Abstract
Bacterial cell wall muramyl dipeptide (MDP) and glucosaminyl-MDP (GMDP) are potent activators of innate immunity. Two receptor targets, NOD2 and YB1, have been reported; we investigated potential overlap of NOD2 and YB1 pathways. Separate knockdown of NOD2 and YB1 demonstrates that both contribute to GMDP induction of NF-κB expression, a marker of innate immunity, although excess YB1 led to induction in the absence of NOD2. YB1 and NOD2 co-migrated on sucrose gradient centrifugation, and GMDP addition led to the formation of higher molecular mass complexes containing both YB1 and NOD2. Co-immunoprecipitation demonstrated a direct interaction between YB1 and NOD2, a major recombinant fragment of NOD2 (NACHT–LRR) bound to YB1, and complex formation was stimulated by GMDP. We also report subcellular colocalization of NOD2 and YB1. Although YB1 may have other binding partners in addition to NOD2, maximal innate immunity activation by muramyl peptides is mediated via an interaction between YB1 and NOD2.
Introduction
Defense against pathogens not only involves Ag-specific immunity, mediated by Abs and specialized immune cells, but also relies on broad-spectrum resistance known as innate immunity. A series of pathogen-specific molecules (PAMPs) produced by microbes and viruses induce a potent ensemble of defense strategies including cytokine release and inflammation, macrophage invasion, and the production of antimicrobial peptides, proteases, reactive oxygen species, and extracellular nets (for review, see Janeway and Medzhitov, 1 Ganz 2 , Brinkmann et al., 3 West et al., 4 and Cui et al. 5 ).
The principal bacterial PAMPs include bacterial LPS, cell-wall muramyl peptides, peptides bearing prokaryotic N-formylmethionyl residues, and unmethylated DNA. These induce innate immune responses by binding to specific PRRs; receptor activation leads to potent induction of innate immune pathways, notably by activation of transcription factor nuclear factor NF-κB that induces the expression of key of cytokines and chemokines involved in innate immunity.
LPS is known to bind to TLR4, 6 and TLR9 recognizes unmethylated CpG residues in bacterial DNA. 7 Three receptors, formyl peptide receptor (FPR)1–3, recognize N-formylated peptides. 8 However, the molecular targets for muramyl peptides have remained contentious.
Muramyl dipeptide (MDP) and glucosaminyl-MDP (GMDP) are potent immunostimulators released from the bacterial cell wall by the action of lysozyme and amidases, and synergize with LPS in the induction of the innate immune response.9–11 MDP is composed of N-acetylmuramic acid linked to the N-terminus of an
Attention initially focused on NOD2, encoded by the CARD15 gene in humans, as a likely sensor for cell-wall peptidoglycans.12–14 NOD2 is widely expressed in innate immune cells, and Nod2 knockout in mice was reported to impair MDP responses15–17 whereas expression of NOD2 can increase MDP signaling.13,18 Nonetheless, there has been debate concerning the identity of the MDP receptor. NOD2 was first reported to lack detectable affinity for immobilized MDP 19 and, although purified NOD2 protein was reported to bind to biotinylated MDP, excess biotinylated MDP retained only about 3% of NOD2 protein. 20 Subsequently it was reported that MDP was able to bind directly to recombinant NOD2 protein produced in insect cells. 21
However, it is open to debate whether NOD2 is the principal binding target of MDP. Although birds appear to lack a NOD2 gene (reviewed in Boyle et al. 22 ), and may respond less well to muramyl peptides, 23 an MDP analog was reported to exert potent immunostimulatory effects in chickens, 24 suggestive of an alternative receptor. In addition, the mitogenic and adjuvant activities of MDP can be separated, 25 and MDP can induce caspase-1 activation and IL-1 induction by a pathway independent of NOD2. 26 Indeed, using a peptide mimetic of GMDP,27,28 we reported expression cloning of GMDP-binding polypeptides from mouse splenocytes: All clones identified multifunctional factor YB1 as a specific target for GMDP. 29
YB1 (also known as Y box binding protein 1, YBX1) is a highly conserved multifunctional factor, and homologs of YB1 are present in bacteria, plants, insects, and vertebrates. 30 In mammals, YB1 acts as a transcription factor/messenger RNA (mRNA) binding factor that modulates the transcription and translation of innate immune genes encoding cytokines and their receptors,30,31 and may also regulate some aspects of Ag-specific immunity. 32 YB1 is secreted from the cell via a non-canonical secretion pathway, 33 and extracellular YB1 is found in the serum of sepsis patients but not of healthy controls. 34 Demonstrating the importance of YB1, mice genetically deficient for YB1 are not viable. 35 More recently, we confirmed specific binding of GMDP to YB1, subcellular colocalization of YB1 and GMDP, and demonstrated that YB1 knockdown impaired the biological response to GMDP. 36
Independent lines of evidence thus suggest that both NOD2 and YB1 may act as receptor targets for muramyl peptides, and we therefore addressed the possibility of overlap between YB1 and NOD2 in the innate immune response of mammalian cells to GMDP. We report here that YB1 interacts with NOD2 in the cellular response to muramyl peptide.
Materials and methods
General experimental procedures
Experimental details and reagents for the background procedures described here are presented in Laman et al. 36 As before, the induction of an innate immune response by GMDP was assessed by quantitative PCR of NF-κB (Nfkb2); Gapdh provided the internal control—as described previously, 36 qPCR of Nfkb2 transcript levels employed primers 5'-ATG CCT GAT TTT GAG GGA CTA TAC CCT GTA-3' and 5'-GAC TTG GTG GGT CTA TTG GAG GCC TTG TC-3' located within the downstream coding region of Nfkb2, and which identify all major splice variants. Gapdh internal control primers were 5'-GAC CAC AGT CCA TGC CAT CA-3' and 5'-TGA AGT CGC AGG AGA CAA CC-3'. For biochemical experiments, GMDP and MDP were synthesized chemically at the Institute for BioOrganic Chemistry.
Antibody (Ab) preparation
Abs to mouse NOD2 were produced by immunizing rabbits subcutaneously with NOD2 peptides KANGLAAFLLQHVRE (peptide 1) and KAEPHNLQIT (peptide 2) conjugated to BSA or OVA. Groups of two rabbits received either 2 mg BSA-peptide 1 or BSA-peptide 2. One wk later animals received the same injections, and after a further week were immunized with 2 mg OVA-peptide 1 or OVA-peptide 2, respectively. Immunization with OVA-conjugated peptides was repeated 1 wk later. After 3d, anti-peptide titers were approximately 1/100 000. Total Ig from combined serum from each pair of animals was purified twice by (NH4)2SO4 precipitation and re-suspended into PBS. For affinity purification, epoxysepharose (0.3 g) was washed in 50 mM NaHCO3 (pH 9.5) and re-suspended in 1 ml of the same buffer containing 4 mg of peptide 1 or peptide 2. After coupling (4h at room temperature, 18℃), the resin was washed, blocked (1 M ethanolamine, pH 8, 1h), and used to prepare affinity columns. Total Ig preparations were applied to the columns at room temperature, washed with PBS, and retained Ig was eluted with 2 ml 200 mM glycine-HCl, pH 2.3. Eluates were immediately neutralized (1 M Tris-HCl, pH 9.0) and dialyzed extensively versus PBS, yielding about 2 ml affinity-purified polyclonal anti-NOD2 peptide Ab (about 2 mg/ml). YB1 Abs were as described previously. 36 The specificity of both Abs was confirmed by detection of bands of the expected molecular mass on Western blotting and by their abrogation in gene-specific knockdown cells (Figure S1 in the Supplementary Data Online).
Knockdown experiments
Specific antisense RNA coding sequences 5'-GGGTCCTCCACGCAATTAC-3' (for YB1) 36 and 5'-TTGATCTCGCTGCGGTGAA-3' (for NOD2) were introduced downstream of a Pol III promoter in plasmid pSIREN-RetroQ-TetH (Clontech Laboratories Inc); mouse WEHI-3 monocytes were stably transfected with the expression plasmids using selection in 250 µg/ml hygromycin.
Sucrose gradient centrifugation
WEHI cells (2 × 107) were lysed in 100 µl 25 mM Tris HCl (pH 6.8), 150 mM NaCl, 5 mM MgCl2 containing 1% Triton X-100 and 1 mM phenylmethanesulfonyl fluoride (PMSF) for 5 min at room temperature. After centrifugation (5 min at 1200 g; 5 min at 5000 g) supernatants were collected and were divided into four portions. These were treated, respectively, with GMDP (0.1 µg/ml), YB1 (1 µg/ml), or GMDP + YB1, and incubated for 0.5h at 37℃. Aliquots (75 µl) of each sample was layered onto 5%–20% sucrose gradients in the same buffer without Triton X-100. The samples were centrifuged for 16h (SW40 rotor, Beckman Instruments), 111,000 g at rav, 4℃. Fractions (400 µl) were collected and analyzed by dot-blotting using biotinylated mouse anti-YB1 monoclonal or biotinylated rabbit anti-NOD2 polyclonal Abs; detection employed streptavidin-peroxidase conjugate and enhanced chemiluminescence detection. Molecular mass markers were IgG (150 kDa) and IgM (950 kDa).
Recombinant NOD2
For cloning and expression of NOD2 NACHT-leucine-rich repeat (LRR) in bacteria, the following primers were used: 5'-CTCCACATATGGTCGCCACCCTGGGCCTGGA273-3' (forward primer NdeI) and 5'-ATGCGGCCGCAAGCAAGAGTCTGGTGTCC-3' (reverse primer 1040) corresponding to amino acids 273–1040 of NOD2 (NACHT-LRR). PCR fragments were cloned into the expression vector pET22b as NdeI–NotI fragments in Escherichia coli strain Rosetta (DE3). Encoded polypeptides were isolated following bacterial disruption (model 300VT Ultrasonic Homogenizer) and Ni chelate chromatography (Ni-NTA agarose, Quiagen 30210) according to suppliers’ recommendations and as described previously for YB1.
36
To measure affinity for YB1 purified proteins (100 nM) were applied to immunoplates, washed (PBST), and different concentrations of YB1 were added as described in the Figure 1 legend. After incubation (1h, 37℃) and washing the plates were incubated with anti-YB1 Abs, washed, and developed using anti-mouse HRP conjugate. Negative control was an extract of total protein from E. coli strain Rosetta containing empty plasmid.
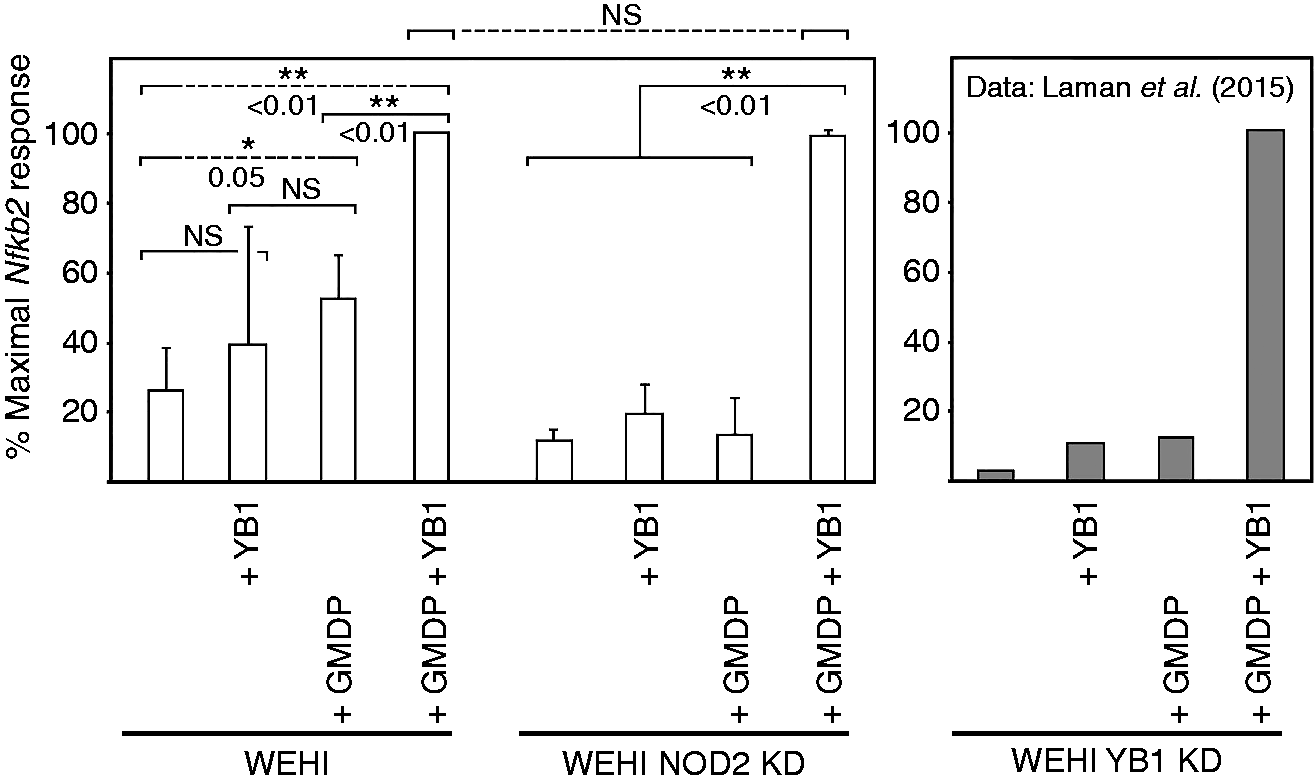
Knockdown demonstrates that both NOD2 and YB1 participate in induction of NF-κB (Nfkb2) expression by GMDP. WEHI cells alone, or knockdown (KD) cells stably transfected with an antisense construct to NOD2, were incubated for 2h at 37℃ in the presence or absence of GMDP (10 µg/ml) and/or recombinant YB1 protein (1 µg/ml), as indicated, and levels of Nfkb2 mRNA were determined. The internal control was Gapdh (not presented). The figure shows means and SDs of band intensities determined from three or more independent replicates; NS: not significant. Values were normalized against maximal stimulation; the maximal level of induction (+GMDP +YB1) was not significantly different in control versus NOD2 antisense transfected cells. The right-hand panel is provided only for comparison and is adapted from Laman et al.
36
Results
Both NOD2 and YB1 are required for the full stimulatory activity of muramyl peptides
We previously reported that knockdown of YB1 impaired the activation of innate immune pathways, as assessed by the induction of NF-κB (Nfkb2) by GMDP (see Figure 1). 36 This is believed to be an accurate proxy for innate immunity because the NF-κB precursor protein was cleaved following stimulation, translocated to the nucleus, and the expression of a key cytokine gene (Cxcr4) was induced. 36 To address potential overlaps between NOD2 and YB1 pathways, we prepared knockdown cell lines for NOD2; knockdown was confirmed by Northern and Western blotting (Supplementary Figure S1). Cells were treated with chemically synthesized GMDP, and qPCR was used to assess the induction of Nfkb2 expression, a marker of innate immunity. As shown in Figure 1, knockdown of NOD2 led to significant down-regulation of NF-κB induction by GMDP, although the response could be restored by excess exogenous YB1; the same result was obtained when GMDP was replaced by chemically synthesized MDP (Supplementary Figure S2), indicating that the YB1 does not distinguish between the two forms of muramyl peptides.
YB1 and NOD2 co-migrate on sucrose gradient centrifugation; GMDP enhances the formation of high molecular mass (MM) complexes
We addressed the possibility that YB1 and NOD2 might both contribute to a joint signaling complex. Splenocyte extracts were incubated with GMDP or YB1, or both agents, and were then resolved by sucrose density-gradient centrifugation. Fractions were separately probed with specific Abs against YB1 and NOD2. As shown in Figure 2, in addition to a peak at around 30 kD, potentially representing YB1 monomer, higher MM peaks were detected in the size range of 100–150 kDa, and these fractions were confirmed to contain both YB1 and NOD2 (Supplementary Figure S3). On treatment with GMDP, a small peak appeared at around 250 kDa that reacted modestly with anti-YB1 Ab, but very weakly with anti-NOD2 Ab. On addition of both GMDP and YB1, new GMDP-dependent high molecular mass complexes appeared in the size range 250–400 kDa that were positive both for YB1 and NOD2. This raised the possibility that, following GMDP stimulation, YB1 might be recruited to a larger complex that also contains NOD2, and that YB1 might bind directly to NOD2.
Sucrose gradient centrifugation (heatmap) of splenocytes pretreated with GMDP and/or YB1 showing higher MM complexes. Cells were pretreated with GMDP (1 µg/ml) or YB1 (1 µg/ml); after cell disruption in the presence of 1% Triton X-100 cell extracts were resolved by sucrose gradient centrifugation. Aliquots were immobilized on membranes and probed with peroxidase-labeled anti-YB1 or anti-NOD2 as indicated, and developed colorimetrically. Intensities of fractions were normalized to peak intensity in each gradient (black) and presented as a heatmap.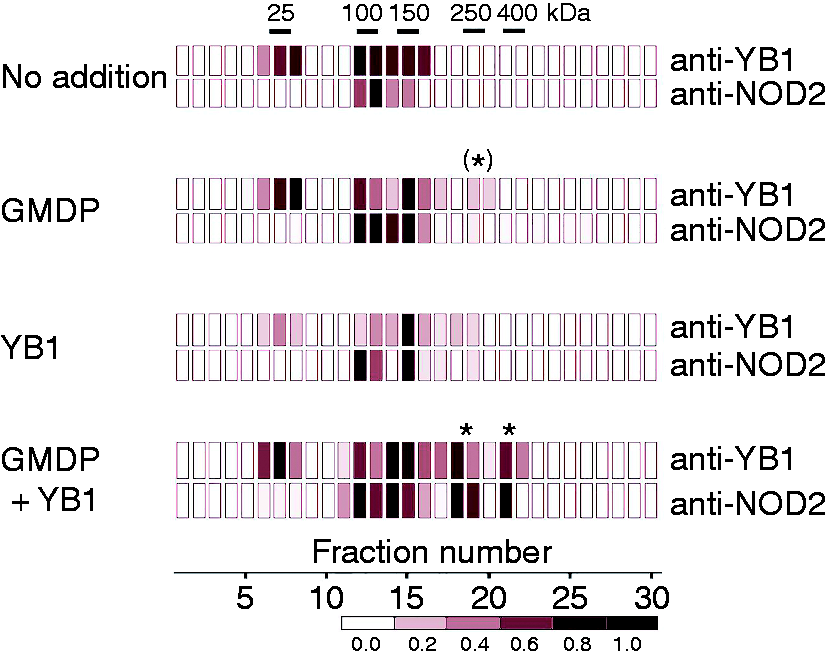
Evidence for a direct interaction: coimmunoprecipitation of YB1 and NOD2
We therefore treated monocytes, macrophages, or splenocytes with GMDP, and recovered protein complexes with either Ab to YB1 or Ab to NOD2. These were resolved by gel electrophoresis and probed with Ab to the other protein. Figure 3 shows that Ab to YB1 retains a protein that reacts with anti-NOD2 Ab and that has an apparent MM (ca. 100 kDa) consistent with NOD2 protein. Conversely, Ab to NOD2 retained a protein that reacts with anti-YB1 Ab and that migrates with an apparent MM (ca 50 kDa) consistent with the migration behavior of YB1 (YB1 migrates abnormally slowly on sodium dodecyl SDS-PAGE
30
). In all cases complex formation was promoted by GMDP, but there was evidence for binding in the absence of GMDP. We conclude that YB1 and NOD2 both contribute to a protein complex, and that GMDP promotes the formation of this complex.
YB1 binds to NOD2. (a) Monocytes immunoprecipitated with anti-YB1 and probed with anti-NOD2. Lane M, molecular mass markers (kDa); 1,2, lysates following GMDP pretreatment at 1 and 10 µg/ml; 3, cell lysate with no addition of GMDP; 4, as in 2, but where the anti-YB1 Ab was omitted. (b) As in (a), but using extracts of macrophages or splenocytes pretreated with GMDP (1 µg/ml) and immunoprecipitated with (lanes 1,3) or without (lanes 2,4) Ab to YB1, and probed with Ab to NOD2. (c) Splenocytes pretreated with GMDP (lane 1), or untreated (lane 2), immunoprecipitated with anti-NOD2 Ab, and Western blotting was performed using anti-YB1 Ab. In lane 3 the primary Ab was omitted.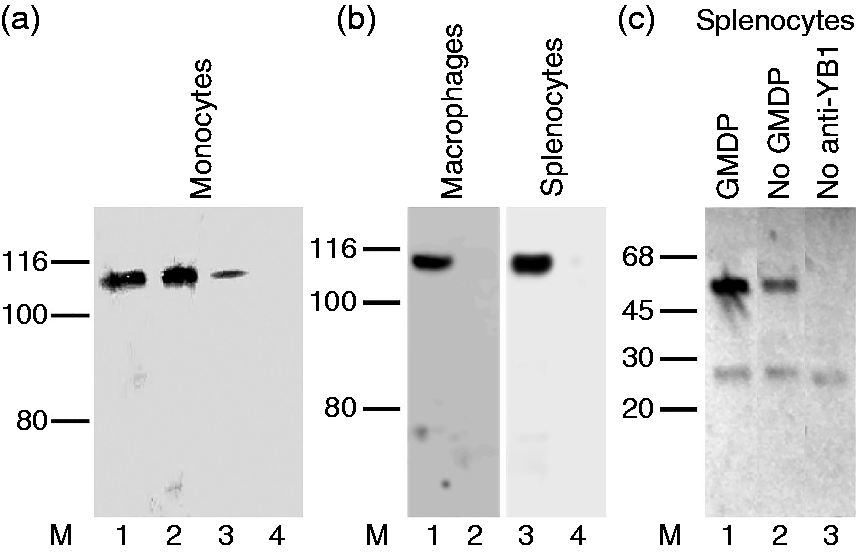
GMDP is present in the YB1/NOD2 complex
The finding that GMDP promotes YB1/NOD2 complex formation raises the question of whether it acts indirectly or whether the molecule itself forms part of a complex containing both polypeptides. We therefore treated splenocytes with fluorescein isothiocyanate (FITC)-labeled GMDP; complexes containing YB1 and/or NOD2 were affinity purified using anti-YB1 or anti-NOD2 Abs, and examined for FITC fluorescence. As shown in Figure 4, both Abs retained fluorescence that was abolished by excess unlabeled GMDP. We conclude that GMDP forms part of the YB1 and NOD2 complex.
Fluorescence of anti-YB1 and anti-NOD2 immune complexes from mouse primary splenocytes treated with FITC-GMDP. Emission intensities were normalized to the mean signal (520 nm) generated with anti-YB1 Ab (100%). Lane 1, fluorescence retention by nonspecific Ab in extracts of splenocytes treated with FITC-GMDP (10 µg/ml). Lanes 2,3, fluorescence of FITC-GMDP (10 µg/ml)-treated splenocytes retained by anti-YB1 and anti-NOD2 Ab, respectively. Lanes 4,5, as in lanes 2,3 but where cells were pre-incubated with excess unlabeled GMDP (10 µg/ml FITC-GMDP + 100 µg/ml GMDP). Values are means ± SD of triplicate replicates. No fluorescence was observed when FITC-GMDP was omitted (not presented).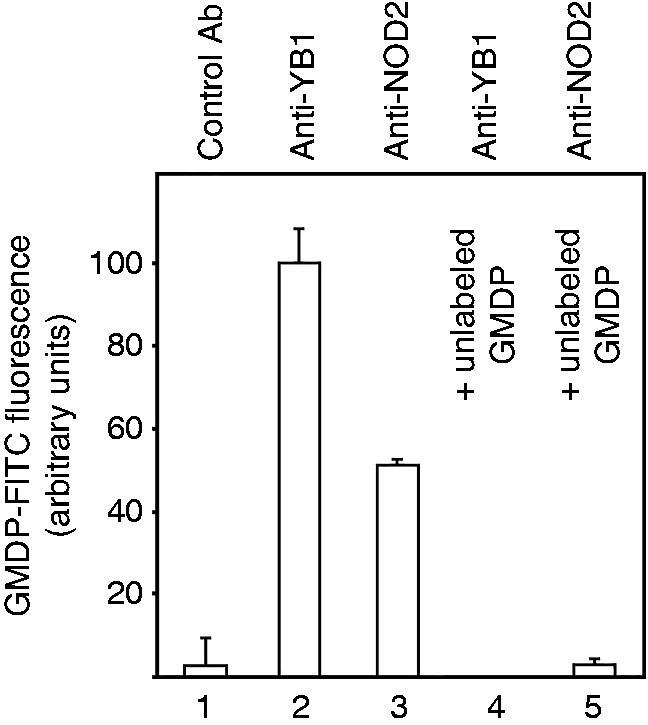
Subcellular colocalization of YB1 and NOD2
We previously demonstrated that YB1 and (FITC)-GMDP colocalize at the subcellular level.
36
Because other cell types were unsuitable for subcellular microscopy, we used primary dendritic cells to address whether YB1 and NOD2 colocalize within the cell. Primary mouse dendritic cells were preincubated with GMDP and then probed with fluorescent (FITC or AlexaFluor 555) Abs against YB1 and NOD2; cells were then visualized by confocal microscopy. Figure 5 reveals prominent colocalization of YB1 with NOD2.
Subcellular colocalization of NOD2 and YB1. Mouse primary dendritic cells treated with GMDP were incubated with Abs against NOD2 or YB1. (a) Hoechst 33342 counterstain (blue); (b) AlexaFluor 555-anti-NOD2; (c) FITC-anti-YB1; (d) superimposition of (a)–(c).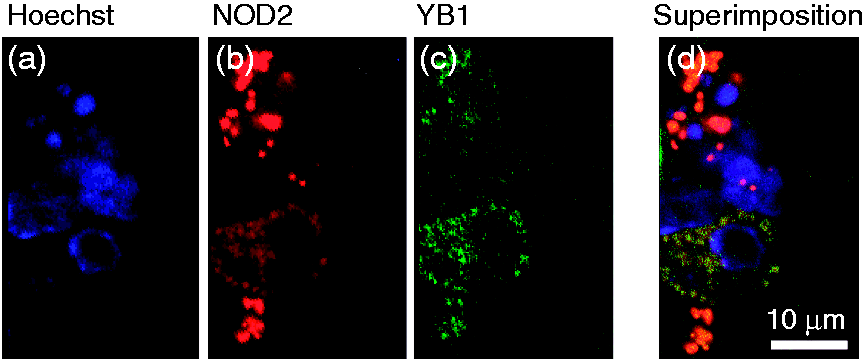
YB1 binds to the NACHT-LRR domain of NOD2
Given evidence for co-migration on sucrose gradient centrifugation, co-immunoprecipitation, and co-localization within the cell, we addressed direct binding of purified YB1 to recombinant NOD2. Because we were unable to express full-length NOD2 in bacteria (data not presented) we cloned the central NACHT (NAIP/CIITA/HET-e/TP-1; nucleotide-binding and oligomerization)-LRR fragment of NOD2 (termed here NODNL); this was expressed in E. coli (Supplementary Figure S4). Both YB1 produced in E. coli
36
and NOD2NL (this work) were purified to homogeneity. The NOD2NL fragment was immobilized on microtiter plates and probed with different concentrations of recombinant YB1 in the presence or absence of GMDP. After washing, the plates were developed with Ab to YB1. Figure 6 demonstrates that YB1 binds directly to NOD2NL. Moreover, the extent of complex formation was increased in the presence of GMDP. We conclude that YB1 binds directly to NOD2, and that complex formation is promoted by GMDP.
The recombinant NACHT-LRR fragment of NOD2 (NOD2NL) purified from an E. coli expression strain (Supplementary Figure S4) was immobilized on a multi-well plate and incubated with different concentrations of recombinant YB1 in the presence or absence of GDMP (10 µg/ml). After washing, the plate was developed with anti-YB1-peroxidase and chemiluminescence detection. Error bars are SD. YB1 concentrations were (nM), 7.5, 15, 30, 60, 120, 240 (log scale) in samples 1–6, respectively. Control extract was from E. coli bearing the same expression plasmid lacking the NOD2NL insert.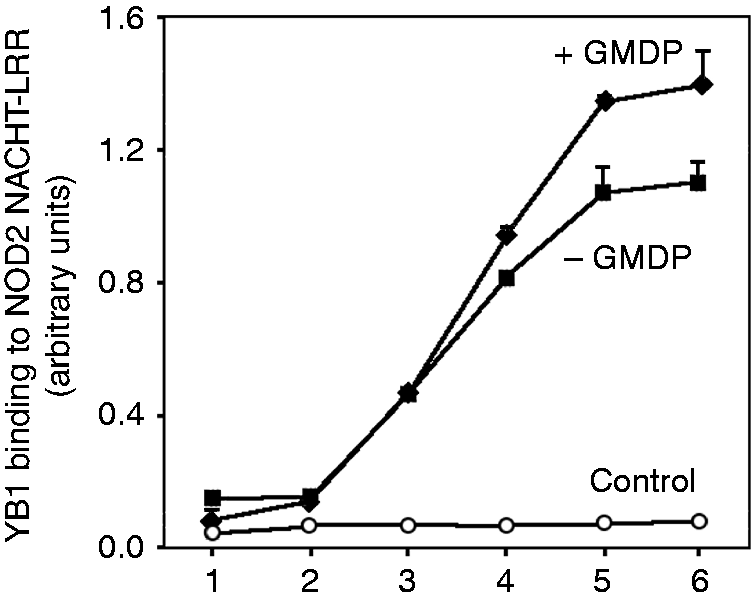
Discussion
This work addresses the specific binding target for muramyl peptides in the induction of the innate immune response in mammals. There is evidence that GMDP binds directly to YB1, 36 but further data argue that NOD2 centrally contributes to the biological activity of muramyl peptides,15–17 potentially through direct binding. 21 We provide data that could explain this dichotomy: We report that YB1 binds to NOD2, and that complex formation is stimulated by chemically synthesized GMDP. Although we suggest that the primary binding site is located on YB1, knockdown experiments indicate that both YB1 and NOD2 are required for a maximal response.
However, despite contradictory data (that muramyl peptides poorly bind to NOD219,20), it was reported that human NOD2 expressed in Sf21 insect cells (derived from the Fall armyworm Spodoptera frugiperda, a moth) directly binds to MDP. 21 We have been unsuccessful in demonstrating binding of GMDP to cloned NOD2 (not presented) but, because we have been unable to express full-length NOD2 in our system (E. coli), this failure per se does not exclude the possibility that GMDP might bind to authentic NOD2. We also note remarkable conservation of YB1 across species including insects (Supplementary Data Online).
Further, we present data that NOD2 may not be absolutely required for complex formation with muramyl peptides and YB1. Knockdown of NOD2 impaired induction of NF-κB expression by GMDP, but this could be overcome by the addition of excess YB1. In addition, although GMDP plus YB1 drove the formation of high MM complexes (250–400 kDa) containing both YB1 and NOD, as revealed by sucrose density-gradient centrifugation, GMDP alone (in the absence of excess YB1) produced high MM complexes that contained endogenous YB1, but little NOD2. Although these experiments do not prove rigorously that YB1 can act independently, they point to the possibility that YB1 may have another binding partner in addition to NOD2. The absence of NOD2 in birds, reptiles, and amphibians 22 (these species all contain YB1 homologs strikingly similar to mammalian YB1), also suggests that other (non-NOD2-dependent) pathways may operate in these species.
In mammals, NOD2 is one of a large family of evolutionarily related NLRs, including NOD1 and NOD3 (NLRC2), with up to 30% sequence identity between pairs; 37 it is possible that YB1 has some affinity for NOD1/3. In the present work it is of note that our anti-NOD2 Abs are unlikely to react with other NOD-like molecules because (i) a single polypeptide was observed on Western blotting (Supplementary Figure S1), (ii) gene expression knockdown of NOD2 abrogated protein detection (Supplementary Figure S1), and (iii) the specific peptides used to prepare anti-NOD2 Ab (“Materials and methods”) are not conserved between NOD2 and other NOD-like molecules (data not presented). Further work using specific Abs against NOD-family members, including NOD1, will be necessary to address whether YB1 interacts with other NLRs.
YB1 may also bind to the NOTCH3 cell surface receptor 38 that, like NOD2, activates NF-κB pathways. 39 Although NOTCH signaling is generally implicated in cell/cell interactions and the organization of organogenesis, NOTCH pathways may also play an important role in innate immunity (reviewed in Shang et al. 40 ). Other potential YB1 binding partners have been reported (for review see Eliseeva et al. 30 ).
We previously raised the issue of the domain of interaction between YB1 and GMDP, 36 now extended to its binding partner NOD2. GMDP is principally an extracellular moiety released from invading bacteria. By contrast, the NOD2 receptor is intracellular, although an association with the inside face of the cell membrane may be required for activity,41,42 specifically the C-terminal LRR domain. Although administered muramyl peptides are rapidly taken up from the circulation, 43 and may enter cells via peptide transporters, membrane vesicles, and bacterial pore-forming toxins/secretion systems (for review see Clarke and Weiser 42 ), YB1 potentially offers a further route. Because YB1 straddles both the extracellular and intracellular domains, being secreted into the extracellular milieu—from where it is efficiently taken up, by a mechanism not yet elucidated—it is therefore possible that extracellular YB1 binding to muramyl peptides might contribute to (or compete with) their cellular uptake.
In sum, we conclude that YB1 binds to NOD2, leading to the formation of higher MM complexes, and this association is promoted by GMDP. A summary schema is presented in Figure 7. Maximal induction of innate immunity, as assessed through induction of NF-κB, requires the integrity both of YB1 and NOD2. It remains to be established whether the major binding contacts of GMDP lie in YB1, in NOD2, or at their interface; this will require crystallization of the ternary complex.
Simplified schema of muramyl peptide activation of innate immunity via YB1 and NOD2. It was previously demonstrated that YB1 + GMDP treatment leads to up-regulation, processing, and nuclear transport of NF-κB2, resulting in gene expression induction of Cxcr4, a component of the innate immune response.
36
A key feature of the NOD2 pathway involves dimerization/multimerization of NOD2 polypeptide;
44
indeed, enforced multimerization of NOD2 alone can drive NF-κB activation.
45
Our sucrose gradient centrifugation data (Figure 2) are consistent with the interpretation that YB1 + GMDP together induce the formation of higher-order complexes. However, NOD2 is not essential for YB1 + muramyl peptide function (Figure 1), suggesting that alternative pathways may be involved. As indicated (*), other NOD-like receptors, including NOD1 and NOD3 (NLRC3), have significant homology to NOD2,
37
and it is possible they may interact with YB1 and/or GMDP/MDP. There are suggestions that the action of muramyl peptides may, in addition, be mediated in part by other members of the NOD-like family, potentially including NALP3 and NALP1 (reviewed in Geddes et al.
46
). Other known binding partners of YB1 (reviewed in Eliseeva et al.
30
) include NOTCH3 that, like NOD2, has been shown to activate NF-κB pathways (see text); it is not yet known whether muramyl peptides modulate YB1 binding to other partners.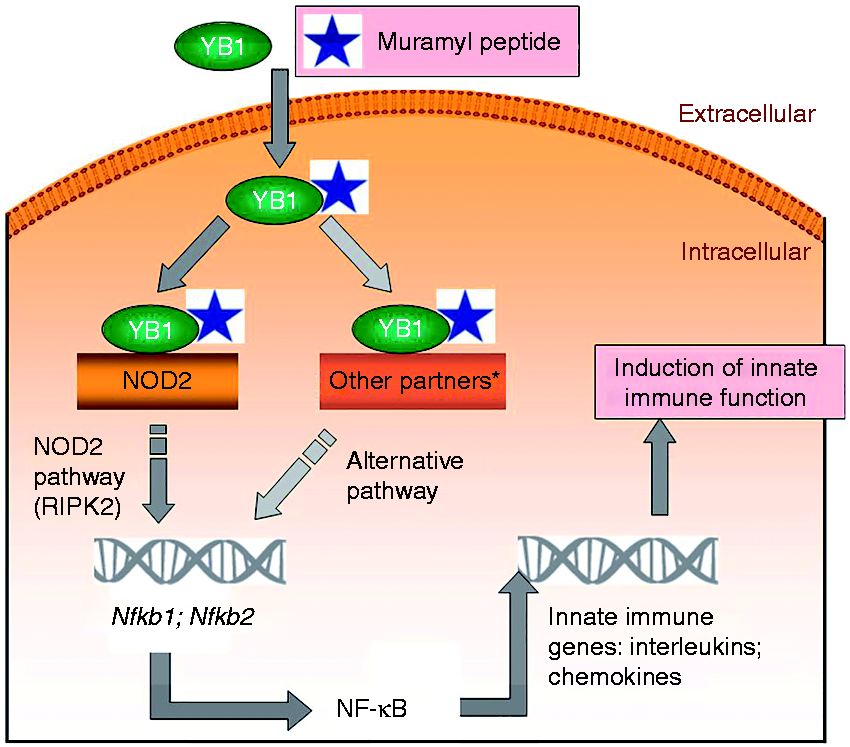
However, there are suggestions that YB1 may have other binding partners, so far unidentified, that could also contribute the innate immune response induced by bacterial muramyl peptides (Figure 7). In addition, the unusual properties of YB1, a conserved polypeptide that exchanges between intracellular and extracellular localizations, could suggest that YB1 might be involved in cellular uptake of muramyl peptides and potentially in their delivery (as a YB1 complex) to NOD2. Moreover, YB1 is up-regulated in response to infection and appears in the circulation (e.g. Hanssen et al. 34 ). YB1 released from sites of infection could thus play an intracrine/endocrine role in sensitizing adjoining tissues to muramyl peptides, a possibility that warrants further investigation.
Footnotes
Acknowledgments
We thank Juergen Haas (Edinburgh) for critical comments on the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Russian Science Foundation (project 14-50-00131). Confocal microscopy experiments (SG) were supported by grant RFBR 16-04-01152.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
