Abstract
ILs, as essential innate immune modulators, are involved in an array of biological processes. In the European rabbit (Oryctolagus cuniculus) IL-1α, IL-1β, IL-2, IL-4, IL-8, IL-10, IL-12A, IL-12B, IL-15 and IL-18 have been implicated in inflammatory processes and in the immune response against rabbit hemorrhagic disease virus and myxoma virus infections. In this study we characterized these ILs in six Lagomorpha species (European rabbit, pygmy rabbit, two cottontail rabbit species, European brown hare and American pika). Overall, these ILs are conserved between lagomorphs, including in their exon/intron structure. Most differences were observed between leporids and American pika. Indeed, when comparing both, some relevant differences were observed in American pika, such as the location of the stop codon in IL-1α and IL-2, the existence of a different transcript in IL8 and the number of cysteine residues in IL-1β. Changes at N-glycosylation motifs were also detected in IL-1, IL-10, IL-12B and IL-15. IL-1α is the protein that presents the highest evolutionary distances, which is in contrast to IL-12A where the distances between lagomorphs are the lowest. For all these ILs, sequences of human and European rabbit are more closely related than between human and mouse or European rabbit and mouse.
Introduction
ILs are polypeptides of low molecular mass involved in several biological activities, including immunity, inflammation, inflammatory diseases, hematopoiesis, oncogenesis and fertility, among others.1–3 In vertebrates, many of these proteins participate in host defense with complementary and conflicting roles in induction, regulation and functioning of the immune system by regulating growth, differentiation, effector functions and survival of cells.4,5 ILs are crucial for the immune response. They are produced as an integral part of the innate immune response, and have the ability to influence the result and nature of adaptive immune response.4–8 Despite these functions, ILs are also considered important therapeutic targets;9–11 therefore, any changes in their sequence or structure may lead to alteration in their normal functioning. In mammals, ILs encoding genes are among the ones with faster evolution,4,6 and 28 of the 46 known ILs present signatures of positive selection. 12
Although being well characterized in most mammalian groups, little is known about ILs in lagomorphs, with the exception of the European rabbit (Oryctolagus cuniculus). The order Lagomorpha comprises two families, Ochotonidae and Leporidae, that diverged about 35 million years ago (mya). 13 While the family Ochotonidae comprise only one genus (Ochotona or pikas), the family Leporidae comprises 11 genera of hares and rabbits that are widely distributed: pygmy rabbit (genus Brachylagus), riverine rabbit (genus Bunolagus), striped rabbit (genus Nesolagus), European rabbit (genus Oryctolagus), Amami rabbit (genus Pentalagus), Bunyoro rabbit (genus Poelagus), red rock rabbit (genus Pronolagus), volcano rabbit (genus Romerolagus), cottontail rabbits (Sylvilagus sp.), hispid hare (genus Caprolagus) and true hares and jackrabbits (Lepus sp.).13–15 The ones that are more closely related to the Oryctolagus genus are Bunolagus, Caprolagus and Pentalagus with divergence times of ∼7, ∼8 and ∼9 mya, respectively. In contrast, Nesolagus, Poelagus and Pronolagus are less related to Oryctolagus, with divergence times of ∼15 mya. 13 The other two leporidae genera with some information for ILs, Sylvilagus and Lepus, diverged from Oryctolagus at ∼12 mya.13,16–18
The European rabbit is one of the most used laboratory animal models for immunologic research, including the study of atherosclerosis, 19 intestinal immunity, 20 arthritis, 21 cancer, 22 Alzheimer’s disease 23 and several viral infections.24–26 In addition, the European rabbit is a key species in the Mediterranean ecosystem, where it is strongly affected by two viral diseases, rabbit hemorrhagic disease (RHD) and myxomatosis.27–32 RHD is caused by a single-stranded RNA virus, the rabbit hemorrhagic disease virus (RHDV), while myxomatosis is caused by the myxoma virus (MYXV), a double-stranded DNA virus.
Rabbit resistance to both viral diseases is highly dependent on the immune response of the host in order to develop an effective adaptive immune response to control these viruses.25,33 ILs are not only important in the European rabbit immune response against RHDV and MYXV infections,24,25 but also in inflammatory processes. 34 For RHDV, IL-1, IL-2, IL-6, IL-8 and IL-10 are among the most important ILs. Indeed, young European rabbits infected with RHDV showed an increase of IL-1 during an early stage of infection until 18 h post-infection, with a decrease to normal values at 24 h of infection, while IL-8 is particularly increased at 24 h after infection, probably owing to an increase of leukocyte migration to the site of infection. 25 In contrast, in infected adult rabbits, IL-10 is significantly increased in the course of the disease.25,26
For MYXV, IL-12, IL-15 and IL-18 have an important anti-viral activity. Indeed, recombinants of myxoma virus expressing human IL-12, despite being similar to the wild type, do not induce myxomatosis in rabbits. 35 IL-15 prevents lethal myxomatosis in the New Zealand rabbit breed through the stimulation of an immune response that leads to the elimination of the viral infection. 36 Poxviruses developed the ability to block IL-18 by interfering with proteins crucial for IL-18 activation or function highlighting the importance of its anti-viral activity.37–39 In contrast, expression of European rabbit IL-4 by recombinant myxoma virus strains increases virus virulence and overcomes genetic resistance in wild rabbits. 40
Considering the important biological role in the European rabbit immune response, we performed a genetic characterization of IL-1, which includes two biologically similar antagonist proteins IL-1α and IL-1β,41–45 IL-2, IL-4, IL-8, IL-10, IL-12A, IL-12B, IL-15 and IL-18 in five Lagomorpha genera (Oryctolagus, Brachylagus, Sylvilagus, Lepus and Ochotona).
Materials and methods
Samples of European rabbit (O. c. cuniculus and O. c. algirus), pygmy rabbit (Brachylagus idahoensis), cottontail rabbits (brush rabbit, Sylvilagus bachmani, and eastern cottontail, Sylvilagus floridanus), European brown hare (Lepus europaeus) and American pika (Ochotona princeps) were provided by the CIBIO Lagomorpha tissue collection. Genomic DNA (gDNA) was extracted using the EasySpin Genomic DNA Minipreps Tissue Kit (Citomed, Torun, Poland) according to the manufacturer’s instructions. Total RNA was extracted by using the RNeasy Mini Kit according to the manufacturer’s instructions (Qiagen, Hilden, Germany) from one specimen of European rabbit, European brown hare, eastern cottontail and American pika. Complementary DNA (cDNA) was synthesized using oligo(dT) as primers and SuperScript III reverse transcriptase (Invitrogen, Carlsbad, CA, USA). The European rabbit and American pika IL sequences were retrieved from public databases (accession numbers are given in bold in Figure 1). PCR amplification was performed with the Multiplex PCR Kit (Qiagen) using several pairs of primers designed according to the retrieved sequences (Supplementary Material Table 1). Sequencing was performed on an ABI PRISM 310 Genetic Analyzer (PE Applied Biosystems, Foster City, CA, USA) and PCR products were sequenced in both directions. The sequences obtained were submitted to GenBank (accession numbers: KT216045-KT216070; KT273911-273919; KT279631-KT279692).
Alignment of the studied ILs for the different lagomorphs: (a) IL-1α; (b) IL-1β; (c) IL-2; (d) IL-4; (e) IL-8; (f) detail of the splicing region of exon 2 of IL-8, with the splicing regions underlined; (g) IL-10; (h) IL-12 A; (i) IL-12B; (j) IL-15; (k) IL-18. GenBank and Ensembl accession numbers are indicated in bold for the retrieved sequences. Positively selected aa are boxed according to Neves et al.,
12
N-Glycosylation sites are shaded in light gray and cysteine residues are shaded in dark gray. (*) Represent stop codons; (–) represent deletions; *1 and *2 represent alleles. The numbering is according to the European rabbit sequences. The signal peptide and indels, indicated as (–), were included in the numbering.
The PHASE program, built into the software DnaSP, 46 was used to reconstruct the haplotype phases of the obtained sequences that were aligned using MUltiple Sequence Comparison by Log-Expectation (MUSCLE) available at http://www.ebi.ac.uk/. 47
Putative N-glycosylation sites were predicted using the NetNGlyc 1.0 server available at http://www.cbs.dtu.dk/services/NetNGlyc/. 48
Predicted splicing sites were determined using the NetGene2 server available at http://www.cbs.dtu.dk/services/NetGene2/.49,50
The number of amino acid (aa) differences per site between sequences was estimated in MEGA6 with the following options: 51 bootstrap method (1000 replicates), P-distance as model and pairwise deletion for gaps/missing data treatment. A maximum likelihood (ML) approach was used to estimate, for each gene, the phylogenetic relationships between the aa sequences. The ML trees were estimated in MEGA6 by using the best-fit nucleotide substitution model predicted by the same software and bootstrap 1000 replicates. In order to simplify the outputs and avoid duplicated sequences, only one sequence from each species was used.
The secondary structure of each IL was predicted using PsiPred (http://bioinf.cs.ucl.ac.uk/psipred/)52,53 and DiAminoacid Neural Network Application (DiANNA) (http://clavius.bc.edu/∼clotelab/DiANNA/). 54 These methods predict protein cysteines that create permanent structural disulfide bonds. PsiPred uses Position Specific Iterated–BLAST (PSI-BLAST) searches of the non-redundant protein sequence database to obtain evolutionary information used to predict the secondary structure of the query protein. DiANNA is a neural network trained to recognize cysteines in an oxidized state (sulfur covalently bonded) telling them apart from those in a reduced state.
Results
Summary of the alterations observed between lagomorph species for the ILs studied.
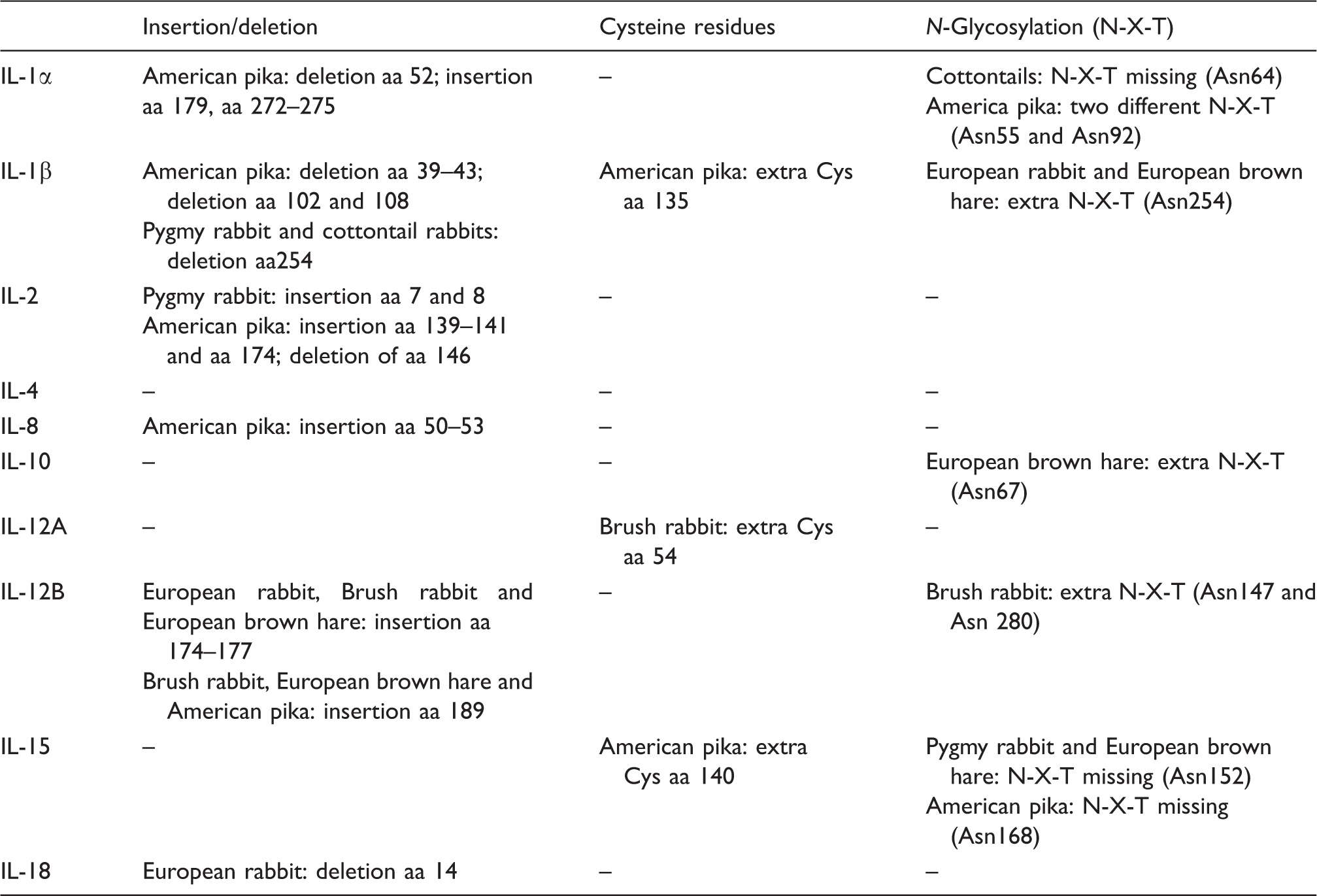

IL-1
IL-1 is composed by two proteins, IL-1α and IL-1β. According to the public databases NCBI, Ensembl and Uniprot, in the European rabbit these proteins are located in tandem in the reverse strand of chromosome 2, while in the American pika they are located in the forward strand.
IL-1α
In lagomorphs, IL-1α is organized into seven exons, with six of them being coding exons. The coding sequence comprises 801 base pairs (bp) that translate into a protein with 267 amino acids (aa). However, for American pika, and like the human IL-1α, the coding sequence has 813 bp translating into a protein with 271 aa (Figure 1a). Some differences existed between lagomorphs: American pika presents an aa deletion at position 52 of the European rabbit and an insertion of five aa at positions 179 and from 272 to 275. All cysteine residues are conserved between lagomorphs (Cys14 and Cys47). Three potential N-glycosylation sites were detected for all leporids (Asn64, Asn103 and Asn144), but Asn64 is absent in the cottontail rabbits studied. For American pika, there are two other potential N-glycosylation sites (Asn55 and Asn92).
IL-1β
In lagomorphs, IL-1β comprises six coding exons, translating into proteins with different aa lengths: 268 aa in the European rabbit and 261 aa in American pika (Figure 1b). In the American pika sequence we observed three deletions: five aa between positions 39 and 43, including the deletion of two cysteine residues that are conserved in all leporids; the two other deletions are located at aa 102 and 108. Between leporids there are nine conserved cysteines at sites 35, 42, 43, 54, 81, 105, 117, 126 and 189. In the American pika sequence, only four cysteines are conserved (Cys81, Cys117, Cys126 and Cys189), and an extra cysteine is found at position 135. All lagomorphs have a putative N-glycosylation site at Asn58; European rabbit and European brown hare have an additional putative N-glycosylation site at Asn254 that had been lost in both the brush rabbit and in the eastern cottontail rabbit due to a deletion of this aa.
IL-2
Regarding the IL-2 sequence, this protein is highly conserved between European rabbit, human, cat and horse. 34 In the European rabbit, IL-2 is located in the reverse strand of chromosome 15, with four coding exons that translate into a protein with 153 aa. In American pika IL-2 also has four exons that translate into a 156aa protein (Figure 1c). Differences between lagomorphs include the insertion of two aa in pygmy rabbit (Met7 and Phe8) and four aa in American pika (Thr139, Ser140, Arg141 and Asn174). In American pika we also observed a deletion of one aa that corresponds to position 146 of European rabbit. All cysteine residues are conserved between lagomorphs (Cys11, Cys94, Cys144 and Cys164). The search of putative N-glycosylation sites identified one site (Asn127) common to all lagomorphs.
IL-4
In the European rabbit, IL-4 is located in chromosome 3 (forward strand). In the European rabbit and American pika, IL-4 is organized into four coding exons that translate into a protein of 147 aa (Figure 1d). No alterations, including insertions/deletions, were observed between IL-4 of lagomorphs. Also, cysteine residues were conserved (Cys13, Cys48, Cys70, Cys89, Cys115 and Cys123) and the same potential N-glycosylation site was detected (Asn62).
IL8
IL-8 from European rabbit is located in the reverse strand of chromosome 15 and it is encoded by four exons that translate into a 101-aa protein. For American pika, the coding sequence available in Ensembl.org (ENSOPRT00000002239) is only known by projection and it is incomplete by not starting with an initiation codon and by missing the stop codon. Nevertheless, we successfully amplified IL-8 for American pika from both gDNA and cDNA (Figure 1e, f). Our results showed that American pika has an insertion of four aa (Gly50, Phe51, Pro52 and Thr53). The cysteine residues are conserved among lagomorph species (Cys48, Cys61, Cys63, Cys93 and Cys109) and there are no putative N-glycosylation sites.
IL-10
IL-10 from European rabbit is located in the forward strand of chromosome 2 and, in both the European rabbit and American pika, it is composed of five coding exons that translate into a protein with 178 aa (Figure 1g). The cysteine residues are conserved between species (Cys8, Cys9 Cys30, Cys80, Cys126 and Cys132). The search for N-glycosylation sites revealed two potential sites in leporids (Asn100 and Asn134). Ans67 was also predicted for European brown hare and American pika has only one of the sites predicted for N-glycosylation (Asn134).
IL-12
IL-12 is a heterodimeric protein composed by two proteins, IL-12A and IL-12B. For European rabbit, and according to the public databases NCBI and Ensembl, the encoding genes are located in the forward strand of chromosome 14 and in the reverse strand of chromosome 3, respectively.
IL-12A
IL-12A from European rabbit is composed of seven coding exons that translate into a protein with 219 aa. For American pika, there are two different coding sequences available in NCBI (XM_012930663.1) and Ensembl (ENSOPRT00000006274). When compared with the European rabbit sequence, the American pika sequence available in NCBI has a start codon 109 aa upstream, while the sequence available in Ensembl is incomplete, as it lacks an initiation codon and aa information in the middle of the sequence. We were not able to amplify IL12A from gDNA or cDNA of American pika (Figure 1h). All cysteine residues are conserved between leporids (Cys2, Cys71, Cys98, Cys119, Cys130, Cys144, Cys157 and Cys230). In addition, brush rabbit presents an extra cysteine at position 54. Four potential N-glycosylation sites were detected for all leporids (Asn73, Asn127, Asn141 and Asn214).
IL-12B
IL-12B from European rabbit and American pika is composed by six coding exons that translate into a protein with 324 and 329 aa, respectively. Despite all the attempts, we were unable to amplify IL-12B from cDNA of any of the lagomorphs studied. Successful amplification from gDNA for all leporids showed that in the middle of exon 4 there is an insertion of four aa (Thr174 to Ala177) that is absent in the predicted European rabbit sequence available in NCBI (XM_002710347.2). However, this insertion is present in human, mouse and rat cDNA (Figure 1i). All cysteine residues (Cys2, Cys50, Cys90, Cys131, Cys142, Cys170, Cys194 and Cys200, Cys275, Cys291, Cys309 and Cys336) and the two potential N-glycosylation sites (Asn135, Asn223) are conserved between lagomorphs. In addition, brush rabbit evidenced two other potential N-glycosylation sites (Asn147 and Asn280).
IL-15
In the European rabbit, IL-15 is located in chromosome 15 (reverse strand) and it is organized into six coding exons that translate into a protein with 162 aa. There are two sequences described for the European rabbit in NCBI, with the accession numbers NM_001082216.2 and XM_008267272.1, being the latter a variant with an insertion of one aa (Gln127) (Figure 1j). For American pika the information available in NCBI and Ensembl is different for the 5’ coding region. Amplification was only successful using the primers constructed according to the American pika IL15 sequence available in Ensembl (ENSOPRT00000008393). All cysteine residues are conserved in lagomorphs (Cys39, Cys42, Cys62, Cys107, Cys114, Cys158 and Cys161), although American pika has an extra cysteine (Cys40). Four potential N-glycosylation sites were detected in lagomorphs, Asn132, Asn143, Asn152 and Asn168, but Asn152 is absent in pygmy rabbit and in European brown hare, and Asn168 is absent in American pika.
IL-18
The European rabbit’s IL-18 is located in the forward strand of chromosome 1 and is organized into five coding exons that translate into a protein with 192 aa. For American pika there are two different predicted sequences available, one from NCBI (XM_004585222.1) and the other from Ensembl (ENSOPRG00000012703). When compared with the other lagomorphs, the American pika sequence from NCBI had an insertion of six aa in the first exon, while exon 5 presented low aa identity and a smaller size. The sequence from Ensembl is incomplete as it lacks an initiation codon and exon 5 (Figure 1k). For this reason, and despite all attempts, we were unable to amplify IL-18 from gDNA or cDNA of American pika. The European rabbit has a deletion at aa 14, whereas all the other leporids have a Gln residue. All cysteine residues are conserved between leporids (Cys16, Cys80, Cys110, Cys118 and Cys169). Our search revealed that Asn135 is the only putative N-glycosylation site.
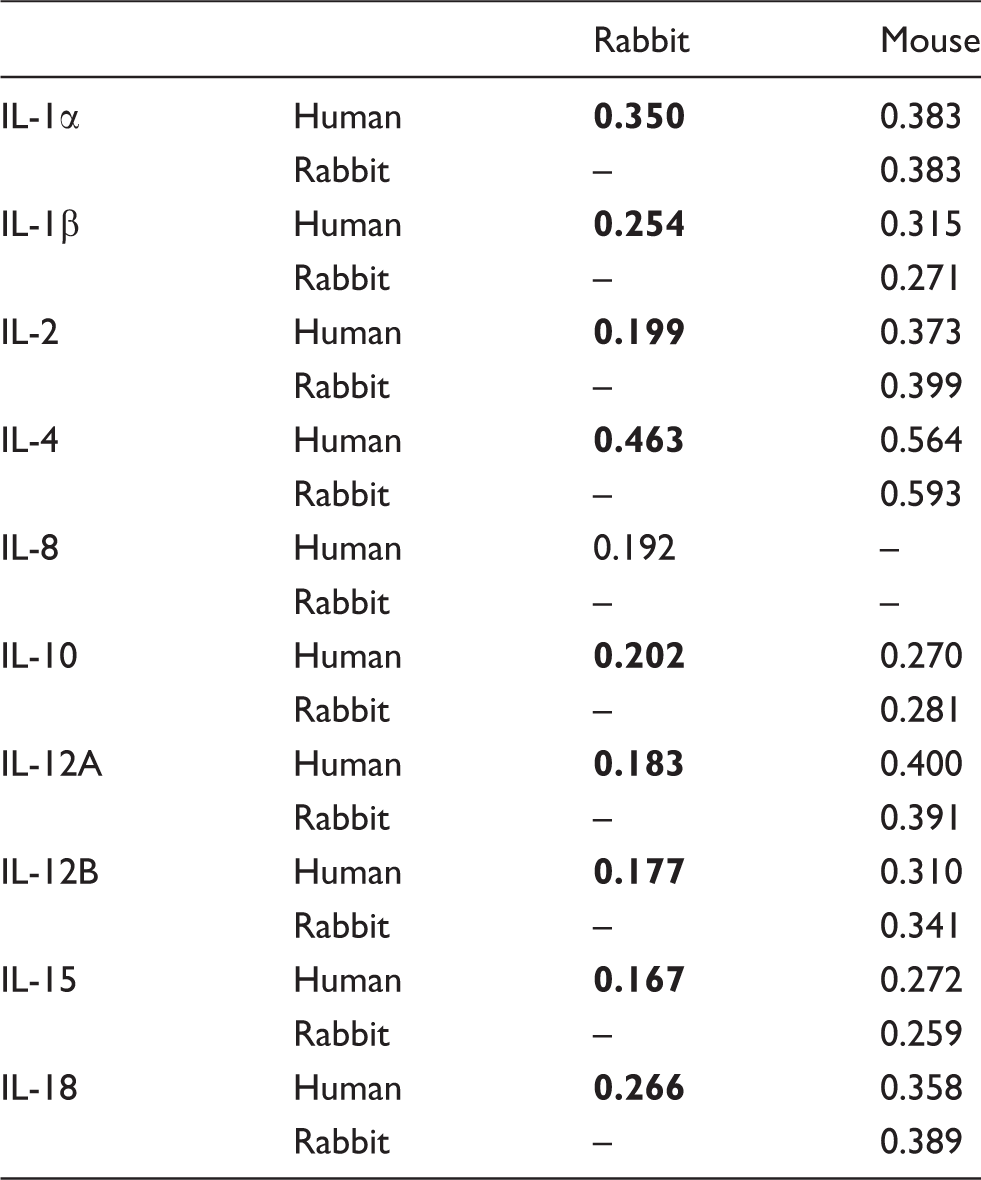
Aa distances between rabbit, mouse and human for the different ILs studied (the lowest values for each IL are in bold).
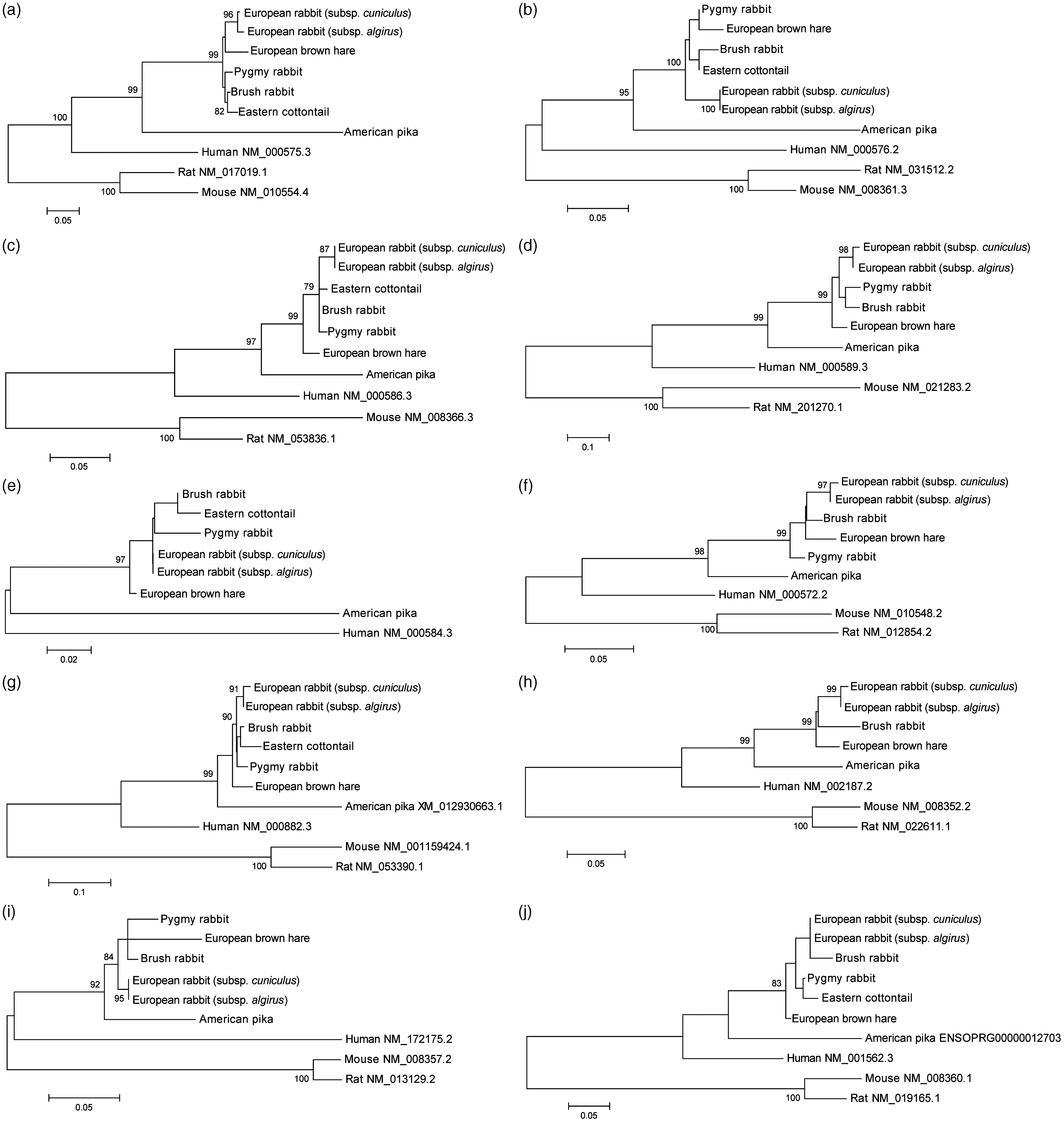
ML trees of the ILs studied: (a) IL-1α; (b) IL-1β; (c) IL-2; (d) IL-4; (e) IL-8; (f) IL-10; (g) IL-12A; (h) IL-12B; (i) IL-15; (j) IL-18. Only bootstrap values ≥ 75% are shown. In order to facilitate visualization, only one sequence/allele of each species was used. GenBank and Ensembl accession numbers are indicated for human, rodents and, in some cases, American pika sequences.
Discussion
ILs are important proteins for the innate and adaptive immune responses. In the European rabbit IL-1α, IL-1β, IL-2, IL-4, IL-8, IL-10, IL-12A, IL-12B, IL-15 and IL-18 have been implicated in the immune response against two highly fatal viral diseases, RHD and myxomatosis, and in inflammatory processes. Recently, we described a mutation in the European rabbit and Amami rabbit (Pentalagus furnessi) IL-6 stop codon that leads to a larger protein with a considerable increase of the number of cysteines. 55 For the ILs included in this study, no significant alterations were observed in any of the stop codons. Interestingly, American pika IL-1α and IL-2 sequences presented an insertion of four and one aa, respectively, in the site where the others lagomorphs have a stop codon.
Some differences were observed between the European rabbit and the American pika sequences reported here and those already available in public databases (Supplementary Material Table 3). The major differences are in American pika IL-8 and in the European rabbit IL-12B, where both sequences have an insertion of four aa. However, the sequences available in public databases are predicted, which might explain the differences observed. The remaining punctual differences are due to single polymorphisms arising from species diversity.
Alternative splicing is an important step for the production of mature mRNA that leads to protein diversity and may occur by exon skipping (38%); alternative 5’ or 3’ spliced sites (26%); intron retention (3%); mutually exclusive exons, alternative promoters or multiple polyadenylation sites (33%).56,57 Exon skipping occurs by cleavage of specific exon–intron motifs located in the 5’ and 3’ termini regions of the intron. These motifs include, in decreasing order of relevance, GT–AG, CG–AG, AC–AT or AT–AC in the 5’ and 3’ regions, respectively.58–60 Studies in the European rabbit reported alternative splicing in several ILs that leads to novel and functional proteins. Indeed, in IL-2 and IL-7 alternative splicing occurs with exclusion of exon 2;34,61 for IL-4 the presence of two variants (δ2, δ3) results from splicing out of exons 2 and 3, respectively; 34 for IL-10 a spliced variant was described with the spliced region occurring between exon 5 and the 3’ UTR region. 34 Beside these variants, abnormal transcripts were also characterized for European rabbit IL-4 and IL-10 (IL-4 int2A, IL-4 int3A, IL-4 int3B and IL-10/C). 34 Although, more recently Mage and Mage suggested that there is no frameshit in exon 2 of IL-4. 62 When comparing American pika IL-8 sequence with the corresponding European rabbit sequence, we observed an insertion of four aa in the 5’ region of exon 2, which is derived from the 3’ region of the preceding intron. This suggests that in American pika the splicing occurred in a CA motif located 12 bp upstream of the motif found in the European rabbit (Figure 1f). These results were further confirmed by analyzing the human and American pika sequences using NetGene2.49,50 Indeed, for human IL-8 the splicing is predicted to occur by NetGene2 at nucleotide position 18 (Figure 1f) (CCCCCAACA|GGTGCAGTTTT; with a 0.77 confidence value), while for American pika the confidence value is of 0.17, with the most likely splicing motif (confidence value of 0.65) being located at nucleotide position 6 (TAATTTTCA|GGTTTCCCCA). Thus, IL-8 from American pika has a different transcript.
Disulfide bonds and glycosylation are important for protein protection against denaturation and proteolytic degradation.
63
Disulfide bonds are formed between the thiol groups of cysteine residues and ensure important roles in folding, stability, function and in the regulation of protein activity, being well conserved between species.63–66 In the studied ILs, and with the exception of IL-1β, the cysteine residues are well conserved in lagomorphs. For American pika IL-1β, and when comparing with other lagomorphs, the number and location of five out of nine cysteines residues is different. The analysis of protein secondary structure using PsiPred52,53 and DiANNA
54
indicated that the loss of cysteine residues reduces the predicted bonds from four in the European rabbit (Cys35-Cys115; Cys42-Cys186; Cys43-Cys123 and Cys54-Cys81) to two in American pika (Cys108-Cys125 and Cys116-Cys179). These differences lead to the formation of an extra helix (Gly84 to Phe89) in American pika (Figure 3).
IL-1β PsiPred sequence analysis results for the (a) European rabbit and (b) American pika (the extra helix is marked with a black circle).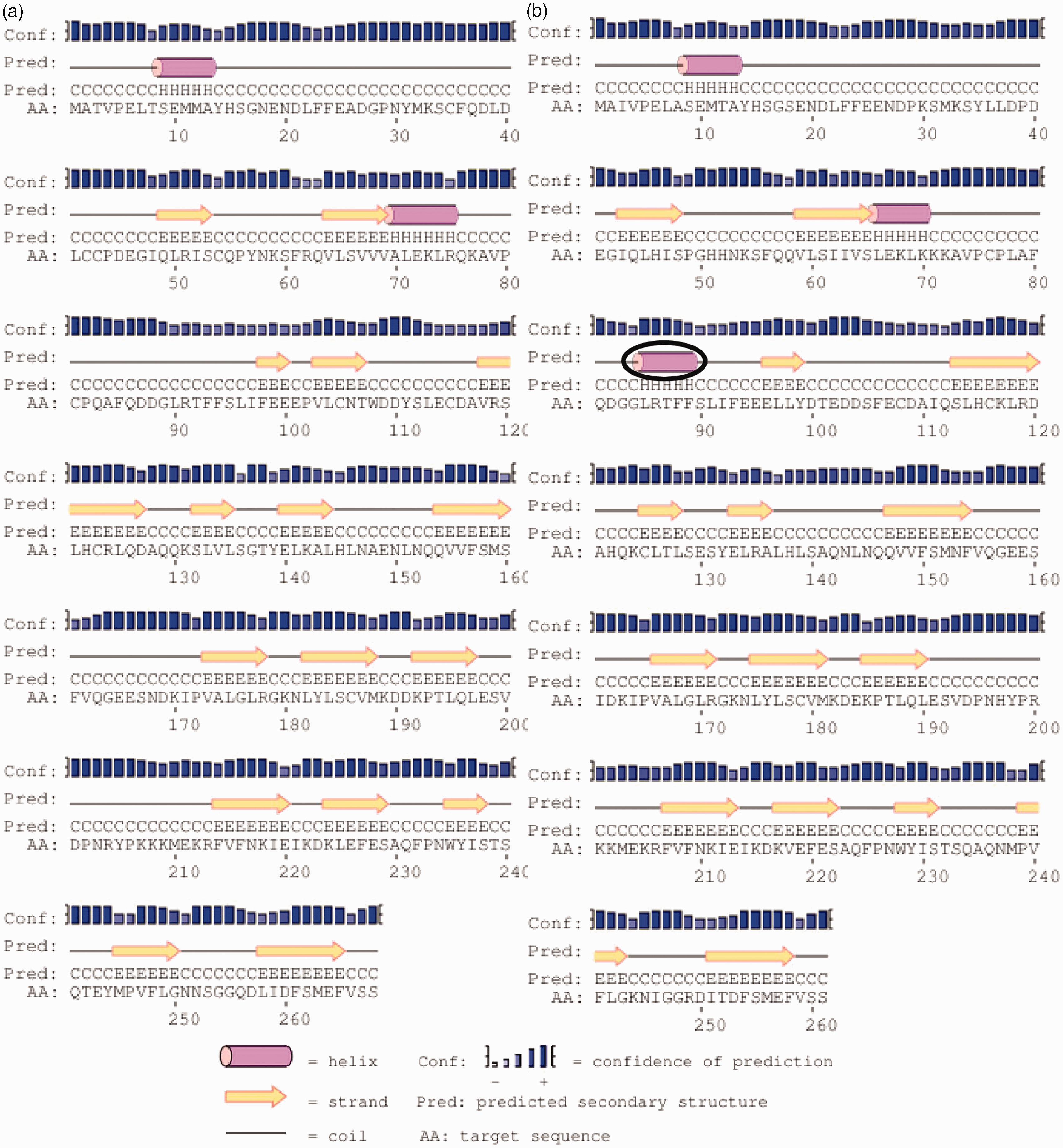
The glycosylation process is also important for several functions that include protein folding and interaction with cell surface receptors.67,68 Thus, glycosylation can lead to protein diversity and modulation of protein properties.67,69–71 Putative N-glycosylation sites prediction showed some differences between lagomorphs, in particular for IL-1α, IL-1β, IL-10, IL-12B and IL-15; however, the consequences of these alterations should be considered in future studies.
Animal models are important tools for the study of human diseases. Historically, the European rabbit was the first animal model for immunological studies. Indeed, some of the foundations of molecular immunology were laid with the use of the European rabbit reviewed in Pinheiro et al. 72 Nevertheless, over the last decade the European rabbit as a research animal model has been replaced by the mouse model due to its smaller size, lower cost, ease of breeding, etc. 73 However, the choice of mouse as the best model to study human diseases has been controversial due to the high variability of results observed between mouse and human, especially when the focus is inflammatory diseases.73–78 A previous study in ILs showed that European rabbit–human sequences are more related than mouse–human or European rabbit–mouse sequences. 34 Overall, the results of the phylogenetic analysis are in agreement with the evolutionary topology described for lagomorphs, 13 where the American pika is the most divergent species. Nevertheless, the ML trees depicted in Figure 2 correspond to gene trees that do not necessarily reflect the true species tree.79,80 Overall, the human IL sequences appear more closely related to those from European rabbit than with mouse and rat. This pattern had been already observed for other molecular markers and might suggest that the European rabbit might be a more appropriate animal model for the study of immunity in humans.34,81–85
Conclusions
In this study we sequenced and characterized ten ILs for six lagomorph species and in the two European rabbit subspecies. As expected, most differences were observed between leporids and American pika. While these differences may lead to alterations in the biological roles of these proteins, the overall genomic organization, the location of the cysteine residues and the presence of N-glycosylation sites is well conserved. In addition, and according to divergence between the European rabbit and human for the ILs studied, the European rabbit might be a more suitable animal model for studies in the human innate immunity.
Footnotes
Acknowledgements
We would like to thank Jeff Wilcox and Dr. Michael Hamilton from Blue Oak Ranch Reserve, University of California, Berkeley, USA, for providing brush rabbit (Sylvilagus bachmani) tissue samples; and Dr. Jay Storz from School of Biological Sciences, University of Nebraska, USA, for the liver sample of Ochotona princeps. We are grateful to Dr. Janet Rachlow, Dr. Lisette Waits and Dr. Caren Goldberg from Department of Fish and Wildlife Sciences, University of Idaho, USA, for providing pygmy rabbit (Brachylagus idahoensis) tissue samples.
Funding
This work is funded by FEDER funds through the Operational Programme for Competitiveness Factors—COMPETE, and by National Funds through Foundation for Science and Technology (FCT) under projects PTDC/BIA-ANM/3963/2012 and FCOMP-01-0124-FEDER-028286. FCT also supported the doctoral grants of Fabiana Neves (ref.: SFRH/BD/81916/2011) and the FCT Investigator grant of Joana Abrantes (ref.: IF/01396/2013). ‘Genomics Applied To Genetic Resources’, co-financed by North Portugal Regional Operational Programme 2007/2013 (ON.2 – O Novo Norte), under the National Strategic Reference Framework (NSRF), through the European Regional Development Fund (ERDF), also supported this work.
Conflict of interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
