Abstract
Actinobacillus pleuropneumoniae (APP), the etiologic agent of porcine pleuropneumonia, forms biofilms on biotic and abiotic surfaces. APP biofilms confers resistance to antibiotics. To our knowledge, no studies have examined the role of APP biofilm in immune evasion and infection persistence. This study was undertaken to (i) investigate biofilm-associated LPS modifications occurring during the switch to biofilm mode of growth; and (ii) characterize pro-inflammatory cytokines expression in porcine pulmonary alveolar macrophages (PAMs) and proliferation in porcine PBMCs challenged with planktonic or biofilm APP cells. Extracted lipid A samples from biofilm and planktonic cultures were analyzed by HPLC high-resolution, accurate mass spectrometry. Biofilm cells displayed significant changes in lipid A profiles when compared with their planktonic counterparts. Furthermore, in vitro experiments were conducted to examine the inflammatory response of PAMs exposed to UV-inactivated APP grown in biofilm or in suspension. Relative mRNA expression of pro-inflammatory genes IL1, IL6, IL8 and MCP1 decreased in PAMs when exposed to biofilm cells compared to planktonic cells. Additionally, the biofilm state reduced PBMCs proliferation. Taken together, APP biofilm cells show a weaker ability to stimulate innate immune cells, which could be due, in part, to lipid A structure modifications.
Introduction
Biofilms are adherent communities of bacteria attached to biological or non-biological surfaces and encased within a self-produced polymeric matrix that protects cells from noxious elements in the surrounding environment.1,2 Biofilm represents a serious therapeutic challenge, as bacteria within a biofilm are usually more resistant to antibiotics and to immune clearance.3–6 Biofilms are ubiquitous and nearly all microorganisms are able to adhere to surfaces and generate biofilms. This includes Actinobacillus pleuropneumoniae, a Gram-negative porcine pathogen and member of the Pasteurellaceae family. 7 A. pleuropneumoniae strains can form biofilms on both abiotic and biotic surfaces.7–10 A. pleuropneumoniae cells within biofilms are 100–30,000 times more resistant to antibiotics than their planktonic counterparts, 11 and might play an important role in the persistence of bacterial infection in swine.
A. pleuropneumoniae is the etiologic agent of porcine pleuropneumonia, a fibrinous, hemorrhagic, highly contagious disease affecting pigs of all ages and leading to high economic losses in the swine industry. 12 Symptoms and signs of porcine pleuropneumonia are depression, loss of appetite, fever and severe breathing difficulties with some coughing. 13 After death, it is common to see blood discharge out of the nasal cavity due to extensive lung tissue damage. 13 The disease in its acute form is associated with high rates of morbidity and mortality.13,14 Lung tissues are damaged by the RTX toxins, ApxI, Apx II, Apx III and Apx IV.15,16 In addition, surface LPS induces an inflammatory response that contributes to necrosis of porcine lung epithelial cell.17–19 Inflammatory cytokines in lung tissue may exacerbate the effect of A. pleuropneumoniae, and add to host-mediated tissue damage.17,19,20 Conversely, subclinical or chronic forms of the disease are asymptomatic, undetectable and less invasive.21,22 Chronic infection plays a role in persistence and transmission of several bacteria.19,20 Numerous studies have also reported that biofilms play a role in immune evasion and bacterial survival.23–25 In A. pleuropneumoniae, most studies have been performed using planktonic cells, despite the important role played by biofilms in the pathogenesis of human and animal chronic infection.26–29
Nothing is known about how A. pleuropneumoniae biofilm interacts with immune cells, and if lipid A undergoes structural modifications in A. pleuropneumoniae biofilms. The purpose of this study was to (i) investigate modifications of lipid A occurring during the switch to the biofilm mode of growth and (ii) characterize the pattern of inflammatory cytokines expression in porcine pulmonary alveolar macrophages (PAMs) and the proliferative response of porcine PBMCs challenged with A. pleuropneumoniae planktonic and biofilm cells.
Materials and methods
Cell line and bacterial strain
A. pleuropneumoniae MBHPP147 (APP147) is a non-hemolytic derivative of the serotype 1 reference strain S4074.
10
APP147 contains deletions in both the apxIC and apxIIC genes. Bacterial strain was cultured in brain heart infusion (BHI) broth or agar (Oxoid Ltd, Basingstoke, UK) supplemented with 15 µg/ml NAD incubated at 37℃ in 5% CO2. Porcine PAM 3D4/21 cell line was purchased from the American Type Culture Collection (ATCC CRL-2843) and was maintained in RPMI 1640 medium (Invitrogen, Eugene, OR, USA) with 10% FBS (heat inactivated) and adjusted to contain 2 mM
Biofilm and planktonic cultures
A. pleuropneumoniae was grown in six-well plates (Costar 3516, Corning, Corning, NY, USA), as previously described. 30 After 4 h in culture, media containing non-attached, planktonic bacteria were removed (the planktonic fraction). Attached bacteria were washed twice with PBS to remove remaining planktonic cells. Attached cells (the biofilm fraction) were scraped off the plate. Both planktonic and biofilm fractions were washed again with PBS, centrifuged and re-suspended in fresh RPMI 1640 medium. Bacterial counts (CFU) and OD were used to normalize bacterial concentrations. Bacteria were killed using UV irradiation (254 nm, 30 min), then plated on BHI agar to verify the loss of viability.
Lipid A isolation
Lipid A molecules were extracted from both planktonic and biofilm bacteria as described by El Hamidi et al. 31 Briefly, bacteria from both planktonic and biofilm fractions were lyophilized. Freeze-dried bacteria (10 mg) were suspended in 200 µl isobutyric acid-1 M ammonium hydroxide mixture (5:3, v:v), and kept for 2 h at 100℃ in a screw-cap test tube under agitation. Suspensions were cooled in ice, and then centrifuged 10 min at 2000 g. Supernatants were diluted with two volumes of water and lyophilized. The dried materials were then washed twice with 200 µl of methanol. Finally, lipid A was extracted from pellets with 100 µl of a mixture of chloroform, methanol and water (3:1.5:0.25, v:v:v).
Mass spectrometry and data processing
Analyses were performed using a Q-Exactive Orbitrap Mass Spectrometer (Thermo Scientific, San Jose, CA, USA), interfaced with a Thermo Scientific UltiMate 3000 Rapid Separation UHPLC system, using a pneumatic assisted heated electrospray ion source (ESI). Chromatography was achieved using an isocratic mobile phase along with a microbore column Thermo Biobasic C8 100 × 1 mm, with a particle size of 5 µm. The mobile phase consisted of methanol and water (fortified with 0.1% of formic acid) at a ratio of 95:5. The flow rate was fixed at 75 µl/min and 2 µl samples were injected. Mass spectrometry (MS) was performed in positive ion mode and operated in scan mode at high resolution and accurate mass (HRAM). Nitrogen was used for sheath and auxiliary gases, set at 10 and 5 arbitrary units, respectively. The heated ESI probe was set to 4000 V and the ion transfer tube temperature was set to 300℃. The scan range was set to m/z 1000–2500. Data were acquired at a resolving power of 140,000 (defined as full width at half maximum), resulting in a scanning rate of 700 ms/scan when using automatic gain control target of 3.0 × 106 and maximum ion injection time of 200 ms. The instrument was calibrated prior to all analyses, and mass accuracy was notably below 1 ppm using Thermo Pierce calibration solution and automated instrument protocol. Differential analyses of planktonic vs. biofilm were performed using Thermo Scientific SIEVE 2.1 software, which executes background subtraction, component detection, peak alignment and differential analyses. Parameters used in SIEVE were as follows: time range 1–10 min, mass range 1000–2500 Da, frame width 10 mDa and retention time width 2 min. SIEVE output was transferred into SIEVE Extractor, an Excel spreadsheet and Lipid Maps databases. SIEVE software measures P-values using a two-tailed Student's t-test. Fold changes and P-values were used to construct a volcano-plot and a Venn diagram analysis. Significance was set a priori to P < 0.01 and only twofold changes were considered biologically significant.
Stimulation of porcine PAMs
PAMs were prepared in six-well plates (Costar 3516) to a final concentration of 1 × 106 viable cells/ml in 1.5 ml of RPMI 1640 medium. Plates were incubated at 37℃ in a humidified incubator 5% CO2 for 24 h. Medium was removed and fresh medium containing planktonic or biofilm bacteria in suspension with a multiplicity of infection (MOI) of 10:1 was added to the wells. 32 As negative control, medium alone was added. Exposed and non-exposed cells were incubated for 1–24 h at 37℃, 5% CO2.
Cytokine mRNA expression
Primers used for quantitative real-time RT-PCR.

IL-8 quantification by ELISA
Levels of IL-8 in cell culture supernatants were measured by sandwich ELISA using pair-matched Abs from R&D Systems (Minneapolis, MN, USA), according to the manufacturer's recommendations. Twofold dilutions of recombinant porcine IL-8 (R&D Systems) were used to generate the standard curves. Sample dilutions giving OD readings in the linear portion of the standard curve were used to quantify the levels of IL-8.
Proliferation of porcine PBMCs
The procedures for animal care followed the guidelines of the Canadian Council on Animal Care and the protocol for blood collection was approved by our Institutional Animal Care Committee (Protocol 13-Rech-1558). Blood was collected from four healthy pigs (3–10 wk of age) into heparinized tubes, and mononuclear cells were isolated using lymphocyte separation medium (Wisent, St-Bruno, QC, Canada) and centrifuged at 400 g for 30 min. The buffy coat containing mononuclear cells was isolated, transferred to a fresh centrifuge tube and washed twice with PBS. The final cell pellet containing PBMCs was re-suspended to a final level of 2 × 106 cells/ml in RPMI 1640 medium supplemented with 25 mM HEPES buffer, 2 mM
Cell proliferation was measured upon completion of the 48-h incubation period using a BrdU-ELISA kit (Millipore, Billerica, MA, USA), as per the manufacturer’s instructions. In brief, kit-provided BrdU labeling solution (20 μl) was added to each well and the plate was incubated at 37℃ for 24 h. Thereafter, the cells were centrifuged at 300 g for 10 min, the labeling solution was removed, and the cells were fixed by addition of fixative/denaturing solution (200 μl/well) and incubation at room temperature for 30 min. Ab conjugate (anti-BrdU solution, 100 μl/well) was then added and the plate was incubated at room temperature (22℃) for 1 h. The cells were then washed twice with washing buffer (200 μl), and a peroxidase goat anti-mouse IgG HRP conjugate (100 μl) was added to each well for 30 min. Again, the cells were washed twice with washing buffer (200 μl). Substrate solution (100 μl) was added to the wells and incubated in the dark at room temperature for 15 min. Then, 100 μl stop solution was added and the absorbance in each well was measured at 450 nm in an automated plate reader (Powerwave; BioTek Instruments, Winooski, VT, USA). Blanks (100 μl culture medium alone) and control wells were included in each experiment.
Statistical analysis
qRT-PCR, ELISA and PBMC proliferation assay data were analyzed statistically using GraphPad Prism software (version 4.03; GraphPad Software Inc., San Diego, CA, USA) by application of Wilcoxon–Mann–Whitney test or one-way ANOVA with Bartlett’s test. P < 0.05 or P < 0.01 were considered to reflect statistically significant differences.
Results
Lipid A structure analysis
Lipid A molecules were extracted following a validated protocol,
31
and their relative abundance in A. pleuropneumoniae planktonic and biofilm cells were analyzed using MS. For planktonic and biofilm cells, a total of 228 distinct molecular entities were observed and compared. Analyses were performed using an untargeted HPLC-HRAM MS lipidomic workflow. After data acquisition, the results were processed using bioinformatics software SIEVE 2.1 to perform retention time alignment and identify specific ions (i.e. m/z) that differed between planktonic and biofilm cell groups. The m/z values for peaks of interest were searched in public metabolite/lipid databases (i.e. METLIN, mzCloud and Lipid MAPS) to obtain putative identifications without success. This is not surprising as the lipid A molecular structure and associated molecules for A. pleuropneumoniae remain unknown and not available in public databases. Following peak alignment and identification from biofilm lipid A (n = 3) and planktonic lipid A (n = 3), relative ion intensities and m/z values were used to study the molecular abundance distribution and modifications between the two cell states. Extracted data were represented using volcano plot with a P-value = 0.01 (confidence interval of 99%) and a twofold change cut-off were assumed to be biologically significant. Figure 1A illustrates the significant variations in lipid A abundance and molecular structures between A. pleuropneumoniae planktonic and biofilm cells with representative chromatograms shown in Figure 1B.
Volcano plot of distribution of A. pleuropneumoniae lipid A structures isolated from biofilm cells (n = 3) or planktonic cells (n = 3). (a) The volcano plot displays the relationship between fold-change and significance between the two groups, using a scatter plot view. The y-axis is the negative log10 of P-values (a higher value indicates greater significance) and the x-axis is log2 fold change or the difference in abundance between two experimental groups. The dashed red line shows where P = 0.01, with points above the line having P < 0.01 and points below the line having P > 0.01. (b) Chromatograms of structures indicate by black arrows in the volcano plot.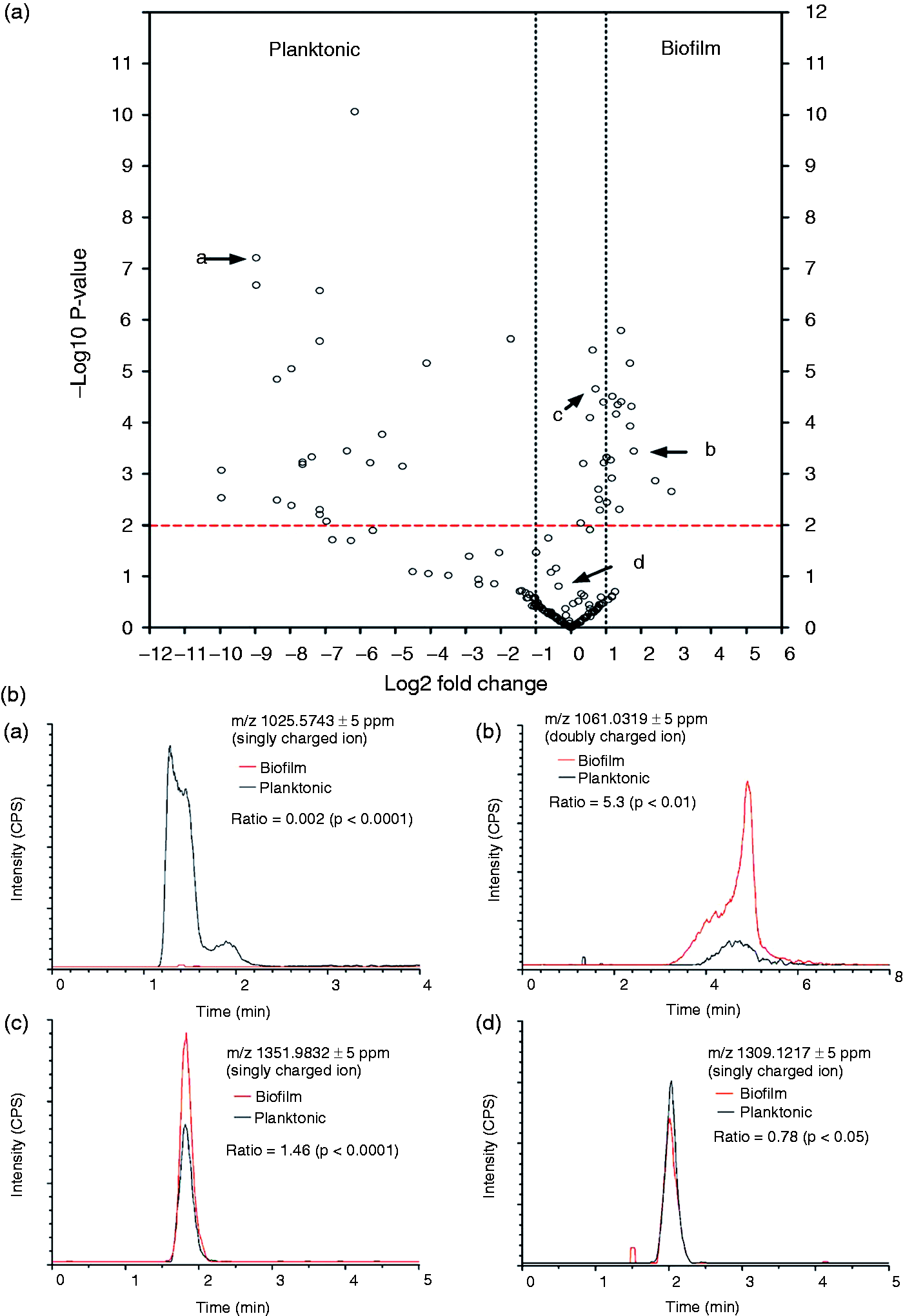
Following an exhaustive analysis, a total of 23 lipid A structures in the planktonic fraction and 17 in the biofilm fraction were found to be significantly different (Figure 2A). Inspection of the detected peaks (m/z) revealed an interesting shift of the lipid A towards higher molecular mass in biofilm cells, as shown in Figure 2B.
Actinobacillus pleuropneumoniae biofilm and planktonic lipid A structures distribution. (a) Venn diagram showing the differential abundance of lipid A structures between planktonic and biofilm cells with twofold change or higher and P-value < 0.01. (b) Distribution of small (1000–2000 Da) and large (2000–4500 Da) molecular mass (MW) lipid A structures in biofilm and planktonic cells.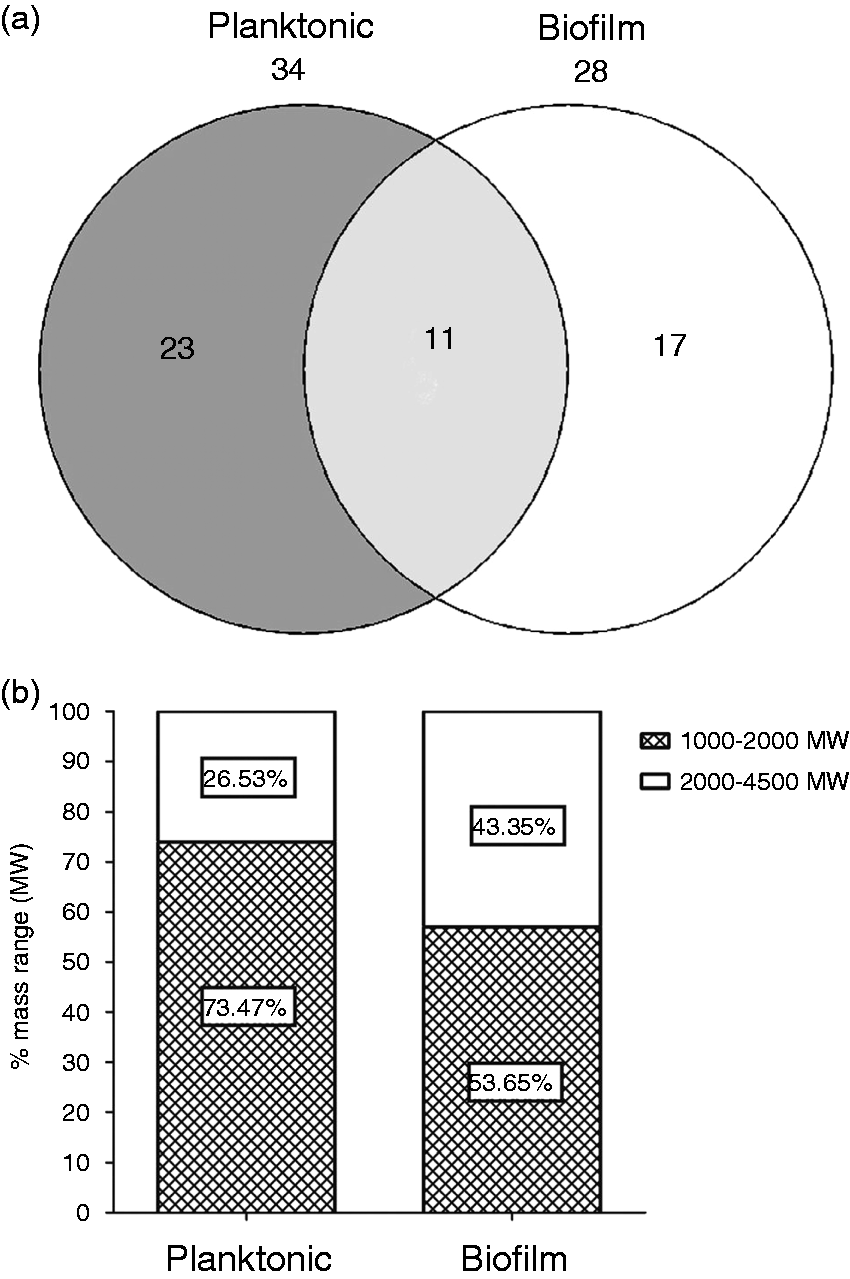
Activation of macrophages
Relative mRNA expression of pro-inflammatory cytokine genes (IL1, IL6, IL8 and MCP1) was measured in PAM cells following exposure to either inactivated A. pleuropneumoniae biofilm or planktonic cells. The expression of all tested cytokines mRNA was up-regulated when PAMs were exposed to A. pleuropneumoniae cells compared with unexposed control cells. qRT-PCR data showed an early expression of all cytokines tested when exposed to planktonic cells (Figure 3). A peak value was reached at 1 h post-stimulation and declined shortly, except for IL-8, which demonstrated a particularly high and sustained expression compared with the other cytokines. Two other peaks at 8 h and 24 h post-stimulation for MCP-1, and one peak for IL-1 at 24 h post-stimulation were observed. These peaks could be the result of an autoactivation process caused by the increase of cytokines in the culture media.
Cytokine expression in porcine pulmonary alveolar macrophages exposed to biofilm (BIOF) or planktonic (PLK) cells of A. pleuropneumoniae (MOI 10:1) for different time periods (1–24 h). Fold changes in mRNA expression of (a) IL-1, (b) IL-6, (c) IL-8 and (d) MCP-1 relative to the uninfected control are presented. The means ± SD from three independent experiments are shown. Statistical analyses were performed using 2–ΔΔCt values and all marked with (*) or (**) were statistically significant (P < 0.05 or P < 0.01, respectively).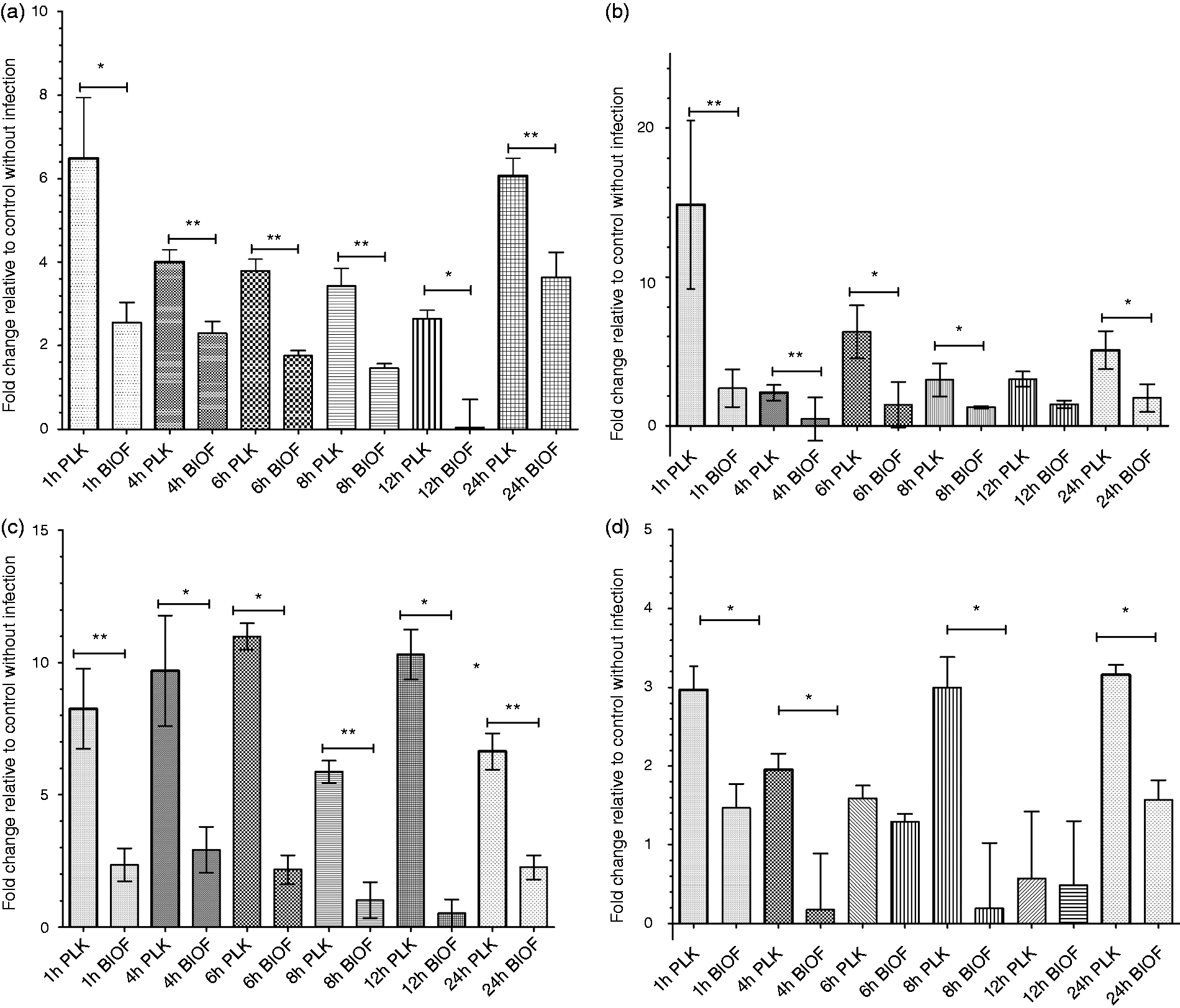
Interestingly, exposure of macrophages to A. pleuropneumoniae biofilm cells appeared to induce significantly lower expression of pro-inflammatory cytokines and chemokines in general compared with macrophages exposed to planktonic cells (P < 0.05) (Figure 3).
The level of IL-8 in the supernatant of PAM cells following exposure to either A. pleuropneumoniae biofilm or planktonic cells was also measured by ELISA at 1 h, 2 h, 4 h and 8 h (Figure 4). Our results demonstrated that the amount of IL-8 tends to increase with time. As expected, levels of IL-8 detected at 4 h and 8 h post-stimulation with biofilm cells were significantly lower compared with levels obtained with planktonic cells (P < 0.05), confirming mRNA results.
IL-8 production by PAMs in response to stimulation by planktonic (PLK) and biofilm (BIOF) cells (MOI 10:1) for different time periods (1–8 h). Data are expressed as mean ± SD (in pg/ml) from at least three independent experiments and results marked with (*) were statistically significant (P < 0.05).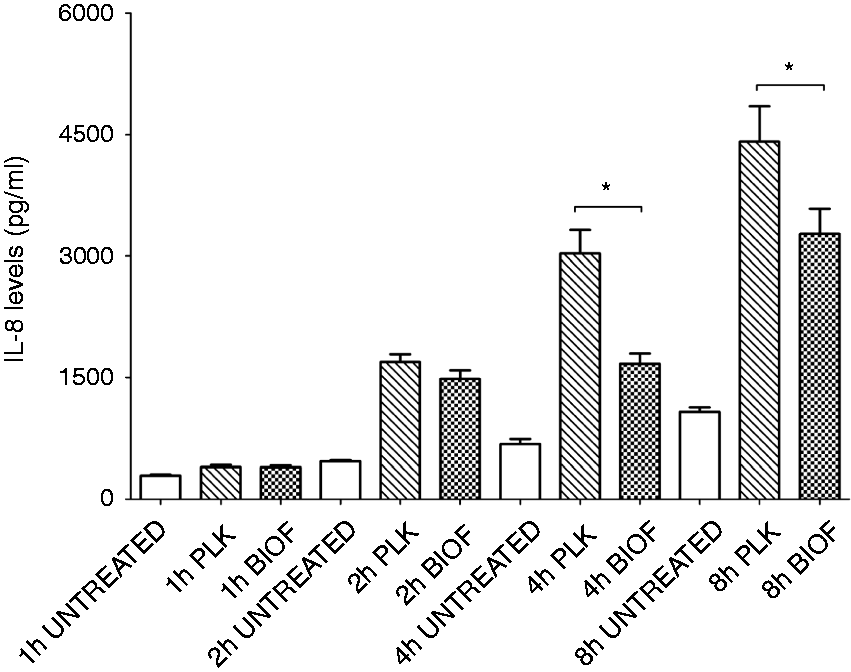
Porcine PBMCs proliferation
As A. pleuropneumoniae biofilm induces less activation of PAMs, we hypothesized that biofilms cells could also modulate the proliferative response of PBMCs. PBMCs were exposed to both biofilm and planktonic cells, and the proliferation was evaluated and compared with unexposed cells (negative controls) (Figure 5). As expected, a significantly higher proliferation was observed when immune cells were exposed to concanavalin A or A. pleuropneumoniae cells (planktonic and biofilm) compared with unexposed cells control (P < 0.05). The proliferative response of PBMCs exposed to planktonic and biofilm cells was then evaluated. Biofilm cells induced a lower response than their planktonic counterparts (P < 0.05) (Figure 5).
Proliferation of porcine peripheral blood monocytes exposed to biofilm and planktonic cells of A. pleuropneumoniae (2 × 107 cells/ml). Proliferation rate in PBMCs stimulated with Con A, planktonic (PLK) and biofilm cells (BIOF) of A. pleuropneumoniae was evaluated using a BrdU-ELISA assay. The means ± SD from three independent experiments are shown. (*) or (**) indicate that results were statistically significant (P < 0.05).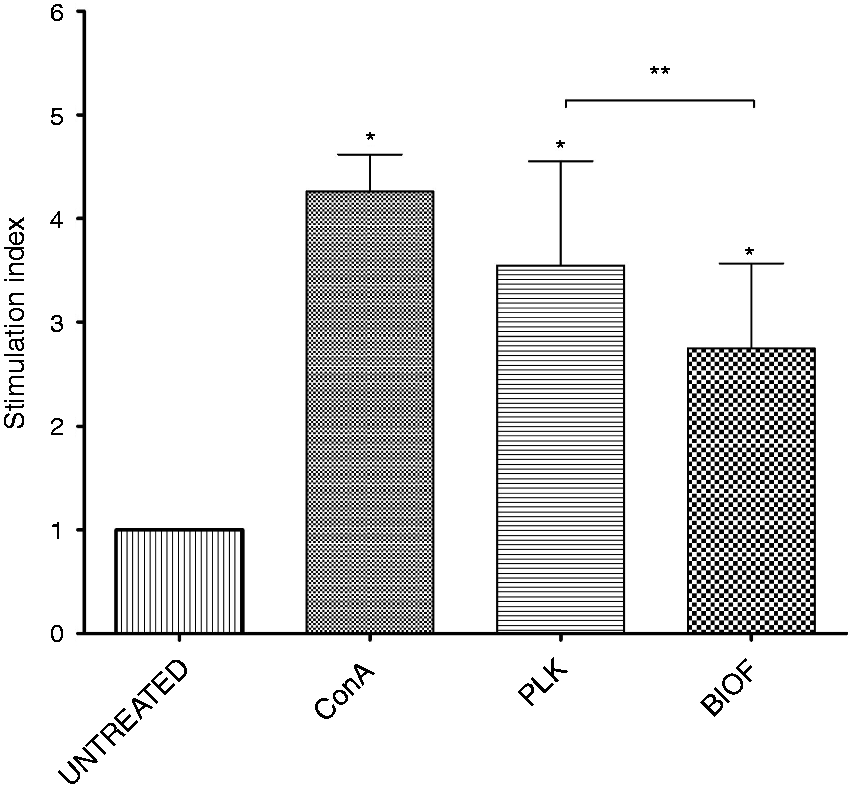
Discussion
LPS has been implicated as one of the major virulence factors in A. pleuropneumoniae and other Gram-negative bacteria.18,36–40 LPS has several functions. First of all, it protects bacteria by establishing a permeability barrier.40–42 The biological activity of LPS stems from its hydrophobic anchor, lipid A, 39 which stimulates the TLR4, found on the surface of monocytes, macrophages and neutrophils.43–45 Lipid A will first bind the co-receptor MD-2, before interacting with the TLR4 receptor.43,44 The formed complex TLR-4/MD-2/LPS subsequently activates transcription factor NF-kB and the pro-inflammatory cytokines IL-1, IL-6 and IL-8.45,46 Several structural modifications in lipid A, such as the loss of phosphate groups and variation in length and number of the fatty acyl chains, have an important role in TLR4 recognition and response.40,45,47–50 Variations in lipid A structure provide bacteria fitness advantages to colonize surfaces, evade the immune system and persist during infections.49,51,52 Some of these variations are associated with chronic infection and biofilm formation.47,51,53
Biofilm bacteria are encased within a polymeric matrix that contains, among other things, polysaccharides, extracellular DNA, proteins and lipids.54–56 They are frequently associated with chronic disease,25,26,57 and represent a serious therapeutic challenge, as bacteria within biofilm exhibit enhanced antibiotic tolerance and are more resistant to clearance by the immune system.3,11,24,57 We recently demonstrated that LPS has an important role in biofilm formation in A. pleuropneumoniae, and that a biofilm matrix exopolysaccharide, poly-N-acetyl-
In this study, using an HPLC-HRAM MS untargeted lipidomic based approach, we demonstrated that the lipid A molecular structure seems to be influenced by the bacterial mode of growth (sessile or planktonic). Planktonic and biofilm cells differed significantly in lipid A structure and quantity, with larger lipid A molecular entities observed in biofilm cells. This constitutes an additional manner in which biofilm cells differ phenotypically from their free-floating counterparts.58–60 However, further analyses will be required to determine molecular identity (i.e. extensive purification and 1H/ 13 C NMR).
Lipid A contributes to cell rigidity by increasing cell wall strength.61,62 During biofilm formation, embedded bacteria may exert pressure on their neighbors, 62 and the increased molecular interactions could favor reactions leading to modified lipid A structures and cell surface remodeling.
Several studies have demonstrated that biofilms can undergo LPS structural modifications that modulate the immune response.47,48,51 Recently, Chalabaev et al. described an interesting biofilm associated-phenotype linked to lipid A structure.
47
They found that among several Gram-negative bacteria, including Escherichia coli and Pseudomonas aeruginosa, lipid A can undergo palmitoylation, and these changes were found to be more abundant in biofilm than in planktonic cells.
47
Lipid A structural modifications, including addition of 4-amino-4-deoxy-
Cytokines like IL-1, IL-6, IL-8 and MCP-1 play important roles in mediating inflammatory response to pathogens. Pro-inflammatory IL-8 and MCP-1 trigger neutrophils migration to the site of the infection. One of our objectives was to see how planktonic and biofilm A. pleuropneumoniae cells interact with immune cells. Our data indicate that biofilm cells trigger a different innate immune response than do planktonic cells. Previous studies have shown that A. pleuropneumoniae infections were followed by increased TNF-α, IL-1 and IL-8 mRNA expression. These cytokines were detected at the periphery of the lung lesion.17,63 Increased inflammatory cytokine levels were also seen in lung lesions from acute porcine pleuropneumonia.17,63
Using IL-10 gene therapy, Morrison et al. demonstrated that this anti-inflammatory cytokine can diminish levels of the pro-inflammatory cytokines IL-1 and TNF-α and consequently reduce lung damage and severity in pleuropneumonia. 20 In this study, we observed that levels of pro-inflammatory cytokines were significantly lower in PAMs exposed to biofilm cells compared with those exposed to planktonic cells. Furthermore, porcine PBMCs exhibited a lower proliferative response when exposed to biofilm vs. planktonic cells. Overall, our data suggest that the biofilm mode of growth mitigates the innate immune response to this pathogen.
The role of biofilm in chronic disease and low-level inflammation response is increasingly recognized. Many studies support this hypothesis.64,65 Daw et al. showed that macrophages infected with of Enterococcus faecalis biofilm cells had lower pro-inflammatory cytokine levels than planktonic cells. 66 Similarly, macrophages infected with Legionella pneumophila biofilm exhibited significantly lower levels of the pro-inflammatory cytokine IL-1β compared with those infected with planktonic bacteria. 67 In this way, biofilm-derived L. pneumophila avoids major inflammatory responses and recruitment of inflammatory cells. 67 Similarly, Thurlow et al. demonstrated that Staphylococcus aureus biofilm evades TLR recognition and alters the immune response to infection. 68 Indeed, several inflammatory signals, including MCP-1, TNF-α and IL-1β, were significantly attenuated in S. aureus biofilm-infected tissues compared with the wound-healing response elicited by insertion of sterile catheter. 68
Fewer lung lesions and lower inflammation and are often associated with the chronic form of pleuropneumoniae,17,20,63 suggesting the involvement of biofilm. A. pleuropneumoniae growing in biofilm may overcome host defenses by modifying its lipid A structure, which enables a lower immune reactivity. However, lipid A structural modifications may not be the sole cause of the immune dysfunction observed here and other bacterial surface components may also be implicated. Further studies will be needed to demonstrate the direct impact of these biofilm associated-lipid A structural modifications in the altered immune response.
The lipid A structure in A. pleuropneumoniae has yet to be elucidated. However, fatty acids in the lipid A of A. pleuropneumoniae serotype 1 were identified, and it has been shown that lipid A contains different proportions of n-dodecanoic acid (0.6%), n-tetradecanoic acid (12.9%), 3-hydroxytetradecanoic acid (64.3%), n-hexadecanoic acid (3.2%) and n-octadecanoic acid (0.6%). 69 Knowing the lipid A structure and the fatty acids arrangement in A. pleuropneumoniae would help to identify the modifications occurring in lipid A during the switch from planktonic to biofilm form and identify their impact on immune response.
This study demonstrates that the A. pleuropneumoniae biofilm mode of growth has an impact on lipid A structure and mitigates the innate immune response. Further investigation must focus on these lipid A modifications associated with lower immune cell stimulation.
Footnotes
Acknowledgements
The mutant strain A. pleuropneumoniae MBHPP147 was kindly provided by Ruud P.A.M Segers (MSD Animal Health, Boxmeer, The Netherlands). We would like to thanks Philip Dominico for reading the manuscript and providing precious advice. Thermo Fisher Scientific provided generous access to a Q-Exactive Orbitrap Mass Spectrometer to FB.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by a Discovery grant from the Natural Sciences and Engineering Research Council of Canada (NSERC grant 003428) to MJ; SH and LM received a scholarship from the Centre de recherche en infectiologie porcine et avicole (CRIPA); the analyses were performed using analytical instruments acquired with an NSERC Research Tools and Instruments grant to FB (439748-2013); partial funding was also provided by the NSERC (grant 342150) to MS.
