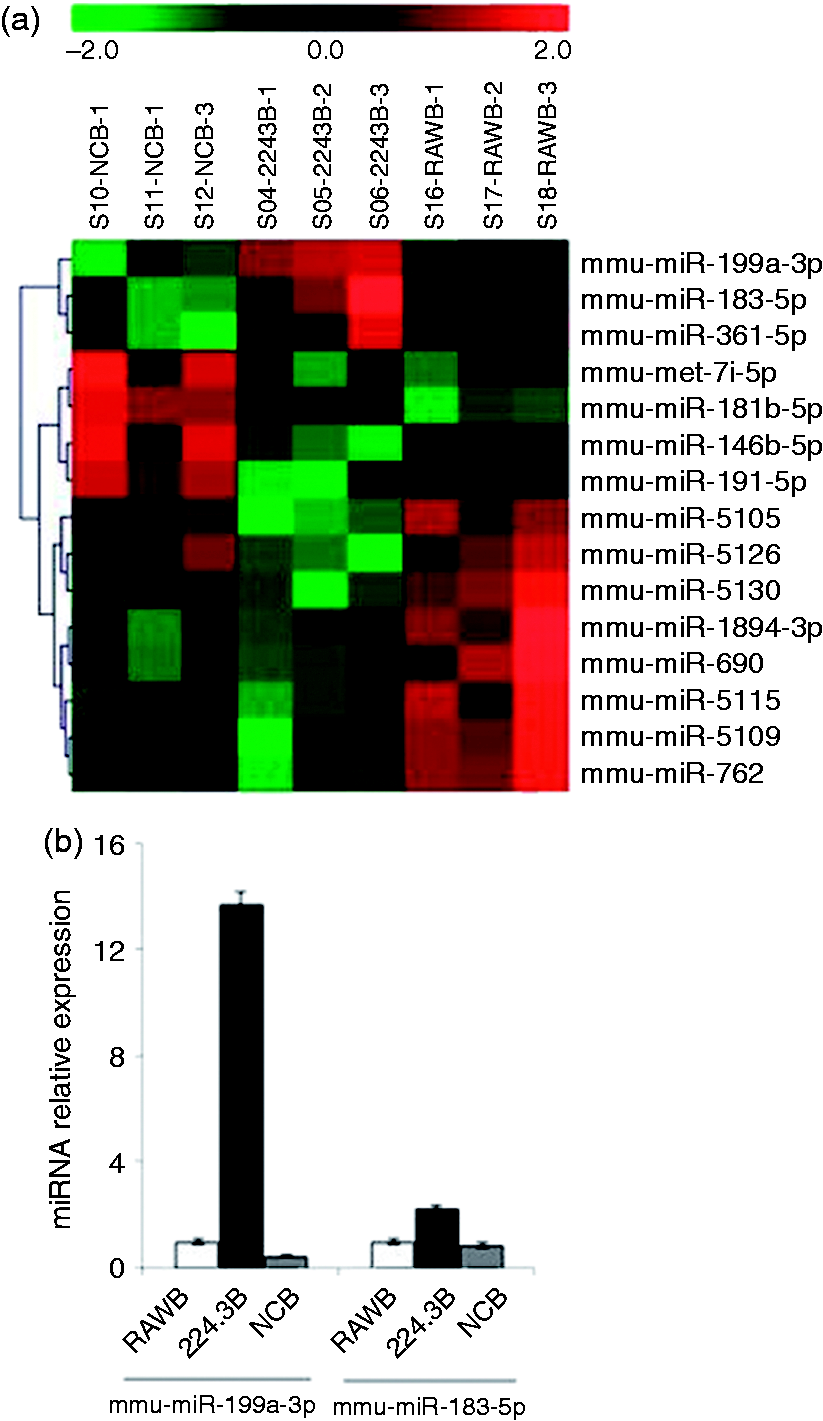
To verify the data from the initial array-based screen, qRT-PCR experiments were performed for mmu-miR-199a-3p and mmu-miR-183-5p, as these had been reported previously. 6 U6 snRNA abundance was used as an internal control. As shown in Figure 1(B), the up-regulation of both mmu-miR-199a-3p and mmu-miR-183-5p was confirmed in the 224.3+ B. melitensis group.
Target gene prediction and GO
Inflammation-related candidate target genes of mmu-miR-199a-3p and mmu-miR-183-5p.

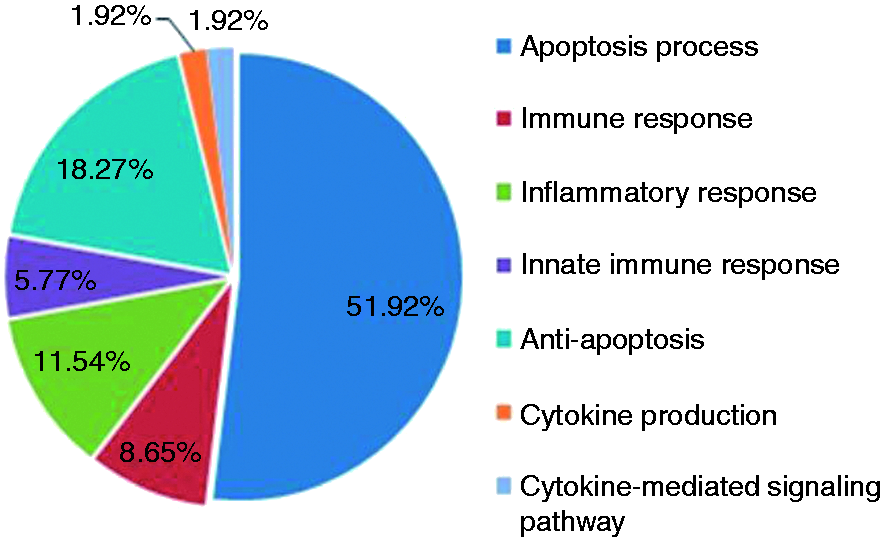
Significantly enriched GO terms among the predicted target genes of mmu-miR-199a3p and mmu-miR-183-5p.
Down-regulation of Cbl-b in 224.3+ B. melitensis group cells
Primers for candidate target genes of miR-199a-3p and miR-183-5p.

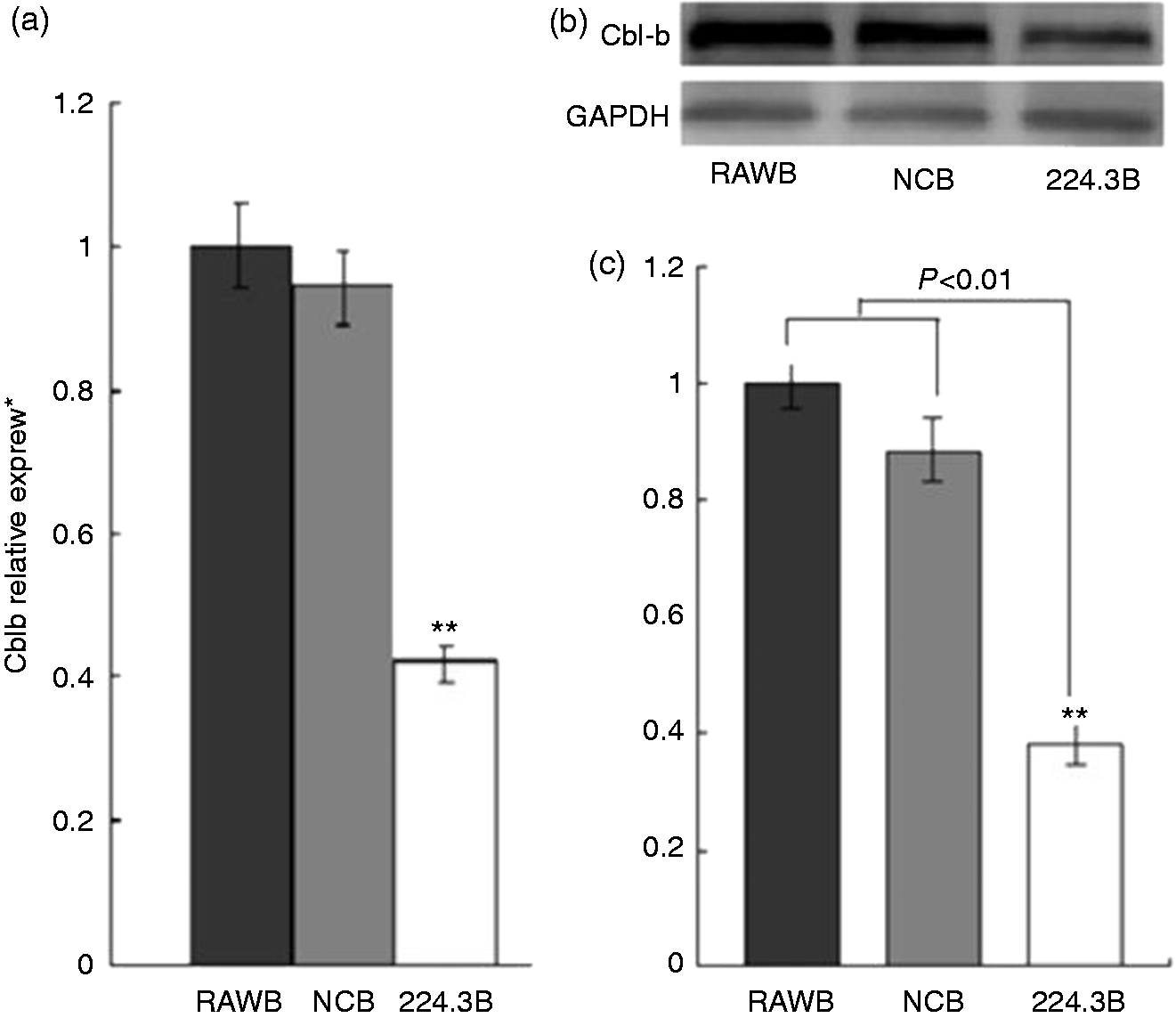
Expression of Cbl-b was down-regulated in 224.3 cells that were stimulated by B. melitensis infection. (a) qRT-PCR analysis. Lane 1, RAW264.7+B. melitensis; lane 2, NC+B. melitensis; lane 3, 224.3+B. melitensis. (b) Western blot analysis, GAPDH was used as loading control. (c) The quantification of the Western blots for protein expression with Bandscan 5.0 (n = 3).
Discussion
Recent reports have provided evidence that miRNA plays an important role in bacterial infections, including Brucella infection.8–13 As an important innate immune molecule, CD14 is involved in the initiation of the inflammatory response to sepsis. However, there are few reports about the relationship between CD14 and the miRNA expression profile of host cells. Here, we are the first to elucidate the role of CD14 in affecting the miRNA expression profile of macrophages after stimulation by Brucella infection. To dissect the mechanisms through which CD14 affects the miRNA expression profile of macrophages after Brucella infection, an initial throughput, array-based screen was performed, followed by qRT-PCR validation. Consistent with the values obtained from the array, up-regulation of mmu-miR-199a-3p and mmu-miR-183-5 was further confirmed with qRT-PCR.
In our study, the targets of mmu-miR-199a-3p and mmu-miR-183-5p were predicted by at least one of four different algorithms (TargetScan, PicTar, miRDB and microRNA.org). More than 1000 different candidate genes were identified (data not shown). Because Brucella stimulates a robust inflammatory response, 15 the significantly enriched GO terms in the predicted target genes of mmu-miR-199a-3p and mmu-miR-183-5p were focused on 104 inflammation-associated candidate genes. Among them, 31 genes predicted by at least two algorithms were selected for further experiment validation. qRT-PCR results indicated that the Cbl-b genes in the cells from 224.3+ B. melitensis group were down-regulated, which was confirmed by reduced protein expression of the gene product in Western blot analysis.
Mammalian Cbl family proteins consist of c-Cbl, Cbl-b and Cbl-3. Cbl-b is an E3 ubiquitin ligase.16–19 In the absence of co-stimulation, Cbl-b–/– T cells show constitutive activation that results in spontaneous autoimmunity or enhanced susceptibility to autoantigen.20,21 Cbl-b has the capacity to regulate Fcγ receptor-mediated phagocytosis. The macrophages from Cbl-b–/– mice demonstrate enhanced Fcγ receptor-mediated tyrosine phosphorylation, phagocytosis and target binding. 22 Cbl-b deficiency could exaggerate high-fat diet-induced insulin resistance through saturated fatty acid-mediated macrophage activation. 23 In vivo, Cbl-b limits the dissemination of Pseudomonas aeruginosa through interaction with the type III-secreted effector exotoxin T. 24 In our study, in the mCD14 knockdown RAW264.7 cells challenged with B. melitensis M5-90, the Cbl-b expression was down-regulated, which suggested that Cbl-b might be implicated in the interaction between Brucella and target cells.
In summary, our present study determined the differential expression of miRNA profiles in CD14-silenced RAW264.7 cells stimulated by B. melitensis infection, where mmu-miR-199a-3p and mmu-miR-183-5p were found to be up-regulated. Bioinformatics analysis revealed that the potential targets, Cbl-b, was altered; this was confirmed by qRT-PCR and Western blot analysis. These findings suggest that CD14 gene silencing alters specific miRNAs which are directly involved in inflammation of macrophages after B. melitensis infection. However, further studies to elucidate the exact function of mmu-miR-199a-3p, mmu-miR-183-5p and Cbl-b are warranted.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was financially supported by the National Natural Science Foundation of China (31460670).
