Abstract
The interactions between chemokines and their receptors are crucial for differentiation and activation of inflammatory cells. CC chemokine ligand 11 (CCL11) binds to CCR3 and to CCR5 that in leporids underwent gene conversion with CCR2. Here, we genetically characterized CCL11 in lagomorphs (leporids and pikas). All lagomorphs have a potentially functional CCL11, and the Pygmy rabbit has a mutation in the stop codon that leads to a longer protein. Other mammals also have mutations at the stop codon that result in proteins with different lengths. By employing maximum likelihood methods, we observed that, in mammals, CCL11 exhibits both signatures of purifying and positive selection. Signatures of purifying selection were detected in sites important for receptor binding and activation. Of the three sites detected as under positive selection, two were located close to the stop codon. Our results suggest that CCL11 is functional in all lagomorphs, and that the signatures of purifying and positive selection in mammalian CCL11 probably reflect the protein’s biological roles.
Introduction
CC chemokine ligand 11 (CCL11), also known as eotaxin-1, is a chemoattractant for eosinophils with an important role in allergic and parasitic inflammation.1–3 First isolated in a guinea pig model of asthma, 4 this protein is located in the monocyte chemoattractant protein region of the CC cluster of several mammals, including human, mouse, rat, rabbit, horse and cow.5,6 In the European rabbit (Oryctolagus cuniculus) genome, CCL11 has been mapped to chromosome 19:23,739,202-23,742,061 on the forward strand, with an Ensembl gene designation of ENSOCU00000005935. CCL11 can exert its functions through the interaction between residues located in its extracellular loops and NH2-terminus and the N-terminus of two receptors, CCR3 and CCR5.7–9
An extensive gene conversion in the CCR5 transmembrane domain has been reported for several species.10–15 In contrast, in some leporids, CCR5 suffered a dramatic change at the second extracellular loop, resulting from a gene conversion event with the paralogous CCR2. This alteration was confirmed in the European rabbit, Riverine rabbit (Bunolagus monticularis) and Amami rabbit (Pentalagus furnessi), but it was not observed in the Eastern cottontail (Sylvilagus floridanus) or in European and Iberian hares (Lepus europaeus and L. granatensis).9,11 In the other Lagomorpha family, Ochotonidae (pikas), this gene conversion is also absent. 16 The most likely evolutionary scenario that could explain this pattern is that the gene conversion event occurred in the ancestor of the Oryctolagus, Bunolagus and Pentalagus genera at ∼8 million yr ago, probably conferring some selective advantage, and thus became fixed in the ancestral population. 16
This CCR5 evolutionary pattern led to the study of the CCR5 ligands in lagomorphs. The study of CCL3, CCL4 and CCL5 revealed that these genes are all functional in leporids and showed evidence of strong purifying selection. 17 CCL14 is functional in the Leporidae family. In the Ochotonidae family, CCL14 is suffering a pseudogenization process with an intact gene in some species, while in others it is a pseudogene. In contrast, among the leporids, CCL8 is pseudogenized in the European rabbit and Riverine rabbit, but functional in Sylvilagus and Lepus.18,19
In chemokine receptors the sites located intracellularly or in the transmembrane domains are involved in signal transduction and dimerization. 20 Consequently, new nucleotide changes that lead to amino acid (aa) alterations tend to be quickly eliminated. In contrast, the aa residues localized in the extracellular domains are evolving under selective pressure resulting from ligand binding and pathogen interactions. Thus, in these regions, the proportion of non-synonymous nucleotide substitutions is expected to be significantly higher than synonymous substitutions. This pattern has been observed in chemokine receptors such as CCR2 and CCR3. 20 The interactions between chemokine ligands and their receptors suggest that the chemokine ligands might also exhibit signatures of positive and purifying selection. Here, we characterized CCL11 in lagomorphs and investigated the selective pressures that have been driving the evolution of this molecule in mammals.
Materials and methods
Primers and conditions used for PCR amplification and sequencing of CCL11 from lagomorphs’ gDNA samples.
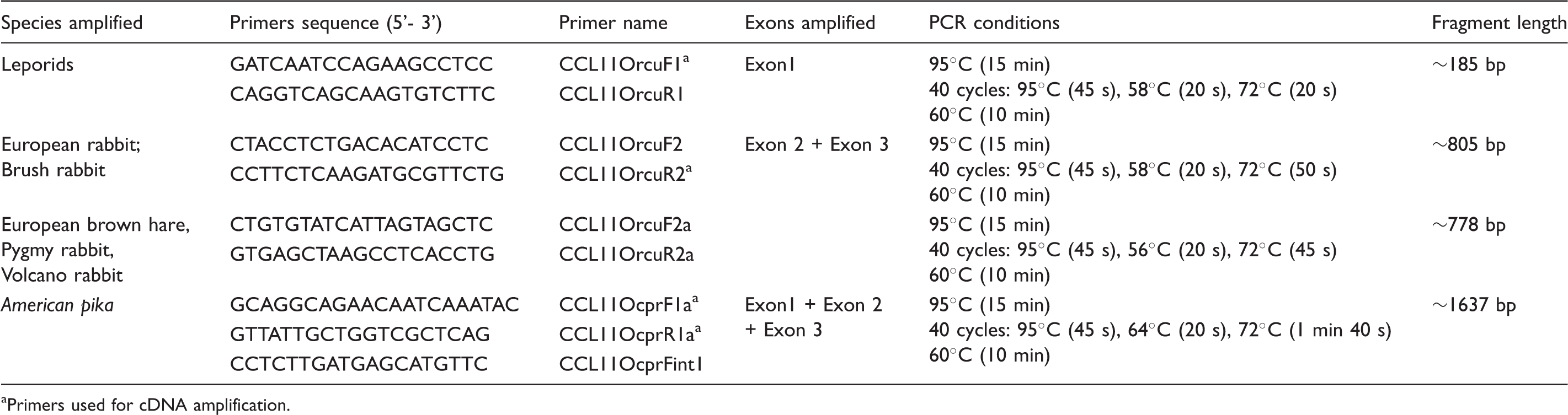
Primers used for cDNA amplification.
Sequences were aligned using MUltiple Sequence Comparison by Log-Expectation (MUSCLE), available at http://www.ebi.ac.uk/. 21 The program PHASE, built into the software DnaSP, 22 was used to reconstruct the haplotype phases of the obtained sequences. Haplotypes were translated using BioEdit. 23
In order to identify which codons of CCL11 are under selection (purifying and positive), we estimated ω, i.e, the ratio of non-synonymous (dN) to synonymous (dS) substitutions in CCL11 orthologs (final dataset of 68 sequences) by employing codon-based maximum likelihood (ML) methods available in the HYPHY package implemented in the DataMonkey webserver:24,25 Single-likelihood ancestor counting (SLAC), fixed-effect likelihood (FEL), internal branch FEL (iFEL), random-effect likelihood (REL) and fast unconstrained Bayesian approximation (FUBAR).25–27 For the first three methods, the P-value was set to ≤ 0.05; for FUBAR we used a P-value ≥ 0.95 and for REL we used a Bayes factor >95. The best-fitting model for nucleotide substitution was determined by the automatic model selection tool available in the webserver. We further used the codon-based ML method (CODEML) implemented in PAML v4.4. 28 An unrooted neighbor-joining tree was constructed using MEGA6, 29 with P-distance as substitution model and the pairwise deletion option for gaps/missing data. The topology of the phylogenetic tree obtained follows the accepted mammalian phylogeny for the major groups. Two pairs of site-based models were compared: M1 (nearly neutral) vs. M2 (selection) and M7 (neutral, β) vs. M8 (selection, β and ω), where M1 and M7 correspond to the null hypothesis and M2 and M8 to the alternative hypothesis by allowing positive selection. A likelihood ratio test with two degrees of freedom determined whether a selection model fit the data better than a neutral model.30,31 Codons under positive selection were identified by using a Bayes Empirical Bayes approach with probability > 95%. As done previously for other immunity genes,32–37 only the codons detected by more than one method were considered as being under selection.
The three-dimensional (3D) structure displaying the interaction of human CCL11 with CCR3 was downloaded from the Protein Data Bank in Europe (PDBeurope), available at http://www.ebi.ac.uk/pdbe/entry/pdb/2MPM, and Discovery Studio 3.5 software (BIOVIA, San Diego, CA, USA) was used to map the sites under selection.
The secondary structures of human, European rabbit and Pygmy rabbit CCL11 were predicted by using PsiPred (http://bioinf.cs.ucl.ac.uk/psipred/).38,39 This software calculates the protein cysteines that create disulfide bonds by using position specific iterated BLAST to obtain evolutionary information that is used to predict the secondary structure of the query protein.
Results and discussion
We amplified and sequenced the CCL11 gene for six lagomorph species: both subspecies of European rabbit, European brown hare, Brush rabbit, Pygmy rabbit, Volcano rabbit and American pika. These sequences were further compared with CCL11 sequences available for other mammals, and some differences were observed (Figure 1). Indeed, at position 40, where human has an Asn, all leporids have a deletion and the American pika has a Lys. There are also some aa changes that are only present in some lagomorph species: Val12 and Met43 in the European rabbit; Thr10 (Pygmy rabbit); His44 (Volcano, Pygmy and Brush rabbits and European brown hare); Phe70 (Volcano rabbit); and Met2, Ser5, Asn53, Leu66, Ser89 (American pika). In addition, we found a Met65 in all CCL11 sequences from the leporids amplified in this work, while the sequence from the European rabbit available in public databases (ENSOCUG00000005935) has an isoleucine. Additionally, and despite the identification of two different transcripts in the American pika sequence available online [XM_004593867.1 and XM_004593868.1 – deletion/insertion of two aa (Asp26 and Ser27, respectively)], we were only able to detect the first transcript. The listed aa alterations lead to changes in charge and polarity (Supplementary data 1), which can induce modifications in protein structure and conformation, and even alter the protein functions.
40
Alignment of CCL11 for several mammalian species. GenBank and Ensembl accession numbers are indicated in bold for the European rabbit and American pika retrieved sequences. Purifying selected aa are shaded in light gray, while positively selected aa are shaded in dark gray; cysteine residues are boxed and human signal peptide is underlined. (*) represent stop codons; (–) represent indels; (▪) above the numbering represent the sites important for the interaction of CCL11 with its receptors9,46–48,57 *1 and *2 represent different alleles. Numbering is according to the human CCL11 sequence and the signal peptide and indels were included in the numbering. Disulfide bonds between side-chain cysteines are represented by a light-gray dashed line. Human (Homo sapiens), European rabbit (O. cuniculus cuniculus and O. cuniculus algirus), European brown hare (L. europaeus), Pygmy rabbit (B. idahoensis), Brush rabbit (S. bachmani), Volcano rabbit (R. diazi) and American pika (O. princeps); mouse (Mus musculus); deer mouse (Peromyscus maniculatus bairdii); rat (Rattus norvegicus); naked mole-rat (Heterocephalus glaber); spalax mole-rat (Nannospalax galili); long-tailed chinchilla (Chinchilla lanigera); Guinea pig (Cavia porcellus); prairie vole (Microtus ochrogaster); lesser Egyptian jerboa (Jaculus jaculus); degu (Octodon degus); Chinese hamster (Cricetulus griseus); Nancy Ma's night monkey (Aotus nancymaae); black-capped squirrel monkey (Saimiri boliviensis boliviensis); green monkey (Chlorocebus sabaeus); Sumatran orangutan (Pongo abelii); common chimpanzee (Pan troglodytes); Pygmy chimpanzee (Pan paniscus); Rhesus macaque (Macaca mulatta); southern pig-tailed macaque (Macaca nemestrina); crab-eating macaque (Macaca fascicularis); Sooty mangabey (Cercocebus atys); Angola black-and-white colobus (Colobus angolensis palliates); drill (Mandrillus leucophaeus); Philippine tarsier (Tarsius syrichta); gray mouse lemur (Microcebus murinus); Coquerel's sifaka (Propithecus coquereli); olive baboon (Papio anubis); northern greater galago (Otolemur garnettii); common marmoset (Callithrix jacchus); northern white-cheeked gibbon (Nomascus leucogenys); cattle (Bos taurus); Arabian camel (Camelus dromedarius); bactrian camel (Camelus bactrianus); wild bactrian camel (Camelus ferus); alpaca (Vicugna pacos); sheep (Ovis aries); wild boar (Sus scrofa); domestic goat (Capra hircus); chiru (Pantholops hodgsonii); lesser hedgehog tenrec (Echinops telfairi); cape golden mole (Chrysochloris asiatica); cape elephant shrew (Elephantulus edwardii); minke whale (Balaenoptera acutorostrata scammoni); killer whale (Orcinus orca); sperm whale (Physeter catodon); baiji (Lipotes vexillifer); horse (Equus caballus); Przewalski's horse (Equus przewalskii); white rhinoceros (Ceratotherium simum simum); black flying fox (Pteropus alecto); David's myotis (Myotis davidii); Florida manatee (Trichechus manatus latirostris); European hedgehog (Erinaceus europaeus); Chinese tree shrew (Tupaia chinensis); nine-banded armadillo (Dasypus novemcinctus).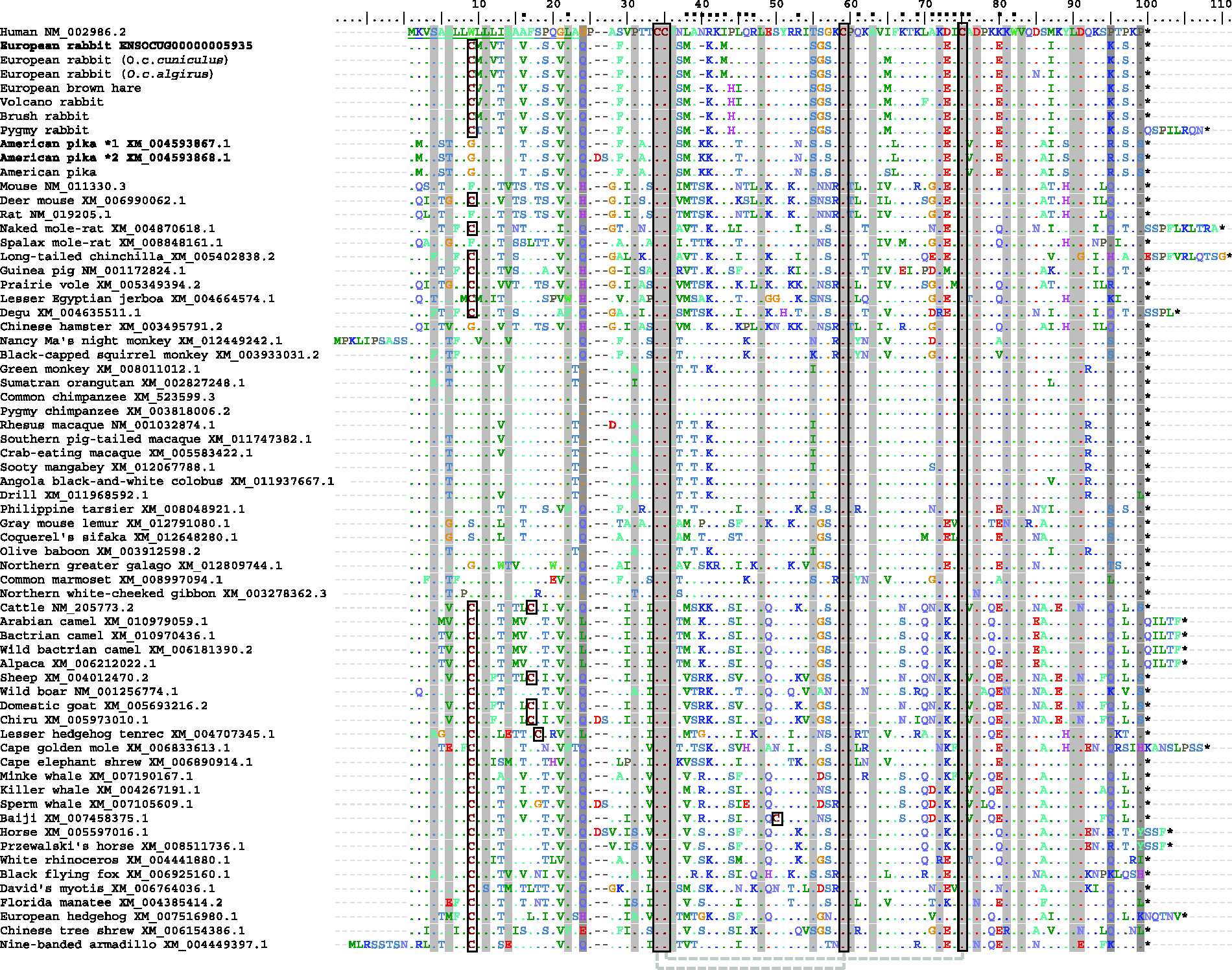
Strikingly, within leporids, the Pygmy rabbit has a mutation in the stop codon leading to an extension of eight aa (Gln100–Asn107) resulting in a protein with 104 aa. This mutation was detected in three different individuals, confirming that this was not a PCR artifact. Other mammals also code for longer proteins, due to mutations in the stop codon (Figure 1), leading to CCL11 proteins ranging from 100 to 108 aa. The protein length is important for the 3D structure, and different sizes may imply different folding patterns, which may affect protein functions. 41 The implications of these mutations in the protein structure are unknown; however PsiPred results do not predict any differences in the secondary structure of these species when compared with humans.
Disulfide bonds between cysteine residues are important for protein structure and function, being involved in an array of biological processes.42–44 For CCL11, the cysteine residues and the predicted disulfide bonds are highly conserved between all the mammals studied (Cys34–Cys59 and Cys35–Cys75). However, some extra cysteines were detected: cattle, sheep, domestic goat and chiru have Cys17 and the lesser hedgehog tenrec has Cys18. In addition, most mammals, with the exception of primates, American pika, mouse, rat, spalax mole-rat and Chinese hamster, have an extra cysteine at position 9. Their location in the signal peptide suggests no role in establishing extra disulfide bonds. In contrast, the Baiji sequence has an extra cysteine at position 50 that can potentially establish a disulfide bond.
Phylogenetic tests of selection. a
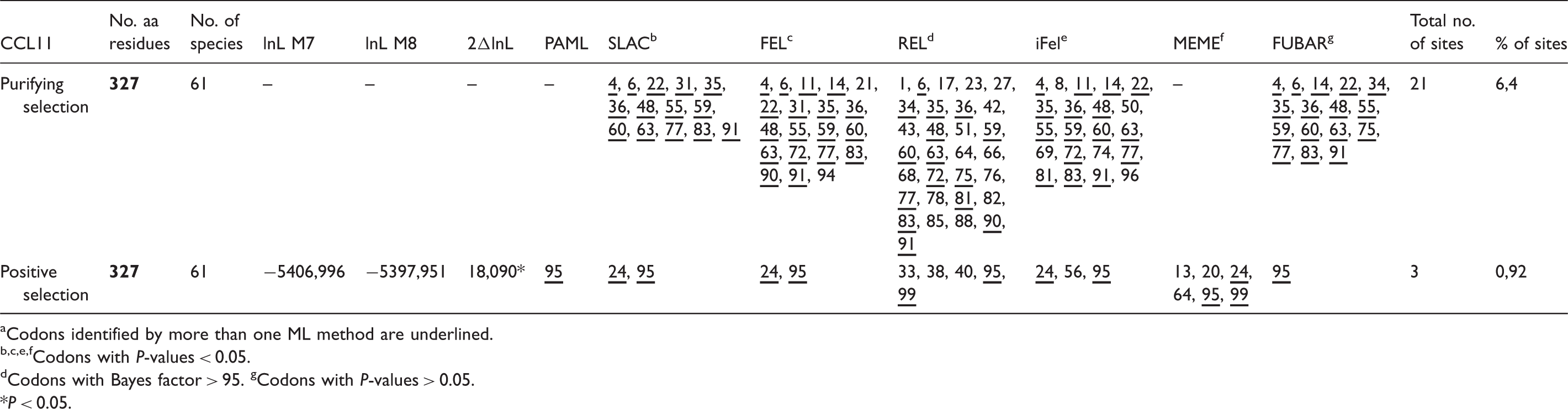
Codons identified by more than one ML method are underlined.
Codons with P-values < 0.05.
Codons with Bayes factor > 95. gCodons with P-values > 0.05.
P < 0.05.
Secretory proteins such as CCL11 only become functional after crossing the membrane and arriving to the appropriate cellular compartment and, consequently, being cleaved by signal peptidases.49,50 The signal peptide plays important roles in targeting and membrane insertion, and after being cleaved can also exert other functions such as protecting cells from being killed by other cells.49–52 Before cleavage, the signal peptide may have also important functions in protein folding and maturation. 52 Therefore, mutations in the signal peptide may interfere with such functions. Indeed, from the 23 aa that compose the CCL11 signal peptide, five were found to be under purifying selection, confirming the importance of their maintenance (Ser4, Ala6, Leu11, Ala14 and Leu22).
Signatures of positive selection in immune system genes tend to be associated with regions where the binding with other molecules occurs (proteins, receptors) and that can alter the proteins’ activity and, consequently, their biological roles.53–56 For CCL11, these regions had not been described, but the aa changes observed as positively selected show different polarity and charge that can cause changes in the protein. In addition, two of the positively selected sites (Pro95 and Pro99) are located in close vicinity to the stop codon that is mutated in several mammalian species, leading to proteins with different sizes.
As the CCL11 predicted secondary structures of Human and the European rabbit obtained in PsiPred were similar (data not shown), we used the human 3D structure available online (CCL11–CCR3 interaction) to locate the sites under selection (Figure 2). This confirmed their relevance for receptor binding. A previous study in lagomorphs identified one site under positive selection in chemokine ligand CCL5,
17
but this was located in the signal peptide that is cleaved in protein maturation.
3D structure of the CCL11–CCR3 complex. CCL11 appears in blue while CCR3 appears in dark gray. The purifying selected aa identified in this study and located near important sites for ligand-receptor binding and interaction are marked in yellow; the positively selected sites are highlighted in light gray.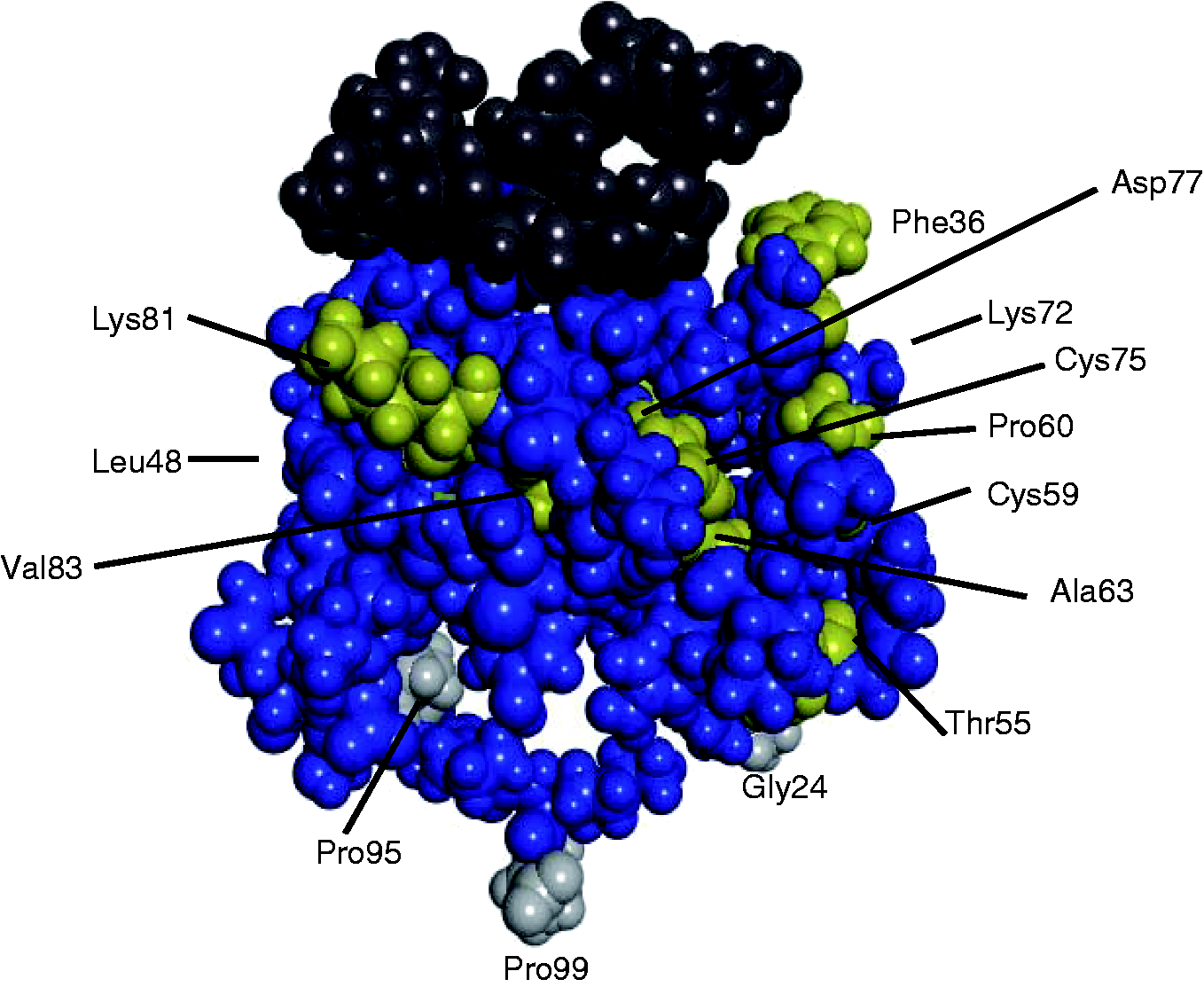
Conclusions
This work describes the detection of codons under positive and purifying selection in CCL11. Purifying selection may result from the proteins’ functional constraints, while an increase in diversity, probably as such mutations are advantageous in the host response against several agents, suggests positive selection. Our results identified 21 codons under purifying selection in sites located in regions important for ligand–receptor binding and activation, and in the signal peptide and three sites under positive selection near signal peptide and the stop codon. We observed that CCL11 is functional in lagomorphs, with Pygmy rabbit having a longer protein owing to a mutation in the stop codon. Further functional studies should evaluate the biological implications of this extension.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work is funded by National Funds through FCT—Foundation for Science and Technology under the Project FCT-ANR/BIA-BIC/0043/2012. FCT also supported the doctoral grants of Fabiana Neves (ref.:SFRH/BD/81916/2011) and the FCT Investigator grant of Joana Abrantes (ref.: IF/01396/2013). ‘Genomics Applied To Genetic Resources’ co-financed by North Portugal Regional Operational Programme 2007/2013 (ON.2 – O Novo Norte), under the National Strategic Reference Framework (NSRF), through the European Regional Development Fund (ERDF), also supported this work.
